

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
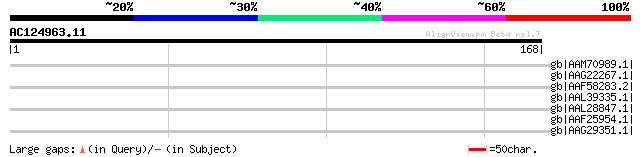
Query= AC124963.11 - phase: 0 /pseudo
(168 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM70989.1| CG8367-PC, isoform C [Drosophila melanogaster] gi... 32 6.8
gb|AAG22267.1| CG8367-PD, isoform D [Drosophila melanogaster] gi... 32 6.8
gb|AAF58283.2| CG8367-PB, isoform B [Drosophila melanogaster] gi... 32 6.8
gb|AAL39335.1| GH24215p [Drosophila melanogaster] 32 6.8
gb|AAL28847.1| LD21089p [Drosophila melanogaster] 32 6.8
gb|AAF25954.1| combgap [Drosophila melanogaster] 32 6.8
gb|AAG29351.1| comb gap splice variant cg14 [Drosophila melanoga... 32 6.8
>gb|AAM70989.1| CG8367-PC, isoform C [Drosophila melanogaster]
gi|24653568|ref|NP_725364.1| CG8367-PC, isoform C
[Drosophila melanogaster]
Length = 629
Score = 32.0 bits (71), Expect = 6.8
Identities = 13/19 (68%), Positives = 17/19 (89%)
Query: 6 GQLQHLESIAKKQQQQQQQ 24
GQ QHL+ +A++QQQQQQQ
Sbjct: 525 GQQQHLQHVAQQQQQQQQQ 543
>gb|AAG22267.1| CG8367-PD, isoform D [Drosophila melanogaster]
gi|21645408|gb|AAM70988.1| CG8367-PA, isoform A
[Drosophila melanogaster] gi|24653564|ref|NP_725363.1|
CG8367-PA, isoform A [Drosophila melanogaster]
gi|24653566|ref|NP_523738.2| CG8367-PD, isoform D
[Drosophila melanogaster] gi|11093540|gb|AAG29350.1|
comb gap [Drosophila melanogaster]
gi|11093543|gb|AAG29352.1| comb gap [Drosophila
melanogaster]
Length = 671
Score = 32.0 bits (71), Expect = 6.8
Identities = 13/19 (68%), Positives = 17/19 (89%)
Query: 6 GQLQHLESIAKKQQQQQQQ 24
GQ QHL+ +A++QQQQQQQ
Sbjct: 567 GQQQHLQHVAQQQQQQQQQ 585
>gb|AAF58283.2| CG8367-PB, isoform B [Drosophila melanogaster]
gi|24653562|ref|NP_725362.1| CG8367-PB, isoform B
[Drosophila melanogaster]
Length = 772
Score = 32.0 bits (71), Expect = 6.8
Identities = 13/19 (68%), Positives = 17/19 (89%)
Query: 6 GQLQHLESIAKKQQQQQQQ 24
GQ QHL+ +A++QQQQQQQ
Sbjct: 668 GQQQHLQHVAQQQQQQQQQ 686
>gb|AAL39335.1| GH24215p [Drosophila melanogaster]
Length = 509
Score = 32.0 bits (71), Expect = 6.8
Identities = 13/19 (68%), Positives = 17/19 (89%)
Query: 6 GQLQHLESIAKKQQQQQQQ 24
GQ QHL+ +A++QQQQQQQ
Sbjct: 405 GQQQHLQHVAQQQQQQQQQ 423
>gb|AAL28847.1| LD21089p [Drosophila melanogaster]
Length = 133
Score = 32.0 bits (71), Expect = 6.8
Identities = 13/19 (68%), Positives = 17/19 (89%)
Query: 6 GQLQHLESIAKKQQQQQQQ 24
GQ QHL+ +A++QQQQQQQ
Sbjct: 29 GQQQHLQHVAQQQQQQQQQ 47
>gb|AAF25954.1| combgap [Drosophila melanogaster]
Length = 650
Score = 32.0 bits (71), Expect = 6.8
Identities = 13/19 (68%), Positives = 17/19 (89%)
Query: 6 GQLQHLESIAKKQQQQQQQ 24
GQ QHL+ +A++QQQQQQQ
Sbjct: 567 GQQQHLQHVAQQQQQQQQQ 585
>gb|AAG29351.1| comb gap splice variant cg14 [Drosophila melanogaster]
Length = 375
Score = 32.0 bits (71), Expect = 6.8
Identities = 13/19 (68%), Positives = 17/19 (89%)
Query: 6 GQLQHLESIAKKQQQQQQQ 24
GQ QHL+ +A++QQQQQQQ
Sbjct: 265 GQQQHLQHVAQQQQQQQQQ 283
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.362 0.162 0.552
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 209,353,984
Number of Sequences: 2540612
Number of extensions: 6310917
Number of successful extensions: 104921
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 104872
Number of HSP's gapped (non-prelim): 21
length of query: 168
length of database: 863,360,394
effective HSP length: 118
effective length of query: 50
effective length of database: 563,568,178
effective search space: 28178408900
effective search space used: 28178408900
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 70 (31.6 bits)
Medicago: description of AC124963.11


