

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124952.2 - phase: 0
(880 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
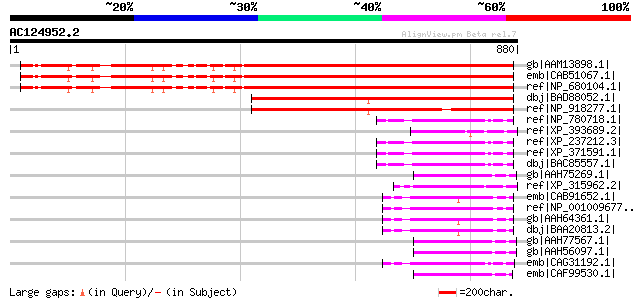
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM13898.1| unknown protein [Arabidopsis thaliana] 681 0.0
emb|CAB51067.1| putative protein [Arabidopsis thaliana] gi|74520... 681 0.0
ref|NP_680104.1| phox (PX) domain-containing protein [Arabidopsi... 681 0.0
dbj|BAD88052.1| phox (PX) domain-containing protein-like [Oryza ... 524 e-147
ref|NP_918277.1| B1156H12.12 [Oryza sativa (japonica cultivar-gr... 489 e-136
ref|NP_780718.1| hypothetical protein LOC241075 [Mus musculus] g... 105 7e-21
ref|XP_393689.2| PREDICTED: similar to CG11534-PA [Apis mellifera] 105 9e-21
ref|XP_237212.3| PREDICTED: similar to RIKEN cDNA 9430067K14 gen... 104 1e-20
ref|XP_371591.1| PREDICTED: hypothetical protein LOC389072 [Homo... 103 2e-20
dbj|BAC85557.1| unnamed protein product [Homo sapiens] 103 2e-20
gb|AAH75269.1| MGC88885 protein [Xenopus tropicalis] gi|52345668... 103 2e-20
ref|XP_315962.2| ENSANGP00000013469 [Anopheles gambiae str. PEST... 103 3e-20
emb|CAB91652.1| unnamed protein product [Homo sapiens] 102 4e-20
ref|NP_001009677.1| pleckstrin homology domain containing, famil... 102 4e-20
gb|AAH64361.1| Pleckstrin homology domain containing, family M (... 102 4e-20
dbj|BAA20813.2| KIAA0356 protein [Homo sapiens] 102 4e-20
gb|AAH77567.1| MGC83535 protein [Xenopus laevis] 102 6e-20
gb|AAH56097.1| Def8-prov protein [Xenopus laevis] 101 9e-20
emb|CAG31192.1| hypothetical protein [Gallus gallus] 101 9e-20
emb|CAF99530.1| unnamed protein product [Tetraodon nigroviridis] 101 9e-20
>gb|AAM13898.1| unknown protein [Arabidopsis thaliana]
Length = 938
Score = 681 bits (1758), Expect = 0.0
Identities = 405/889 (45%), Positives = 542/889 (60%), Gaps = 81/889 (9%)
Query: 20 SDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDD 79
S G S S ES+ ERYCSA NS +GTP S+ S+ F D +F+
Sbjct: 13 SPGSSLHYSSCGESEFERYCSA-NSALGTP----SMCSSTGPFQDSEFENFSLGPSLVKL 67
Query: 80 SNRKSFQYGSSGLELYGDEGD----ELSMTGLDSSELIGNNGIEESDGNENGGEVGEREI 135
S+ + G G+ + + G S GL++ GN I+ +GG E++
Sbjct: 68 SSLDMSRLGDRGIHFFDEGGSCNGRSSSAPGLNT----GNVNIDMCGDLMDGGATIEKDS 123
Query: 136 ETEEEE-----EEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEE 190
+ E+ ++E S+GDDS+ + G S Y SR++ +E
Sbjct: 124 GIDREDGSSIDDDEHSDGDDSLSDSGD------------------HSRNYVSRNLQFQKE 165
Query: 191 GEVRNENPLLMNSSVAFGSHDLDDFLLQNGP-VSVVSDLFYNPRESNNRVEEHGVS---- 245
+ N+NP L+NSS AFG++D D+F L+ V D + E G S
