

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124966.3 + phase: 0
(470 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
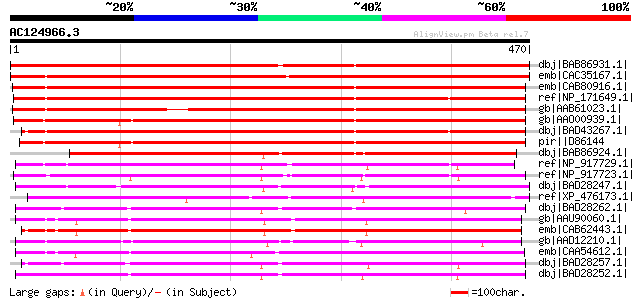
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB86931.1| glucosyltransferase-13 [Vigna angularis] 642 0.0
emb|CAC35167.1| arbutin synthase [Rauvolfia serpentina] gi|28380... 512 e-144
emb|CAB80916.1| putative flavonol glucosyltransferase [Arabidops... 484 e-135
ref|NP_171649.1| UDP-glucoronosyl/UDP-glucosyl transferase famil... 456 e-127
gb|AAB61023.1| Similar to UTP-Glucose Glucosyltransferase; coded... 451 e-125
gb|AAO00939.1| Putative UTP-glucose glucosyltransferase [Arabido... 448 e-124
dbj|BAD43267.1| putative flavonol 3-o-glucosyltransferase [Arabi... 446 e-124
pir||D86144 protein probable UTP-glucose glucosyltransferase [im... 439 e-122
dbj|BAB86924.1| glucosyltransferase-6 [Vigna angularis] 417 e-115
ref|NP_917729.1| putative UTP-glucose glucosyltransferase [Oryza... 327 5e-88
ref|NP_917723.1| putative UTP-glucose glucosyltransferase [Oryza... 322 2e-86
dbj|BAD28247.1| putative Hydroquinone glucosyltransferase [Oryza... 316 1e-84
ref|XP_476173.1| hypothetical protein [Oryza sativa (japonica cu... 311 4e-83
dbj|BAD28262.1| putative Hydroquinone glucosyltransferase [Oryza... 306 9e-82
gb|AAU90060.1| At3g50740 [Arabidopsis thaliana] gi|15146272|gb|A... 305 2e-81
emb|CAB62443.1| UTP-glucose glucosyltransferase-like protein [Ar... 301 3e-80
gb|AAD12210.1| putative flavonol 3-O-glucosyltransferase [Arabid... 301 3e-80
emb|CAA54612.1| UTP-glucose glucosyltransferase [Manihot esculen... 300 5e-80
dbj|BAD28257.1| putative Hydroquinone glucosyltransferase [Oryza... 291 3e-77
dbj|BAD28252.1| putative Hydroquinone glucosyltransferase [Oryza... 291 3e-77
>dbj|BAB86931.1| glucosyltransferase-13 [Vigna angularis]
Length = 559
Score = 642 bits (1656), Expect = 0.0
Identities = 314/470 (66%), Positives = 379/470 (79%), Gaps = 5/470 (1%)
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPS 60
MEK V+IAVVPG G+SHLVPILQFSKRL+QLHPNFHVTC IP+ GSPS+A+KS+L+TLP
Sbjct: 95 MEKRVNIAVVPGPGFSHLVPILQFSKRLLQLHPNFHVTCLIPSFGSPSSASKSVLETLPP 154
Query: 61 NINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
NI FL PV P DLPQG +E+QI T+T SLP +HQ LKS+ + P VALV D+F+ E
Sbjct: 155 NIESIFLEPVKPEDLPQGVAIETQIQFTVTLSLPSIHQALKSITSKTPFVALVADSFAFE 214
Query: 121 VLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF 180
L+ +E N+LSYIYFPSAATTL+W Y+ KLD+ETSCEYRD+ EP+KIPGCVP+HGRD
Sbjct: 215 ALDFAEEFNLLSYIYFPSAATTLSWYLYVLKLDKETSCEYRDLPEPVKIPGCVPIHGRDL 274
Query: 181 LSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPI 240
+ AQDRSSQ YK FL + S DG+ +NSF EIE GP+ A+KEEG P V+PVGPI
Sbjct: 275 NNQAQDRSSQVYKLFLQRAQRFCSVDGIFINSFFEIETGPIRALKEEGRGYPQVFPVGPI 334
Query: 241 IETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLW 300
++T GDDA GLECL WLDKQ+ SVLYVSFGSGGTL+QEQ+ ELA GLELSN KFLW
Sbjct: 335 VQT----GDDAKGLECLTWLDKQEDGSVLYVSFGSGGTLTQEQVNELAYGLELSNHKFLW 390
Query: 301 VLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGF 360
V+R PSS + A YL A+ +D L FLP GFLERTKE+G V+ SWAPQIQ+L+ +S+GGF
Sbjct: 391 VVREPSSLAFDA-YLRAQRSVDPLHFLPDGFLERTKEQGMVVPSWAPQIQVLAHSSIGGF 449
Query: 361 LTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAK 420
LTHCGWNS LESV++GVPLITWPLFAEQ+MNAV+LSEGLKVG+R V+ENG+VERVE+ K
Sbjct: 450 LTHCGWNSVLESVMNGVPLITWPLFAEQRMNAVVLSEGLKVGVRPRVSENGLVERVEIVK 509
Query: 421 VIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWRNLV 470
VIK LME +EG ++ M+ELK+AASNA+K DGSSTKT+S++ KW +LV
Sbjct: 510 VIKCLMEEEEGGEMHKRMEELKQAASNALKADGSSTKTLSELVQKWESLV 559
>emb|CAC35167.1| arbutin synthase [Rauvolfia serpentina]
gi|28380078|sp|Q9AR73|HQGT_RAUSE Hydroquinone
glucosyltransferase (Arbutin synthase)
Length = 470
Score = 512 bits (1319), Expect = e-144
Identities = 248/470 (52%), Positives = 333/470 (70%), Gaps = 3/470 (0%)
Query: 1 MEKTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPS 60
ME T HIA+VP G HL+P+++F+KRLV H NF VT IPT G A KS L LP+
Sbjct: 1 MEHTPHIAMVPTPGMGHLIPLVEFAKRLVLRH-NFGVTFIIPTDGPLPKAQKSFLDALPA 59
Query: 61 NINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
+N+ LPPV+ +DLP +E++I LT+T SLP++ +K+L L ALVVD F +
Sbjct: 60 GVNYVLLPPVSFDDLPADVRIETRICLTITRSLPFVRDAVKTLLATTKLAALVVDLFGTD 119
Query: 121 VLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF 180
++ E + YI++P+ A L+ F+LPKLD+ SCEYRD+ EP++IPGC+P+HG+DF
Sbjct: 120 AFDVAIEFKVSPYIFYPTTAMCLSLFFHLPKLDQMVSCEYRDVPEPLQIPGCIPIHGKDF 179
Query: 181 LSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPI 240
L AQDR + AYK L K A+G++VN+F ++E GPL A++EE PPVYP+GP+
Sbjct: 180 LDPAQDRKNDAYKCLLHQAKRYRLAEGIMVNTFNDLEPGPLKALQEEDQGKPPVYPIGPL 239
Query: 241 IETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLW 300
I ++ S D ECL WLD Q SVL++SFGSGG +S Q +ELALGLE+S +FLW
Sbjct: 240 IRADSSSKVD--DCECLKWLDDQPRGSVLFISFGSGGAVSHNQFIELALGLEMSEQRFLW 297
Query: 301 VLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGF 360
V+R+P+ ++A Y S +N D L +LP GFLERTK + ++ SWAPQ +ILS S GGF
