

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124966.7 + phase: 0 /pseudo
(1084 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
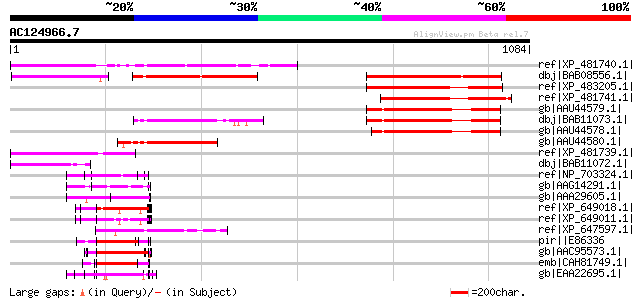
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_481740.1| unknown protein [Oryza sativa (japonica cultiva... 385 e-105
dbj|BAB08556.1| unnamed protein product [Arabidopsis thaliana] g... 357 2e-96
ref|XP_483205.1| unknown protein [Oryza sativa (japonica cultiva... 322 6e-86
ref|XP_481741.1| unknown protein [Oryza sativa (japonica cultiva... 315 5e-84
gb|AAU44579.1| hypothetical protein AT5G48090 [Arabidopsis thali... 283 2e-74
dbj|BAB11073.1| unnamed protein product [Arabidopsis thaliana] g... 283 2e-74
gb|AAU44578.1| hypothetical protein AT5G48090 [Arabidopsis thali... 281 6e-74
gb|AAU44580.1| hypothetical protein AT5G48090 [Arabidopsis thali... 190 2e-46
ref|XP_481739.1| unknown protein [Oryza sativa (japonica cultiva... 136 4e-30
dbj|BAB11072.1| unnamed protein product [Arabidopsis thaliana] g... 102 7e-20
ref|NP_703324.1| glutamic acid-rich protein (garp) [Plasmodium f... 99 1e-18
gb|AAG14291.1| glutamic acid-rich protein [Plasmodium falciparum] 93 6e-17
gb|AAA29605.1| glutamic acid-rich protein [Plasmodium falciparum... 92 1e-16
ref|XP_649018.1| hypothetical protein 354.t00010 [Entamoeba hist... 91 2e-16
ref|XP_649011.1| hypothetical protein 354.t00008 [Entamoeba hist... 91 2e-16
ref|XP_647597.1| hypothetical protein DDB0189820 [Dictyostelium ... 89 6e-16
pir||E86336 hypothetical protein F14O10.11 - Arabidopsis thalian... 89 8e-16
gb|AAC95573.1| orf 48 [Ateline herpesvirus 3] gi|11361081|pir||T... 88 2e-15
emb|CAH81749.1| conserved hypothetical protein [Plasmodium chaba... 87 2e-15
gb|EAA22695.1| hypothetical protein [Plasmodium yoelii yoelii] 87 3e-15
>ref|XP_481740.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|38637134|dbj|BAD03388.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|38424060|dbj|BAD01750.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 585
Score = 385 bits (988), Expect = e-105
Identities = 230/601 (38%), Positives = 321/601 (53%), Gaps = 69/601 (11%)
Query: 1 MASSDEEGEIVPDSVDGYWFENDKAEFVSLSSLTLLWSVNDVTCNSEAKVFLRGTTDNGL 60
M SSD++ E +V+ Y+F +D VS L + + + + V+LRG TD GL
Sbjct: 1 MMSSDDDLEPQLKAVENYYFVDDNDVPVSFDVLPFQFDAAEGVASFKKDVYLRGFTDGGL 60
Query: 61 QKIHKQIIGWRFELSFEQPEISVLLREKYWMTLLKPSKCFENTIRSVLVTVYWLHCVKWK 120
QK++KQ++ W+ L + PEI+VL E W+ LLKP +E TIRSVL+TV LH V+ +
Sbjct: 61 QKVYKQVVAWKLVLDGDSPEIAVLSTEGSWIALLKPRPSYEETIRSVLITVEMLHFVRRR 120
Query: 121 PEESRASILVKVLKEFSSFDITPSENDVLCHMALISEAVKRDTDLTKSKDVHTTKKLKVI 180
P +S + + F F + P E+D H LI +RD DL S+ + K K++
Sbjct: 121 PTDSEKDMWDHLYGVFERFVVRPLEDDFANHQNLIKLFAQRDPDLANSQVLQVFIKDKIM 180
Query: 181 VESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEE 240
E+ N++ + D ++E DI +E D+++E
Sbjct: 181 ---------------------EKTNEVGSNNLDNKREPDI--KQEPDIKQE--------- 208
Query: 241 DPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDIGYDTVCAIC 300
P D +E ++VE D +D D E DEE +D+VCAIC
Sbjct: 209 -PVAAGDEME----------EIVEEGIPDAPSNDDDDDEE-----DEEDGDLFDSVCAIC 252
Query: 301 DNGGEILPCEGRCLRSFHATLEDGRDSLCASLGYTRTEVNAFPNFYCENCKHKKHQCFAC 360
DNGGE+L CEG C+RSFHA + DG DS CA+LGYT+ EV A NF C+NC HK+HQCF C
Sbjct: 253 DNGGELLCCEGPCMRSFHAKIRDGEDSYCATLGYTKAEVKALKNFVCKNCDHKQHQCFVC 312
Query: 361 GKLGSSDESSNPEVFPCVTANCGHYYHPECVARLLYPGIDIGQEEMRKRIIIEKTFVCPL 420
G+L SD N +VF C A CGH+YHP CVA+LL+P EM K+I+ +F CP+
Sbjct: 313 GELEPSD-GPNAKVFLCNNATCGHFYHPRCVAQLLHPNSRNEASEMEKKIMAGFSFTCPV 371
Query: 421 HICSLCRKGENRNVHDLQFAMCRRCPKAYHRKCLPKEISFTYDYYTGIEMRAWDNLLDKR 480
H C C+ E+R LQFA+CRRCP++YHRKCLP+EISF GI RAW+ L KR
Sbjct: 372 HWCFHCKGLEDRTQEPLQFAVCRRCPRSYHRKCLPREISFEDINTQGIITRAWE--LSKR 429
Query: 481 ILMYCMNHKIVLELGTPARDHLIFPNKEVKRKIISTESLHREKDAIPLKKFFEDLLPNKT 540
IL+YC++H+I L++GTP RDH+ FP H EK A KK ++L K
Sbjct: 430 ILIYCLDHEIDLDIGTPPRDHIKFP--------------HVEKSAYSAKKKVKELAEKKR 475
Query: 541 LKPKMTIKERVGLQMGGSSKVMEKICSKQDTHMSTG-PVYFDRARKYLKVETMSGSNRSL 599
++ V + +K+ EK +K D G F+ + K +T S R+
Sbjct: 476 ---RICDDSYVSEPLQKRAKLNEKFNAKGDKSKKAGVKSEFEEVLESEKKKTRSLKKRTQ 532
Query: 600 P 600
P
Sbjct: 533 P 533
>dbj|BAB08556.1| unnamed protein product [Arabidopsis thaliana]
gi|15240501|ref|NP_200350.1| hydroxyproline-rich
glycoprotein family protein [Arabidopsis thaliana]
Length = 1332
Score = 357 bits (915), Expect = 2e-96
Identities = 173/284 (60%), Positives = 213/284 (74%), Gaps = 3/284 (1%)
Query: 745 ALQKLDEGCGIEEAKAICEPEIIRQLFTWQKQLKIYLAPFLHGRRYTSFGRHFTKIDKLK 804
AL+KL+EG IE+AKA+CEPE++ Q+ W+ +LK+YLAPFLHG RYTSFGRHFT +KL+
Sbjct: 660 ALKKLEEGGNIEDAKAVCEPEVLSQILKWKDKLKVYLAPFLHGARYTSFGRHFTNPEKLQ 719
Query: 805 EIVDRLHWYVQSGDTVLDFCCGANDFSCLMKSKLEQTGKLCSFKNYDLFQAKNDFNFEKR 864
+IVDRLHWY GD ++DFCCG+NDFSCLM +KLE+TGK C +KNYDLF AKN+FNFE++
Sbjct: 720 QIVDRLHWYADDGDMIVDFCCGSNDFSCLMNAKLEETGKKCLYKNYDLFPAKNNFNFERK 779
Query: 865 DWMSVQAEELPHGSNLIIGLNPPFGVRGSLANKFIDKALTFKPKLLILIVPKVT-RSQST 923
DWM+V +EL GS LI+GLNPPFGV SLANKFI KAL F+PK+LILIVP T R Q
Sbjct: 780 DWMTVSKDELEPGSKLIMGLNPPFGVNASLANKFITKALEFRPKILILIVPPETERFQFP 839
Query: 924 VEKQAGIYISICLSASNINQENTNVSTNFYLARLDRKKAGYNLIWADEEICSGKSFYLPG 983
A +Y SI L ++ + V + +L RLD+KK+ Y LIW D+ SG SFYLPG
Sbjct: 840 SISSAPLYHSITLIYRLLSL--SLVKSITFLNRLDKKKSSYVLIWEDKTFLSGNSFYLPG 897
Query: 984 SVDTRDKQLEDWNLKPPPLYLWSRPDWTAWHMQIARMHRHIKYD 1027
SV+ DKQLEDWNL PPPL LWSR D+ A H +IA H H+ D
Sbjct: 898 SVNEEDKQLEDWNLVPPPLSLWSRSDFAAKHKKIAEKHCHLSRD 941
Score = 270 bits (690), Expect = 2e-70
Identities = 135/265 (50%), Positives = 173/265 (64%), Gaps = 10/265 (3%)
Query: 256 PEEEN--DMVESAEEDPEEENDKSDGEGVLNLDEEQDIGYDTVCAICDNGGEILPCEGRC 313
P+E+N D +ED +D+ + LD+E D +++VCAICDNGGEIL CEG C
Sbjct: 188 PDEDNAKDDFIVGDEDTYVASDEDE------LDDEDDDFFESVCAICDNGGEILCCEGSC 241
Query: 314 LRSFHATLEDGRDSLCASLGYTRTEVNAFPNFYCENCKHKKHQCFACGKLGSSDESSN-P 372
LRSFHAT +DG DSLC SLG+ + +V A ++C NC+HK HQCF C LGSSD SS
Sbjct: 242 LRSFHATKKDGEDSLCDSLGFNKMQVEAIQKYFCPNCEHKIHQCFICKNLGSSDNSSGAA 301
Query: 373 EVFPCVTANCGHYYHPECVARLLYPGIDIGQEEMRKRIIIEKTFVCPLHICSLCRKGENR 432
EVF CV+A CG++YHP CV R L G + + E +R II + CPLH CS+C GE +
Sbjct: 302 EVFQCVSATCGYFYHPHCVTRRLRLG-NKEESEALERQIIAGEYTCPLHKCSVCENGEVK 360
Query: 433 NVHDLQFAMCRRCPKAYHRKCLPKEISFTYDYYTGIEMRAWDNLLDKRILMYCMNHKIVL 492
+LQFA+CRRCPK+YHRKCLP+EISF I RAWD LL R+L+YC H+I
Sbjct: 361 TDSNLQFAVCRRCPKSYHRKCLPREISFEDIEDEDILTRAWDGLLHNRVLIYCQEHEIDE 420
Query: 493 ELGTPARDHLIFPNKEVKRKIISTE 517
EL TP RDH+ FP E ++ + +
Sbjct: 421 ELLTPVRDHVKFPFTEEQKVFVKEQ 445
Score = 143 bits (361), Expect = 3e-32
Identities = 78/211 (36%), Positives = 125/211 (58%), Gaps = 9/211 (4%)
Query: 5 DEEGEIVPDSVDGYWFENDKAEFVSLSSLTLLWSVNDVTCNSEAKVFLRGTTDNGLQKIH 64
+EE VP S Y+FE+D E VS + L + WSV + S +LRG +DNGL +H
Sbjct: 8 EEEDFSVPQSASNYYFEDDDKEPVSFARLPIQWSVEEKVDGSGLGFYLRGRSDNGLLPLH 67
Query: 65 KQIIGWRFELSFEQPEISVLLREKYWMTLLKPSKCFENTIRSVLVTVYWLHCVKWKPEES 124
K + WR++LS QPEISVL ++ W+ L +P K + IR+VLVT++ + ++ P+ S
Sbjct: 68 KLVKAWRYDLSNFQPEISVLTKDNIWIKLEEPRKSYGELIRTVLVTLHSIQFLRRNPQAS 127
Query: 125 RASILVKVLKEFSSFDITPSENDVLCHMALISEAVKRDTDLTKSKDV---HTTKKLKVIV 181
++ K+ + S+D+ PS+ND++ H+ LI+EA KRD +L SK + T K K +
Sbjct: 128 EKALWEKLTRSLRSYDVKPSQNDLVDHIGLIAEAAKRDRNLANSKFILAFLTKKPTKRRL 187
Query: 182 ESEENPE------QENDIVESEEEDPEEEND 206
E+N + E+ V S+E++ ++E+D
Sbjct: 188 PDEDNAKDDFIVGDEDTYVASDEDELDDEDD 218
>ref|XP_483205.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|42407765|dbj|BAD08911.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 694
Score = 322 bits (824), Expect = 6e-86
Identities = 155/285 (54%), Positives = 191/285 (66%), Gaps = 36/285 (12%)
Query: 745 ALQKLDEGCGIEEAKAICEPEIIRQLFTWQKQLKIYLAPFLHGRRYTSFGRHFTKIDKLK 804
AL L+ G IE+ K++C P + QL W+ +L IYLAPFLHG RYTS+GRHFTK+DKL+
Sbjct: 136 ALHMLENGASIEDVKSVCAPSDLFQLARWKNKLNIYLAPFLHGMRYTSYGRHFTKLDKLE 195
Query: 805 EIVDRLHWYVQSGDTVLDFCCGANDFSCLMKSKLEQTGKLCSFKNYDLFQAKNDFNFEKR 864