Sbjct: 166 AKDENDNPFLINSSTAFGTNDWDEFELEATELVDTQFDFSGFEKRDKGCTESEGTSTDLF 225
Query: 246 SVRKEEKDSLIVNDEVEETK----------DIGDREA-LEEVRDRDRDRDRDRDRDAPVS 294
SV ++ ++ ++ EE + D+GD A +E++R RD D A V
Sbjct: 226 SVALQKLPDVVQAEKGEEHENVTVSTRHAPDVGDFSANIEDIRSRDFG-----DLSAEVK 280
Query: 295 CEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLD-- 352
V L ++P + + G S+ + N+ S D + LD+
Sbjct: 281 TLVVRQS-------LVTDEPLRGSCL--GNSQTEDRPVVMNNLQSCSAD---DVLDITPT 328
Query: 353 --NFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSFKNL-----KPIVLPSNGGTRKILER 405
E S CD DDVS+ +H + D K +P++ N +
Sbjct: 329 ELGIEDSSGGVCDLDDDVSSG--LLHESSEDGKQSNPFGECTSEPLLASQNSDMPSSRD- 385
Query: 406 SSTSTNVLEKSHVISKIEDSELSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQS 465
S TN + ++ K E++EL++FYD+ V +MEEILL+S +S RFS ++MF Q S
Sbjct: 386 SHPVTNASKVTYTQPKKENTELNDFYDDFVHDMEEILLDSGESSGVRFSKNDKMFQLQLS 445
Query: 466 VPSRDGGLTASTSSTDDACLLVKRPRRIDRIEVVGARQKRGDVSFSERLVGVKEYTVYKI 525
+P+RDGG TA+TS DD+ V + RIDR+EVVG +QK+GDVS SERLVGVKEYTVY I
Sbjct: 446 LPNRDGGQTATTSGLDDSSPTVSQRFRIDRVEVVGVKQKKGDVSLSERLVGVKEYTVYVI 505
Query: 526 KVWSGKDQWEVEKRYRDFLTLHRCMKSLFNEQGWTLPLPWSSVEKESKIFRSASLDIIAK 585
+VWSGKD+WE+E+RYRDF +L+R + SLF +QGWTLP PW+SVE+ES+ S + +A+
Sbjct: 506 RVWSGKDKWEIERRYRDFYSLYRRLTSLFADQGWTLPTPWTSVERESRKIFGTSPNAVAE 565
Query: 586 RSVLIQECLQSILCSRFFSSPPRALVWFLSPEDSHPSSPVSNSPVSLSSFTRGENIRNSS 645
R+VLIQ+CL S+L SRFF + P AL+ FLSP+D++ +S +S VS ++ + SS
Sbjct: 566 RTVLIQDCLNSVLQSRFFPTLPNALLRFLSPQDAYANSSGLDSIVSPTASAAIDAATTSS 625
Query: 646 TWGKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCEY 705
++G TIS IV+I +KS +QLLEAQH+ CAGCHR+FDDG + + DFV+ GWGKPRLCEY
Sbjct: 626 SYGNTISFIVDIRPHKSVKQLLEAQHYICAGCHRYFDDGATLVRDFVKALGWGKPRLCEY 685
Query: 706 TGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLVS 765
TG LFCSSCHTN+ AVLPA VLHHWDF YPVSQLAKSYLDSIHE PMLCV+AVNPFL S
Sbjct: 686 TGHLFCSSCHTNDMAVLPATVLHHWDFNRYPVSQLAKSYLDSIHEQPMLCVSAVNPFLSS 745
Query: 766 KVPALLHVMSVRKKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKGVF 825
KVPAL H+MS+RK+I MLPYVRCPF++++ KG+ +RRYLLES++FFALRDLIDLSKG F
Sbjct: 746 KVPALNHIMSIRKRITIMLPYVRCPFQKTLYKGLSSRRYLLESSEFFALRDLIDLSKGPF 805
Query: 826 AALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
AALP +VETV RKILEHIT+QCLVCCDVG+PC+ARQ C D SSLIFPFQ
Sbjct: 806 AALPAIVETVRRKILEHITEQCLVCCDVGVPCNARQACDDTSSLIFPFQ 854
>emb|CAB51067.1| putative protein [Arabidopsis thaliana] gi|7452052|pir||T13009
hypothetical protein T24C20.80 - Arabidopsis thaliana
Length = 1998
Score = 681 bits (1757), Expect = 0.0
Identities = 405/889 (45%), Positives = 541/889 (60%), Gaps = 81/889 (9%)
