

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124963.1 + phase: 1 /pseudo
(826 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
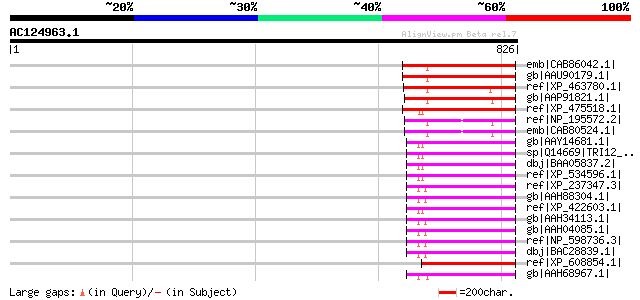
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB86042.1| putative protein [Arabidopsis thaliana] gi|15242... 236 3e-60
gb|AAU90179.1| unknown protein [Oryza sativa (japonica cultivar-... 227 1e-57
ref|XP_463780.1| putative HECT ubiquitin-protein ligase 3 [Oryza... 204 1e-50
gb|AAP91821.1| HECT ubiquitin-protein ligase 3 [Arabidopsis thal... 191 8e-47
ref|XP_475518.1| unknown protein [Oryza sativa (japonica cultiva... 187 9e-46
ref|NP_195572.2| HECT-domain-containing protein / ubiquitin-tran... 181 1e-43
emb|CAB80524.1| putative protein [Arabidopsis thaliana] gi|44671... 178 7e-43
gb|AAY14681.1| unknown [Homo sapiens] 150 1e-34
sp|Q14669|TRI12_HUMAN Thyroid receptor interacting protein 12 (T... 150 1e-34
dbj|BAA05837.2| KIAA0045 [Homo sapiens] 150 1e-34
ref|XP_534596.1| PREDICTED: similar to KIAA0045 [Canis familiaris] 149 3e-34
ref|XP_237347.3| PREDICTED: similar to thyroid hormone receptor ... 149 4e-34
gb|AAH88304.1| Gtl6_predicted protein [Rattus norvegicus] 149 4e-34
ref|XP_422603.1| PREDICTED: similar to Thyroid receptor interact... 147 1e-33
gb|AAH34113.1| Trip12 protein [Mus musculus] 146 2e-33
gb|AAH04085.1| Trip12 protein [Mus musculus] 146 2e-33
ref|NP_598736.3| thyroid hormone receptor interactor 12 [Mus mus... 146 2e-33
dbj|BAC28839.1| unnamed protein product [Mus musculus] 146 2e-33
ref|XP_608854.1| PREDICTED: similar to thyroid hormone receptor ... 144 1e-32
gb|AAH68967.1| MGC83258 protein [Xenopus laevis] 143 2e-32
>emb|CAB86042.1| putative protein [Arabidopsis thaliana] gi|15242560|ref|NP_195908.1|
HECT-domain-containing protein / ubiquitin-transferase
family protein / armadillo/beta-catenin-like
repeat-containing protein [Arabidopsis thaliana]
gi|11357909|pir||T48309 hypothetical protein F9G14.190 -
Arabidopsis thaliana
Length = 1502
Score = 236 bits (601), Expect = 3e-60
Identities = 119/206 (57%), Positives = 149/206 (71%), Gaps = 22/206 (10%)
Query: 641 AFRNSKIEDLCLDFSLPGYDQTFLLPFSIY*ITNLYSF---------------------- 678
+F +KIEDLCL+F+LPGY L P+S + NL +
Sbjct: 1294 SFHGTKIEDLCLEFALPGYTDYDLAPYSANDMVNLDNLEEYIKGIVNATVCNGIQKQVEA 1353
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F GFNQVF IEHLR+F+EEELE +LCGE + ++ N++L H KFDHGYT+ SPP+ LL+
Sbjct: 1354 FRSGFNQVFSIEHLRIFNEEELETMLCGECDLFSMNEVLDHIKFDHGYTSSSPPVEYLLQ 1413
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCANYL 798
IL EF+ E++RAF+QFVT SPRLP GGLASL PKLT+V+K + +DTDLPSVMTCANYL
Sbjct: 1414 ILHEFDREQQRAFLQFVTGSPRLPHGGLASLSPKLTIVRKHGSDSSDTDLPSVMTCANYL 1473
Query: 799 KLPPYSSKERMKQKLLYAITEGRGCF 824
KLPPYSSKE+MK+KL+YAITEG+G F
Sbjct: 1474 KLPPYSSKEKMKEKLIYAITEGQGSF 1499
Score = 38.9 bits (89), Expect = 0.70
Identities = 17/34 (50%), Positives = 28/34 (82%), Gaps = 1/34 (2%)
Query: 1 VLILALQIVELILQKFFSDKFIKLFIEEGVYFAI 34
V+++ALQ+ E++L+K+ D F+ FI+EGV+FAI
Sbjct: 503 VIVVALQVAEVLLEKY-RDTFLNSFIKEGVFFAI 535
>gb|AAU90179.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 523
Score = 227 bits (578), Expect = 1e-57
Identities = 114/206 (55%), Positives = 150/206 (72%), Gaps = 22/206 (10%)