Sbjct: 298 VVRSPNDKIANATYFSIQNQNDALAYLPEGFLERTKGRCLLVPSWAPQTEILSHGSTGGF 357
Query: 361 LTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAK 420
LTHCGWNS LESVV+GVPLI WPL+AEQKMNAV+L+EGLKV LR ENG++ RVE+A
Sbjct: 358 LTHCGWNSILESVVNGVPLIAWPLYAEQKMNAVMLTEGLKVALRPKAGENGLIGRVEIAN 417
Query: 421 VIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWRNLV 470
+K LMEG+EG+K R+ MK+LK+AAS A+ +DGSSTK ++++A KW N +
Sbjct: 418 AVKGLMEGEEGKKFRSTMKDLKDAASRALSDDGSSTKALAELACKWENKI 467
>emb|CAB80916.1| putative flavonol glucosyltransferase [Arabidopsis thaliana]
gi|14532902|gb|AAK64133.1| putative flavonol
glucosyltransferase [Arabidopsis thaliana]
gi|13430700|gb|AAK25972.1| putative flavonol
glucosyltransferase [Arabidopsis thaliana]
gi|21537114|gb|AAM61455.1| putative flavonol
glucosyltransferase [Arabidopsis thaliana]
gi|15234056|ref|NP_192016.1|
UDP-glucoronosyl/UDP-glucosyl transferase family protein
[Arabidopsis thaliana] gi|25286798|pir||B85014 probable
flavonol glucosyltransferase [imported] - Arabidopsis
thaliana gi|28380085|sp|Q9M156|HQGT_ARATH Probable
hydroquinone glucosyltransferase (Arbutin synthase)
Length = 480
Score = 484 bits (1247), Expect = e-135
Identities = 243/466 (52%), Positives = 323/466 (69%), Gaps = 3/466 (0%)
Query: 3 KTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNI 62
KT H+A++P G HL+P+++F+KRLV LH VT I G PS A +++L +LPS+I
Sbjct: 5 KTPHVAIIPSPGMGHLIPLVEFAKRLVHLH-GLTVTFVIAGEGPPSKAQRTVLDSLPSSI 63
Query: 63 NHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPL-VALVVDAFSVEV 121
+ FLPPV+ DL T +ES+I LT+T S P L + S + L ALVVD F +
Sbjct: 64 SSVFLPPVDLTDLSSSTRIESRISLTVTRSNPELRKVFDSFVEGGRLPTALVVDLFGTDA 123
Query: 122 LNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFL 181
++ E ++ YI++P+ A L++ +LPKLDE SCE+R++ EP+ +PGCVP+ G+DFL
Sbjct: 124 FDVAVEFHVPPYIFYPTTANVLSFFLHLPKLDETVSCEFRELTEPLMLPGCVPVAGKDFL 183
Query: 182 SIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPII 241
AQDR AYK L K A+G+LVN+F E+E + A++E G D PPVYPVGP++
Sbjct: 184 DPAQDRKDDAYKWLLHNTKRYKEAEGILVNTFFELEPNAIKALQEPGLDKPPVYPVGPLV 243
Query: 242 ETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWV 301
+ ECL WLD Q SVLYVSFGSGGTL+ EQ+ ELALGL S +FLWV
Sbjct: 244 NIGKQEAKQTEESECLKWLDNQPLGSVLYVSFGSGGTLTCEQLNELALGLADSEQRFLWV 303
Query: 302 LRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFL 361
+R+PS ++S+ Y + + D L FLP GFLERTK++GFVI WAPQ Q+L+ S GGFL
Sbjct: 304 IRSPSGIANSS-YFDSHSQTDPLTFLPPGFLERTKKRGFVIPFWAPQAQVLAHPSTGGFL 362
Query: 362 THCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKV 421
THCGWNSTLESVV G+PLI WPL+AEQKMNAVLLSE ++ LR ++G+V R EVA+V
Sbjct: 363 THCGWNSTLESVVSGIPLIAWPLYAEQKMNAVLLSEDIRAALRPRAGDDGLVRREEVARV 422
Query: 422 IKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
+K LMEG+EG+ +RN MKELKEAA +K+DG+STK +S +ALKW+
Sbjct: 423 VKGLMEGEEGKGVRNKMKELKEAACRVLKDDGTSTKALSLVALKWK 468
>ref|NP_171649.1| UDP-glucoronosyl/UDP-glucosyl transferase family protein
[Arabidopsis thaliana] gi|25286824|pir||G86144
hypothetical protein F6F3.22 [imported] - Arabidopsis
thaliana gi|9665137|gb|AAF97321.1| Similar to
UTP-glucose glucosyltransferases [Arabidopsis thaliana]
Length = 481
Score = 456 bits (1172), Expect = e-127
Identities = 227/465 (48%), Positives = 322/465 (68%), Gaps = 4/465 (0%)
Query: 4 TVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNIN 63
T H+A++P G HL+P+++ +KRL+ H F VT IP PS A +S+L +LPS+I
Sbjct: 6 TPHVAIIPSPGIGHLIPLVELAKRLLDNH-GFTVTFIIPGDSPPSKAQRSVLNSLPSSIA 64
Query: 64 HTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVA-LVVDAFSVEVL 122
FLPP + +D+P +E++I LT+T S P L + SL+ E L A LVVD F +
Sbjct: 65 SVFLPPADLSDVPSTARIETRISLTVTRSNPALRELFGSLSAEKRLPAVLVVDLFGTDAF 124
Query: 123 NIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLS 182
++ E ++ YI++ S A L + +LPKLDE SCE+R++ EP+ IPGCVP+ G+DF+
Sbjct: 125 DVAAEFHVSPYIFYASNANVLTFLLHLPKLDETVSCEFRELTEPVIIPGCVPITGKDFVD 184
Query: 183 IAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPIIE 242
QDR ++YK L VK A+G+LVNSF+++E + ++E D PPVY +GP++
Sbjct: 185 PCQDRKDESYKWLLHNVKRFKEAEGILVNSFVDLEPNTIKIVQEPAPDKPPVYLIGPLVN 244
Query: 243 TETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVL 302
+ + D + +CL WLD Q SVLYVSFGSGGTL+ EQ +ELALGL S +FLWV+
Sbjct: 245 SGSHDADVNDEYKCLNWLDNQPFGSVLYVSFGSGGTLTFEQFIELALGLAESGKRFLWVI 304
Query: 303 RAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLT 362
R+PS +SS+ Y + ++ D FLP GFL+RTKEKG V+ SWAPQ QIL+ S+GGFLT
Sbjct: 305 RSPSGIASSS-YFNPQSRNDPFSFLPQGFLDRTKEKGLVVGSWAPQAQILTHTSIGGFLT 363
Query: 363 HCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKVI 422
HCGWNS+LES+V+GVPLI WPL+AEQKMNA+LL + + LRA + E+G+V R EVA+V+
Sbjct: 364 HCGWNSSLESIVNGVPLIAWPLYAEQKMNALLLVD-VGAALRARLGEDGVVGREEVARVV 422
Query: 423 KYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
K L+EG+EG +R MKELKE + +++DG STK++++++LKW+
Sbjct: 423 KGLIEGEEGNAVRKKMKELKEGSVRVLRDDGFSTKSLNEVSLKWK 467
>gb|AAB61023.1| Similar to UTP-Glucose Glucosyltransferase; coded for by A.