Query: 20 SDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDD 79
S G S S ES+ ERYCSA NS +GTP S+ S+ F D +F+
Sbjct: 1073 SPGSSLHYSSCGESEFERYCSA-NSALGTP----SMCSSTGPFQDSEFENFSLGPSLVKL 1127
Query: 80 SNRKSFQYGSSGLELYGDEGD----ELSMTGLDSSELIGNNGIEESDGNENGGEVGEREI 135
S+ + G G+ + + G S GL++ GN I+ +GG E++
Sbjct: 1128 SSLDMSRLGDRGIHFFDEGGSCNGRSSSAPGLNT----GNVNIDMCGDLMDGGATIEKDS 1183
Query: 136 ETEEEE-----EEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEE 190
+ E+ ++E S+GDDS+ + G S Y SR++ +E
Sbjct: 1184 GIDREDGSSIDDDEHSDGDDSLSDSGD------------------HSRNYVSRNLQFQKE 1225
Query: 191 GEVRNENPLLMNSSVAFGSHDLDDFLLQNGP-VSVVSDLFYNPRESNNRVEEHGVS---- 245
+ N+NP L+NSS AFG++D D+F L+ V D + E G S
Sbjct: 1226 AKDENDNPFLINSSTAFGTNDWDEFELEATELVDTQFDFSGFEKRDKGCTESEGTSTDLF 1285
Query: 246 SVRKEEKDSLIVNDEVEETK----------DIGDREA-LEEVRDRDRDRDRDRDRDAPVS 294
SV ++ ++ ++ EE + D+GD A +E++R RD D A V
Sbjct: 1286 SVALQKLPDVVQAEKGEEHENVTVSTRHAPDVGDFSANIEDIRSRDFG-----DLSAEVK 1340
Query: 295 CEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLD-- 352
V L ++P + + G S+ + N+ S D + LD+
Sbjct: 1341 TLVVRQS-------LVTDEPLRGSCL--GNSQTEDRPVVMNNLQSCSAD---DVLDITPT 1388
Query: 353 --NFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSFKNL-----KPIVLPSNGGTRKILER 405
E S CD DDVS+ +H + D K +P++ N +
Sbjct: 1389 ELGIEDSSGGVCDLDDDVSSG--LLHESSEDGKQSNPFGECTSEPLLASQNSDMPSSRD- 1445
Query: 406 SSTSTNVLEKSHVISKIEDSELSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQS 465
S TN + ++ K E++EL++FYD+ V +MEEILL+S +S RFS ++MF Q S
Sbjct: 1446 SHPVTNASKVTYTQPKKENTELNDFYDDFVHDMEEILLDSGESSGVRFSKNDKMFQLQLS 1505
Query: 466 VPSRDGGLTASTSSTDDACLLVKRPRRIDRIEVVGARQKRGDVSFSERLVGVKEYTVYKI 525
+P+RDGG TA+TS DD+ V + RIDR+EVVG +QK+GDVS SERLVGVKEYTVY I
Sbjct: 1506 LPNRDGGQTATTSGLDDSSPTVSQRFRIDRVEVVGVKQKKGDVSLSERLVGVKEYTVYVI 1565
Query: 526 KVWSGKDQWEVEKRYRDFLTLHRCMKSLFNEQGWTLPLPWSSVEKESKIFRSASLDIIAK 585
+VWSGKD+WE+E+RYRDF +L+R + SLF +QGWTLP PW+SVE+ES+ S + +A+
Sbjct: 1566 RVWSGKDKWEIERRYRDFYSLYRRLTSLFADQGWTLPTPWTSVERESRKIFGTSPNAVAE 1625
Query: 586 RSVLIQECLQSILCSRFFSSPPRALVWFLSPEDSHPSSPVSNSPVSLSSFTRGENIRNSS 645
R+VLIQ+CL S+L SRFF + P AL+ FLSP+D++ +S +S VS + + SS
Sbjct: 1626 RTVLIQDCLNSVLQSRFFPTLPNALLRFLSPQDAYANSSGLDSIVSPTGSAAIDAATTSS 1685
Query: 646 TWGKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCEY 705
++G TIS IV+I +KS +QLLEAQH+ CAGCHR+FDDG + + DFV+ GWGKPRLCEY
Sbjct: 1686 SYGNTISFIVDIRPHKSVKQLLEAQHYICAGCHRYFDDGATLVRDFVKALGWGKPRLCEY 1745
Query: 706 TGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLVS 765
TG LFCSSCHTN+ AVLPA VLHHWDF YPVSQLAKSYLDSIHE PMLCV+AVNPFL S