Query: 641 AFRNSKIEDLCLDFSLPGYDQTFL-LPFSIY*IT--NLYSF------------------- 678
++R +IEDL ++F+LPGY + L L S+ ++ NL +
Sbjct: 315 SYRGCRIEDLAIEFALPGYPEYVLSLENSLDNVSADNLEQYVSFVVDATIRSGIARQLEA 374
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F GFN+VFP+ L+VFSE+ELE +LCGE ++W F L+ H KFDHGYT+ SPP++NLLE
Sbjct: 375 FKSGFNEVFPLSMLQVFSEDELERLLCGEQDTWDFAKLVDHIKFDHGYTSSSPPVINLLE 434
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCANYL 798
++QEF +RRAF+QF+T SPRLPPGGLA+L+PKLTVV+K + N D DLPSVMTCANYL
Sbjct: 435 VIQEFEGHQRRAFLQFITGSPRLPPGGLAALNPKLTVVRKHNSNEADDDLPSVMTCANYL 494
Query: 799 KLPPYSSKERMKQKLLYAITEGRGCF 824
KLPPYSSK++M++KLLYAITEG+G F
Sbjct: 495 KLPPYSSKDKMREKLLYAITEGQGSF 520
>ref|XP_463780.1| putative HECT ubiquitin-protein ligase 3 [Oryza sativa (japonica
cultivar-group)] gi|41053228|dbj|BAD08189.1| putative
HECT ubiquitin-protein ligase 3 [Oryza sativa (japonica
cultivar-group)] gi|41052894|dbj|BAD07806.1| putative
HECT ubiquitin-protein ligase 3 [Oryza sativa (japonica
cultivar-group)]
Length = 1781
Score = 204 bits (518), Expect = 1e-50
Identities = 110/217 (50%), Positives = 136/217 (61%), Gaps = 34/217 (15%)
Query: 642 FRNSKIEDLCLDFSLPGYDQTFLLPFSIY*ITNLYSF----------------------F 679
FR + IEDLCLDF+LPGY L I N+Y+ F
Sbjct: 1562 FRGTPIEDLCLDFTLPGYPDYILKEGEENTIVNIYNLEEYVTLVVDATVKSGIMRQVEAF 1621
Query: 680 LVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEI 739
GFNQVF I L++FS EEL+ ++CG W + L+ + KFDHGYTA+SP I+NLLEI
Sbjct: 1622 RSGFNQVFDISSLKIFSPEELDYLICGRREIWEPDSLVDNIKFDHGYTAKSPAIVNLLEI 1681
Query: 740 LQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYN------------HTDTD 787
+ EF E++ AF QFVT +PRLPPGGLA+L+PKLT+V+K + D D
Sbjct: 1682 MAEFTPEQQHAFCQFVTGAPRLPPGGLAALNPKLTIVRKHPSSAVNTSNIAGVTESADDD 1741
Query: 788 LPSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
LPSVMTCANYLKLPPYS+KE M++KLLYAI EGRG F
Sbjct: 1742 LPSVMTCANYLKLPPYSTKEVMRKKLLYAILEGRGSF 1778
>gb|AAP91821.1| HECT ubiquitin-protein ligase 3 [Arabidopsis thaliana]
gi|42627861|tpe|CAE30362.1| TPA: KAKTUS protein
[Arabidopsis thaliana] gi|42570183|ref|NP_849567.2|
HECT-domain-containing protein / ubiquitin-transferase
family protein [Arabidopsis thaliana]
Length = 1888
Score = 191 bits (485), Expect = 8e-47
Identities = 107/214 (50%), Positives = 132/214 (61%), Gaps = 32/214 (14%)
Query: 643 RNSKIEDLCLDFSLPGYDQTFLLPFS-IY*ITNLYSF-------------------FLVG 682
R +IEDL L+F+LPGY + L I ITNL + F G
Sbjct: 1672 RGCRIEDLSLEFTLPGYPEYILRSGDEIVDITNLEEYISLVVDATVKRGVTRQIEAFRSG 1731
Query: 683 FNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQE 742
FNQVF I L++F+ EL+ +LCG W L +H KFDHGY A+SP I+NLLEI+ E
Sbjct: 1732 FNQVFDITSLQIFTPSELDYLLCGRRELWEVETLAEHIKFDHGYNAKSPAIINLLEIMGE 1791
Query: 743 FNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHT------------DTDLPS 790
+++RAF QFVT +PRLPPGGLA L+PKLT+V+K S + D DLPS
Sbjct: 1792 LTADQQRAFCQFVTGAPRLPPGGLAVLNPKLTIVRKHSSTSSAAANGAGASETADDDLPS 1851
Query: 791 VMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
VMTCANYLKLPPYS+KE M +KLLYAI EG+G F
Sbjct: 1852 VMTCANYLKLPPYSTKEIMYKKLLYAINEGQGSF 1885
>ref|XP_475518.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|45642735|gb|AAS72363.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 1321