thaliana cDNA T46230; coded for by A. thaliana cDNA
H76538; coded for by A. thaliana cDNA H76290
[Arabidopsis thaliana] gi|7433902|pir||T01732
UTP-glucose glucosyltransferase homolog A_IG002N01.15 -
Arabidopsis thaliana
Length = 462
Score = 451 bits (1160), Expect = e-125
Identities = 233/466 (50%), Positives = 309/466 (66%), Gaps = 21/466 (4%)
Query: 3 KTVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNI 62
KT H+A++P G HL+P+++F+KRLV LH VT I G PS A +++L +LPS+I
Sbjct: 5 KTPHVAIIPSPGMGHLIPLVEFAKRLVHLH-GLTVTFVIAGEGPPSKAQRTVLDSLPSSI 63
Query: 63 NHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPL-VALVVDAFSVEV 121
+ FLPPV+ DL T +ES+I LT+T S P L + S + L ALVVD F +
Sbjct: 64 SSVFLPPVDLTDLSSSTRIESRISLTVTRSNPELRKVFDSFVEGGRLPTALVVDLFGTDA 123
Query: 122 LNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFL 181
++ E ++ YI++P+ A L ++ EP+ +PGCVP+ G+DFL
Sbjct: 124 FDVAVEFHVPPYIFYPTTANVL------------------ELTEPLMLPGCVPVAGKDFL 165
Query: 182 SIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPII 241
AQDR AYK L K A+G+LVN+F E+E + A++E G D PPVYPVGP++
Sbjct: 166 DPAQDRKDDAYKWLLHNTKRYKEAEGILVNTFFELEPNAIKALQEPGLDKPPVYPVGPLV 225
Query: 242 ETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWV 301
+ ECL WLD Q SVLYVSFGSGGTL+ EQ+ ELALGL S +FLWV
Sbjct: 226 NIGKQEAKQTEESECLKWLDNQPLGSVLYVSFGSGGTLTCEQLNELALGLADSEQRFLWV 285
Query: 302 LRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFL 361
+R+PS ++S+ Y + + D L FLP GFLERTK++GFVI WAPQ Q+L+ S GGFL
Sbjct: 286 IRSPSGIANSS-YFDSHSQTDPLTFLPPGFLERTKKRGFVIPFWAPQAQVLAHPSTGGFL 344
Query: 362 THCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKV 421
THCGWNSTLESVV G+PLI WPL+AEQKMNAVLLSE ++ LR ++G+V R EVA+V
Sbjct: 345 THCGWNSTLESVVSGIPLIAWPLYAEQKMNAVLLSEDIRAALRPRAGDDGLVRREEVARV 404
Query: 422 IKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
+K LMEG+EG+ +RN MKELKEAA +K+DG+STK +S +ALKW+
Sbjct: 405 VKGLMEGEEGKGVRNKMKELKEAACRVLKDDGTSTKALSLVALKWK 450
>gb|AAO00939.1| Putative UTP-glucose glucosyltransferase [Arabidopsis thaliana]
gi|15223392|ref|NP_171646.1|
UDP-glucoronosyl/UDP-glucosyl transferase family protein
[Arabidopsis thaliana] gi|17065184|gb|AAL32746.1|
Putative UTP-glucose glucosyltransferase [Arabidopsis
thaliana]
Length = 480
Score = 448 bits (1152), Expect = e-124
Identities = 226/466 (48%), Positives = 312/466 (66%), Gaps = 5/466 (1%)
Query: 4 TVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNIN 63
T HIA++P G HL+P ++ +KRLVQ H F VT I SPS A +S+L +LPS+I
Sbjct: 6 TPHIAIMPSPGMGHLIPFVELAKRLVQ-HDCFTVTMIISGETSPSKAQRSVLNSLPSSIA 64
Query: 64 HTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQ--GLKSLAKEIPLVALVVDAFSVEV 121
FLPP + +D+P +E++ +LT+T S P L + G S K +P V LVVD F +
Sbjct: 65 SVFLPPADLSDVPSTARIETRAMLTMTRSNPALRELFGSLSTKKSLPAV-LVVDMFGADA 123
Query: 122 LNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFL 181
++ + ++ YI++ S A L++ +LPKLD+ SCE+R + EP+KIPGCVP+ G+DFL
Sbjct: 124 FDVAVDFHVSPYIFYASNANVLSFFLHLPKLDKTVSCEFRYLTEPLKIPGCVPITGKDFL 183
Query: 182 SIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPII 241
QDR+ AYK L K A G+LVNSF+++E + A++E D P VYP+GP++
Sbjct: 184 DTVQDRNDDAYKLLLHNTKRYKEAKGILVNSFVDLESNAIKALQEPAPDKPTVYPIGPLV 243
Query: 242 ETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWV 301
T + + + + CL+WLD Q SVLY+SFGSGGTL+ EQ ELA+GL S +F+WV
Sbjct: 244 NTSSSNVNLEDKFGCLSWLDNQPFGSVLYISFGSGGTLTCEQFNELAIGLAESGKRFIWV 303
Query: 302 LRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFL 361
+R+PS SS+ Y + ++ D FLP GFL+RTKEKG V+ SWAPQ+QIL+ S GFL
Sbjct: 304 IRSPSEIVSSS-YFNPHSETDPFSFLPIGFLDRTKEKGLVVPSWAPQVQILAHPSTCGFL 362
Query: 362 THCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKV 421
THCGWNSTLES+V+GVPLI WPLFAEQKMN +LL E + LR E+GIV R EV +V
Sbjct: 363 THCGWNSTLESIVNGVPLIAWPLFAEQKMNTLLLVEDVGAALRIHAGEDGIVRREEVVRV 422
Query: 422 IKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
+K LMEG+EG+ + N +KELKE + +DG S+K+ ++ LKW+
Sbjct: 423 VKALMEGEEGKAIGNKVKELKEGVVRVLGDDGLSSKSFGEVLLKWK 468
>dbj|BAD43267.1| putative flavonol 3-o-glucosyltransferase [Arabidopsis thaliana]
Length = 468
Score = 446 bits (1147), Expect = e-124
Identities = 225/458 (49%), Positives = 318/458 (69%), Gaps = 6/458 (1%)
Query: 11 PGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHTFLPPV 70
PG+G HL+P+++ +KRL+ H F VT IP PS A +S+L +LPS+I FLPP
Sbjct: 2 PGIG--HLIPLVELAKRLLDNH-GFTVTFIIPGDSPPSKAQRSVLNSLPSSIASVFLPPA 58
Query: 71 NPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVA-LVVDAFSVEVLNIGKELN 129
+ +D+P +E++I LT+T S P L + SL+ E L A LVVD F + ++ E +
Sbjct: 59 DLSDVPSTARIETRISLTVTRSNPALRELFGSLSAEKRLPAVLVVDLFGTDAFDVAAEFH 118
Query: 130 MLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIAQDRSS 189
+ YI++ S A L + +LPKLDE SCE+R++ EP+ IPGCVP+ G+DF+ QDR
Sbjct: 119 VSPYIFYASNANVLTFLLHLPKLDETVSCEFRELTEPVIIPGCVPITGKDFVDPCQDRKD 178
Query: 190 QAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPIIETETKSGD 249
++YK L VK A+G+LVNSF+++E + ++E D PPVY +GP++ + + D
Sbjct: 179 ESYKWLLHNVKRFKEAEGILVNSFVDLEPNTIKIVQEPAPDKPPVYLIGPLVNSGSHDAD 238
Query: 250 DANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSS 309
+ +CL WLD Q SVLYVSFGSGGTL+ EQ +ELALGL S +FLWV+R+PS +
Sbjct: 239 VNDEYKCLNWLDNQPFGSVLYVSFGSGGTLTFEQFIELALGLAESGKRFLWVIRSPSGIA 298
Query: 310 SSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWNST 369