Sbjct: 1746 TGHLFCSSCHTNDMAVLPATVLHHWDFNRYPVSQLAKSYLDSIHEQPMLCVSAVNPFLSS 1805
Query: 766 KVPALLHVMSVRKKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKGVF 825
KVPAL H+MS+RK+I MLPYVRCPF++++ KG+ +RRYLLES++FFALRDLIDLSKG F
Sbjct: 1806 KVPALNHIMSIRKRITIMLPYVRCPFQKTLYKGLSSRRYLLESSEFFALRDLIDLSKGPF 1865
Query: 826 AALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
AALP +VETV RKILEHIT+QCLVCCDVG+PC+ARQ C D SSLIFPFQ
Sbjct: 1866 AALPAIVETVRRKILEHITEQCLVCCDVGVPCNARQACDDTSSLIFPFQ 1914
>ref|NP_680104.1| phox (PX) domain-containing protein [Arabidopsis thaliana]
Length = 938
Score = 681 bits (1757), Expect = 0.0
Identities = 405/889 (45%), Positives = 541/889 (60%), Gaps = 81/889 (9%)
Query: 20 SDGLSRRSSFGAESDSERYCSASNSMMGTPNTSMSIRSAVTVFHDFSDIDFASSSRTFDD 79
S G S S ES+ ERYCSA NS +GTP S+ S+ F D +F+
Sbjct: 13 SPGSSLHYSSCGESEFERYCSA-NSALGTP----SMCSSTGPFQDSEFENFSLGPSLVKL 67
Query: 80 SNRKSFQYGSSGLELYGDEGD----ELSMTGLDSSELIGNNGIEESDGNENGGEVGEREI 135
S+ + G G+ + + G S GL++ GN I+ +GG E++
Sbjct: 68 SSLDMSRLGDRGIHFFDEGGSCNGRSSSAPGLNT----GNVNIDMCGDLMDGGATIEKDS 123
Query: 136 ETEEEE-----EEEFSEGDDSMFNYGSGCDGDNENEFYSMKGENVSSEFYSSRSVSLYEE 190
+ E+ ++E S+GDDS+ + G S Y SR++ +E
Sbjct: 124 GIDREDGSSIDDDEHSDGDDSLSDSGD------------------HSRNYVSRNLQFQKE 165
Query: 191 GEVRNENPLLMNSSVAFGSHDLDDFLLQNGP-VSVVSDLFYNPRESNNRVEEHGVS---- 245
+ N+NP L+NSS AFG++D D+F L+ V D + E G S
Sbjct: 166 AKDENDNPFLINSSTAFGTNDWDEFELEATELVDTQFDFSGFEKRDKGCTESEGTSTDLF 225
Query: 246 SVRKEEKDSLIVNDEVEETK----------DIGDREA-LEEVRDRDRDRDRDRDRDAPVS 294
SV ++ ++ ++ EE + D+GD A +E++R RD D A V
Sbjct: 226 SVALQKLPDVVQAEKGEEHENVTVSTRHAPDVGDFSANIEDIRSRDFG-----DLSAEVK 280
Query: 295 CEVQCADNSPDLDLLPEEDPQKSLNITDGGSEGKGNRYSKNDEAGASGDAQRENLDLD-- 352
V L ++P + + G S+ + N+ S D + LD+
Sbjct: 281 TLVVRQS-------LVTDEPLRGSCL--GNSQTEDRPVVMNNLQSCSAD---DVLDITPT 328
Query: 353 --NFEFKSDQFCDNRDDVSTSNVSVHVENVDAKSFKNL-----KPIVLPSNGGTRKILER 405
E S CD DDVS+ +H + D K +P++ N +
Sbjct: 329 ELGIEDSSGGVCDLDDDVSSG--LLHESSEDGKQSNPFGECTSEPLLASQNSDMPSSRD- 385
Query: 406 SSTSTNVLEKSHVISKIEDSELSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQS 465
S TN + ++ K E++EL++FYD+ V +MEEILL+S +S RFS ++MF Q S
Sbjct: 386 SHPVTNASKVTYTQPKKENTELNDFYDDFVHDMEEILLDSGESSGVRFSKNDKMFQLQLS 445
Query: 466 VPSRDGGLTASTSSTDDACLLVKRPRRIDRIEVVGARQKRGDVSFSERLVGVKEYTVYKI 525
+P+RDGG TA+TS DD+ V + RIDR+EVVG +QK+GDVS SERLVGVKEYTVY I
Sbjct: 446 LPNRDGGQTATTSGLDDSSPTVSQRFRIDRVEVVGVKQKKGDVSLSERLVGVKEYTVYVI 505
Query: 526 KVWSGKDQWEVEKRYRDFLTLHRCMKSLFNEQGWTLPLPWSSVEKESKIFRSASLDIIAK 585