+IVDRL WY++SGDTV+DFCCG+NDFS L+K KLE + K C +KN+DL Q KNDFNFE+R
Sbjct: 196 QIVDRLQWYIESGDTVVDFCCGSNDFSLLLKEKLEASEKSCFYKNFDLIQPKNDFNFERR 255
Query: 865 DWMSVQAEELPHGSNLIIGLNPPFGVRGSLANKFIDKALTFKPKLLILIVPKVTRSQSTV 924
DWM+VQ +ELP G LI+GLNPPFG + SLAN+FI+KALTFKPKL+ILIVPK T
Sbjct: 256 DWMTVQPDELPTGCRLIMGLNPPFGFKASLANQFINKALTFKPKLIILIVPKETE----- 310
Query: 925 EKQAGIYISICLSASNINQENTNVSTNFYLARLDRKKAGYNLIWADEEICSGKSFYLPGS 984
RLDRK Y LIW D +GKSFYLPGS
Sbjct: 311 -------------------------------RLDRKYPPYELIWEDSHQLAGKSFYLPGS 339
Query: 985 VDTRDKQLEDWNLKPPPLYLWSRPDWTAWHMQIARMHRHIKYDAY 1029
+D +K +E WN+ PPPL LWSR DW H +IA+ HI + +
Sbjct: 340 LDADNKIMEQWNMSPPPLSLWSRSDWARKHKEIAKTMGHISKNVW 384
>ref|XP_481741.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|38637135|dbj|BAD03389.1| unknown protein [Oryza sativa
(japonica cultivar-group)] gi|38424061|dbj|BAD01751.1|
unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 653
Score = 315 bits (807), Expect = 5e-84
Identities = 153/278 (55%), Positives = 188/278 (67%), Gaps = 42/278 (15%)
Query: 774 QKQLKIYLAPFLHGRRYTSFGRHFTKIDKLKEIVDRLHWYVQSGDTVLDFCCGANDFSCL 833
Q +L++YLAPF+HG RYTSFGRHFTK +KL EI ++LHWYVQ GD ++DF CG NDFS
Sbjct: 9 QNKLRVYLAPFIHGMRYTSFGRHFTKKEKLIEIAEKLHWYVQPGDMIVDFSCGTNDFSQF 68
Query: 834 MKSKLEQTGKLCSFKNYDLFQAKNDFNFEKRDWMSVQAEELPHGSNLIIGLNPPFGVRGS 893
MK KL++ GK C+FKNYD+ Q KN F+FEKRDWM+V+ +ELPHGS LI+GLNPPFG +
Sbjct: 69 MKEKLDKVGKRCNFKNYDVIQPKNSFSFEKRDWMTVRQKELPHGSKLIMGLNPPFGPKAM 128
Query: 894 LANKFIDKALTFKPKLLILIVPKVTRSQSTVEKQAGIYISICLSASNINQENTNVSTNFY 953
LANKFIDKALTFKPKL+ILIVPK
Sbjct: 129 LANKFIDKALTFKPKLIILIVPKEAE---------------------------------- 154
Query: 954 LARLDRKKAGYNLIWADEEICSGKSFYLPGSVDTRDKQLEDWNLKPPPLYLWSRPDWTAW 1013
RLDRK+ Y+L+W D++ SGKSFYLPGS+D DKQ++ WN PPPLYLWSRPDWT
Sbjct: 155 --RLDRKQQPYDLVWEDDQRLSGKSFYLPGSLDVSDKQIDQWNKSPPPLYLWSRPDWTQK 212
Query: 1014 HMQIARMHRHIKYDAYNNNKGKKVTNYLME----ENHD 1047
H +IA H H K + +++N+ V YL E +NHD
Sbjct: 213 HKRIAEQHGHTKANVFSHNEEDLV--YLFEDRATQNHD 248
>gb|AAU44579.1| hypothetical protein AT5G48090 [Arabidopsis thaliana]
gi|60547933|gb|AAX23930.1| hypothetical protein At5g48090
[Arabidopsis thaliana]
Length = 364
Score = 283 bits (725), Expect = 2e-74
Identities = 145/280 (51%), Positives = 179/280 (63%), Gaps = 40/280 (14%)
Query: 745 ALQKLDEGCGIEEAKAICEPEIIRQLFTWQKQLKIYLAPFLHGRRYTSFGRHFTKIDKLK 804
AL+ +EG +A+A+ +P+ + QL +K+L+I +PFLHG RYTSFGRHFT +KLK
Sbjct: 90 ALKMFEEGRD-RDARALFDPDSLLQLMKHKKKLEI--SPFLHGMRYTSFGRHFTNPEKLK 146
Query: 805 EIVDRLHWYVQSGDTVLDFCCGANDFSCLMKSKLEQTGKLCSFKNYDLFQAKNDFNFEKR 864
EIV+RLHWYV++GDTV+DFCCG+NDFSCLMK KL +TGK+C +KN DL KN+FNFE R
Sbjct: 147 EIVERLHWYVENGDTVVDFCCGSNDFSCLMKEKLMETGKICFYKNLDLIPPKNNFNFEMR 206
Query: 865 DWMSVQAEELPHGSNLIIGLNPPFGVRGSLANKFIDKALTFKPKLLILIVPKVTRSQSTV 924
DW+SV+ EELP GS LI+GLNPPFG + SLAN FI KAL FKPK+LILIVP T+ +
Sbjct: 207 DWLSVKEEELPDGSQLIMGLNPPFGYKASLANTFIKKALEFKPKILILIVPSETKRVDAI 266
Query: 925 EKQAGIYISICLSASNINQENTNVSTNFYLARLDRKKAGYNLIWADEEICSGKSFYLPGS 984
+ Y LIW D + +G SFYLPGS
Sbjct: 267 D-------------------------------------DYELIWEDRNLLAGMSFYLPGS 289
Query: 985 VDTRDKQLEDWNLKPPPLYLWSRPDWTAWHMQIARMHRHI 1024
VD DK +E WN PPLYLWSR D + H A HI
Sbjct: 290 VDVNDKTIEQWNNITPPLYLWSRRDLSRSHKTTAVQQGHI 329
>dbj|BAB11073.1| unnamed protein product [Arabidopsis thaliana]
gi|15238893|ref|NP_199620.1| expressed protein
[Arabidopsis thaliana]
Length = 636
Score = 283 bits (725), Expect = 2e-74
Identities = 145/280 (51%), Positives = 179/280 (63%), Gaps = 40/280 (14%)
Query: 745 ALQKLDEGCGIEEAKAICEPEIIRQLFTWQKQLKIYLAPFLHGRRYTSFGRHFTKIDKLK 804
AL+ +EG +A+A+ +P+ + QL +K+L+I +PFLHG RYTSFGRHFT +KLK
Sbjct: 362 ALKMFEEGRD-RDARALFDPDSLLQLMKHKKKLEI--SPFLHGMRYTSFGRHFTNPEKLK 418
Query: 805 EIVDRLHWYVQSGDTVLDFCCGANDFSCLMKSKLEQTGKLCSFKNYDLFQAKNDFNFEKR 864
EIV+RLHWYV++GDTV+DFCCG+NDFSCLMK KL +TGK+C +KN DL KN+FNFE R
Sbjct: 419 EIVERLHWYVENGDTVVDFCCGSNDFSCLMKEKLMETGKICFYKNLDLIPPKNNFNFEMR 478
Query: 865 DWMSVQAEELPHGSNLIIGLNPPFGVRGSLANKFIDKALTFKPKLLILIVPKVTRSQSTV 924
DW+SV+ EELP GS LI+GLNPPFG + SLAN FI KAL FKPK+LILIVP T+ +
Sbjct: 479 DWLSVKEEELPDGSQLIMGLNPPFGYKASLANTFIKKALEFKPKILILIVPSETKRVDAI 538
Query: 925 EKQAGIYISICLSASNINQENTNVSTNFYLARLDRKKAGYNLIWADEEICSGKSFYLPGS 984
+ Y LIW D + +G SFYLPGS
Sbjct: 539 D-------------------------------------DYELIWEDRNLLAGMSFYLPGS 561
Query: 985 VDTRDKQLEDWNLKPPPLYLWSRPDWTAWHMQIARMHRHI 1024
VD DK +E WN PPLYLWSR D + H A HI
Sbjct: 562 VDVNDKTIEQWNNITPPLYLWSRRDLSRSHKTTAVQQGHI 601
Score = 191 bits (486), Expect = 9e-47
Identities = 112/292 (38%), Positives = 160/292 (54%), Gaps = 39/292 (13%)
Query: 258 EENDMVESAEEDPEEENDKSDGEGVLNLDEEQDIGYDTVCAICDNGGEILPCEGRCLRSF 317
E++ +VE+ + EEN S + D E ++ +D VC+ICDNGG +L CEG CLRSF
Sbjct: 2 EQDSIVENMTD---EENSSSSDD-----DSEANLQFDPVCSICDNGGYVLCCEGSCLRSF 53
Query: 318 HATLEDGRDSLCASLGYT-RTEVNAFPNFYCENCKHKKHQCFACGKLGSSDESSNPEVFP 376
H T+ DG ++ C SLG+T +T++ A + C NC +K+HQC+ACG+LGSSDE+ + +VFP
Sbjct: 54 HPTIADGIETECESLGFTDKTQIQALGTYLCNNCLYKQHQCYACGELGSSDENFSQQVFP 113
Query: 377 CVTANCGHYYHPECVARLLYPGIDIGQEEMRKRIIIEKTFVCPLHICSLCRKGENRNVHD 436
C +NCGH+YHPECVARLL EE++ +I F CPLH C LC E++N
Sbjct: 114 CSASNCGHFYHPECVARLLCADDQNKAEELQAKIAARDCFACPLHTCKLCNMSEDKN--- 170
Query: 437 LQFAMCRRCPKAYHRKCLPKEISFTYDYYTGI-------EMRAWDNL-----LDKRILMY 484
Q+A C Y + T + Y GI MR L L L
Sbjct: 171 -QYA----CILLYADVA---QQLITENVYLGILLLNSTLTMRHCKELGKVCFLTTAFLYI 222
Query: 485 CMNHKIVLE-------LGTPARDHLIFPNKEVKRKIISTESLHREKDAIPLK 529
HK++ + TPARDHL+FP+ +R+ S ++D + ++
Sbjct: 223 VSFHKLLNRAHEIDGLILTPARDHLVFPDVSGQRRTRSHGLAQFKEDVLGME 274
>gb|AAU44578.1| hypothetical protein AT5G48090 [Arabidopsis thaliana]
Length = 272
Score = 281 bits (720), Expect = 6e-74
Identities = 141/268 (52%), Positives = 173/268 (63%), Gaps = 39/268 (14%)
Query: 757 EAKAICEPEIIRQLFTWQKQLKIYLAPFLHGRRYTSFGRHFTKIDKLKEIVDRLHWYVQS 816
+A+A+ +P+ + QL +K+L+I +PFLHG RYTSFGRHFT +KLKEIV+RLHWYV++
Sbjct: 9 DARALFDPDSLLQLMKHKKKLEI--SPFLHGMRYTSFGRHFTNPEKLKEIVERLHWYVEN 66
Query: 817 GDTVLDFCCGANDFSCLMKSKLEQTGKLCSFKNYDLFQAKNDFNFEKRDWMSVQAEELPH 876
GDTV+DFCCG+NDFSCLMK KL +TGK+C +KN DL KN+FNFE RDW+SV+ EELP
Sbjct: 67 GDTVVDFCCGSNDFSCLMKEKLMETGKICFYKNLDLIPPKNNFNFEMRDWLSVKEEELPD 126
Query: 877 GSNLIIGLNPPFGVRGSLANKFIDKALTFKPKLLILIVPKVTRSQSTVEKQAGIYISICL 936
GS LI+GLNPPFG + SLAN FI KAL FKPK+LILIVP T+ ++
Sbjct: 127 GSQLIMGLNPPFGYKASLANTFIKKALEFKPKILILIVPSETKRVDAID----------- 175
Query: 937 SASNINQENTNVSTNFYLARLDRKKAGYNLIWADEEICSGKSFYLPGSVDTRDKQLEDWN 996
Y LIW D + +G SFYLPGSVD DK +E WN
Sbjct: 176 --------------------------DYELIWEDRNLLAGMSFYLPGSVDVNDKTIEQWN 209
Query: 997 LKPPPLYLWSRPDWTAWHMQIARMHRHI 1024
PPLYLWSR D + H A HI
Sbjct: 210 NITPPLYLWSRRDLSRSHKTTAVQQGHI 237
>gb|AAU44580.1| hypothetical protein AT5G48090 [Arabidopsis thaliana]
Length = 268
Score = 190 bits (483), Expect = 2e-46
Identities = 93/217 (42%), Positives = 135/217 (61%), Gaps = 17/217 (7%)
Query: 225 EEDLEKENDMV-------ESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKS 277
+EDL K ++ ++ + E D ++ A+ P E++ +VE+ + EEN S
Sbjct: 4 DEDLTKSKFLITFLGKTSQTTPIEVELPTDHLQDAQT-PMEQDSIVENMTD---EENSSS 59
Query: 278 DGEGVLNLDEEQDIGYDTVCAICDNGGEILPCEGRCLRSFHATLEDGRDSLCASLGYT-R 336
+ D E ++ +D VC+ICDNGG +L CEG CLRSFH T+ DG ++ C SLG+T +
Sbjct: 60 SDD-----DSEANLQFDPVCSICDNGGYVLCCEGSCLRSFHPTIADGIETECESLGFTDK 114
Query: 337 TEVNAFPNFYCENCKHKKHQCFACGKLGSSDESSNPEVFPCVTANCGHYYHPECVARLLY 396
T++ A + C NC +K+HQC+ACG+LGSSDE+ + +VFPC +NCGH+YHPECVARLL
Sbjct: 115 TQIQALGTYLCNNCLYKQHQCYACGELGSSDENFSQQVFPCSASNCGHFYHPECVARLLC 174
Query: 397 PGIDIGQEEMRKRIIIEKTFVCPLHICSLCRKGENRN 433
EE++ +I F CPLH C LC E++N
Sbjct: 175 ADDQNKAEELQAKIAARDCFACPLHTCKLCNMSEDKN 211
>ref|XP_481739.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|38637133|dbj|BAD03387.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|38424059|dbj|BAD01749.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 657
Score = 136 bits (342), Expect = 4e-30
Identities = 86/262 (32%), Positives = 136/262 (51%), Gaps = 9/262 (3%)