Score = 187 bits (476), Expect = 9e-46
Identities = 101/206 (49%), Positives = 132/206 (64%), Gaps = 22/206 (10%)
Query: 641 AFRNSKIEDLCLDFSLPGYDQTFLLP----------------FSIY*------ITNLYSF 678
+++N ++EDLCLDF+LPG + L+P SI I+N
Sbjct: 1113 SYKNVRLEDLCLDFTLPGNPEYELVPGGSEKMVTLDNLEEYVSSIVDATLKSGISNQIEA 1172
Query: 679 FLVGFNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLE 738
F G N+VF ++ LR+FSE+E+E ILCGE +SW N L H FD+GY A S +++ LE
Sbjct: 1173 FKAGINKVFALKTLRLFSEDEMERILCGEQDSWASNKLEDHINFDYGYDANSASVISFLE 1232
Query: 739 ILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHTDTDLPSVMTCANYL 798
IL+EF E++RAF+ F T +P+LP GGLASLDPKLTVV+K D +LPSV TC ++
Sbjct: 1233 ILREFGREDQRAFLHFTTGAPQLPLGGLASLDPKLTVVRKQCDGKVDNELPSVNTCRHFF 1292
Query: 799 KLPPYSSKERMKQKLLYAITEGRGCF 824
KLPPYSSKE M+QKL YAI EG G F
Sbjct: 1293 KLPPYSSKEIMRQKLKYAIKEGLGSF 1318
>ref|NP_195572.2| HECT-domain-containing protein / ubiquitin-transferase family protein
[Arabidopsis thaliana]
Length = 1794
Score = 181 bits (458), Expect = 1e-43
Identities = 104/214 (48%), Positives = 129/214 (59%), Gaps = 35/214 (16%)
Query: 643 RNSKIEDLCLDFSLPGYDQTFLLPFS-IY*ITNLYSF-------------------FLVG 682
R +IEDL L+F+LPGY + L I ITNL + F G
Sbjct: 1581 RGCRIEDLSLEFTLPGYPEYILRSGDEIVDITNLEEYISLVVDATVKRGVTRQIEAFRSG 1640
Query: 683 FNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQE 742
FNQVF I L++F+ EL+ +LCG W L +H KFDHGY A+SP I+N I+ E
Sbjct: 1641 FNQVFDITSLQIFTPSELDYLLCGRRELWEVETLAEHIKFDHGYNAKSPAIIN---IMGE 1697
Query: 743 FNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHT------------DTDLPS 790
+++RAF QFVT +PRLPPGGLA L+PKLT+V+K S + D DLPS
Sbjct: 1698 LTADQQRAFCQFVTGAPRLPPGGLAVLNPKLTIVRKHSSTSSAAANGAGASETADDDLPS 1757
Query: 791 VMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
VMTCANYLKLPPYS+KE M +KLLYAI EG+G F
Sbjct: 1758 VMTCANYLKLPPYSTKEIMYKKLLYAINEGQGSF 1791
>emb|CAB80524.1| putative protein [Arabidopsis thaliana] gi|4467147|emb|CAB37516.1|
putative protein [Arabidopsis thaliana]
gi|7485881|pir||T05688 hypothetical protein F20M13.160 -
Arabidopsis thaliana
Length = 757
Score = 178 bits (451), Expect = 7e-43
Identities = 102/214 (47%), Positives = 129/214 (59%), Gaps = 35/214 (16%)
Query: 643 RNSKIEDLCLDFSLPGYDQTFLLPFS-IY*ITNLYSF-------------------FLVG 682
R +IEDL L+F+LPGY + L I ITNL + F G
Sbjct: 544 RGCRIEDLSLEFTLPGYPEYILRSGDEIVDITNLEEYISLVVDATVKRGVTRQIEAFRSG 603
Query: 683 FNQVFPIEHLRVFSEEELELILCGEPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQE 742
FNQVF I L++F+ EL+ +LCG W L +H KFDHGY A+SP I+N +
Sbjct: 604 FNQVFDITSLQIFTPSELDYLLCGRRELWEVETLAEHIKFDHGYNAKSPAIIN---VCYP 660
Query: 743 FNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKISYNHT------------DTDLPS 790
+++++RAF QFVT +PRLPPGGLA L+PKLT+V+K S + D DLPS
Sbjct: 661 SSSDQQRAFCQFVTGAPRLPPGGLAVLNPKLTIVRKHSSTSSAAANGAGASETADDDLPS 720
Query: 791 VMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
VMTCANYLKLPPYS+KE M +KLLYAI EG+G F
Sbjct: 721 VMTCANYLKLPPYSTKEIMYKKLLYAINEGQGSF 754
>gb|AAY14681.1| unknown [Homo sapiens]
Length = 1299
Score = 150 bits (380), Expect = 1e-34
Identities = 89/203 (43%), Positives = 120/203 (58%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ ++ + F GF
Sbjct: 1094 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVSRQFDSFRDGFE 1153
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1154 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1213
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1214 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1273
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1274 DYSSIEIMREKLLIAAREGQQSF 1296
>sp|Q14669|TRI12_HUMAN Thyroid receptor interacting protein 12 (TRIP12)
gi|10863903|ref|NP_004229.1| thyroid hormone receptor
interactor 12 [Homo sapiens]
Length = 1992
Score = 150 bits (380), Expect = 1e-34
Identities = 89/203 (43%), Positives = 120/203 (58%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ ++ + F GF
Sbjct: 1787 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVSRQFDSFRDGFE 1846
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1847 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1906
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1907 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1966
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1967 DYSSIEIMREKLLIAAREGQQSF 1989
>dbj|BAA05837.2| KIAA0045 [Homo sapiens]
Length = 2005
Score = 150 bits (380), Expect = 1e-34
Identities = 89/203 (43%), Positives = 120/203 (58%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ ++ + F GF
Sbjct: 1800 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVSRQFDSFRDGFE 1859
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1860 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1919
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1920 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1979
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M++KLL A EG+ F
Sbjct: 1980 DYSSIEIMREKLLIAAREGQQSF 2002
>ref|XP_534596.1| PREDICTED: similar to KIAA0045 [Canis familiaris]
Length = 2019
Score = 149 bits (377), Expect = 3e-34
Identities = 88/203 (43%), Positives = 120/203 (58%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ ++ + F GF
Sbjct: 1814 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVSRQFDSFRDGFE 1873
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1874 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1933
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1934 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1993
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS + M++KLL A EG+ F
Sbjct: 1994 DYSSIDIMREKLLIAAREGQQSF 2016
>ref|XP_237347.3| PREDICTED: similar to thyroid hormone receptor interactor 12 [Rattus
norvegicus]
Length = 2034
Score = 149 bits (376), Expect = 4e-34
Identities = 89/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + F GF
Sbjct: 1829 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVCRQFDSFRDGFE 1888
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1889 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1948
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1949 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 2008
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M+ KLL A EG+ F
Sbjct: 2009 DYSSIEIMRDKLLIAAREGQQSF 2031
>gb|AAH88304.1| Gtl6_predicted protein [Rattus norvegicus]
Length = 462
Score = 149 bits (376), Expect = 4e-34
Identities = 89/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + F GF
