

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124958.16 + phase: 0
(248 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
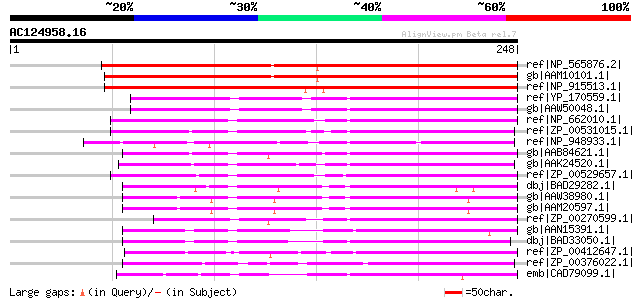
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_565876.2| SOUL heme-binding family protein [Arabidopsis t... 300 2e-80
gb|AAM10101.1| unknown protein [Arabidopsis thaliana] gi|4895187... 299 5e-80
ref|NP_915513.1| P0529H11.19 [Oryza sativa (japonica cultivar-gr... 273 2e-72
ref|YP_170559.1| hypothetical protein FTT1651 [Francisella tular... 133 5e-30
gb|AAW50048.1| hypothetical protein FTT1651 [synthetic construct] 133 5e-30
ref|NP_662010.1| hypothetical protein CT1119 [Chlorobium tepidum... 119 1e-25
ref|ZP_00531015.1| SOUL heme-binding protein [Chlorobium phaeoba... 117 3e-25
ref|NP_948933.1| hypothetical protein RPA3595 [Rhodopseudomonas ... 105 1e-21
gb|AAB84621.1| unknown [Methanothermobacter thermautotrophicus s... 104 2e-21
gb|AAK24520.1| conserved hypothetical protein [Caulobacter cresc... 102 1e-20
ref|ZP_00529657.1| SOUL heme-binding protein [Chlorobium phaeoba... 100 5e-20
dbj|BAD29282.1| SOUL heme-binding protein-like [Oryza sativa (ja... 100 5e-20
gb|AAW38980.1| At3g10130 [Arabidopsis thaliana] gi|6143864|gb|AA... 99 1e-19
gb|AAM20597.1| unknown protein [Arabidopsis thaliana] 99 1e-19
ref|ZP_00270599.1| hypothetical protein Rrub02000642 [Rhodospiri... 93 6e-18
gb|AAN15391.1| unknown protein [Arabidopsis thaliana] gi|1747381... 87 4e-16
dbj|BAD33050.1| SOUL heme-binding protein-like [Oryza sativa (ja... 85 2e-15
ref|ZP_00412647.1| SOUL heme-binding protein [Arthrobacter sp. F... 77 3e-13
ref|ZP_00376022.1| hypothetical protein ELI1263 [Erythrobacter l... 74 5e-12
emb|CAD79099.1| conserved hypothetical protein [Rhodopirellula b... 69 2e-10
>ref|NP_565876.2| SOUL heme-binding family protein [Arabidopsis thaliana]
Length = 225
Score = 300 bits (769), Expect = 2e-80
Identities = 152/217 (70%), Positives = 173/217 (79%), Gaps = 15/217 (6%)
Query: 46 IMGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYI 105
IMGMVFGKI VETPKY VTK+ YEIR Y P+VAAEVTYD S+FKG+KDGGF +LA YI
Sbjct: 10 IMGMVFGKIAVETPKYTVTKSGDGYEIREYPPAVAAEVTYDASEFKGDKDGGFQLLAKYI 69
Query: 106 GALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGE--------------RN 151
G G P+N KPEKIAMTAPVITK EKIAMTAPV+TK SE+ E R
Sbjct: 70 GVFGKPENEKPEKIAMTAPVITK-EGEKIAMTAPVITKESEKIEMTSPVVTKEGGGEGRK 128
Query: 152 KMVTMQFILPSSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRL 211
K+VTMQF+LPS Y+KAEEAP+PTDERVVI+EEG RKYGV+KFSG+AS+ VV EKV+KL
Sbjct: 129 KLVTMQFLLPSMYKKAEEAPRPTDERVVIKEEGGRKYGVIKFSGIASESVVSEKVKKLSS 188