Query: 1 MASSDEEGEIVPDSVDGYWFENDKAEFVSLSSLTLLWSVNDVTCNSEAKVFLRGTTDNGL 60
M SSD++ E +V+ Y+F +D VS L + + + + V+LRG TD GL
Sbjct: 1 MMSSDDDLEPQLKAVENYYFVDDNDVPVSFDVLPFQFDAAEGVASFKKDVYLRGFTDGGL 60
Query: 61 QKIHKQIIGWRFELSFEQPEISVLLREKYWMTLLKPSKCFENTIRSVLVTVYWLHCVKWK 120
QK++KQ++ W+ L + PEI+VL E W+ LLKP +E TIRSVL+TV LH V+ +
Sbjct: 61 QKVYKQVVAWKLVLDGDSPEIAVLSTEGSWIALLKPRPSYEETIRSVLITVEMLHFVRRR 120
Query: 121 PEESRASILVKVLKEFSSFDITPSENDVLCHMALISEAVKRDTDLTKSKDVHTTKKLKVI 180
P +S + + F F + P E+D H LI +RD DL S+ + K K++
Sbjct: 121 PTDSEKDMWDHLYGVFERFVVRPLEDDFANHQNLIKLFAQRDPDLANSQVLQVFIKDKIM 180
Query: 181 VESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEE 240
++ E V S D + E DI + E D +QE E E++ +E + +
Sbjct: 181 EKTNE--------VGSNNLDNKREPDI-KQEPDIKQEPVAAGDEMEEIVEEGIPDAPSND 231
Query: 241 DPEEENDMVESAEEDPEEENDM 262
D ++E D+V E+ + +D+
Sbjct: 232 DDDDEEDLVYLFEDRATQNHDV 253
>dbj|BAB11072.1| unnamed protein product [Arabidopsis thaliana]
gi|15238892|ref|NP_199619.1| expressed protein
[Arabidopsis thaliana]
Length = 227
Score = 102 bits (254), Expect = 7e-20
Identities = 66/172 (38%), Positives = 89/172 (51%), Gaps = 19/172 (11%)
Query: 1 MASSDEEGEIVPDSVDGYWFENDKAEFVSLSSLTLLWSVNDVTCNSEA-KVFLRGTTDNG 59
M S+EEGEI+PD VD Y+F + E V L L W+ D +S VFLRG D+G
Sbjct: 1 MEWSEEEGEILPDFVDNYYFIDQSEELVKFCDLPLKWNSVDYDDDSSGLSVFLRGNIDSG 60
Query: 60 LQKIHKQIIGWRFELSF--EQPEISVLLREKYWMTLLKPSKCFENTIRSVLVTVYWLHCV 117
+ K + GW F LS E P+I VLL +W+TL P K +E IR+ LVT+ +LH V
Sbjct: 61 DESFSKLVKGWNFYLSSVDEHPKIEVLLHGLHWITLQSPRKSYETLIRTSLVTLQFLHFV 120
Query: 118 KWKPEESRASILVKVLKEFSSFDITPSENDVLCHMALISEAVKRDTDLTKSK 169
K P+ + + L++ +D +KRD DLTKSK
Sbjct: 121 KRDPDVASGDVW-SSLQKVHGYD---------------WNTMKRDEDLTKSK 156
>ref|NP_703324.1| glutamic acid-rich protein (garp) [Plasmodium falciparum 3D7]
gi|6594249|emb|CAB63561.1| glutamic acid-rich protein
(garp) [Plasmodium falciparum 3D7]
Length = 673
Score = 98.6 bits (244), Expect = 1e-18
Identities = 59/164 (35%), Positives = 93/164 (55%), Gaps = 4/164 (2%)
Query: 118 KWKPEESRASILVKVLKEFSSFDITPSENDVLCHMALISEA-VKRDTDLTKSKDVHTTKK 176
K K EE SI +++ ++ P ++ MA I EA +++ + K +D K
Sbjct: 497 KSKVEEKNLSIQEQLIGTIGRVNVVPRRDNHKKKMAKIEEAELQKQKHVDKEEDKKEESK 556
Query: 177 LKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVE 236
+V ES+E E E ++ E EEE+ EEE + E EE+ E+E D VE +E+D E++ D E
Sbjct: 557 -EVEEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEDEVEEDEDDAEEDEDDAE 615
Query: 237 SAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGE 280
E+D EE++D E ++D EE++D E +ED EEE D+ + E
Sbjct: 616 EDEDDAEEDDDDAEEDDDDAEEDDD--EDEDEDEEEEEDEEEEE 657
Score = 66.2 bits (160), Expect = 6e-09
Identities = 36/113 (31%), Positives = 67/113 (58%), Gaps = 3/113 (2%)
Query: 156 SEAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEE-ENDIVESEEDP 214
S+ V+ D + + + ++ + E EE E+E D VE +E+D EE E+D E E+D
Sbjct: 562 SKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEDEVEEDEDDAEEDEDDAEEDEDDA 621
Query: 215 EQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAE 267
E+++D E +++D E+++D E +ED EEE D E E + + + ++ ++A+
Sbjct: 622 EEDDDDAEEDDDDAEEDDD--EDEDEDEEEEEDEEEEEESEKKIKRNLRKNAK 672
Score = 53.5 bits (127), Expect = 4e-05
Identities = 33/120 (27%), Positives = 67/120 (55%), Gaps = 12/120 (10%)
Query: 171 VHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEK 230
+ T ++ V+ + + ++ I E+E + + V+ EED ++E+ VE E +++++
Sbjct: 512 IGTIGRVNVVPRRDNHKKKMAKIEEAELQKQKH----VDKEEDKKEESKEVEEESKEVQE 567
Query: 231 ENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
+ + VE EE+ EEE EE+ EEE + E E++ EE+ D ++ E + +E++D
Sbjct: 568 DEEEVEEDEEEEEEE-------EEEEEEEEEEEEEEEDEVEEDEDDAE-EDEDDAEEDED 619
Score = 45.4 bits (106), Expect = 0.010
Identities = 33/122 (27%), Positives = 61/122 (49%), Gaps = 7/122 (5%)
Query: 178 KVIVESEENPEQENDIVESEEEDPEEE--NDIVESEEDPEQEND---IVESEEEDLEKEN 232
K+I E E+ + VE + +E+ I P ++N + + EE +L+K+
Sbjct: 484 KLIDEPEQLTLMDKSKVEEKNLSIQEQLIGTIGRVNVVPRRDNHKKKMAKIEEAELQKQK 543
Query: 233 DMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDIG 292
+ EED +EE+ VE ++ +E+ + VE EE+ EEE ++ + E +EE ++
Sbjct: 544 HV--DKEEDKKEESKEVEEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEDEVE 601
Query: 293 YD 294
D
Sbjct: 602 ED 603
Score = 37.4 bits (85), Expect = 2.8
Identities = 25/126 (19%), Positives = 56/126 (43%), Gaps = 11/126 (8%)
Query: 181 VESEENPEQEND---IVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVES 237
V N ++ND I E ++ED ++E + ++ ++ + E+E +EK+ E
Sbjct: 228 VHETSNDTKDNDKENISEDKKEDHQQEEMLKTLDKKERKQKEKEMKEQEKIEKKKKKQEE 287
Query: 238 AEEDPEEENDMVESAEEDPEEENDMVE--------SAEEDPEEENDKSDGEGVLNLDEEQ 289
E+ +E+ + +E ++E +M + +E+ E++ K D E + +
Sbjct: 288 KEKKKQEKERKKQEKKERKQKEKEMKKQKKIEKERKKKEEKEKKKKKHDKENEETMQQPD 347
Query: 290 DIGYDT 295
+T
Sbjct: 348 QTSEET 353
Score = 36.2 bits (82), Expect = 6.1
Identities = 33/146 (22%), Positives = 57/146 (38%), Gaps = 27/146 (18%)
Query: 167 KSKDVHTTKKLKVIVESEENPEQENDIVESEEED-----PEEENDIVESEEDPEQENDIV 221
K K+ KK K E+EE +Q + E + P D+ EE E E+
Sbjct: 323 KKKEEKEKKKKKHDKENEETMQQPDQTSEETNNEIMVPLPSPLTDVTTPEEHKEGEH--- 379
Query: 222 ESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEE--------- 272
EEE E E+ E EE+ +EE E + + + ++D +
Sbjct: 380 -KEEEHKEGEHKEGEHKEEEHKEEEHKKEEHKSKEHKSKGKKDKGKKDKGKHKKAKKEKV 438
Query: 273 ---------ENDKSDGEGVLNLDEEQ 289
E++ DG ++NL++++
Sbjct: 439 KKHVVKNVIEDEDKDGVEIINLEDKE 464
Score = 35.8 bits (81), Expect = 8.0
Identities = 24/120 (20%), Positives = 56/120 (46%), Gaps = 7/120 (5%)
Query: 160 KRDTDLTKSKDVHTTKKLKVIVESE----ENPEQENDIVESEEEDPEEENDIVESEEDPE 215
K + K +D + LK + + E E +E + +E +++ EE+ + +E +
Sbjct: 240 KENISEDKKEDHQQEEMLKTLDKKERKQKEKEMKEQEKIEKKKKKQEEKEKKKQEKERKK 299
Query: 216 QENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEEND 275
QE + +E++++K+ + + ++ E+E + + D E E M + + E N+
Sbjct: 300 QEKKERKQKEKEMKKQKKIEKERKKKEEKEK---KKKKHDKENEETMQQPDQTSEETNNE 356
>gb|AAG14291.1| glutamic acid-rich protein [Plasmodium falciparum]
Length = 682
Score = 92.8 bits (229), Expect = 6e-17
Identities = 64/175 (36%), Positives = 92/175 (52%), Gaps = 17/175 (9%)
Query: 118 KWKPEESRASILVKVLKEFSSFDITPSENDVLCHMALISEAVKRDTDLTKSKDVHTTKKL 177
K K EE SI +++ ++ P ++ MA I EA +L K K H K+
Sbjct: 501 KSKVEEKNLSIQEQLIGTIGRVNVVPRRDNHKKKMAKIEEA-----ELQKQK--HVDKEE 553
Query: 178 KVIVESEENPEQENDIVESEEE--DPEEENDIVESEEDPEQENDIVESEEEDLEKENDMV 235
ES+E E+ ++ E EEE + EEE + E EE+ E+E D VE +E+D E++ D
Sbjct: 554 DKKEESKEVEEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEDEVEEDEDDAEEDEDDA 613
Query: 236 ESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
E E+D E+ D E E+D EE++D AEED +EE D D E DEE+D
Sbjct: 614 EEDEDDAGEDEDDAEEDEDDAEEDDD---DAEEDDDEEEDVEDEE-----DEEED 660
Score = 61.6 bits (148), Expect = 1e-07
Identities = 37/124 (29%), Positives = 68/124 (54%), Gaps = 16/124 (12%)
Query: 171 VHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEK 230
+ T ++ V+ + + ++ I E+E + + V+ EED ++E+ VE E +++++
Sbjct: 516 IGTIGRVNVVPRRDNHKKKMAKIEEAELQKQKH----VDKEEDKKEESKEVEEESKEVQE 571
Query: 231 ENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
+ + VE EE+ EEE + EE+ EEE D VE E+D EE+ D ++ ++E D
Sbjct: 572 DEEEVEEDEEEEEEEEE-----EEEEEEEEDEVEEDEDDAEEDEDDAE-------EDEDD 619
Query: 291 IGYD 294
G D
Sbjct: 620 AGED 623
Score = 40.0 bits (92), Expect = 0.42
Identities = 30/135 (22%), Positives = 62/135 (45%), Gaps = 7/135 (5%)
Query: 178 KVIVESEENPEQENDIVESEEEDPEEEN-------DIVESEEDPEQENDIVESEEEDLEK 230
K+I E E+ + VE + +E+ ++V ++ +++ +E E +K
Sbjct: 488 KLIDEPEQLTLMDKSKVEEKNLSIQEQLIGTIGRVNVVPRRDNHKKKMAKIEEAELQKQK 547
Query: 231 ENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
D E +E+ +E + + +ED EE + E EE+ EEE ++ + + V +++ +
Sbjct: 548 HVDKEEDKKEESKEVEEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEDEVEEDEDDAE 607
Query: 291 IGYDTVCAICDNGGE 305
D D+ GE
Sbjct: 608 EDEDDAEEDEDDAGE 622
Score = 37.7 bits (86), Expect = 2.1