SS+ Y + ++ D FLP GFL+RTKEKG V+ SWAPQ QIL+ S+GGFLTHCGWNS+
Sbjct: 299 SSS-YFNPQSRNDPFSFLPQGFLDRTKEKGLVVGSWAPQAQILTHTSIGGFLTHCGWNSS 357
Query: 370 LESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKVIKYLMEGD 429
LES+V+GVPLI WPL+AEQKMNA+LL + + LRA + E+G+V R EVA+V+K L+EG+
Sbjct: 358 LESIVNGVPLIAWPLYAEQKMNALLLVD-VGAALRARLGEDGVVGREEVARVVKGLIEGE 416
Query: 430 EGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
EG +R MKELKE + +++DG STK++++++LKW+
Sbjct: 417 EGNAVRKKMKELKEGSVRVLRDDGFSTKSLNEVSLKWK 454
>pir||D86144 protein probable UTP-glucose glucosyltransferase [imported] -
Arabidopsis thaliana gi|9665140|gb|AAF97324.1| Putative
UTP-glucose glucosyltransferase [Arabidopsis thaliana]
Length = 469
Score = 439 bits (1130), Expect = e-122
Identities = 222/460 (48%), Positives = 307/460 (66%), Gaps = 5/460 (1%)
Query: 10 VPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHTFLPP 69
+P G HL+P ++ +KRLVQ H F VT I SPS A +S+L +LPS+I FLPP
Sbjct: 1 MPSPGMGHLIPFVELAKRLVQ-HDCFTVTMIISGETSPSKAQRSVLNSLPSSIASVFLPP 59
Query: 70 VNPNDLPQGTTMESQILLTLTNSLPYLHQ--GLKSLAKEIPLVALVVDAFSVEVLNIGKE 127
+ +D+P +E++ +LT+T S P L + G S K +P V LVVD F + ++ +
Sbjct: 60 ADLSDVPSTARIETRAMLTMTRSNPALRELFGSLSTKKSLPAV-LVVDMFGADAFDVAVD 118
Query: 128 LNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIAQDR 187
++ YI++ S A L++ +LPKLD+ SCE+R + EP+KIPGCVP+ G+DFL QDR
Sbjct: 119 FHVSPYIFYASNANVLSFFLHLPKLDKTVSCEFRYLTEPLKIPGCVPITGKDFLDTVQDR 178
Query: 188 SSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPIIETETKS 247
+ AYK L K A G+LVNSF+++E + A++E D P VYP+GP++ T + +
Sbjct: 179 NDDAYKLLLHNTKRYKEAKGILVNSFVDLESNAIKALQEPAPDKPTVYPIGPLVNTSSSN 238
Query: 248 GDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSS 307
+ + CL+WLD Q SVLY+SFGSGGTL+ EQ ELA+GL S +F+WV+R+PS
Sbjct: 239 VNLEDKFGCLSWLDNQPFGSVLYISFGSGGTLTCEQFNELAIGLAESGKRFIWVIRSPSE 298
Query: 308 SSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWN 367
SS+ Y + ++ D FLP GFL+RTKEKG V+ SWAPQ+QIL+ S GFLTHCGWN
Sbjct: 299 IVSSS-YFNPHSETDPFSFLPIGFLDRTKEKGLVVPSWAPQVQILAHPSTCGFLTHCGWN 357
Query: 368 STLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKVIKYLME 427
STLES+V+GVPLI WPLFAEQKMN +LL E + LR E+GIV R EV +V+K LME
Sbjct: 358 STLESIVNGVPLIAWPLFAEQKMNTLLLVEDVGAALRIHAGEDGIVRREEVVRVVKALME 417
Query: 428 GDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
G+EG+ + N +KELKE + +DG S+K+ ++ LKW+
Sbjct: 418 GEEGKAIGNKVKELKEGVVRVLGDDGLSSKSFGEVLLKWK 457
>dbj|BAB86924.1| glucosyltransferase-6 [Vigna angularis]
Length = 414
Score = 417 bits (1072), Expect = e-115
Identities = 221/410 (53%), Positives = 282/410 (67%), Gaps = 9/410 (2%)
Query: 55 LQTLPSNINHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVV 114
L++LPSNIN TFLPP+N DLPQ QI L + S+P + L+SL PLVAL+
Sbjct: 1 LKSLPSNINCTFLPPINKRDLPQDALSVVQIQLAVFQSIPSIQHALRSLLSTTPLVALIA 60
Query: 115 DAFSVEVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVP 174
D F+ E L I KEL +LSY+YFP +A ++ +LP L ++ SCEYRD E + IPGCVP
Sbjct: 61 DLFANEALEIAKELKLLSYVYFPHSAMAVSVFLHLPSLHQQISCEYRDHKEAVNIPGCVP 120
Query: 175 LHGRDFLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEG-----G 229
+ GRD S QDRS+ AYK L K LS A G +VNSF +IE A++E
Sbjct: 121 IQGRDLPSHFQDRSTLAYKLILDRCKRLSHAHGFIVNSFSKIEESCERALQEHNRVSSSS 180
Query: 230 DNPPVYPVGPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELAL 289
+ VY +GP ++T S +D G EC+ WL+ Q+ SVLYVSFGSGGTLSQ+Q+ ELA
Sbjct: 181 KSSGVYLIGPNVQTG--SSNDPKGSECVNWLENQEAKSVLYVSFGSGGTLSQQQMNELAF 238
Query: 290 GLELSNTKFLWVLRAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQI 349
GLELS KFLWV+RAPS S+ A YL A +D D LQFLP+GFLERTK +GFV+ SWAPQ
Sbjct: 239 GLELSGEKFLWVVRAPSDSADGA-YLGASSD-DPLQFLPNGFLERTKGRGFVVRSWAPQT 296
Query: 350 QILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNE 409
QIL S GGFLTHCGWNS LES+V GVP++ WPLFAEQ+ NAVLL+EG+KV LR N+
Sbjct: 297 QILGHVSTGGFLTHCGWNSALESIVLGVPMVAWPLFAEQRTNAVLLTEGVKVALRPKFND 356
Query: 410 NGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTI 459
+GI ER E+A+VIK LM G+EG + +++L++AA+ A++E G T+
Sbjct: 357 SGIAEREEIAEVIKGLMVGEEGRLIPGRIEKLRDAAAEALEEHGPPQGTL 406
>ref|NP_917729.1| putative UTP-glucose glucosyltransferase [Oryza sativa (japonica
cultivar-group)] gi|11034680|dbj|BAB17182.1| arbutin
synthase-like [Oryza sativa (japonica cultivar-group)]
Length = 507
Score = 327 bits (838), Expect = 5e-88
Identities = 186/463 (40%), Positives = 270/463 (58%), Gaps = 18/463 (3%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHT 65
H+ ++ G HL+P+ + ++R+V+ + F T T S + + S +LP +I+
Sbjct: 16 HVVLLASPGAGHLLPVAELARRIVE-YDGFTATIVTHTNFSSAEHS-STFSSLPPSISIA 73
Query: 66 FLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVV-DAFSVEVLNI 124
LP V+ +DLP +E++IL + +LP+L L+SL VA+ + D S L +
Sbjct: 74 ALPEVSVDDLPADARVETRILTVVRRALPHLRDLLRSLLDSPAGVAVFLSDLLSPRALAV 133
Query: 125 GKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIA 184
EL + Y++ S L + P LD T+CE+RD+ P+ +PGCVPLHG D +
Sbjct: 134 AAELGIPRYVFCTSNLMCLTSFLHNPVLDRTTTCEFRDLPGPVLLPGCVPLHGSDLVDPV 193
Query: 185 QDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKE--EGGDNPPVYPVGPIIE 242
QDR++ Y+ + ADG LVN+F +E A KE + G PP Y VGP +
Sbjct: 194 QDRANPVYRLVIEMGLDYLRADGFLVNTFDAMEHDTAVAFKELSDKGVYPPAYAVGPFVR 253
Query: 243 TET-KSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWV 301