+VWSGKD+WE+E+RYRDF +L+R + SLF +QGWTLP PW+SVE+ES+ S + +A+
Sbjct: 506 RVWSGKDKWEIERRYRDFYSLYRRLTSLFADQGWTLPTPWTSVERESRKIFGTSPNAVAE 565
Query: 586 RSVLIQECLQSILCSRFFSSPPRALVWFLSPEDSHPSSPVSNSPVSLSSFTRGENIRNSS 645
R+VLIQ+CL S+L SRFF + P AL+ FLSP+D++ +S +S VS + + SS
Sbjct: 566 RTVLIQDCLNSVLQSRFFPTLPNALLRFLSPQDAYANSSGLDSIVSPTGSAAIDAATTSS 625
Query: 646 TWGKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCEY 705
++G TIS IV+I +KS +QLLEAQH+ CAGCHR+FDDG + + DFV+ GWGKPRLCEY
Sbjct: 626 SYGNTISFIVDIRPHKSVKQLLEAQHYICAGCHRYFDDGATLVRDFVKALGWGKPRLCEY 685
Query: 706 TGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLVS 765
TG LFCSSCHTN+ AVLPA VLHHWDF YPVSQLAKSYLDSIHE PMLCV+AVNPFL S
Sbjct: 686 TGHLFCSSCHTNDMAVLPATVLHHWDFNRYPVSQLAKSYLDSIHEQPMLCVSAVNPFLSS 745
Query: 766 KVPALLHVMSVRKKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKGVF 825
KVPAL H+MS+RK+I MLPYVRCPF++++ KG+ +RRYLLES++FFALRDLIDLSKG F
Sbjct: 746 KVPALNHIMSIRKRITIMLPYVRCPFQKTLYKGLSSRRYLLESSEFFALRDLIDLSKGPF 805
Query: 826 AALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
AALP +VETV RKILEHIT+QCLVCCDVG+PC+ARQ C D SSLIFPFQ
Sbjct: 806 AALPAIVETVRRKILEHITEQCLVCCDVGVPCNARQACDDTSSLIFPFQ 854
>dbj|BAD88052.1| phox (PX) domain-containing protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 678
Score = 524 bits (1349), Expect = e-147
Identities = 267/470 (56%), Positives = 344/470 (72%), Gaps = 15/470 (3%)
Query: 420 SKIEDSELSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQSVPSRDGGLTASTSS 479
+++++ + + +D++VQEME ILL S + + R + Q+ RDG TASTS
Sbjct: 108 TQVDNLDDNHLFDDMVQEMEHILLNSGEPHESASFTDYRANNSSQAHHFRDGSTTASTSG 167
Query: 480 TDDACL--LVKRPRRIDRIEVVGARQKRGDVSFSERLVGVKEYTVYKIKVWSGKDQWEVE 537
TDDA + L P +ID +EVVGA+Q+ GDVSF ER+VGV+EYTVY +KV SG+D WE+E
Sbjct: 168 TDDAYVYPLPHHPSKIDWVEVVGAKQRTGDVSFGERMVGVREYTVYLLKVKSGEDDWEIE 227
Query: 538 KRYRDFLTLHRCMKSLFNEQGWTLPLPWSSVEKES-KIFRSASLDIIAKRSVLIQECLQS 596
+RYR+F L++ +K LF E+G++LP W +VEKES K+F +AS DI+ +RS LIQ+CL S
Sbjct: 228 RRYREFYALYQQLKLLFAEKGFSLPPAWRNVEKESSKLFGNASPDIVNERSSLIQDCLCS 287
Query: 597 ILCSRFFSSPPRALVWFLSPEDSH----------PSS--PVSNSPVSLSSFTRGENIRNS 644
+L S + P LV FLSP P S +S+ S S G + ++S
Sbjct: 288 LLVSSYPFGTPTPLVSFLSPGSPAYEYSLLKTLIPRSLQRLSSDSHSKGSSCNGTSHKDS 347
Query: 645 STWGKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCE 704
++ GKTISL+VE KSTRQLLE QH+ CAGCHRH D G + + + VQT GW KPR C
Sbjct: 348 ASMGKTISLVVEDRPRKSTRQLLELQHYNCAGCHRHLDAGRTMLQEIVQTIGWNKPRFCA 407
Query: 705 YTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLV 764
YTGQLFC+SCHTN+TAVLPA+VLHHWDF+ YP+SQLAK+YLDSI++ PMLCV+AVNPFL