Query: 212 SLERDGFKVIGDFLLGRYNPPWTLPMFRTNEVMIPIE 248
LE+DGFK+ GDF+L RYNPPWTLP FRTNEVMIP+E
Sbjct: 189 HLEKDGFKITGDFVLARYNPPWTLPPFRTNEVMIPVE 225
>gb|AAM10101.1| unknown protein [Arabidopsis thaliana] gi|4895187|gb|AAD32774.1|
unknown protein [Arabidopsis thaliana]
gi|15451064|gb|AAK96803.1| Unknown protein [Arabidopsis
thaliana] gi|13877685|gb|AAK43920.1| Unknown protein
[Arabidopsis thaliana] gi|25360399|pir||D84799
hypothetical protein At2g37970 [imported] - Arabidopsis
thaliana
Length = 215
Score = 299 bits (765), Expect = 5e-80
Identities = 151/216 (69%), Positives = 172/216 (78%), Gaps = 15/216 (6%)
Query: 47 MGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIG 106
MGMVFGKI VETPKY VTK+ YEIR Y P+VAAEVTYD S+FKG+KDGGF +LA YIG
Sbjct: 1 MGMVFGKIAVETPKYTVTKSGDGYEIREYPPAVAAEVTYDASEFKGDKDGGFQLLAKYIG 60
Query: 107 ALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGE--------------RNK 152
G P+N KPEKIAMTAPVITK EKIAMTAPV+TK SE+ E R K
Sbjct: 61 VFGKPENEKPEKIAMTAPVITK-EGEKIAMTAPVITKESEKIEMTSPVVTKEGGGEGRKK 119
Query: 153 MVTMQFILPSSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLS 212
+VTMQF+LPS Y+KAEEAP+PTDERVVI+EEG RKYGV+KFSG+AS+ VV EKV+KL
Sbjct: 120 LVTMQFLLPSMYKKAEEAPRPTDERVVIKEEGGRKYGVIKFSGIASESVVSEKVKKLSSH 179
Query: 213 LERDGFKVIGDFLLGRYNPPWTLPMFRTNEVMIPIE 248
LE+DGFK+ GDF+L RYNPPWTLP FRTNEVMIP+E
Sbjct: 180 LEKDGFKITGDFVLARYNPPWTLPPFRTNEVMIPVE 215
>ref|NP_915513.1| P0529H11.19 [Oryza sativa (japonica cultivar-group)]
gi|20805177|dbj|BAB92846.1| SOUL heme-binding
protein-like [Oryza sativa (japonica cultivar-group)]
Length = 216
Score = 273 bits (699), Expect = 2e-72
Identities = 138/216 (63%), Positives = 163/216 (74%), Gaps = 14/216 (6%)
Query: 47 MGMVFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIG 106
MGMV GKI VETPK+EV T YE+R Y P V AEVTYDP++ KG++DGGF VL NYIG
Sbjct: 1 MGMVLGKITVETPKHEVLHTGAGYEVRKYPPCVVAEVTYDPAEMKGDRDGGFTVLGNYIG 60
Query: 107 ALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTK-------------SSEEGERNK- 152
ALGNPQNTKPEKI MTAPVIT G E IAMTAPV+T ++EE + K
Sbjct: 61 ALGNPQNTKPEKIDMTAPVITSGEPESIAMTAPVITSGEPEPVAMTAPVITAEERSQGKG 120
Query: 153 MVTMQFILPSSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLS 212
+TMQF+LPS Y K EEAP+PTDERVV+R+ GERKYGVV+FSG+ D+VVKEK E L+ +
Sbjct: 121 QMTMQFLLPSKYSKVEEAPRPTDERVVLRQVGERKYGVVRFSGLTGDKVVKEKAEWLKAA 180
Query: 213 LERDGFKVIGDFLLGRYNPPWTLPMFRTNEVMIPIE 248
LE+DGF V G F+L RYNPP+TLP RTNEVM+P+E
Sbjct: 181 LEKDGFTVKGPFVLARYNPPFTLPPLRTNEVMVPVE 216
>ref|YP_170559.1| hypothetical protein FTT1651 [Francisella tularensis subsp.
tularensis Schu 4] gi|56605155|emb|CAG46284.1| conserved
hypothetical protein [Francisella tularensis subsp.