+ + K+ +DA C+ WLD Q SVLYV GSGGTLS EQ E+A GLE S +FLWV
Sbjct: 254 SPSGKAANDA----CIRWLDDQPDGSVLYVCLGSGGTLSTEQTAEVAAGLEASGQRFLWV 309
Query: 302 LRAPSSSSSSAGYLSAENDID----TLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSV 357
+R PS +A Y S D D +LP GFLERTK G + WAPQ++IL+ +V
Sbjct: 310 VRYPSDKDKTASYFSVSGDGDGEDSPTNYLPEGFLERTKGTGLAVPMWAPQVEILNHRAV 369
Query: 358 GGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLR---ASVNENGIVE 414
GGF++HCGWNSTLE+V GVP++ WPL+AEQ+MNAV+LS + GL ++ E+G+V
Sbjct: 370 GGFVSHCGWNSTLETVAAGVPMVAWPLYAEQRMNAVMLSSS-RAGLALRPSNAREDGVVT 428
Query: 415 RVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTK 457
R EVA V + L+ G++G R +EL+EAA+ A + G ++
Sbjct: 429 RDEVAAVARELITGEKGAAARRKARELREAAAKATRAPGGPSR 471
>ref|NP_917723.1| putative UTP-glucose glucosyltransferase [Oryza sativa (japonica
cultivar-group)] gi|15623919|dbj|BAB67976.1| arbutin
synthase-like [Oryza sativa (japonica cultivar-group)]
gi|11034674|dbj|BAB17176.1| arbutin synthase-like [Oryza
sativa (japonica cultivar-group)]
Length = 480
Score = 322 bits (825), Expect = 2e-86
Identities = 191/478 (39%), Positives = 268/478 (55%), Gaps = 22/478 (4%)
Query: 4 TVHIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNA-TKSILQTLPSNI 62
T H+ ++P G H+ P Q + RL H T I T + S A S L +LP+ +
Sbjct: 8 TPHVVLLPSPGAGHVAPAAQLAARLATHHG---CTATIVTYTNLSTARNSSALASLPTGV 64
Query: 63 NHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKS-LAKEIP--LVALVVDAFSV 119
T LP V+ +DLP +E++I + +LP+L + L S L P + L+ D
Sbjct: 65 TATALPEVSLDDLPADARIETRIFAVVRRTLPHLRELLLSFLGSSSPAGVTTLLTDMLCP 124
Query: 120 EVLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRD 179
L + EL + Y++F S L Y P+L T+CE RD+ EP+ +PGCVPLHG D
Sbjct: 125 AALAVAAELGIPRYVFFTSNLLCLTTLLYTPELATTTACECRDLPEPVVLPGCVPLHGAD 184
Query: 180 FLSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKE--EGGDNPPVYPV 237
+ Q+R++ Y+ + ADG L+N+F +E L A KE + G PP Y V
Sbjct: 185 LIDPVQNRANPVYQLMVELGLDYLLADGFLINTFDAMEHDTLVAFKELSDKGVYPPAYAV 244
Query: 238 GPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTK 297
GP++ + T AN + C+ WLD+Q SVLYV GSGGTLS Q ELA GLE S +
Sbjct: 245 GPLVRSPTSEA--ANDV-CIRWLDEQPDGSVLYVCLGSGGTLSVAQTAELAAGLEASGQR 301
Query: 298 FLWVLRAPSSSSSSAGYLSA----ENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILS 353
FLWV+R PS SA Y +ND D + +LP GF+ERTK G + WAPQ+++L+
Sbjct: 302 FLWVVRFPSDKDVSASYFGTNDRGDND-DPMSYLPEGFVERTKGAGLAVPLWAPQVEVLN 360
Query: 354 PNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRA----SVNE 409
+VGGFL+HCGWNSTLE+ GVP + WPLFAEQKMNAV+LS +VGL A ++
Sbjct: 361 HRAVGGFLSHCGWNSTLEAASAGVPTLAWPLFAEQKMNAVMLSSE-RVGLAALRVRPDDD 419
Query: 410 NGIVERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
G+V R EVA ++ LM G +G +EL+ AA+ A G + ++ + +W+
Sbjct: 420 RGVVTREEVASAVRELMAGKKGAAAWKKARELRAAAAVASAPGGPQHQALAGMVGEWK 477
>dbj|BAD28247.1| putative Hydroquinone glucosyltransferase [Oryza sativa (japonica
cultivar-group)] gi|50251522|dbj|BAD28883.1| putative
Hydroquinone glucosyltransferase [Oryza sativa (japonica
cultivar-group)]
Length = 484
Score = 316 bits (809), Expect = 1e-84
Identities = 188/473 (39%), Positives = 271/473 (56%), Gaps = 19/473 (4%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHT 65
H+ ++ G HL+P+ + ++RLV H I +L P+ ++L +LP+++
Sbjct: 19 HVVLMASPGAGHLIPLAELARRLVSDHGFAVTVVTIASLSDPATDA-AVLSSLPASVATA 77
Query: 66 FLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNIG 125
LPPV +DLP S + + S+P+L + L P A+V D F L +
Sbjct: 78 VLPPVALDDLPADIGFGSVMFELVRRSVPHL----RPLVVGSPAAAIVCDFFGTPALALA 133
Query: 126 KELNMLSYIYFPSAATTLAWCFYLPKL-DEETSCEYRDILEPIKIPGCVPLHGRDFLSIA 184
EL + Y++FP++ + ++ + +L D + EYRD+ +P+ +PGC PL D
Sbjct: 134 AELGVPGYVFFPTSISFISVVRSVVELHDGAAAGEYRDLPDPLVLPGCAPLRHGDIPDGF 193
Query: 185 QDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEG--GDNPPVYPVGPIIE 242
+D + Y + L + ADG LVNSF E+E G A + +G G PPVY VGP +
Sbjct: 194 RDSADPVYAYVLEEGRRYGGADGFLVNSFPEMEPGAAEAFRRDGENGAFPPVYLVGPFVR 253
Query: 243 TETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVL 302
+S +DA+ CL WLD+Q SV+YVSFGSGG LS EQ ELA GLE+S +FLWV+
Sbjct: 254 P--RSDEDADESACLEWLDRQPAGSVVYVSFGSGGALSVEQTRELAAGLEMSGHRFLWVV 311
Query: 303 RAPSSSS--SSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGF 360
R P SS G S N +D FLP GF+ERT +G + SWAPQ+++L+ + F
Sbjct: 312 RMPRKGGLLSSMG-ASYGNPMD---FLPEGFVERTNGRGLAVASWAPQVRVLAHPATAAF 367
Query: 361 LTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRAS-VNENGIVERVEVA 419
++HCGWNS LESV GVP+I WPL AEQKMNA +L+E V L S V G+V R EVA
Sbjct: 368 VSHCGWNSALESVSSGVPMIAWPLHAEQKMNAAILTEVAGVALPLSPVAPGGVVSREEVA 427
Query: 420 KVIKYLME-GDEGEKLRNNMKELK-EAASNAVKEDGSSTKTISQIALKWRNLV 470
+K LM+ G++G R +EL+ AA+ A DG+S + + ++A KW+N V
Sbjct: 428 AAVKELMDPGEKGSAARRRARELQAAAAARAWSPDGASRRALEEVAGKWKNAV 480
>ref|XP_476173.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|48843758|gb|AAT47017.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 490
Score = 311 bits (796), Expect = 4e-83
Identities = 180/463 (38%), Positives = 261/463 (55%), Gaps = 16/463 (3%)
Query: 17 HLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQT-LPSNINHTFLPPVNPND- 74
HL+P + ++RLV H F PS ++ + L ++ LP P D
Sbjct: 34 HLIPFAELARRLVADHGLAATLLFASARSPPSEQYLAVAASVLAEGVDLVALPAPAPADA 93
Query: 75 LPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNIGKELNMLSYI 134