Sbjct: 257 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVCRQFDSFRDGFE 316
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 317 SVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 376
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 377 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 436
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS E M+ KLL A EG+ F
Sbjct: 437 DYSSIEIMRDKLLIAAREGQQSF 459
>ref|XP_422603.1| PREDICTED: similar to Thyroid receptor interacting protein 12
(TRIP12) [Gallus gallus]
Length = 2043
Score = 147 bits (372), Expect = 1e-33
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*----------------ITNLYSFFLVGFN 684
+EDL LDF+LPG+ L P +I+ + + F GF
Sbjct: 1838 VEDLGLDFTLPGFPNIELKKGGKDTPVTIHNLEEYLRLVIFWALNEGVARQFDSFRDGFE 1897
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EELE +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1898 SVFPLSHLQYFYPEELEQLLCGSKTDTWDAKTLMECCRPDHGYTHDSRAVKYLFEILSSF 1957
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
++E++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1958 DSEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 2017
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YS+ E M++KLL A EG+ F
Sbjct: 2018 DYSTIEIMREKLLIAAREGQQSF 2040
>gb|AAH34113.1| Trip12 protein [Mus musculus]
Length = 1103
Score = 146 bits (369), Expect = 2e-33
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + F GF
Sbjct: 898 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVCRQFDSFRDGFE 957
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 958 SVFPLCHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1017
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1018 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 1077
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS + M+ KLL A EG+ F
Sbjct: 1078 DYSSIDIMRDKLLIAAREGQQSF 1100
>gb|AAH04085.1| Trip12 protein [Mus musculus]
Length = 589
Score = 146 bits (369), Expect = 2e-33
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + F GF
Sbjct: 384 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVCRQFDSFRDGFE 443
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 444 SVFPLCHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 503
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 504 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 563
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS + M+ KLL A EG+ F
Sbjct: 564 DYSSIDIMRDKLLIAAREGQQSF 586
>ref|NP_598736.3| thyroid hormone receptor interactor 12 [Mus musculus]
Length = 2026
Score = 146 bits (369), Expect = 2e-33
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + F GF
Sbjct: 1821 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVCRQFDSFRDGFE 1880
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 1881 SVFPLCHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1940
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 1941 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 2000
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS + M+ KLL A EG+ F
Sbjct: 2001 DYSSIDIMRDKLLIAAREGQQSF 2023
>dbj|BAC28839.1| unnamed protein product [Mus musculus]
Length = 357
Score = 146 bits (369), Expect = 2e-33