Identities = 25/122 (20%), Positives = 57/122 (46%), Gaps = 12/122 (9%)
Query: 181 VESEENPEQEND---IVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVES 237
V N ++ND I E ++ED ++E + ++ ++ + E+E +EK+ E
Sbjct: 234 VHETSNDTKDNDKENISEDKKEDHQQEEMLKTLDKKERKQKEKEMKEQEKIEKKKKKQEE 293
Query: 238 AEEDPEEENDMVESAEEDPEEENDMV---------ESAEEDPEEENDKSDGEGVLNLDEE 288
E+ +E+ + +E ++E +M + EE ++++DK + E + D+
Sbjct: 294 KEKKKQEKERKKQEKKERKQKEKEMKKQKKIEKERKKKEEKEKKKHDKENEETMQQPDQT 353
Query: 289 QD 290
+
Sbjct: 354 SE 355
>gb|AAA29605.1| glutamic acid-rich protein [Plasmodium falciparum]
gi|120943|sp|P13816|GARP_PLAFF Glutamic acid-rich
protein precursor gi|627054|pir||A54514 glutamic
acid-rich protein precursor - malaria parasite
(Plasmodium falciparum)
Length = 678
Score = 91.7 bits (226), Expect = 1e-16
Identities = 59/182 (32%), Positives = 92/182 (50%), Gaps = 8/182 (4%)
Query: 118 KWKPEESRASILVKVLKEFSSFDITPSENDVLCHMALISEAV--------KRDTDLTKSK 169
K K EE SI +++ ++ P ++ MA I EA K + +SK
Sbjct: 497 KSKVEEKNLSIQEQLIGTIGRVNVVPRRDNHKKKMAKIEEAELQKQKHVDKEEDKKEESK 556
Query: 170 DVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLE 229
+V K E E ++E + E EEE+ EEE + E EE+ E+E D E +E+D E
Sbjct: 557 EVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAE 616
Query: 230 KENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQ 289
++ D E E+D EE++D + EED +E+ D E EE+ EEE ++S+ + NL +
Sbjct: 617 EDEDDAEEDEDDAEEDDDEEDDDEEDDDEDEDEDEEDEEEEEEEEEESEKKIKRNLRKNA 676
Query: 290 DI 291
I
Sbjct: 677 KI 678
Score = 51.2 bits (121), Expect = 2e-04
Identities = 27/84 (32%), Positives = 47/84 (55%)
Query: 211 EEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDP 270
EE Q+ V+ EE+ E+ ++ E ++E E+E ++ E EE+ EEE + E EE+
Sbjct: 535 EEAELQKQKHVDKEEDKKEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEE 594
Query: 271 EEENDKSDGEGVLNLDEEQDIGYD 294
EEE ++ + E + ++E D D
Sbjct: 595 EEEEEEEEEEEDEDEEDEDDAEED 618
Score = 38.1 bits (87), Expect = 1.6
Identities = 46/200 (23%), Positives = 87/200 (43%), Gaps = 33/200 (16%)
Query: 155 ISEAVKRDTDLTKSKDVHTTKKLKVIVESEEN---PEQENDIVESEEEDPE------EEN 205
I+ + +T+L K+KD ++ + + E +E P ND +++ + E N
Sbjct: 45 ITGRLLNETELEKNKDDNSKSETLLKEEKDEKDDVPTTSNDNLKNAHNNNEISSSTDPTN 104
Query: 206 DIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESA--------EEDPE 257
I +++D E D + ++E K++ + ++D +E+ D E ++ +
Sbjct: 105 IINVNDKDNENSVDKKKDKKEKKHKKDKKEKKEKKDKKEKKDKKEKKHKKEKKHKKDKKK 164
Query: 258 EENDMVESAEEDPEEENDKSDGEGVLNLDEEQDIGYDTVCAICDN--GGEILPC------ 309
+EN V S + + + + G NLDEE V I +N GG +L
Sbjct: 165 KENSEVMSLYKTGQHKPKNATEHGEENLDEEM------VSEINNNAQGGLLLSSPYQYRE 218
Query: 310 EGRC--LRSFHATLEDGRDS 327
+G C + S H T D +D+
Sbjct: 219 QGGCGIISSVHETSNDTKDN 238
Score = 37.4 bits (85), Expect = 2.8
Identities = 25/126 (19%), Positives = 56/126 (43%), Gaps = 11/126 (8%)
Query: 181 VESEENPEQEND---IVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVES 237
V N ++ND I E ++ED ++E + ++ ++ + E+E +EK+ E
Sbjct: 228 VHETSNDTKDNDKENISEDKKEDHQQEEMLKTLDKKERKQKEKEMKEQEKIEKKKKKQEE 287
Query: 238 AEEDPEEENDMVESAEEDPEEENDMVE--------SAEEDPEEENDKSDGEGVLNLDEEQ 289
E+ +E+ + +E ++E +M + +E+ E++ K D E + +
Sbjct: 288 KEKKKQEKERKKQEKKERKQKEKEMKKQKKIEKERKKKEEKEKKKKKHDKENEETMQQPD 347
Query: 290 DIGYDT 295
+T
Sbjct: 348 QTSEET 353
Score = 36.2 bits (82), Expect = 6.1
Identities = 33/146 (22%), Positives = 57/146 (38%), Gaps = 27/146 (18%)
Query: 167 KSKDVHTTKKLKVIVESEENPEQENDIVESEEED-----PEEENDIVESEEDPEQENDIV 221
K K+ KK K E+EE +Q + E + P D+ EE E E+
Sbjct: 323 KKKEEKEKKKKKHDKENEETMQQPDQTSEETNNEIMVPLPSPLTDVTTPEEHKEGEH--- 379
Query: 222 ESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEE--------- 272
EEE E E+ E EE+ +EE E + + + ++D +
Sbjct: 380 -KEEEHKEGEHKEGEHKEEEHKEEEHKKEEHKSKEHKSKGKKDKGKKDKGKHKKAKKEKV 438
Query: 273 ---------ENDKSDGEGVLNLDEEQ 289
E++ DG ++NL++++
Sbjct: 439 KKHVVKNVIEDEDKDGVEIINLEDKE 464
Score = 35.8 bits (81), Expect = 8.0
Identities = 24/120 (20%), Positives = 56/120 (46%), Gaps = 7/120 (5%)
Query: 160 KRDTDLTKSKDVHTTKKLKVIVESE----ENPEQENDIVESEEEDPEEENDIVESEEDPE 215
K + K +D + LK + + E E +E + +E +++ EE+ + +E +
Sbjct: 240 KENISEDKKEDHQQEEMLKTLDKKERKQKEKEMKEQEKIEKKKKKQEEKEKKKQEKERKK 299
Query: 216 QENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEEND 275
QE + +E++++K+ + + ++ E+E + + D E E M + + E N+
Sbjct: 300 QEKKERKQKEKEMKKQKKIEKERKKKEEKEK---KKKKHDKENEETMQQPDQTSEETNNE 356
>ref|XP_649018.1| hypothetical protein 354.t00010 [Entamoeba histolytica HM-1:IMSS]
gi|56465359|gb|EAL43632.1| hypothetical protein
354.t00010 [Entamoeba histolytica HM-1:IMSS]
Length = 277
Score = 90.9 bits (224), Expect = 2e-16
Identities = 51/143 (35%), Positives = 83/143 (57%), Gaps = 2/143 (1%)
Query: 148 VLCHMALISEAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDI 207
VL +ALI+ A + +L + D+ T+ V + EEN +++++ EEED EE+ +
Sbjct: 3 VLLLLALIAFAFAEENELDNTFDIDFTELDNVELFEEENNDEDDEFQLDEEEDDEEDEED 62
Query: 208 VESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAE 267
E EED E E D E +EED E E D + +E+ EE+ + E E++ +EE++ E E
Sbjct: 63 EEDEEDEEDEED--EEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDE 120
Query: 268 EDPEEENDKSDGEGVLNLDEEQD 290
ED E+E D+ D E + ++E+D
Sbjct: 121 EDEEDEEDEEDEEDEEDEEDEED 143
Score = 85.5 bits (210), Expect = 9e-15
Identities = 53/157 (33%), Positives = 87/157 (54%), Gaps = 8/157 (5%)
Query: 137 SSFDITPSENDVLCHMALISEAVKRDTD---LTKSKDVHTTKKLKVIVESEENPEQENDI 193
++FDI +E D ++ L E + D L + +D ++ + E EE+ E E D
Sbjct: 22 NTFDIDFTELD---NVELFEEENNDEDDEFQLDEEEDDEEDEEDEEDEEDEEDEEDEEDE 78
Query: 194 VESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAE 253
+ E+E+ EE+ + E EED E E D E +EED E E D + +E+ EE+ + E E
Sbjct: 79 EDEEDEEDEEDEEDEEDEEDEEDEED--EEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEE 136
Query: 254 EDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
++ +EE++ E EED E+E D+ D E +L+EE++
Sbjct: 137 DEEDEEDEEDEEDEEDEEDEEDEEDEEDEYDLEEEEE 173
Score = 79.3 bits (194), Expect = 6e-13
Identities = 40/110 (36%), Positives = 67/110 (60%), Gaps = 2/110 (1%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE+ E E D E +EED E+E D + E++ ++E++ E +EED E E D + +E+
Sbjct: 61 EDEEDEEDEED--EEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEE 118
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDI 291
EE+ + E E++ +EE++ E EED E+E D+ D E + ++E D+
Sbjct: 119 DEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEYDL 168
Score = 74.3 bits (181), Expect = 2e-11
Identities = 44/110 (40%), Positives = 63/110 (57%), Gaps = 6/110 (5%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE+ E E D + E+E+ EE+ + E EED E E D E +EED E E D E EED
Sbjct: 88 EDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEED--EEDEEDEEDEED--EEDEED 143
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDI 291
E+E D E EED E+E D + + + EEE+D + E + +++D+
Sbjct: 144 EEDEED--EEDEEDEEDEEDEEDEYDLEEEEEDDDAVAESLFAEFDDEDL 191
Score = 72.8 bits (177), Expect = 6e-11
Identities = 45/122 (36%), Positives = 67/122 (54%), Gaps = 12/122 (9%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE+ E E D + E+E+ EE+ + E EED E E D E EE++ ++E++ E EED
Sbjct: 91 EDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEED-EEDEEDEEDEEDEEDEEDEED 149
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPE---------EENDKSDGEGVLNLDEEQDIG 292
E+E D E EED E+E D+ E E+D ++ D D + L+ D+E D+
Sbjct: 150 EEDEED--EEDEEDEEDEYDLEEEEEDDDAVAESLFAEFDDEDLLDSDEGLDDDDELDLD 207
Query: 293 YD 294
D
Sbjct: 208 DD 209
Score = 63.5 bits (153), Expect = 4e-08
Identities = 40/118 (33%), Positives = 65/118 (54%), Gaps = 4/118 (3%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE+ E E D + E+E+ EE+ + E EED E E D + E+E+ E++ + EE+
Sbjct: 112 EDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEYDLEEE 171
Query: 242 PEEENDMVES--AEEDPEEENDMVESAEEDPE--EENDKSDGEGVLNLDEEQDIGYDT 295
E+++ + ES AE D E+ D E ++D E ++D D E ++E+D DT
Sbjct: 172 EEDDDAVAESLFAEFDDEDLLDSDEGLDDDDELDLDDDDEDLEEADFFEDEEDEEDDT 229