Sbjct: 408 YTGQLFCASCHTNDTAVLPAKVLHHWDFSLYPISQLAKAYLDSIYDQPMLCVSAVNPFLF 467
Query: 765 SKVPALLHVMSVRKKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKGV 824
+KVPALL++MS+RKKI MLP V+CPFR SI +G+G RRYLL+ NDFFALRDL+DLSKG
Sbjct: 468 AKVPALLNIMSIRKKIAAMLPCVQCPFRNSIFRGLGARRYLLDGNDFFALRDLVDLSKGA 527
Query: 825 FAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
FAALPV V+T+S +IL HIT+QCLVC D G+PC+ARQ C DP +LIFPFQ
Sbjct: 528 FAALPVKVQTISNRILLHITEQCLVCYDSGVPCAARQACDDPLALIFPFQ 577
>ref|NP_918277.1| B1156H12.12 [Oryza sativa (japonica cultivar-group)]
Length = 812
Score = 489 bits (1258), Expect = e-136
Identities = 255/470 (54%), Positives = 331/470 (70%), Gaps = 30/470 (6%)
Query: 420 SKIEDSELSEFYDEVVQEMEEILLESMDSPAARFSVGNRMFDPQQSVPSRDGGLTASTSS 479
+++++ + + +D++VQEME ILL S + + R + Q+ RDG TASTS
Sbjct: 158 TQVDNLDDNHLFDDMVQEMEHILLNSGEPHESASFTDYRANNSSQAHHFRDGSTTASTSG 217
Query: 480 TDDACL--LVKRPRRIDRIEVVGARQKRGDVSFSERLVGVKEYTVYKIKVWSGKDQWEVE 537
TDDA + L P +ID +EVVGA+Q+ GDVSF ER+VGV+EYTVY +KV SG+D WE+E
Sbjct: 218 TDDAYVYPLPHHPSKIDWVEVVGAKQRTGDVSFGERMVGVREYTVYLLKVKSGEDDWEIE 277
Query: 538 KRYRDFLTLHRCMKSLFNEQGWTLPLPWSSVEKES-KIFRSASLDIIAKRSVLIQECLQS 596
+RYR+F L++ +K LF E+G++LP W +VEKES K+F +AS DI+ +RS LIQ+CL S
Sbjct: 278 RRYREFYALYQQLKLLFAEKGFSLPPAWRNVEKESSKLFGNASPDIVNERSSLIQDCLCS 337
Query: 597 ILCSRFFSSPPRALVWFLSPEDSH----------PSS--PVSNSPVSLSSFTRGENIRNS 644
+L S + P LV FLSP P S +S+ S S G + ++S
Sbjct: 338 LLVSSYPFGTPTPLVSFLSPGSPAYEYSLLKTLIPRSLQRLSSDSHSKGSSCNGTSHKDS 397
Query: 645 STWGKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCE 704
++ GKTISL+VE KSTRQLLE QH+ CAGCHRH D G + + + VQT GW KPR C
Sbjct: 398 ASMGKTISLVVEDRPRKSTRQLLELQHYNCAGCHRHLDAGRTMLQEIVQTIGWNKPRFCA 457
Query: 705 YTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLV 764
YTGQLFC+SCHTN+TAVLPA+VLHHWDF+ YP+SQLAK+YLDSI++
Sbjct: 458 YTGQLFCASCHTNDTAVLPAKVLHHWDFSLYPISQLAKAYLDSIYD-------------- 503
Query: 765 SKVPALLHVMSVRKKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKGV 824
+VPALL++MS+RKKI MLP V+CPFR SI +G+G RRYLL+ NDFFALRDL+DLSKG
Sbjct: 504 -QVPALLNIMSIRKKIAAMLPCVQCPFRNSIFRGLGARRYLLDGNDFFALRDLVDLSKGA 562
Query: 825 FAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
FAALPV V+T+S +IL HIT+QCLVC D G+PC+ARQ C DP +LIFPFQ
Sbjct: 563 FAALPVKVQTISNRILLHITEQCLVCYDSGVPCAARQACDDPLALIFPFQ 612
>ref|NP_780718.1| hypothetical protein LOC241075 [Mus musculus]
gi|38649074|gb|AAH63098.1| RIKEN cDNA 9430067K14 gene
[Mus musculus] gi|26330330|dbj|BAC28895.1| unnamed
protein product [Mus musculus]
Length = 761
Score = 105 bits (262), Expect = 7e-21