tularensis SCHU S4] gi|54114407|gb|AAV29837.1|
NT02FT0503 [synthetic construct]
Length = 208
Score = 133 bits (334), Expect = 5e-30
Identities = 75/189 (39%), Positives = 106/189 (55%), Gaps = 9/189 (4%)
Query: 60 KYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTKPEKI 119
KY K ++ IRIYAP A+VT S +K + GF L YI N + I
Sbjct: 29 KYTNIKKDDNFSIRIYAPLTEAQVTVQDSDYKSAVNKGFGYLFRYITGA----NIAKQDI 84
Query: 120 AMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTDERVV 179
MTAPV + S++KI MTAPV+ K G+ N T+ F+LP+ Y E APKPT+ +V
Sbjct: 85 QMTAPVKIEQSSQKIQMTAPVMVK----GDTNNEWTIAFVLPAQYT-LENAPKPTNNKVK 139
Query: 180 IREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTLPMFR 239
+ E+ E K V+ FSG + + KL+ ++ + ++++G YNPPWT+P R
Sbjct: 140 LVEKPETKMAVITFSGFLDKDTIDSNTTKLKAWVKANNYQIVGQPEAAGYNPPWTIPFLR 199
Query: 240 TNEVMIPIE 248
TNEVMIPI+
Sbjct: 200 TNEVMIPIK 208
>gb|AAW50048.1| hypothetical protein FTT1651 [synthetic construct]
Length = 243
Score = 133 bits (334), Expect = 5e-30
Identities = 75/189 (39%), Positives = 106/189 (55%), Gaps = 9/189 (4%)
Query: 60 KYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTKPEKI 119
KY K ++ IRIYAP A+VT S +K + GF L YI N + I
Sbjct: 55 KYTNIKKDDNFSIRIYAPLTEAQVTVQDSDYKSAVNKGFGYLFRYITGA----NIAKQDI 110
Query: 120 AMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTDERVV 179
MTAPV + S++KI MTAPV+ K G+ N T+ F+LP+ Y E APKPT+ +V
Sbjct: 111 QMTAPVKIEQSSQKIQMTAPVMVK----GDTNNEWTIAFVLPAQYT-LENAPKPTNNKVK 165
Query: 180 IREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTLPMFR 239
+ E+ E K V+ FSG + + KL+ ++ + ++++G YNPPWT+P R
Sbjct: 166 LVEKPETKMAVITFSGFLDKDTIDSNTTKLKAWVKANNYQIVGQPEAAGYNPPWTIPFLR 225
Query: 240 TNEVMIPIE 248
TNEVMIPI+
Sbjct: 226 TNEVMIPIK 234
>ref|NP_662010.1| hypothetical protein CT1119 [Chlorobium tepidum TLS]
gi|21647087|gb|AAM72352.1| lipoprotein, putative
[Chlorobium tepidum TLS]
Length = 215
Score = 119 bits (297), Expect = 1e-25
Identities = 72/199 (36%), Positives = 105/199 (52%), Gaps = 10/199 (5%)
Query: 50 VFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALG 109
V GK P YE+ K +E+R Y P V AE D + GF LA YI
Sbjct: 18 VLGKREAAEPPYELLKHDGAFEVRRYGPMVIAETILDEKSYSAASGKGFNRLAGYIFG-- 75
Query: 110 NPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
+N I+MTAPV+ + S+EKI+MTAPV+ + + G +M F+LP + +
Sbjct: 76 --KNRSKTSISMTAPVLQERSSEKISMTAPVLQQPQKGGW-----SMAFVLPEGFT-LQS 127
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRY 229
AP+P D V +RE VV FSG+ S +++ +L+ L++ G++ + + L Y
Sbjct: 128 APEPLDPEVKLRELPPSTIAVVTFSGLHSAANLEKYSRQLQAWLKKQGYRALSEPKLASY 187
Query: 230 NPPWTLPMFRTNEVMIPIE 248
+PPWT+P R NEV I IE
Sbjct: 188 DPPWTIPFLRRNEVQIRIE 206
>ref|ZP_00531015.1| SOUL heme-binding protein [Chlorobium phaeobacteroides BS1]
gi|67915333|gb|EAM64656.1| SOUL heme-binding protein
[Chlorobium phaeobacteroides BS1]
Length = 205