Score = 60.8 bits (146), Expect = 2e-07
Identities = 43/119 (36%), Positives = 66/119 (55%), Gaps = 14/119 (11%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKEND----MVES 237
E EE+ E E D + E+E+ EE+ + E EED E E D E +E DLE+E + + ES
Sbjct: 124 EDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEED--EEDEYDLEEEEEDDDAVAES 181
Query: 238 --AEEDPEEENDMVESAEEDPE----EENDMVESAE--EDPEEENDKSDGEGVLNLDEE 288
AE D E+ D E ++D E ++++ +E A+ ED E+E D ++ E L +E
Sbjct: 182 LFAEFDDEDLLDSDEGLDDDDELDLDDDDEDLEEADFFEDEEDEEDDTEEEEEYELTQE 240
Score = 54.7 bits (130), Expect = 2e-05
Identities = 36/125 (28%), Positives = 62/125 (48%), Gaps = 19/125 (15%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEED-------------- 227
E EE+ E E D + E+E+ EE+ + E EED E E D+ E EE+D
Sbjct: 130 EDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEYDLEEEEEDDDAVAESLFAEFDDE 189
Query: 228 --LEKENDMVESAEEDPEEENDMVESA---EEDPEEENDMVESAEEDPEEENDKSDGEGV 282
L+ + + + E D +++++ +E A E++ +EE+D E E + +E + V
Sbjct: 190 DLLDSDEGLDDDDELDLDDDDEDLEEADFFEDEEDEEDDTEEEEEYELTQEWSEQAKSEV 249
Query: 283 LNLDE 287
LD+
Sbjct: 250 HPLDK 254
Score = 45.4 bits (106), Expect = 0.010
Identities = 31/103 (30%), Positives = 55/103 (53%), Gaps = 17/103 (16%)
Query: 182 ESEENPEQENDIVESEEEDPE---------EENDIVESEEDPEQENDI-VESEEEDLEKE 231
E EE+ E E D+ E EE+D ++ D+++S+E + ++++ ++ ++EDLE+
Sbjct: 157 EDEEDEEDEYDLEEEEEDDDAVAESLFAEFDDEDLLDSDEGLDDDDELDLDDDDEDLEEA 216
Query: 232 NDMV-ESAEEDPEEENDMVESAEEDPEEEN------DMVESAE 267
+ E EED EE + E +E E+ D +ESAE
Sbjct: 217 DFFEDEEDEEDDTEEEEEYELTQEWSEQAKSEVHPLDKIESAE 259
Score = 37.7 bits (86), Expect = 2.1
Identities = 23/74 (31%), Positives = 42/74 (56%), Gaps = 5/74 (6%)
Query: 180 IVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAE 239
+++S+E + ++++ ++++ EE D E EED +E+D E EE +L +E +E
Sbjct: 191 LLDSDEGLDDDDELDLDDDDEDLEEADFFEDEED--EEDDTEEEEEYELTQEWSEQAKSE 248
Query: 240 EDPEEENDMVESAE 253
P D +ESAE
Sbjct: 249 VHP---LDKIESAE 259
>ref|XP_649011.1| hypothetical protein 354.t00008 [Entamoeba histolytica HM-1:IMSS]
gi|56465355|gb|EAL43629.1| hypothetical protein
354.t00008 [Entamoeba histolytica HM-1:IMSS]
Length = 274
Score = 90.9 bits (224), Expect = 2e-16
Identities = 51/143 (35%), Positives = 83/143 (57%), Gaps = 2/143 (1%)
Query: 148 VLCHMALISEAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDI 207
VL +ALI+ A + +L + D+ T+ V + EEN +++++ EEED EE+ +
Sbjct: 3 VLLLLALIAFAFAEENELDNTFDIDFTELDNVELFEEENNDEDDEFQLDEEEDDEEDEED 62
Query: 208 VESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAE 267
E EED E E D E +EED E E D + +E+ EE+ + E E++ +EE++ E E
Sbjct: 63 EEDEEDEEDEED--EEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDE 120
Query: 268 EDPEEENDKSDGEGVLNLDEEQD 290
ED E+E D+ D E + ++E+D
Sbjct: 121 EDEEDEEDEEDEEDEEDEEDEED 143
Score = 85.1 bits (209), Expect = 1e-14
Identities = 51/154 (33%), Positives = 85/154 (55%), Gaps = 5/154 (3%)
Query: 137 SSFDITPSENDVLCHMALISEAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVES 196
++FDI +E D ++ L E + D + + ++ + E EE+ E E D +
Sbjct: 22 NTFDIDFTELD---NVELFEEENNDEDDEFQLDEEEDDEEDEEDEEDEEDEEDEEDEEDE 78
Query: 197 EEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDP 256
E+E+ EE+ + E EED E E D E +EED E E D + +E+ EE+ + E E++
Sbjct: 79 EDEEDEEDEEDEEDEEDEEDEED--EEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEE 136
Query: 257 EEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
+EE++ E EED E+E D+ D E +L+EE++
Sbjct: 137 DEEDEEDEEDEEDEEDEEDEEDEEDEYDLEEEEE 170
Score = 74.3 bits (181), Expect = 2e-11
Identities = 44/110 (40%), Positives = 63/110 (57%), Gaps = 6/110 (5%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE+ E E D + E+E+ EE+ + E EED E E D E +EED E E D E EED
Sbjct: 85 EDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEED--EEDEEDEEDEED--EEDEED 140
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDI 291
E+E D E EED E+E D + + + EEE+D + E + +++D+
Sbjct: 141 EEDEED--EEDEEDEEDEEDEEDEYDLEEEEEDDDAVAESLFAEFDDEDL 188
Score = 72.8 bits (177), Expect = 6e-11
Identities = 45/122 (36%), Positives = 67/122 (54%), Gaps = 12/122 (9%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE+ E E D + E+E+ EE+ + E EED E E D E EE++ ++E++ E EED
Sbjct: 88 EDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEED-EEDEEDEEDEEDEEDEEDEED 146
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPE---------EENDKSDGEGVLNLDEEQDIG 292
E+E D E EED E+E D+ E E+D ++ D D + L+ D+E D+
Sbjct: 147 EEDEED--EEDEEDEEDEYDLEEEEEDDDAVAESLFAEFDDEDLLDSDEGLDDDDELDLD 204
Query: 293 YD 294
D
Sbjct: 205 DD 206
Score = 63.5 bits (153), Expect = 4e-08
Identities = 40/118 (33%), Positives = 65/118 (54%), Gaps = 4/118 (3%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE+ E E D + E+E+ EE+ + E EED E E D + E+E+ E++ + EE+
Sbjct: 109 EDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEYDLEEE 168
Query: 242 PEEENDMVES--AEEDPEEENDMVESAEEDPE--EENDKSDGEGVLNLDEEQDIGYDT 295
E+++ + ES AE D E+ D E ++D E ++D D E ++E+D DT
Sbjct: 169 EEDDDAVAESLFAEFDDEDLLDSDEGLDDDDELDLDDDDEDLEEADFFEDEEDEEDDT 226
Score = 60.8 bits (146), Expect = 2e-07
Identities = 43/119 (36%), Positives = 66/119 (55%), Gaps = 14/119 (11%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKEND----MVES 237
E EE+ E E D + E+E+ EE+ + E EED E E D E +E DLE+E + + ES
Sbjct: 121 EDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEED--EEDEYDLEEEEEDDDAVAES 178
Query: 238 --AEEDPEEENDMVESAEEDPE----EENDMVESAE--EDPEEENDKSDGEGVLNLDEE 288
AE D E+ D E ++D E ++++ +E A+ ED E+E D ++ E L +E
Sbjct: 179 LFAEFDDEDLLDSDEGLDDDDELDLDDDDEDLEEADFFEDEEDEEDDTEEEEEYELTQE 237
Score = 54.7 bits (130), Expect = 2e-05
Identities = 36/125 (28%), Positives = 62/125 (48%), Gaps = 19/125 (15%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEED-------------- 227
E EE+ E E D + E+E+ EE+ + E EED E E D+ E EE+D
Sbjct: 127 EDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEYDLEEEEEDDDAVAESLFAEFDDE 186
Query: 228 --LEKENDMVESAEEDPEEENDMVESA---EEDPEEENDMVESAEEDPEEENDKSDGEGV 282
L+ + + + E D +++++ +E A E++ +EE+D E E + +E + V
Sbjct: 187 DLLDSDEGLDDDDELDLDDDDEDLEEADFFEDEEDEEDDTEEEEEYELTQEWSEQAKSEV 246
Query: 283 LNLDE 287
LD+
Sbjct: 247 HPLDK 251
Score = 45.4 bits (106), Expect = 0.010
Identities = 31/103 (30%), Positives = 55/103 (53%), Gaps = 17/103 (16%)
Query: 182 ESEENPEQENDIVESEEEDPE---------EENDIVESEEDPEQENDI-VESEEEDLEKE 231
E EE+ E E D+ E EE+D ++ D+++S+E + ++++ ++ ++EDLE+
Sbjct: 154 EDEEDEEDEYDLEEEEEDDDAVAESLFAEFDDEDLLDSDEGLDDDDELDLDDDDEDLEEA 213
Query: 232 NDMV-ESAEEDPEEENDMVESAEEDPEEEN------DMVESAE 267
+ E EED EE + E +E E+ D +ESAE
Sbjct: 214 DFFEDEEDEEDDTEEEEEYELTQEWSEQAKSEVHPLDKIESAE 256
Score = 37.7 bits (86), Expect = 2.1
Identities = 23/74 (31%), Positives = 42/74 (56%), Gaps = 5/74 (6%)
Query: 180 IVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAE 239
+++S+E + ++++ ++++ EE D E EED +E+D E EE +L +E +E
Sbjct: 188 LLDSDEGLDDDDELDLDDDDEDLEEADFFEDEED--EEDDTEEEEEYELTQEWSEQAKSE 245
Query: 240 EDPEEENDMVESAE 253
P D +ESAE
Sbjct: 246 VHP---LDKIESAE 256
>ref|XP_647597.1| hypothetical protein DDB0189820 [Dictyostelium discoideum]
gi|60475597|gb|EAL73532.1| hypothetical protein
DDB0189820 [Dictyostelium discoideum]
Length = 688
Score = 89.4 bits (220), Expect = 6e-16
Identities = 71/292 (24%), Positives = 123/292 (41%), Gaps = 42/292 (14%)
Query: 180 IVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQEN------DIVESEEEDLEKEND 233
I + N N+ S+ +D+ S D + E+ + + E++ +K +
Sbjct: 15 INNNNNNNNNNNNNNNSKNSKNFNTSDLYNSVSDYKVESITEFQINQINMEQQSTKKRSA 74
Query: 234 MVESAEEDPE---EENDMVESAEEDPEEENDMVE---SAEEDPEEENDKSDGEGVLNLDE 287
SA EE++++ ++ N+ S+++ N+ +D +
Sbjct: 75 STLSAPPSTSSSGEEDNIINKNNKNNNNNNNSTNNNNSSKKKKRISNETNDDD-----KP 129
Query: 288 EQDIGYDTVCAICDNGGEILPCEGRCLRSFHATLEDGRDSLCASLGYTR--TEVNAFPNF 345
++ + VC C+ GE+L C+G CLRSFH + R+ S T ++ +
Sbjct: 130 KRPRKNEAVCTFCEKPGELLMCDGLCLRSFHISCLKARNLYNPSSSSISPVTTIDGTVRW 189
Query: 346 YCENCKHKKHQCFACGKLGSSDESSNPEVFPCVTANCGHYYHPECVARLLYPGIDIGQEE 405