Identities = 64/240 (26%), Positives = 116/240 (47%), Gaps = 24/240 (10%)
Query: 638 GENIRNSSTWGKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGW 697
G +R + T S + I + S + L AQ CAGC R N
Sbjct: 479 GRELRKNKRQSVTTSFL-SILTTLSLERGLTAQSFKCAGCQRSIGLSN------------ 525
Query: 698 GKPRLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVT 757
GK ++C Y+G +CSSCH +++ ++PAR++H+WD + Y VS+ AK +L+ ++E P++ +
Sbjct: 526 GKAKVCNYSGWYYCSSCHVDDSFLIPARIVHNWDTSKYKVSKQAKEFLEYVYEEPLIDIQ 585
Query: 758 AVNPFLVSKVPALLHVMSVRKKIGTMLPYV---RCPFRRSINKGVGNRRYLLESNDFFAL 814
NP L L V+ +R+++ ++ Y+ R + + + R YLL+ ++L
Sbjct: 586 QENPMLYLHAEPLATVVRLRQRLKSLRAYLFSCRAAVAEDLRRRIFPREYLLQQIHLYSL 645
Query: 815 RDLIDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
DL + +G A P + + + K C +C G C + + +++PF+
Sbjct: 646 ADLQQVIEGKLA--PFLGKVI--KFATAHVYSCSLCSQKGFIC----EICNNGEILYPFE 697
>ref|XP_393689.2| PREDICTED: similar to CG11534-PA [Apis mellifera]
Length = 483
Score = 105 bits (261), Expect = 9e-21
Identities = 63/189 (33%), Positives = 102/189 (53%), Gaps = 14/189 (7%)
Query: 697 WGKPRLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCV 756
W +PRLC+YTG +C CH N V+PARV+ +WD PVS++A L + E +L +
Sbjct: 246 WIEPRLCDYTGLYYCQRCHWNTAMVIPARVIRNWDMESRPVSRVAAQLLTLLEERSVLLL 305
Query: 757 TAVNPFLVSKVPALLHVMSVRKKIGTMLPY-VRCPFRRSINKG----VGNRRYLLESNDF 811
+NP L + VP L V +R+++ M Y V CP + N+G +G R +++E++
Sbjct: 306 EDLNPKLFTLVPDLSVVKRMREEMQMMKRYLVLCP--EANNQGLPWKIGIRTHMIENSGN 363
Query: 812 FALRDLIDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIF 871
++++DLIDL+ G+ L + + HIT+QC +C G C C + +I+
Sbjct: 364 YSIKDLIDLNNGI---LLEEIRAAYDTMHTHITEQCELCKARGHLCEL---CGN-DEIIY 416
Query: 872 PFQVCICQC 880
P+ C
Sbjct: 417 PWDASSISC 425
>ref|XP_237212.3| PREDICTED: similar to RIKEN cDNA 9430067K14 gene [Rattus
norvegicus]
Length = 761
Score = 104 bits (259), Expect = 1e-20
Identities = 64/240 (26%), Positives = 115/240 (47%), Gaps = 24/240 (10%)
Query: 638 GENIRNSSTWGKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGW 697
G +R + T S + I + S + L AQ CAGC R N
Sbjct: 479 GHELRKNKRQSVTTSFL-SILTTLSLERGLTAQSFKCAGCQRPIGLSN------------ 525
Query: 698 GKPRLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVT 757
GK ++C Y+G +CSSCH +++ ++PAR++H+WD + Y VS+ AK +L+ ++E P++ +
Sbjct: 526 GKAKVCNYSGWYYCSSCHVDDSFLIPARIVHNWDTSKYKVSKQAKEFLEYVYEEPLIDIQ 585
Query: 758 AVNPFLVSKVPALLHVMSVRKKIGTMLPYV---RCPFRRSINKGVGNRRYLLESNDFFAL 814
NP L L V +R+++ ++ Y+ R + + + R YLL+ ++L
Sbjct: 586 QENPMLYLHAEPLATVARLRQRLKSLRAYLFSCRAAVAEDLRRRIFPREYLLQQIHLYSL 645
Query: 815 RDLIDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