Identities = 88/203 (43%), Positives = 118/203 (57%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNL----------------YSFFLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + F GF
Sbjct: 152 VEDLGLDFTLPGFPNIELKKGGKDIPVTIHNLEEYLRLVIFWALNEGVCRQFDSFRDGFE 211
Query: 685 QVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S + L EIL F
Sbjct: 212 SVFPLCHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDSRAVKFLFEILSSF 271
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D LPSVMTC NYLKLP
Sbjct: 272 DNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFLPSVMTCVNYLKLP 331
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS + M+ KLL A EG+ F
Sbjct: 332 DYSSIDIMRDKLLIAAREGQQSF 354
>ref|XP_608854.1| PREDICTED: similar to thyroid hormone receptor interactor 12,
partial [Bos taurus]
Length = 309
Score = 144 bits (363), Expect = 1e-32
Identities = 77/156 (49%), Positives = 102/156 (65%), Gaps = 3/156 (1%)
Query: 672 ITNLYSFFLVGFNQVFPIEHLRVFSEEELELILCG-EPNSWTFNDLLKHFKFDHGYTARS 730
++ + F GF VFP+ HL+ F EEL+ +LCG + ++W L++ + DHGYT S
Sbjct: 151 VSRQFDSFRDGFESVFPLSHLQYFYPEELDQLLCGSKADTWDAKTLMECCRPDHGYTHDS 210
Query: 731 PPIMNLLEILQEFNNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDL 788
+ L EIL F+NE++R F+QFVT SPRLP GG SL+P LT+V+K S + D L
Sbjct: 211 RAVKFLFEILSSFDNEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFESTENPDDFL 270
Query: 789 PSVMTCANYLKLPPYSSKERMKQKLLYAITEGRGCF 824
PSVMTC NYLKLP YSS E M++KLL A EG+ F
Sbjct: 271 PSVMTCVNYLKLPDYSSLEIMREKLLMAAREGQQSF 306
>gb|AAH68967.1| MGC83258 protein [Xenopus laevis]
Length = 2027
Score = 143 bits (361), Expect = 2e-32
Identities = 86/203 (42%), Positives = 116/203 (56%), Gaps = 25/203 (12%)
Query: 647 IEDLCLDFSLPGYDQTFL------LPFSIY*ITNLYSF----------------FLVGFN 684
+EDL LDF+LPG+ L +P +I+ + + F GF
Sbjct: 1822 VEDLGLDFTLPGFPNIELKKGGKDVPVTIHNLEDYVRLVIYWALNEGVSRQLDSFRDGFE 1881
Query: 685 QVFPIEHLRVFSEEELELILCGE-PNSWTFNDLLKHFKFDHGYTARSPPIMNLLEILQEF 743
VFP+ HL+ F EEL+ +LCG + W L++ + DHGYT S + L EIL F
Sbjct: 1882 SVFPLNHLQYFYPEELDQLLCGSRADPWDVKTLMECCRPDHGYTHDSRAVKFLFEILSSF 1941
Query: 744 NNEERRAFVQFVTRSPRLPPGGLASLDPKLTVVQKI--SYNHTDTDLPSVMTCANYLKLP 801
+ E++R F+QFVT SPRLP GG SL+P LT+V+K + + D LPSVMTC NYLKLP
Sbjct: 1942 DKEQQRLFLQFVTGSPRLPVGGFRSLNPPLTIVRKTFEATENPDDFLPSVMTCVNYLKLP 2001
Query: 802 PYSSKERMKQKLLYAITEGRGCF 824
YSS + M++KLL A EG+ F
Sbjct: 2002 DYSSIDVMREKLLMAAREGQQSF 2024
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.358 0.160 0.582
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,186,913,500
Number of Sequences: 2540612
Number of extensions: 43630544
Number of successful extensions: 198909
Number of sequences better than 10.0: 703
Number of HSP's better than 10.0 without gapping: 538
Number of HSP's successfully gapped in prelim test: 165
Number of HSP's that attempted gapping in prelim test: 196986
Number of HSP's gapped (non-prelim): 947
length of query: 826
length of database: 863,360,394
effective HSP length: 137
effective length of query: 689
effective length of database: 515,296,550
effective search space: 355039322950
effective search space used: 355039322950
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 80 (35.4 bits)
Medicago: description of AC124963.1