C +C ++ CF+C K G ++ C CG +YH +CVA +
Sbjct: 190 ECNDCVSSQNSCFSCKKRG----IIGIDLMKCKVHQCGKFYHYKCVA-----------DY 234
Query: 406 MRKRIIIEKT--FVCPLHICSLCR-KGENRNVHDLQFAMCRRCPKAYHRKCL 454
++I KT F CPLH CS+C G+ + Q C RCP AYH C+
Sbjct: 235 KLAKLINTKTPRFNCPLHYCSVCEVSGDGK-----QSVHCFRCPTAYHVICM 281
>pir||E86336 hypothetical protein F14O10.11 - Arabidopsis thaliana
gi|9558597|gb|AAF88160.1| Contains weak similarity to an
unknown protein T23E18.5 gi|6573728 from Arabidopsis
thaliana BAC T23E18 gb|AC009978. EST gb|W43440 comes
from this gene
Length = 409
Score = 89.0 bits (219), Expect = 8e-16
Identities = 54/154 (35%), Positives = 83/154 (53%), Gaps = 5/154 (3%)
Query: 139 FDITPSENDVLCHMALISEAVKRDTDLTKSKD--VHTTKKLKVIVESEENPEQENDIVES 196
+D+ + VLC ++ + D+D K VH + V E EE E+E + E
Sbjct: 234 YDMEANSWRVLCSTKWVNPS---DSDFYKFTPSMVHVGEGTIVQAEEEEEEEEEEEEEEE 290
Query: 197 EEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDP 256
EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+ EEE + E EE+
Sbjct: 291 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 350
Query: 257 EEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ EEE ++ + E +EE++
Sbjct: 351 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 384
Score = 85.9 bits (211), Expect = 7e-15
Identities = 45/112 (40%), Positives = 66/112 (58%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 285 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 344
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDIGY 293
EEE + E EE+ EEE + E EE+ EEE ++ + E +EE++ Y
Sbjct: 345 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKKY 396
Score = 85.1 bits (209), Expect = 1e-14
Identities = 44/109 (40%), Positives = 65/109 (59%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 279 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 338
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ EEE + E EE+ EEE ++ + E +EE++
Sbjct: 339 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 387
Score = 85.1 bits (209), Expect = 1e-14
Identities = 44/109 (40%), Positives = 65/109 (59%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 282 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 341
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ EEE + E EE+ EEE ++ + E +EE++
Sbjct: 342 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 390
Score = 85.1 bits (209), Expect = 1e-14
Identities = 44/109 (40%), Positives = 65/109 (59%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 278 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 337
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ EEE + E EE+ EEE ++ + E +EE++
Sbjct: 338 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 386
Score = 85.1 bits (209), Expect = 1e-14
Identities = 44/109 (40%), Positives = 65/109 (59%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 281 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 340
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ EEE + E EE+ EEE ++ + E +EE++
Sbjct: 341 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 389
Score = 85.1 bits (209), Expect = 1e-14
Identities = 44/109 (40%), Positives = 65/109 (59%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 284 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 343
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ EEE + E EE+ EEE ++ + E +EE++
Sbjct: 344 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 392
Score = 85.1 bits (209), Expect = 1e-14
Identities = 44/109 (40%), Positives = 65/109 (59%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 280 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 339
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ EEE + E EE+ EEE ++ + E +EE++
Sbjct: 340 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 388
Score = 85.1 bits (209), Expect = 1e-14
Identities = 44/109 (40%), Positives = 65/109 (59%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 283 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 342
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ EEE + E EE+ EEE ++ + E +EE++
Sbjct: 343 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 391
Score = 85.1 bits (209), Expect = 1e-14
Identities = 44/109 (40%), Positives = 65/109 (59%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 277 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 336
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ EEE + E EE+ EEE ++ + E +EE++
Sbjct: 337 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 385
Score = 69.3 bits (168), Expect = 7e-10
Identities = 35/82 (42%), Positives = 50/82 (60%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 317 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 376
Query: 242 PEEENDMVESAEEDPEEENDMV 263
EEE + E EE+ EE+ M+
Sbjct: 377 EEEEEEEEEEEEEEEEEKKYMI 398
Score = 66.6 bits (161), Expect = 4e-09
Identities = 34/87 (39%), Positives = 53/87 (60%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EE+ E+E + E EEE+ E+E + E EE+
Sbjct: 318 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 377
Query: 242 PEEENDMVESAEEDPEEENDMVESAEE 268
EEE + E EE+ E++ +V+ ++
Sbjct: 378 EEEEEEEEEEEEEEEEKKYMIVKKKKK 404
>gb|AAC95573.1| orf 48 [Ateline herpesvirus 3] gi|11361081|pir||T42963 hypothetical
protein 48 - ateline herpesvirus 3 (strain 73)
gi|9631239|ref|NP_048021.1| orf 48 [Ateline herpesvirus
3]
Length = 792
Score = 87.8 bits (216), Expect = 2e-15
Identities = 48/109 (44%), Positives = 64/109 (58%), Gaps = 2/109 (1%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E D E EEE+ EE+ + E EED E E D E +EED E E D + EE+
Sbjct: 479 EEEEEEEEEEDEEEEEEEEDEEDEEEEEDEEDEEDEED--EEDEEDEEDEEDEEDEEEEE 536
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ EEE + E EED E+E +K D E + +E++D
Sbjct: 537 DEEEEEDEEDEEEEEEEEEEEEEEDEEDEEDEEEKEDEEEKEDEEEKED 585
Score = 84.3 bits (207), Expect = 2e-14
Identities = 44/131 (33%), Positives = 74/131 (55%)
Query: 160 KRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQEND 219
K + + KD KK E EE+ E+E++ E EEE+ E+E + E E++ ++E +
Sbjct: 446 KDSEEEAEDKDEEENKKKGDGEEDEEDEEEEDEEEEEEEEEEEDEEEEEEEEDEEDEEEE 505
Query: 220 IVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDG 279
E +EED E E D + +E+ EE+ + E EE+ +EE++ E EE+ EEE D+ D
Sbjct: 506 EDEEDEEDEEDEEDEEDEEDEEDEEDEEEEEDEEEEEDEEDEEEEEEEEEEEEEEDEEDE 565
Query: 280 EGVLNLDEEQD 290
E ++E++
Sbjct: 566 EDEEEKEDEEE 576
Score = 84.0 bits (206), Expect = 3e-14
Identities = 44/109 (40%), Positives = 66/109 (60%), Gaps = 2/109 (1%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E D + EEE+ EE+ + E EED E E D E +EED E+E D E +E+
Sbjct: 488 EDEEEEEEEEDEEDEEEEEDEEDEEDEEDEEDEEDEED--EEDEEDEEEEEDEEEEEDEE 545
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EE+ +EE++ E +ED EE+ D+ + E + +E++D
Sbjct: 546 DEEEEEEEEEEEEEEDEEDEEDEEEKEDEEEKEDEEEKEDEEDEEEKED 594
Score = 81.6 bits (200), Expect = 1e-13
Identities = 42/109 (38%), Positives = 62/109 (56%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E E D E E+E+ EE+ + E EED E E D + EEE+ E+E + E EE+
Sbjct: 491 EEEEEEEDEEDEEEEEDEEDEEDEEDEEDEEDEEDEEDEEDEEEEEDEEEEEDEEDEEEE 550
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E EED E+E + + E++ EEE + + E DE+++
Sbjct: 551 EEEEEEEEEEDEEDEEDEEEKEDEEEKEDEEEKEDEEDEEEKEDDEDEE 599
Score = 80.9 bits (198), Expect = 2e-13
Identities = 42/109 (38%), Positives = 65/109 (59%), Gaps = 1/109 (0%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE+ E E D + E+E+ EE+ + E EED E+E D E EEE+ E+E + E EED
Sbjct: 506 EDEEDEEDEEDEEDEEDEEDEEDEEDEEEEEDEEEEED-EEDEEEEEEEEEEEEEEDEED 564
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
E+E + + E++ EEE + E EE ++E+++ + EG D+E D
Sbjct: 565 EEDEEEKEDEEEKEDEEEKEDEEDEEEKEDDEDEEEEEEGEEKEDDEDD 613
Score = 80.9 bits (198), Expect = 2e-13
Identities = 44/109 (40%), Positives = 65/109 (59%), Gaps = 7/109 (6%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E E D + E+E+ EE+ + E EED E+E D E E+E+ E+E + E EE+