Score = 117 bits (293), Expect = 3e-25
Identities = 73/198 (36%), Positives = 104/198 (51%), Gaps = 11/198 (5%)
Query: 50 VFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALG 109
V GK + P Y + K +EIR Y + AE D S ++ GF LA YI
Sbjct: 18 VLGKRTADEPGYSIVKKDGAFEIREYDAMIIAETLLDGS-YRSTSGKGFSKLAKYIFG-- 74
Query: 110 NPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
N EKIAMTAPV+ + EKI+MTAPV+ + + G + KM F++P+ Y +
Sbjct: 75 --SNVGSEKIAMTAPVLQEAEGEKISMTAPVIQEKA--GTKWKMA---FVMPAEYT-LQN 126
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRY 229
PKP D ++IRE RK V++SG+ S++ + KL LE+ G K + Y
Sbjct: 127 LPKPVDPDILIREVPARKVASVRYSGLHSEKNIANWSAKLTEWLEKQGVKAVSVPRSASY 186
Query: 230 NPPWTLPMFRTNEVMIPI 247
+PPWT+P R NE+ I +
Sbjct: 187 DPPWTIPFLRRNEIHIDV 204
>ref|NP_948933.1| hypothetical protein RPA3595 [Rhodopseudomonas palustris CGA009]
gi|39650513|emb|CAE29036.1| conserved hypothetical
protein [Rhodopseudomonas palustris CGA009]
Length = 209
Score = 105 bits (261), Expect = 1e-21
Identities = 77/213 (36%), Positives = 117/213 (54%), Gaps = 15/213 (7%)
Query: 37 HFTLSHSRSIMGMVFGKIGVETPKYEVTKTTQD-YEIRIYAPSVAAEVTYDPSQFKGNKD 95
++T+ + S++G+V ++ E P Y V D EIR YAP VAAEV + +GN D
Sbjct: 7 YYTVLVAESVLGIVGLRL-YEEPAYTVLDRPSDTIEIRRYAPRVAAEVDLER---RGNAD 62
Query: 96 G-GFMVLANYIGALGNPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMV 154
G F +L NYI + E++AMT PV A KIAMTAPV T + + +M
Sbjct: 63 GQAFTLLFNYIAGANRGGSGTSERVAMTVPVDVARPA-KIAMTAPVETATQD-----RMT 116
Query: 155 TMQFILPSSYEKAEEAPKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLE 214
M+F LP+++ A+ APKP+DERV I E+ ++FSG D ++E+ ++L +L
Sbjct: 117 RMRFFLPATFT-ADTAPKPSDERVQIVTVPEQTIATLRFSGTGRD--LREREQQLIAALA 173
Query: 215 RDGFKVIGDFLLGRYNPPWTLPMFRTNEVMIPI 247
++ +G Y+ P+TLP R NE + +
Sbjct: 174 NTPWQPVGAPYGLFYDAPFTLPFVRRNEAAVEV 206
>gb|AAB84621.1| unknown [Methanothermobacter thermautotrophicus str. Delta H]
gi|15678143|ref|NP_275258.1| hypothetical protein MTH115
[Methanothermobacter thermautotrophicus str. Delta H]
gi|7482353|pir||B69020 hypothetical protein MTH115 -
Methanobacterium thermoautotrophicum (strain Delta H)
Length = 189
Score = 104 bits (260), Expect = 2e-21
Identities = 63/195 (32%), Positives = 109/195 (55%), Gaps = 9/195 (4%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
VE+P+Y V +EIR Y + A+V + S F+ GF +LANYI N +
Sbjct: 2 VESPEYTVELKDGKFEIRRYPGYILAQVDVEAS-FRDAMVIGFSILANYIFG----GNRR 56
Query: 116 PEKIAMTAPV--ITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
E++ MT+PV + GS+E+I M PV + ++ + K + F +PSSY E P+P
Sbjct: 57 KEELPMTSPVTGVNLGSSERIPMKVPVTEEVPDDADSGKY-RISFTMPSSYT-LETLPEP 114
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPW 233
D+R+ REE ++++ +FSG + ++ +++ +L+ LER+ + +F++ +YN P
Sbjct: 115 LDDRIRFREEKDQRFAAYRFSGRVNSDMAAQRIAELKEWLERNSIEPRSNFIIAQYNHPA 174
Query: 234 TLPMFRTNEVMIPIE 248
R NEV++ I+