LP ++ + + S+P + +SLA PL ALVVD + +EL + Y+
Sbjct: 94 LPGDASVRERAAHAVARSVPRVRDVARSLAATAPLAALVVDMIGAPARAVAEELGVPFYM 153
Query: 135 YFPSAATTLAWCFYLPKLDEETSC---EYRDILEPIKIPGCVPLHGRDF-LSIAQDRSSQ 190
+F S L+ +LP LD + + E+RD EPI++PGCVP+H D S+ DRSS
Sbjct: 154 FFTSPWMLLSLFLHLPSLDADAARAGGEHRDATEPIRLPGCVPIHAHDLPSSMLADRSSA 213
Query: 191 AYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNPPVYPVGPIIETETKSGDD 250
Y L + + ADGVLVN+F E+E P +G PPV+ VGP+I T + +
Sbjct: 214 TYAGLLAMARDAARADGVLVNTFRELE--PAIGDGADGVKLPPVHAVGPLIWTRPVAMER 271
Query: 251 ANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSSSSS 310
+ ECL+WL++Q SV+YVSFGSGGTL+ +Q ELALGLELS +F+W ++ P +S
Sbjct: 272 DH--ECLSWLNQQPRGSVVYVSFGSGGTLTWQQTAELALGLELSQHRFIWAIKRPDQDTS 329
Query: 311 SAGYLSAEN---DIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLTHCGWN 367
S + N + + + FLP GF+ERT+ G ++ SWAPQ IL S+G FLTHCGWN
Sbjct: 330 SGAFFGTANSRGEEEGMDFLPEGFIERTRGVGLLVPSWAPQTSILGHASIGCFLTHCGWN 389
Query: 368 STLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENGIVERVEVAKVIKYLME 427
STLESV +GVP+I WPL+AEQKMNA ++ KV +R +V + E+A IK +M+
Sbjct: 390 STLESVSNGVPMIAWPLYAEQKMNAAMMEVQAKVAIRINVGNERFIMNEEIANTIKRVMK 449
Query: 428 GDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWRNLV 470
G+E E L+ + EL + A A+ S ++Q+ W++ V
Sbjct: 450 GEEAEMLKMRIGELNDKAVYALSRGCS---ILAQVTHVWKSTV 489
>dbj|BAD28262.1| putative Hydroquinone glucosyltransferase [Oryza sativa (japonica
cultivar-group)]
Length = 489
Score = 306 bits (784), Expect = 9e-82
Identities = 173/469 (36%), Positives = 265/469 (55%), Gaps = 17/469 (3%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHT 65
H+ ++ G HL+P+ + ++RL H L +P +A ++L +LP+++
Sbjct: 26 HVVLLASPGAGHLIPLAELARRLADHHGVAPTLVTFADLDNP-DARSAVLSSLPASVATA 84
Query: 66 FLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNIG 125
LP V +D+P +E + + SLP+L L+S+ ALV D F L++
Sbjct: 85 TLPAVPLDDIPADAGLERMLFEVVHRSLPHLRVLLRSIGST---AALVPDFFCAAALSVA 141
Query: 126 KELNMLSYIYFPSAATTLAWCFYLPKL-DEETSCEYRDILEPIKIPGCVPLHGRDFLSIA 184
EL + YI+FP++ T L +L D + EY + +P+++PG V L +F
Sbjct: 142 AELGVPGYIFFPTSITALYLMRRTVELHDFAAAGEYHALPDPLELPGGVSLRTAEFPEAF 201
Query: 185 QDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKE--EGGDNPPVYPVGPIIE 242
+D ++ Y + +L A G L NSF E+E + K+ E G PP YPVGP +
Sbjct: 202 RDSTAPVYGQLVETGRLYRGAAGFLANSFYELEPAAVEDSKKAAEKGTFPPAYPVGPFVR 261
Query: 243 TETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVL 302
+ S D+A CL WLD Q SV++VSFGS G LS EQ ELA GLE+S +FLWV+
Sbjct: 262 S---SSDEAGESACLEWLDLQPAGSVVFVSFGSFGVLSVEQTRELAAGLEMSGHRFLWVV 318
Query: 303 RAPSSSSSSAGYLSAENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGFLT 362
R PS + + + + +D D L ++P GFLERT+ +G + +WAPQ+++LS + F++
Sbjct: 319 RMPSLNDA---HRNGGHDEDPLAWVPDGFLERTRGRGLAVAAWAPQVRVLSHPATAAFVS 375
Query: 363 HCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRASVNENG----IVERVEV 418
HCGWNSTLESV GVP+I WPL +EQ+MNAV+L E + + LR E +V R E+
Sbjct: 376 HCGWNSTLESVATGVPMIAWPLHSEQRMNAVVLEESVGMALRPRAREEDVGGTVVRRGEI 435
Query: 419 AKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
A +K +MEG++G +R +EL++AA +GSS + + +A KW+
Sbjct: 436 AVAVKEVMEGEKGHGVRRRARELQQAAGRVWSPEGSSRRALEVVAGKWK 484
>gb|AAU90060.1| At3g50740 [Arabidopsis thaliana] gi|15146272|gb|AAK83619.1|
AT3g50740/T3A5_120 [Arabidopsis thaliana]
gi|18409172|ref|NP_566938.1|
UDP-glucoronosyl/UDP-glucosyl transferase family protein
[Arabidopsis thaliana]
Length = 487
Score = 305 bits (781), Expect = 2e-81
Identities = 174/472 (36%), Positives = 283/472 (59%), Gaps = 22/472 (4%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLP----SN 61
H+A+ G H++P+++ KRL H F VT F+ L + + + +S P +
Sbjct: 7 HVAMFASPGMGHIIPVIELGKRLAGSH-GFDVTIFV--LETDAASAQSQFLNSPGCDAAL 63
Query: 62 INHTFLPPVNPNDLPQGTTMES-QILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVE 120
++ LP + + L + ++L+ + ++P + ++ + + AL+VD F ++
Sbjct: 64 VDIVGLPTPDISGLVDPSAFFGIKLLVMMRETIPTIRSKIEEMQHKP--TALIVDLFGLD 121
Query: 121 VLNIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDF 180
+ +G E NML+YI+ S A LA + P LD++ E+ +P+ +PGC P+ D
Sbjct: 122 AIPLGGEFNMLTYIFIASNARFLAVALFFPTLDKDMEEEHIIKKQPMVMPGCEPVRFEDT 181
Query: 181 LSIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGG----DNPPVYP 236
L D +SQ Y+ F+PF + + DG++VN++ ++E L ++++ PVYP
Sbjct: 182 LETFLDPNSQLYREFVPFGSVFPTCDGIIVNTWDDMEPKTLKSLQDPKLLGRIAGVPVYP 241
Query: 237 VGPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNT 296
+GP+ S + L+ WL+KQ SVLY+SFGSGG+LS +Q+ ELA GLE+S
Sbjct: 242 IGPLSRPVDPSKTNHPVLD---WLNKQPDESVLYISFGSGGSLSAKQLTELAWGLEMSQQ 298
Query: 297 KFLWVLRAPSSSSSSAGYLSAENDI---DTLQFLPSGFLERTKEKGFVITSWAPQIQILS 353
+F+WV+R P S+ + YLSA + T +LP GF+ RT E+GF+++SWAPQ +IL+
Sbjct: 299 RFVWVVRPPVDGSACSAYLSANSGKIRDGTPDYLPEGFVSRTHERGFMVSSWAPQAEILA 358
Query: 354 PNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRA-SVNENGI 412
+VGGFLTHCGWNS LESVV GVP+I WPLFAEQ MNA LL+E L V +R+ + G+
Sbjct: 359 HQAVGGFLTHCGWNSILESVVGGVPMIAWPLFAEQMMNATLLNEELGVAVRSKKLPSEGV 418
Query: 413 VERVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGS-STKTISQIA 463
+ R E+ +++ +M +EG ++R +K+LKE A+ ++ DG + +++S+IA
Sbjct: 419 ITRAEIEALVRKIMVEEEGAEMRKKIKKLKETAAESLSCDGGVAHESLSRIA 470
>emb|CAB62443.1| UTP-glucose glucosyltransferase-like protein [Arabidopsis thaliana]