Sbjct: 500 EDEEEEEDEEDEEDEEDEEDEEDEEDEEDEEDEEEEEDEEEEEDEEDEEEEEEEEEEEEE 559
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
+EE++ E +ED EE+ D E +ED E+E +K D E DEE++
Sbjct: 560 EDEEDEEDEEEKEDEEEKED--EEEKEDEEDEEEKEDDE-----DEEEE 601
Score = 80.1 bits (196), Expect = 4e-13
Identities = 39/109 (35%), Positives = 67/109 (60%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE ++E++ E +EED E+E D + E++ ++E++ E EEED E+E D + EE+
Sbjct: 492 EEEEEEDEEDEEEEEDEEDEEDEEDEEDEEDEEDEEDEEDEEEEEDEEEEEDEEDEEEEE 551
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE + E E++ +EE E +ED EE+ D+ D E + ++E++
Sbjct: 552 EEEEEEEEEDEEDEEDEEEKEDEEEKEDEEEKEDEEDEEEKEDDEDEEE 600
Score = 79.3 bits (194), Expect = 6e-13
Identities = 41/111 (36%), Positives = 69/111 (61%), Gaps = 2/111 (1%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEED--PEQENDIVESEEEDLEKENDMVESAE 239
E EE+ E+E D + EEE+ EEE + E EED E+E + E +E++ EKE++ E +
Sbjct: 533 EEEEDEEEEEDEEDEEEEEEEEEEEEEEDEEDEEDEEEKEDEEEKEDEEEKEDEEDEEEK 592
Query: 240 EDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
ED E+E + E E++ +E+++ E E+D EEE+++ + E + DEE +
Sbjct: 593 EDDEDEEEEEEGEEKEDDEDDEEEEDEEDDEEEEDEEEEDEDEEDEDEEDE 643
Score = 79.3 bits (194), Expect = 6e-13
Identities = 45/109 (41%), Positives = 63/109 (57%), Gaps = 4/109 (3%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE+ E E D + EEE+ EEE + E EE+ E+E + E EEED E E D E +E+
Sbjct: 518 EDEEDEEDEEDEEDEEEEEDEEEEEDEEDEEEEEEEEE--EEEEEDEEDEEDEEEKEDEE 575
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
+E+ + E E++ E+E+D E EE+ EE+ D D E DEE D
Sbjct: 576 EKEDEEEKEDEEDEEEKEDDEDEEEEEEGEEKEDDEDDEE--EEDEEDD 622
Score = 79.3 bits (194), Expect = 6e-13
Identities = 45/109 (41%), Positives = 65/109 (59%), Gaps = 2/109 (1%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E E D E EEE+ EEE + E EED E++ D E E+E+ EKE++ E +ED
Sbjct: 536 EDEEEEEDEEDEEEEEEEEEEEEEEDEEDEEDEEEKEDEEEKEDEE-EKEDEEDEEEKED 594
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
E+E + E EE ++E+D E EED EEE D+ + + ++E+D
Sbjct: 595 DEDEEEE-EEGEEKEDDEDDEEEEDEEDDEEEEDEEEEDEDEEDEDEED 642
Score = 79.0 bits (193), Expect = 8e-13
Identities = 45/125 (36%), Positives = 69/125 (55%), Gaps = 12/125 (9%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E E D + EE++ EEE + E +ED E E + + E+E+ E+E + E E+D
Sbjct: 554 EEEEEEEDEEDEEDEEEKEDEEEKEDEEEKEDEEDEEEKEDDEDEEEEEEGEEKEDDEDD 613
Query: 242 PEEENDMVESAEEDPEE--ENDMVESAEEDPEEENDKSDGEGVLNLDEEQD--------- 290
EEE++ + EED EE E++ E E++ EE+ D+ D EG + D + D
Sbjct: 614 EEEEDEEDDEEEEDEEEEDEDEEDEDEEDEDEEDEDEEDEEGDRDRDADADRDGGDGDGD 673
Query: 291 -IGYD 294
+GYD
Sbjct: 674 GVGYD 678
Score = 78.2 bits (191), Expect = 1e-12
Identities = 43/134 (32%), Positives = 74/134 (55%)
Query: 157 EAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQ 216
++VK KD + K E+++ + E D + EEED EEE + E E++ E+
Sbjct: 434 KSVKSGEGKKSEKDSEEEAEDKDEEENKKKGDGEEDEEDEEEEDEEEEEEEEEEEDEEEE 493
Query: 217 ENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDK 276
E + E +EE+ E E D + +E+ EE+ + E E++ EEE++ E EED EEE ++
Sbjct: 494 EEEEDEEDEEEEEDEEDEEDEEDEEDEEDEEDEEDEEDEEEEEDEEEEEDEEDEEEEEEE 553
Query: 277 SDGEGVLNLDEEQD 290
+ E + ++E+D
Sbjct: 554 EEEEEEEDEEDEED 567
Score = 78.2 bits (191), Expect = 1e-12
Identities = 41/109 (37%), Positives = 66/109 (59%), Gaps = 2/109 (1%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E EEE+ EEE + E EED E E D E E+E+ +++ + E E++
Sbjct: 532 EEEEEDEEEEEDEEDEEEEEEEEEE--EEEEDEEDEEDEEEKEDEEEKEDEEEKEDEEDE 589
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
E+E+D E EE+ EE+ D + EE+ EE++++ + E + DEE +
Sbjct: 590 EEKEDDEDEEEEEEGEEKEDDEDDEEEEDEEDDEEEEDEEEEDEDEEDE 638
Score = 78.2 bits (191), Expect = 1e-12
Identities = 41/109 (37%), Positives = 66/109 (59%), Gaps = 1/109 (0%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE+ E+E D E E+E+ EEE + E EE+ E E D + EE++ E+E + E +ED
Sbjct: 527 EDEEDEEEEEDEEEEEDEEDEEEEEEEEEEEEEEDEEDEEDEEEKEDEEEKED-EEEKED 585
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
E+E + + +E+ EEE + E E+D EEE+++ D E +E++D
Sbjct: 586 EEDEEEKEDDEDEEEEEEGEEKEDDEDDEEEEDEEDDEEEEDEEEEDED 634
Score = 78.2 bits (191), Expect = 1e-12
Identities = 41/111 (36%), Positives = 67/111 (59%), Gaps = 2/111 (1%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E +E E++ + E EEE+ +EE++ E EE+ E+E + E EE++ EKE++ + EE+
Sbjct: 523 EEDEEDEEDEEEEEDEEEEEDEEDEEEEEEEEEEEEEEDEEDEEDEEEKEDEEEKEDEEE 582
Query: 242 PEEENDMVESAEEDPEEENDMVESAE--EDPEEENDKSDGEGVLNLDEEQD 290
E+E D E +++ EEE + E E ED EEE D+ D E + +EE +
Sbjct: 583 KEDEEDEEEKEDDEDEEEEEEGEEKEDDEDDEEEEDEEDDEEEEDEEEEDE 633
Score = 77.4 bits (189), Expect = 2e-12
Identities = 46/113 (40%), Positives = 64/113 (55%), Gaps = 5/113 (4%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E + E E+E+ EE+ + E EE+ E E + E EE++ EKE+D E EE+
Sbjct: 545 EDEEEEEEEEEEEEEEDEEDEEDEEEKEDEEEKEDEEE-KEDEEDEEEKEDDEDEEEEEE 603
Query: 242 PEE----ENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EE E+D E EED EEE D E E++ +E+ + D E DEE D
Sbjct: 604 GEEKEDDEDDEEEEDEEDDEEEEDEEEEDEDEEDEDEEDEDEEDEDEEDEEGD 656
Score = 75.1 bits (183), Expect = 1e-11
Identities = 44/129 (34%), Positives = 75/129 (58%), Gaps = 3/129 (2%)
Query: 162 DTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIV 221
D ++ + K V + + K +SEE E +++ ++ D EE+ + E E++ E+E +
Sbjct: 427 DKEVGEEKSVKSGEGKKSEKDSEEEAEDKDEEENKKKGDGEEDEEDEEEEDEEEEEEEEE 486
Query: 222 ESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEG 281
E +EE+ E+E D E EE+ +EE++ E EED E+E D E EED EEE D+ + E
Sbjct: 487 EEDEEEEEEEED-EEDEEEEEDEEDEEDEEDEEDEEDEED--EEDEEDEEEEEDEEEEED 543
Query: 282 VLNLDEEQD 290
+ +EE++
Sbjct: 544 EEDEEEEEE 552
Score = 70.9 bits (172), Expect = 2e-10
Identities = 38/114 (33%), Positives = 66/114 (57%), Gaps = 1/114 (0%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E E+ E+E++ E E+ED E+E + E EE + E+D E +EED E+E D E +ED
Sbjct: 576 EKEDEEEKEDEEDEEEKEDDEDEEEEEEGEEKEDDEDDEEEEDEEDDEEEEDE-EEEDED 634
Query: 242 PEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDIGYDT 295
E+E++ E E++ EE+ + + D + + DG+GV ++++ G D+
Sbjct: 635 EEDEDEEDEDEEDEDEEDEEGDRDRDADADRDGGDGDGDGVGYDYKDEEKGTDS 688
Score = 56.6 bits (135), Expect = 4e-06
Identities = 41/138 (29%), Positives = 69/138 (49%), Gaps = 7/138 (5%)
Query: 157 EAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEE-EDPEEENDIVESEEDPE 215
E K D + + ++ ++ + E +E+ E+E + E E+ ED EEE D E+D E
Sbjct: 568 EEEKEDEEEKEDEEEKEDEEDEEEKEDDEDEEEEEEGEEKEDDEDDEEEED---EEDDEE 624
Query: 216 QENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEED---PEEENDMVESAEEDPEE 272
+E++ E E+E+ E E D E E++ +EE D A+ D + + D V +D E+
Sbjct: 625 EEDEEEEDEDEEDEDEEDEDEEDEDEEDEEGDRDRDADADRDGGDGDGDGVGYDYKDEEK 684
Query: 273 ENDKSDGEGVLNLDEEQD 290
D E N E++D
Sbjct: 685 GTDSYKNETEYNNKEQKD 702
Score = 55.8 bits (133), Expect = 7e-06
Identities = 40/113 (35%), Positives = 57/113 (50%), Gaps = 6/113 (5%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E ++ N ++E EE++ V+S E + E D E E+ E+EN EED
Sbjct: 412 EKNRKRKKPNGFRVGDKEVGEEKS--VKSGEGKKSEKDSEEEAEDKDEEENKKKGDGEED 469
Query: 242 P--EEENDMVESAEEDPE--EENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EEE D E EE+ E EE + E EED EEE D+ D E + ++E+D
Sbjct: 470 EEDEEEEDEEEEEEEEEEEDEEEEEEEEDEEDEEEEEDEEDEEDEEDEEDEED 522
Score = 52.0 bits (123), Expect = 1e-04
Identities = 31/103 (30%), Positives = 51/103 (49%), Gaps = 2/103 (1%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDP--EQENDIVESEEEDLEKENDMVESAE 239
E EE + E+D E +EED EEE D E +ED E E D E +E++ ++E D A+
Sbjct: 603 EGEEKEDDEDDEEEEDEEDDEEEEDEEEEDEDEEDEDEEDEDEEDEDEEDEEGDRDRDAD 662
Query: 240 EDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGV 282
D + + + D ++E +S + + E N + E +
Sbjct: 663 ADRDGGDGDGDGVGYDYKDEEKGTDSYKNETEYNNKEQKDENL 705
Score = 51.2 bits (121), Expect = 2e-04
Identities = 33/102 (32%), Positives = 56/102 (54%), Gaps = 4/102 (3%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E +E+ E+E D + EEE+ EEE D E +ED E++ D + +EED E + D A+ D