Sbjct: 175 VPGFLRKNEVLVKID 189
>gb|AAK24520.1| conserved hypothetical protein [Caulobacter crescentus CB15]
gi|16126788|ref|NP_421352.1| hypothetical protein CC2549
[Caulobacter crescentus CB15] gi|25360398|pir||D87565
conserved hypothetical protein CC2549 [imported] -
Caulobacter crescentus
Length = 208
Score = 102 bits (254), Expect = 1e-20
Identities = 70/194 (36%), Positives = 100/194 (51%), Gaps = 11/194 (5%)
Query: 54 IGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQN 113
+ VE P ++V D+++R Y V AEVT Q K + GF +LA YI N
Sbjct: 24 MAVEEPVFKVVLHEGDFDVRDYPALVVAEVTVSGDQ-KQAANRGFRLLAGYIFG----GN 78
Query: 114 TKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKP 173
+ IAMTAPV + + IAMTAPV T++ G+ ++F +PS Y E P+P
Sbjct: 79 RTRQSIAMTAPVAQAPAGQTIAMTAPV-TQTQSAGQW----VVRFTMPSRYS-LEALPEP 132
Query: 174 TDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPW 233
D +V +R + V++FSG+A + V+ K L+ L + G L +YN PW
Sbjct: 133 NDPQVKLRLIPPSRLAVLRFSGLAGADTVEVKTADLKKRLSAHQLQATGPATLAQYNTPW 192
Query: 234 TLPMFRTNEVMIPI 247
T R NEVMIP+
Sbjct: 193 TPWFMRRNEVMIPV 206
>ref|ZP_00529657.1| SOUL heme-binding protein [Chlorobium phaeobacteroides DSM 266]
gi|67774427|gb|EAM34113.1| SOUL heme-binding protein
[Chlorobium phaeobacteroides DSM 266]
Length = 198
Score = 100 bits (248), Expect = 5e-20
Identities = 63/199 (31%), Positives = 101/199 (50%), Gaps = 11/199 (5%)
Query: 50 VFGKIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALG 109
V GK + P + + K +E+R Y +V AE D + K F LA YI
Sbjct: 11 VVGKRSADEPSFTLQKKDGVFEVRHYGRTVYAETVVDGAYAK-TSGVAFSRLAGYIFG-- 67
Query: 110 NPQNTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
+N +KI MTAPV+ + + KI MTAPV+ + +G M F++P + E
Sbjct: 68 --KNRAKQKIPMTAPVLQEPVSLKIPMTAPVLQEKKGDGW-----LMSFVMPDG-SRLET 119
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRY 229
P+P D V +RE R V+ ++G+ S++ +++ L+ + + G++ I + Y
Sbjct: 120 LPEPLDPAVKLREAEGRSVAVIGYAGLHSEKNIRKYAGLLKEWIGKKGYRAISEPRAASY 179
Query: 230 NPPWTLPMFRTNEVMIPIE 248
+PPWT+P R NEV I +E
Sbjct: 180 DPPWTIPFLRRNEVQIDVE 198
>dbj|BAD29282.1| SOUL heme-binding protein-like [Oryza sativa (japonica
cultivar-group)] gi|50251400|dbj|BAD28427.1| SOUL
heme-binding protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 287
Score = 100 bits (248), Expect = 5e-20
Identities = 69/202 (34%), Positives = 106/202 (52%), Gaps = 19/202 (9%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQ---FKGNKDGGFMVLANYIGALGNPQ 112
+ET + V K +YEIR AE T F G+ F VLA+Y+ +
Sbjct: 90 LETVPFRVLKREAEYEIREVESYYVAETTMPGRSGFDFNGSSQS-FNVLASYLFG----K 144
Query: 113 NTKPEKIAMTAPVITKGS---AEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
NT E++ MT PV T+ EK+ MT PV+TK S + KM F++PS Y +
Sbjct: 145 NTTSEQMEMTTPVFTRKGEPDGEKMDMTTPVITKKSANENKWKM---SFVMPSKY--GPD 199
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDG-FKVIGDFL--L 226