gi|4835225|emb|CAB42903.1| UTP-glucose
glucosyltransferase like protein [Arabidopsis thaliana]
gi|7433913|pir||T08395 UTP-glucose
glucosyltransferase-like protein - Arabidopsis thaliana
Length = 478
Score = 301 bits (771), Expect = 3e-80
Identities = 174/467 (37%), Positives = 282/467 (60%), Gaps = 24/467 (5%)
Query: 11 PGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLP----SNINHTF 66
PG+G H++P+++ KRL H F VT F+ L + + + +S P + ++
Sbjct: 5 PGMG--HIIPVIELGKRLAGSH-GFDVTIFV--LETDAASAQSQFLNSPGCDAALVDIVG 59
Query: 67 LPPVNPNDLPQGTTMES-QILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNIG 125
LP + + L + ++L+ + ++P + ++ + + AL+VD F ++ + +G
Sbjct: 60 LPTPDISGLVDPSAFFGIKLLVMMRETIPTIRSKIEEMQHKP--TALIVDLFGLDAIPLG 117
Query: 126 KELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLSIAQ 185
E NML+YI+ S A LA + P LD++ E+ +P+ +PGC P+ D L
Sbjct: 118 GEFNMLTYIFIASNARFLAVALFFPTLDKDMEEEHIIKKQPMVMPGCEPVRFEDTLETFL 177
Query: 186 DRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGG----DNPPVYPVGPII 241
D +SQ Y+ F+PF + + DG++VN++ ++E L ++++ PVYP+GP+
Sbjct: 178 DPNSQLYREFVPFGSVFPTCDGIIVNTWDDMEPKTLKSLQDPKLLGRIAGVPVYPIGPLS 237
Query: 242 ETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWV 301
S + L+ WL+KQ SVLY+SFGSGG+LS +Q+ ELA GLE+S +F+WV
Sbjct: 238 RPVDPSKTNHPVLD---WLNKQPDESVLYISFGSGGSLSAKQLTELAWGLEMSQQRFVWV 294
Query: 302 LRAPSSSSSSAGYLSAENDI---DTLQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVG 358
+R P S+ + YLSA + T +LP GF+ RT E+GF+++SWAPQ +IL+ +VG
Sbjct: 295 VRPPVDGSACSAYLSANSGKIRDGTPDYLPEGFVSRTHERGFMVSSWAPQAEILAHQAVG 354
Query: 359 GFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRA-SVNENGIVERVE 417
GFLTHCGWNS LESVV GVP+I WPLFAEQ MNA LL+E L V +R+ + G++ R E
Sbjct: 355 GFLTHCGWNSILESVVGGVPMIAWPLFAEQMMNATLLNEELGVAVRSKKLPSEGVITRAE 414
Query: 418 VAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGS-STKTISQIA 463
+ +++ +M +EG ++R +K+LKE A+ ++ DG + +++S+IA
Sbjct: 415 IEALVRKIMVEEEGAEMRKKIKKLKETAAESLSCDGGVAHESLSRIA 461
>gb|AAD12210.1| putative flavonol 3-O-glucosyltransferase [Arabidopsis thaliana]
gi|42570280|ref|NP_849978.2|
UDP-glucoronosyl/UDP-glucosyl transferase family protein
[Arabidopsis thaliana] gi|25286819|pir||H84565 probable
flavonol 3-O-glucosyltransferase [imported] -
Arabidopsis thaliana
Length = 470
Score = 301 bits (771), Expect = 3e-80
Identities = 187/471 (39%), Positives = 281/471 (58%), Gaps = 25/471 (5%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPS-NATKSILQTLPSNINH 64
H +V G HL+PIL+ RL + N HVT T GS S T++I I
Sbjct: 5 HALLVASPGLGHLIPILELGNRLSSVL-NIHVTILAVTSGSSSPTETEAIHAAAARTICQ 63
Query: 65 -TFLPPVNPNDLPQ-GTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVL 122
T +P V+ ++L + T+ +++++ + P + +K L K P V ++VD E++
Sbjct: 64 ITEIPSVDVDNLVEPDATIFTKMVVKMRAMKPAVRDAVK-LMKRKPTV-MIVDFLGTELM 121
Query: 123 NIGKELNMLS-YIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFL 181
++ ++ M + Y+Y P+ A LA YLP LD EY DI EP+KIPGC P+ ++ +
Sbjct: 122 SVADDVGMTAKYVYVPTHAWFLAVMVYLPVLDTVVEGEYVDIKEPLKIPGCKPVGPKELM 181
Query: 182 SIAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKEEGGDNP----PVYPV 237
DRS Q YK + + +DGVLVN++ E++ L+A++E+ + PVYP+
Sbjct: 182 ETMLDRSGQQYKECVRAGLEVPMSDGVLVNTWEELQGNTLAALREDEELSRVMKVPVYPI 241
Query: 238 GPIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTK 297
GPI+ T + D N + WLD+Q+ SV++V GSGGTL+ EQ VELALGLELS +
Sbjct: 242 GPIVRTN-QHVDKPNSI--FEWLDEQRERSVVFVCLGSGGTLTFEQTVELALGLELSGQR 298
Query: 298 FLWVLRAPSSSSSSAGYLSA--ENDIDTLQFLPSGFLERTKEKGFVITSWAPQIQILSPN 355
F+WVLR P+S YL A +D LP GFL+RT+ G V+T WAPQ++ILS
Sbjct: 299 FVWVLRRPAS------YLGAISSDDEQVSASLPEGFLDRTRGVGIVVTQWAPQVEILSHR 352
Query: 356 SVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRAS-VNENGIVE 414
S+GGFL+HCGW+S LES+ GVP+I WPL+AEQ MNA LL+E + V +R S + ++
Sbjct: 353 SIGGFLSHCGWSSALESLTKGVPIIAWPLYAEQWMNATLLTEEIGVAVRTSELPSERVIG 412
Query: 415 RVEVAKVIKYLM--EGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIA 463
R EVA +++ +M E +EG+K+R +E++ ++ A +DGSS ++ + A
Sbjct: 413 REEVASLVRKIMAEEDEEGQKIRAKAEEVRVSSERAWSKDGSSYNSLFEWA 463
>emb|CAA54612.1| UTP-glucose glucosyltransferase [Manihot esculenta]
gi|542015|pir||S41951 UTP-glucose glucosyltransferase -
cassava gi|2501494|sp|Q40287|UFO5_MANES Flavonol
3-O-glucosyltransferase 5 (UDP-glucose flavonoid
3-O-glucosyltransferase 5)
Length = 487
Score = 300 bits (769), Expect = 5e-80
Identities = 178/470 (37%), Positives = 270/470 (56%), Gaps = 16/470 (3%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATK-SILQTL--PSNI 62
HI ++ G HL+P+L+ KR+V L NF VT F+ +GS ++A + +L++ P
Sbjct: 11 HIVLLSSPGLGHLIPVLELGKRIVTLC-NFDVTIFM--VGSDTSAAEPQVLRSAMTPKLC 67
Query: 63 NHTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVL 122
LPP N + L L L + + S K P A++VD F E L
Sbjct: 68 EIIQLPPPNISCLIDPEATVCTRLFVLMREIRPAFRAAVSALKFRP-AAIIVDLFGTESL 126
Query: 123 NIGKELNMLSYIYFPSAATTLAWCFYLPKLDEETSCEYRDILEPIKIPGCVPLHGRDFLS 182
+ KEL + Y+Y S A LA Y+P LD+E E+ EP+KIPGC P+ + +
Sbjct: 127 EVAKELGIAKYVYIASNAWFLALTIYVPILDKEVEGEFVLQKEPMKIPGCRPVRTEEVVD 186
Query: 183 IAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIE---MGPLSAMKEEGG-DNPPVYPVG 238
DR++Q Y + + +ADG+L+N++ +E G L +K G PV+P+G
Sbjct: 187 PMLDRTNQQYSEYFRLGIEIPTADGILMNTWEALEPTTFGALRDVKFLGRVAKVPVFPIG 246
Query: 239 PIIETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKF 298