Sbjct: 608 EDDEDDEEEEDEEDDEEEEDEEEEDEDEEDED-EEDEDEEDEDEEDEEGDRDRDADADRD 666
Query: 242 -PEEENDMVESAEEDPEEENDMVESAEE--DPEEENDKSDGE 280
+ + D V +D E+ D ++ E + E++++ GE
Sbjct: 667 GGDGDGDGVGYDYKDEEKGTDSYKNETEYNNKEQKDENLHGE 708
Score = 47.4 bits (111), Expect = 0.003
Identities = 34/126 (26%), Positives = 59/126 (45%), Gaps = 4/126 (3%)
Query: 167 KSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEE 226
K K K K I ++E ++I + ++ N +++ +E + E +
Sbjct: 383 KYKGDGANDKDKSIDKNESEGGDHSEINREKNRKRKKPNGFRVGDKEVGEEKSVKSGEGK 442
Query: 227 DLEKENDMVESAEEDPEEENDMVESAEEDP--EEENDMVESAEEDPEEENDKSDGEGVLN 284
EK+++ E AE+ EEEN EED EEE D E EE+ EE+ ++ + E
Sbjct: 443 KSEKDSE--EEAEDKDEEENKKKGDGEEDEEDEEEEDEEEEEEEEEEEDEEEEEEEEDEE 500
Query: 285 LDEEQD 290
+EE++
Sbjct: 501 DEEEEE 506
>emb|CAH81749.1| conserved hypothetical protein [Plasmodium chabaudi]
Length = 391
Score = 87.4 bits (215), Expect = 2e-15
Identities = 48/116 (41%), Positives = 69/116 (59%), Gaps = 4/116 (3%)
Query: 177 LKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVE 236
+ +I E EE+ E+E + + EEE+ EEE D E EE+ E E + E EEED E+E E
Sbjct: 1 MPLIEEGEEDDEEEEEEEDEEEEEEEEEEDEEEEEEEDEDEEEEEEEEEEDDEEE----E 56
Query: 237 SAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDIG 292
++D EEE + E +E+ EEE D E EE+ EEEN++ + E DEE++ G
Sbjct: 57 EEDDDEEEEEEEEEEDDEEEEEEEDDEEEEEEEEEEENEEKEEENEEEEDEEEEEG 112
Score = 84.0 bits (206), Expect = 3e-14
Identities = 55/139 (39%), Positives = 74/139 (52%), Gaps = 20/139 (14%)
Query: 152 MALISEAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQENDIVESEEEDPEEENDIVESE 211
M LI E + D + + +D E EE E+E D E EEED +EE + E E
Sbjct: 1 MPLIEEGEEDDEEEEEEED-----------EEEEEEEEEEDEEEEEEEDEDEEEEEEEEE 49
Query: 212 EDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPE 271
ED E+E E E++D E+E + E +E+ EEE D E EE+ EEEN E EE+ E
Sbjct: 50 EDDEEE----EEEDDDEEEEEEEEEEDDEEEEEEEDDEEEEEEEEEEEN---EEKEEENE 102
Query: 272 EENDKSDGEGVLNLDEEQD 290
EE D+ + EG DE +D
Sbjct: 103 EEEDEEEEEG--EKDETED 119
Score = 68.2 bits (165), Expect = 1e-09
Identities = 34/85 (40%), Positives = 53/85 (62%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E EE E+E+D E EE+D EEE + E E+D E+E + + EEE+ E+E + E EE+
Sbjct: 42 EEEEEEEEEDDEEEEEEDDDEEEEEEEEEEDDEEEEEEEDDEEEEEEEEEEENEEKEEEN 101
Query: 242 PEEENDMVESAEEDPEEENDMVESA 266
EEE++ E E+D E+ +++ A
Sbjct: 102 EEEEDEEEEEGEKDETEDPYLLKDA 126
>gb|EAA22695.1| hypothetical protein [Plasmodium yoelii yoelii]
Length = 2959
Score = 87.0 bits (214), Expect = 3e-15
Identities = 51/135 (37%), Positives = 81/135 (59%), Gaps = 4/135 (2%)
Query: 157 EAVKRDTDLTKSKDVHTTKKLKVIVESEENPEQEN-DIVESEEEDPEEENDIVESEEDPE 215
E+++ + +L + KD ++++ E+EE E+E D E EEE+ EEE +I E EE+ E
Sbjct: 1072 ESIRSEDELEEEKDAEEEQEVEAENETEEEEEEEEEDDEEIEEEEEEEEEEIEEEEEEKE 1131
Query: 216 QENDIVESEEEDLEKENDMVESAEE-DPEEENDMVESAEEDPEEENDMVESAEEDPEEEN 274
E + +E EEE++E+E + E EE + EEE + E EE+ +EE + E EE+ EEE
Sbjct: 1132 IEEEEIEEEEEEIEEEEEEEEGGEEKEGEEEEEGGEEEEEEEDEEGEEEEEGEEEEEEEG 1191
Query: 275 DKSDGEGVLNLDEEQ 289
+S E + DEE+
Sbjct: 1192 IESKQE--IEEDEEE 1204
Score = 72.0 bits (175), Expect = 1e-10
Identities = 53/178 (29%), Positives = 80/178 (44%), Gaps = 2/178 (1%)
Query: 120 KPEESRASILVKVLKEFSSFDITPSENDVLCHMALISEAVKRDTDLTKSKDVHTTKKLKV 179
K S I + L+ S D E D + +E + + + +D ++ +
Sbjct: 1058 KNGSSEQEIYEETLESIRSEDELEEEKDAEEEQEVEAENETEEEEEEEEEDDEEIEEEEE 1117
Query: 180 IVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAE 239
E EE E+E + E EEE+ EEE + +E EE+ E+ + E EEE+ E + E E
Sbjct: 1118 --EEEEEIEEEEEEKEIEEEEIEEEEEEIEEEEEEEEGGEEKEGEEEEEGGEEEEEEEDE 1175
Query: 240 EDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDIGYDTVC 297
E EEE E EE E + ++ E EE +EE+ D N+ EE I D C
Sbjct: 1176 EGEEEEEGEEEEEEEGIESKQEIEEDEEETEKEESIAEDSHMENNICEENSIEEDEEC 1233
Score = 70.9 bits (172), Expect = 2e-10
Identities = 44/118 (37%), Positives = 67/118 (56%), Gaps = 11/118 (9%)
Query: 181 VESEENPEQENDIVES------EEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDM 234
+E E+N E +I E E++ EEE D E E++ E EN+ E EEE E+E+D
Sbjct: 1054 IEQEKNGSSEQEIYEETLESIRSEDELEEEKD-AEEEQEVEAENETEEEEEE--EEEDD- 1109
Query: 235 VESAEEDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQDIG 292
E EE+ EEE + +E EE+ E E + +E EE+ EEE ++ +G +EE++ G
Sbjct: 1110 -EEIEEEEEEEEEEIEEEEEEKEIEEEEIEEEEEEIEEEEEEEEGGEEKEGEEEEEGG 1166
Score = 68.6 bits (166), Expect = 1e-09
Identities = 49/159 (30%), Positives = 86/159 (53%), Gaps = 13/159 (8%)
Query: 136 FSSFDITPSENDVLCHMALISEAVKRDTDLTKSKDVHTTKKLKV----IVESEENPEQEN 191
F S + S+N V+ + + +TD+ + K+ + +++ + SE+ E+E
Sbjct: 1025 FYSANTEGSDNFVVYKDIVETVPDNDETDIEQEKNGSSEQEIYEETLESIRSEDELEEEK 1084
Query: 192 DIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEEDPEEENDMVES 251
D EE++ E EN+ E EE+ E++++ +E EEE+ E+E +E EE+ E E + +E
Sbjct: 1085 DA--EEEQEVEAENETEEEEEEEEEDDEEIEEEEEEEEEE---IEEEEEEKEIEEEEIEE 1139
Query: 252 AEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
EE+ EEE + E EE E E ++ GE +EE+D
Sbjct: 1140 EEEEIEEEEEEEEGGEEK-EGEEEEEGGE---EEEEEED 1174
Score = 64.3 bits (155), Expect = 2e-08
Identities = 42/120 (35%), Positives = 63/120 (52%), Gaps = 12/120 (10%)
Query: 181 VESEENPEQENDIVESEEEDP--------EEENDIVESEEDPEQENDIVESEEEDLEKEN 232
+E EE E+E +I E EEE+ EEE E EE+ ++E + E EE + E+E
Sbjct: 1132 IEEEEIEEEEEEIEEEEEEEEGGEEKEGEEEEEGGEEEEEEEDEEGE--EEEEGEEEEEE 1189
Query: 233 DMVESAE--EDPEEENDMVESAEEDPEEENDMVESAEEDPEEENDKSDGEGVLNLDEEQD 290
+ +ES + E+ EEE + ES ED EN++ E + +EE + G G N +E D
Sbjct: 1190 EGIESKQEIEEDEEETEKEESIAEDSHMENNICEENSIEEDEECYNNSGSGDKNREESND 1249
Score = 62.8 bits (151), Expect = 6e-08
Identities = 42/131 (32%), Positives = 74/131 (56%), Gaps = 8/131 (6%)
Query: 177 LKVIVESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVE 236
++ + +++E ++ SE+E EE + + SE++ E+E D EE+++E EN+ E
Sbjct: 1043 VETVPDNDETDIEQEKNGSSEQEIYEETLESIRSEDELEEEKDA--EEEQEVEAENETEE 1100
Query: 237 SAEEDPEEENDMVESAEEDPEEENDMVESAEEDP--EEENDKSDGEGVLNLDEEQDIGYD 294
EE+ EEE+D E EE+ EEE + +E EE+ EEE + + E + +EE++ G +
Sbjct: 1101 --EEEEEEEDD--EEIEEEEEEEEEEIEEEEEEKEIEEEEIEEEEEEIEEEEEEEEGGEE 1156
Query: 295 TVCAICDNGGE 305
+ GGE
Sbjct: 1157 KEGEEEEEGGE 1167
Score = 56.6 bits (135), Expect = 4e-06
Identities = 37/99 (37%), Positives = 55/99 (55%), Gaps = 3/99 (3%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVESAEED 241
E +E E+E E EEE+ EE + E EE+ E+E + E E+ E+E + ES ED
Sbjct: 1155 EEKEGEEEEEGGEEEEEEEDEEGEEEEEGEEEEEEEGIESKQEIEEDEEETEKEESIAED 1214
Query: 242 PEEENDMVE--SAEEDPEEENDMVESAEEDPEEENDKSD 278
EN++ E S EED E N+ S +++ EE ND ++
Sbjct: 1215 SHMENNICEENSIEEDEECYNNS-GSGDKNREESNDTNE 1252
Score = 51.2 bits (121), Expect = 2e-04
Identities = 41/119 (34%), Positives = 60/119 (49%), Gaps = 11/119 (9%)
Query: 182 ESEENPEQENDIVESEEEDPEEENDIVESEEDPEQENDIVESEEEDLEKENDMVE--SAE 239
E EE E+ + E EEE+ EE + + E+ E+E + ES ED EN++ E S E
Sbjct: 1169 EEEEEDEEGEEEEEGEEEEEEEGIESKQEIEEDEEETEKEESIAEDSHMENNICEENSIE 1228
Query: 240 EDPEEENDMVESAEEDPEEENDMVE---SAEEDPEEENDKS-----DGEGVLNLDEEQD 290
ED E N+ S +++ EE ND E S +E N++S D +GV DE +
Sbjct: 1229 EDEECYNNS-GSGDKNREESNDTNEKNSSNSRTIDESNNRSSDKYGDDQGVKGDDERSE 1286
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,972,516,433
Number of Sequences: 2540612
Number of extensions: 90470439
Number of successful extensions: 1112480
Number of sequences better than 10.0: 13863
Number of HSP's better than 10.0 without gapping: 5604
Number of HSP's successfully gapped in prelim test: 9097
Number of HSP's that attempted gapping in prelim test: 596494
Number of HSP's gapped (non-prelim): 146708
length of query: 1084
length of database: 863,360,394
effective HSP length: 139
effective length of query: 945
effective length of database: 510,215,326
effective search space: 482153483070
effective search space used: 482153483070
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 81 (35.8 bits)
Medicago: description of AC124966.7