P P D V I+E + V FSG+ +D+ + ++ +LR +L++D F+V D + +
Sbjct: 200 LPLPKDPSVTIKEVPAKIVAVAAFSGLVTDDDISQRESRLRETLQKDSQFRVKDDSVVEI 259
Query: 227 GRYNPPWTLPMFRTNEVMIPIE 248
+YNPP+TLP R NE+ + ++
Sbjct: 260 AQYNPPFTLPFTRRNEIALEVK 281
>gb|AAW38980.1| At3g10130 [Arabidopsis thaliana] gi|6143864|gb|AAF04411.1| unknown
protein [Arabidopsis thaliana]
gi|15228209|ref|NP_187624.1| SOUL heme-binding family
protein [Arabidopsis thaliana]
Length = 309
Score = 98.6 bits (244), Expect = 1e-19
Identities = 70/202 (34%), Positives = 104/202 (50%), Gaps = 19/202 (9%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGG---FMVLANYIGALGNPQ 112
+ET + V T YEIR P AE T P + + G F VLA Y+ +
Sbjct: 114 LETMNFRVLFRTDKYEIRQVEPYFVAE-TIMPGETGFDSYGASKSFNVLAEYLFG----K 168
Query: 113 NTKPEKIAMTAPVITK---GSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
NT EK+ MT PV+T+ EK+ MT PV+T +++ + +M F++PS Y
Sbjct: 169 NTIKEKMEMTTPVVTRKVQSVGEKMEMTTPVITSKAKDQNQWRM---SFVMPSKY--GSN 223
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGD---FLL 226
P P D V I++ + VV FSG +DE ++ + +LR +L+ D + D F +
Sbjct: 224 LPLPKDPSVKIQQVPRKIVAVVAFSGYVTDEEIERRERELRRALQNDKKFRVRDGVSFEV 283
Query: 227 GRYNPPWTLPMFRTNEVMIPIE 248
+YNPP+TLP R NEV + +E
Sbjct: 284 AQYNPPFTLPFMRRNEVSLEVE 305
>gb|AAM20597.1| unknown protein [Arabidopsis thaliana]
Length = 309
Score = 98.6 bits (244), Expect = 1e-19
Identities = 70/202 (34%), Positives = 104/202 (50%), Gaps = 19/202 (9%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGG---FMVLANYIGALGNPQ 112
+ET + V T YEIR P AE T P + + G F VLA Y+ +
Sbjct: 114 LETMNFRVLFRTDKYEIRQVEPYFVAE-TIMPGETGFDSYGASKSFNVLAEYLFG----K 168
Query: 113 NTKPEKIAMTAPVITK---GSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEE 169
NT EK+ MT PV+T+ EK+ MT PV+T +++ + +M F++PS Y
Sbjct: 169 NTIKEKMEMTTPVVTRKVQSVGEKMEMTTPVITSKAKDQNQWRM---SFVMPSKY--GSN 223
Query: 170 APKPTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGD---FLL 226
P P D V I++ + VV FSG +DE ++ + +LR +L+ D + D F +
Sbjct: 224 LPLPKDPSVKIQQVPRKIVAVVAFSGYVTDEEIERRERELRRALQNDKKFRVRDGVSFEV 283
Query: 227 GRYNPPWTLPMFRTNEVMIPIE 248
+YNPP+TLP R NEV + +E
Sbjct: 284 AQYNPPFTLPFMRRNEVSLEVE 305
>ref|ZP_00270599.1| hypothetical protein Rrub02000642 [Rhodospirillum rubrum]
Length = 179
Score = 93.2 bits (230), Expect = 6e-18
Identities = 63/181 (34%), Positives = 91/181 (49%), Gaps = 16/181 (8%)
Query: 71 EIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTKPEKIAMTAPV---IT 127
EIR Y P VAAEV S G + F +L YI NT + + MT PV
Sbjct: 10 EIRHYGPRVAAEVAARHSGGAGERTHAFRLLFAYITGA----NTARQNLPMTKPVGVGAV 65
Query: 128 KGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTDERVVIREEGERK 187
G+++++AMT PV T + +QF LP+ A+ AP P+D RV +R+ +
Sbjct: 66 GGASQRLAMTIPVATGAG--------AALQFFLPAGLT-AQTAPVPSDPRVTLRDIAAQD 116
Query: 188 YGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTLPMFRTNEVMIPI 247
V+ FSG V + +LR SL G+ G+ + Y+PP++LP R NEV +P+
Sbjct: 117 MAVLGFSGFRHGIEVDRRKAQLRQSLTASGWTASGEAVAYFYDPPFSLPFLRRNEVAVPV 176
Query: 248 E 248
E
Sbjct: 177 E 177
>gb|AAN15391.1| unknown protein [Arabidopsis thaliana] gi|17473811|gb|AAL38336.1|
unknown protein [Arabidopsis thaliana]
gi|22326906|ref|NP_197514.2| SOUL heme-binding family
protein [Arabidopsis thaliana]
Length = 378
Score = 87.0 bits (214), Expect = 4e-16
Identities = 62/194 (31%), Positives = 98/194 (49%), Gaps = 26/194 (13%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
+ETPKY++ K T +YE+R Y P + E D K + GF +A YI +N+
Sbjct: 205 LETPKYQILKRTANYEVRNYEPFIVVETIGD----KLSGSSGFNNVAGYIFG----KNST 256
Query: 116 PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
EKI MT PV T+ + +++ V++Q ++PS + + P P +
Sbjct: 257 MEKIPMTTPVFTQTTDTQLSSD----------------VSVQIVIPSGKDLSS-LPMPNE 299
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPW-T 234
E+V +++ VKFSG +++VV+ K +LR SL +DG + +L RYN P T
Sbjct: 300 EKVNLKKLEGGFAAAVKFSGKPTEDVVQAKENELRSSLSKDGLRAKKGCMLARYNDPGRT 359
Query: 235 LPMFRTNEVMIPIE 248
NEV+I +E
Sbjct: 360 WNFIMRNEVIIWLE 373
>dbj|BAD33050.1| SOUL heme-binding protein-like [Oryza sativa (japonica
cultivar-group)] gi|50725456|dbj|BAD32927.1| SOUL
heme-binding protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 381
Score = 84.7 bits (208), Expect = 2e-15
Identities = 64/190 (33%), Positives = 89/190 (46%), Gaps = 26/190 (13%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
+ETPKY + K T +YEIR Y P + E D K GF + YI +N
Sbjct: 210 IETPKYLILKRTANYEIRSYPPFLIVEAKGD----KLTGSSGFNNVTGYIFG----KNAS 261
Query: 116 PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
E IAMT PV T+ S +K++ V++Q +LP + + + P P
Sbjct: 262 SETIAMTTPVFTQASDDKLSD-----------------VSIQIVLPMNKD-LDSLPAPNT 303
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTL 235
E V +R+ V KFSG +E+V +K ++LR L +D K LL RYN P T
Sbjct: 304 EAVNLRKVEGGIAAVKKFSGRPKEEIVIQKEKELRSQLLKDVLKPQHGCLLARYNDPRTQ 363
Query: 236 PMFRTNEVMI 245
NEV+I
Sbjct: 364 SFIMRNEVLI 373
>ref|ZP_00412647.1| SOUL heme-binding protein [Arthrobacter sp. FB24]
gi|66869123|gb|EAL96491.1| SOUL heme-binding protein
[Arthrobacter sp. FB24]
Length = 201
Score = 77.4 bits (189), Expect = 3e-13
Identities = 63/193 (32%), Positives = 98/193 (50%), Gaps = 10/193 (5%)
Query: 57 ETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTKP 116
E ++V + D+E+R Y AEV + F + F +L YI GN NT
Sbjct: 3 EQQPFDVVQRFPDFEVRRYPGHAVAEVKVK-APFDSAGNAAFRLLFGYIS--GN--NTAR 57
Query: 117 EKIAMTAPVI-TKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
E ++MTAPV+ + + K+AMT PVV +S G+ +V F+LP+S A AP P +
Sbjct: 58 ESVSMTAPVLQSPAPSRKLAMTTPVV-QSGALGDSEFVVA--FVLPASITAAT-APVPNN 113
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTL 235
+V IR V+ FSG ++ +++ L+ +L + G K +G R++PP+
Sbjct: 114 PQVEIRAVPGSVAAVLGFSGRGTEAAFEKRNSVLQEALAQAGLKPVGAPRFARFDPPFKP 173