P+ ++G + E L WLD+Q SV+YVSFGSGGTLS EQ++ELA GLE S +F
Sbjct: 247 PL---RRQAGPCGSNCELLDWLDQQPKESVVYVSFGSGGTLSLEQMIELAWGLERSQQRF 303
Query: 299 LWVLRAPSSSSSSAGYLSAENDIDTLQ-FLPSGFLERTKEKGFVITSWAPQIQILSPNSV 357
+WV+R P+ + A + + + D + + P GFL R + G V+ W+PQI I+S SV
Sbjct: 304 IWVVRQPTVKTGDAAFFTQGDGADDMSGYFPEGFLTRIQNVGLVVPQWSPQIHIMSHPSV 363
Query: 358 GGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLR-ASVNENGIVERV 416
G FL+HCGWNS LES+ GVP+I WP++AEQ+MNA LL+E L V +R ++ +V+R
Sbjct: 364 GVFLSHCGWNSVLESITAGVPIIAWPIYAEQRMNATLLTEELGVAVRPKNLPAKEVVKRE 423
Query: 417 EVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKW 466
E+ ++I+ +M +EG ++R ++ELK++ A+ E GSS +S + +W
Sbjct: 424 EIERMIRRIMVDEEGSEIRKRVRELKDSGEKALNEGGSSFNYMSALGNEW 473
>dbj|BAD28257.1| putative Hydroquinone glucosyltransferase [Oryza sativa (japonica
cultivar-group)]
Length = 498
Score = 291 bits (745), Expect = 3e-77
Identities = 175/473 (36%), Positives = 254/473 (52%), Gaps = 25/473 (5%)
Query: 11 PGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNATKSILQTLPSNINHTFLPPV 70
PG G HL+P+ + ++ L H L P +A ++L +LP+ + LP V
Sbjct: 26 PGAG--HLIPLAELARWLADHHGVAPTLVTFADLEHP-DARSAVLSSLPATVATATLPAV 82
Query: 71 NPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLNIGKELNM 130
+DLP +E + + SLP L L+S A L ALV D F L + EL +
Sbjct: 83 PLDDLPADAGLERTLFEVVHRSLPNLRALLRSAAS---LAALVPDIFCAAALPVAAELGV 139
Query: 131 LSYIYFPSAATTLAWCFYLPKL-DEETSCEYRDILEPIKIPGCVPLHGRDFLSIAQDRSS 189
Y++ P++ L+ +L D + E R + +P+++PG V L + +D ++
Sbjct: 140 PGYVFVPTSLAALSLMRRTVELHDGAAAGEQRALPDPLELPGGVSLRNAEVPRGFRDSTT 199
Query: 190 QAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKE--EGGDNPPVYPVGPIIETETKS 247
Y L +L A G L NSF E+E + K+ E G PP YPVGP + + S
Sbjct: 200 PVYGQLLATGRLYRRAAGFLANSFYELEPAAVEEFKKAAERGTFPPAYPVGPFVRS---S 256
Query: 248 GDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLWVLRAPSS 307
D+A CL WLD Q SV++VSFGS GTLS EQ ELA GLE+S +FLWV+R PS
Sbjct: 257 SDEAGESACLEWLDLQPAGSVVFVSFGSAGTLSVEQTRELAAGLEMSGHRFLWVVRMPSF 316
Query: 308 SSSSAGYLSAENDIDT-------LQFLPSGFLERTKEKGFVITSWAPQIQILSPNSVGGF 360
+ S + D D L +LP GFLERT +G + +WAPQ+++LS + F
Sbjct: 317 NGESFAFGKGAGDEDDHRVHDDPLAWLPDGFLERTSGRGLAVAAWAPQVRVLSHPATAAF 376
Query: 361 LTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLRA------SVNENGIVE 414
++HCGWNSTLESV GVP+I WPL AEQ +NAV+L E + V +R V +V
Sbjct: 377 VSHCGWNSTLESVAAGVPMIAWPLHAEQTVNAVVLEESVGVAVRPRSWEEDDVIGGAVVT 436
Query: 415 RVEVAKVIKYLMEGDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIALKWR 467
R E+A +K +MEG++G +R +EL++A +GSS + + ++A KW+
Sbjct: 437 REEIAAAVKEVMEGEKGRGMRRRARELQQAGGRVWSPEGSSRRALEEVAGKWK 489
>dbj|BAD28252.1| putative Hydroquinone glucosyltransferase [Oryza sativa (japonica
cultivar-group)]
Length = 495
Score = 291 bits (745), Expect = 3e-77
Identities = 181/483 (37%), Positives = 266/483 (54%), Gaps = 26/483 (5%)
Query: 6 HIAVVPGVGYSHLVPILQFSKRLVQLHPNFHVTCFIPTLGSPSNA-TKSILQTLP-SNIN 63
H+ +V H++P+ + ++RLV H L + +A + ++L +LP S++
Sbjct: 10 HVVLVASPCAGHVMPMAELARRLVAFHGCAATLVTFSGLAASLDAHSAAVLASLPASSVA 69
Query: 64 HTFLPPVNPNDLPQGTTMESQILLTLTNSLPYLHQGLKSLAKEIPLVALVVDAFSVEVLN 123
LP V +D+P + I + SLP L Q L+S+ + ALV D F VL+
Sbjct: 70 AVTLPEVTLDDVPADANFGTLIFELVRRSLPNLRQFLRSIGGGV--AALVSDFFCGVVLD 127
Query: 124 IGKELNMLSYIYFPSAATTLAWCFYLPKL-DEETSCEYRDILEPIKIPGCVPLHGRDFLS 182
+ EL + Y++ PS +LA+ ++ D EYRD+ +P+++ G V + D
Sbjct: 128 LAVELGVPGYVFVPSNTASLAFMRRFVEVHDGAAPGEYRDLPDPLRLAGDVTIRVADMPD 187
Query: 183 IAQDRSSQAYKHFLPFVKLLSSADGVLVNSFLEIEMGPLSAMKE--EGGDNPPVYPVGPI 240
DRS+ + L V+ ADG LVNSF E+E + K E G PPVYPVGP
Sbjct: 188 GYLDRSNPVFWQLLEEVRRYRRADGFLVNSFAEMESTIVEEFKTAAEQGAFPPVYPVGPF 247
Query: 241 IETETKSGDDANGLECLAWLDKQQPCSVLYVSFGSGGTLSQEQIVELALGLELSNTKFLW 300
+ D+A L CL WLD+Q SV++VSFGS G LS EQ ELA GLE+S FLW
Sbjct: 248 VRP---CSDEAGELACLEWLDRQPAGSVVFVSFGSAGMLSVEQTRELAAGLEMSGHGFLW 304
Query: 301 VLRAPSSSSSSAGYLSAE------------NDIDTLQFLPSGFLERTKEKGFVITSWAPQ 348
V+R PS S + + +D D L +LP GFLERT +G + SWAPQ
Sbjct: 305 VVRMPSHDGESYDFATDHRNDDEEDRDGGGHDDDPLAWLPDGFLERTSGRGLAVASWAPQ 364
Query: 349 IQILSPNSVGGFLTHCGWNSTLESVVHGVPLITWPLFAEQKMNAVLLSEGLKVGLR---A 405
+++LS + F++HCGWNS LESV GVP++ WPL+AEQK+NAV+L+E V LR A
Sbjct: 365 VRVLSHPATAAFVSHCGWNSALESVSAGVPMVPWPLYAEQKVNAVILTEVAGVALRPAAA 424
Query: 406 SVNENGIVERVEVAKVIKYLME-GDEGEKLRNNMKELKEAASNAVKEDGSSTKTISQIAL 464
+G+V R EVA ++ LM+ G++G R +E++ AA+ A G+S + + ++A
Sbjct: 425 RGGVDGVVTREEVAAAVEELMDPGEKGSAARRRAREMQAAAARARSPGGASHRELDEVAG 484
Query: 465 KWR 467
KW+
Sbjct: 485 KWK 487
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.135 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 800,978,826
Number of Sequences: 2540612
Number of extensions: 34187345
Number of successful extensions: 88490
Number of sequences better than 10.0: 1231
Number of HSP's better than 10.0 without gapping: 1154
Number of HSP's successfully gapped in prelim test: 77
Number of HSP's that attempted gapping in prelim test: 85509
Number of HSP's gapped (non-prelim): 1472
length of query: 470
length of database: 863,360,394
effective HSP length: 132
effective length of query: 338
effective length of database: 527,999,610
effective search space: 178463868180
effective search space used: 178463868180
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC124966.3