Query: 236 PMFRTNEVMIPIE 248
R NEV+ IE
Sbjct: 174 WFLRKNEVVQDIE 186
>ref|ZP_00376022.1| hypothetical protein ELI1263 [Erythrobacter litoralis HTCC2594]
gi|60737243|gb|EAL75500.1| hypothetical protein ELI1263
[Erythrobacter litoralis HTCC2594]
Length = 214
Score = 73.6 bits (179), Expect = 5e-12
Identities = 61/192 (31%), Positives = 91/192 (46%), Gaps = 12/192 (6%)
Query: 56 VETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQNTK 115
VE YE + E+R Y P + A+ + + + GF LA+YI A P
Sbjct: 31 VEEQAYERIASDGVIELRQYEPMIIAQTIHAGPRERALA-AGFRRLADYIFAEDRPG--- 86
Query: 116 PEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPKPTD 175
+IAMT+PV+ + AE IAMTAPV+ ++G +F++P Y A P
Sbjct: 87 -AEIAMTSPVL-QDQAEAIAMTAPVM----QDGVGQGAWRTRFVMPRQYTMATLPAAP-- 138
Query: 176 ERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKVIGDFLLGRYNPPWTL 235
+ + ++E R + FSG A E + + LR +E +GF+VIG Y+ P
Sbjct: 139 DYIQLQEVPTRTVAAITFSGRAGSEELGRQERALREWIETNGFEVIGGAEYAFYDAPMVP 198
Query: 236 PMFRTNEVMIPI 247
R NEVMI +
Sbjct: 199 GPLRRNEVMIEV 210
>emb|CAD79099.1| conserved hypothetical protein [Rhodopirellula baltica SH 1]
gi|32476962|ref|NP_869956.1| hypothetical protein
RB11397 [Rhodopirellula baltica SH 1]
Length = 207
Score = 68.6 bits (166), Expect = 2e-10
Identities = 55/201 (27%), Positives = 88/201 (43%), Gaps = 30/201 (14%)
Query: 53 KIGVETPKYEVTKTTQDYEIRIYAPSVAAEVTYDPSQFKGNKDGGFMVLANYIGALGNPQ 112
+ G E+ +Y+V ++ ++E+R Y P + T +G +DG FM L YI
Sbjct: 32 RAGYESAEYKVIESDGNFEVREY-PDLMLVATSTKIDAQG-RDGSFMKLFRYISGA---- 85
Query: 113 NTKPEKIAMTAPVITKGSAEKIAMTAPVVTKSSEEGERNKMVTMQFILPSSYEKAEEAPK 172
N +KI+MT PV E + + V M F++P E P
Sbjct: 86 NESEQKISMTTPVFM------------------ENDKADSEVQMGFVMPKEVA-VEGVPS 126
Query: 173 PTDERVVIREEGERKYGVVKFSGVASDEVVKEKVEKLRLSLERDGFKV-----IGDFLLG 227
PT V +R+ ++ V++FSG + ++ KE KLR +E G
Sbjct: 127 PTGADVDVRKRSGGRFAVLRFSGRLNKKLAKESETKLRTWMESKGLAADDSPEASGVESA 186
Query: 228 RYNPPWTLPMFRTNEVMIPIE 248
Y+PP+T R NEV+I ++
Sbjct: 187 SYDPPFTPGPLRRNEVLIRLK 207
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.315 0.133 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 423,054,171
Number of Sequences: 2540612
Number of extensions: 17783983
Number of successful extensions: 35551
Number of sequences better than 10.0: 114
Number of HSP's better than 10.0 without gapping: 46
Number of HSP's successfully gapped in prelim test: 68
Number of HSP's that attempted gapping in prelim test: 35403
Number of HSP's gapped (non-prelim): 121
length of query: 248
length of database: 863,360,394
effective HSP length: 125
effective length of query: 123
effective length of database: 545,783,894
effective search space: 67131418962
effective search space used: 67131418962
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 73 (32.7 bits)
Medicago: description of AC124958.16