DL + +G A P + + + K C +C G C + + +++PF+
Sbjct: 646 ADLQQVIEGKLA--PFLGKVI--KFATAHVYSCSLCSQKGFIC----EICNNGEILYPFE 697
>ref|XP_371591.1| PREDICTED: hypothetical protein LOC389072 [Homo sapiens]
Length = 717
Score = 103 bits (257), Expect = 2e-20
Identities = 63/240 (26%), Positives = 115/240 (47%), Gaps = 24/240 (10%)
Query: 638 GENIRNSSTWGKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGW 697
G +R + T S + I + S + L AQ CAGC R N
Sbjct: 479 GHELRKNKRQSVTTSFL-SILTTLSLERGLTAQSFKCAGCQRSIGLSN------------ 525
Query: 698 GKPRLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVT 757
GK ++C Y+G +CSSCH +++ ++PAR++H+WD + Y VS+ AK +L+ ++E P++ +
Sbjct: 526 GKAKVCNYSGWYYCSSCHVDDSFLIPARIVHNWDTSKYKVSKQAKEFLEYVYEEPLIDIQ 585
Query: 758 AVNPFLVSKVPALLHVMSVRKKIGTMLPYV---RCPFRRSINKGVGNRRYLLESNDFFAL 814
N L L V+ +R+++ ++ Y+ R + + + R YLL+ ++L
Sbjct: 586 QENAMLYHHAEPLAAVLRLRQRLKSLRAYLFSCRAAVAEDLRRRIFPREYLLQQIHLYSL 645
Query: 815 RDLIDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
DL + +G A P + + + K C +C G C + + +++PF+
Sbjct: 646 ADLQQVIEGKLA--PFLGKVI--KFATSHVYSCSLCSQKGFIC----EICNNGEILYPFE 697
>dbj|BAC85557.1| unnamed protein product [Homo sapiens]
Length = 443
Score = 103 bits (257), Expect = 2e-20
Identities = 63/240 (26%), Positives = 115/240 (47%), Gaps = 24/240 (10%)
Query: 638 GENIRNSSTWGKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGW 697
G +R + T S + I + S + L AQ CAGC R N
Sbjct: 205 GHELRKNKRQSVTTSFL-SILTTLSLERGLTAQSFKCAGCQRSIGLSN------------ 251
Query: 698 GKPRLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVT 757
GK ++C Y+G +CSSCH +++ ++PAR++H+WD + Y VS+ AK +L+ ++E P++ +
Sbjct: 252 GKAKVCNYSGWYYCSSCHVDDSFLIPARIVHNWDTSKYKVSKQAKEFLEYVYEEPLIDIQ 311
Query: 758 AVNPFLVSKVPALLHVMSVRKKIGTMLPYV---RCPFRRSINKGVGNRRYLLESNDFFAL 814
N L L V+ +R+++ ++ Y+ R + + + R YLL+ ++L
Sbjct: 312 QENAMLYHHAEPLAAVLRLRQRLKSLRAYLFSCRAAVAEDLRRRIFPREYLLQQIHLYSL 371
Query: 815 RDLIDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
DL + +G A P + + + K C +C G C + + +++PF+
Sbjct: 372 ADLQQVIEGKLA--PFLGKVI--KFATSHVYSCSLCSQKGFIC----EICNNGEILYPFE 423
>gb|AAH75269.1| MGC88885 protein [Xenopus tropicalis]
gi|52345668|ref|NP_001004881.1| MGC88885 protein
[Xenopus tropicalis]
Length = 443
Score = 103 bits (257), Expect = 2e-20
Identities = 59/184 (32%), Positives = 97/184 (52%), Gaps = 12/184 (6%)
Query: 701 RLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVN 760
R C+YTGQ +C SCH N+ AV+PAR +H+WDF + VS+ + YL + P+L + +N
Sbjct: 230 RQCDYTGQYYCISCHWNDIAVIPARAIHNWDFEPHKVSRCSMRYLALMLGRPVLKLREIN 289
Query: 761 PFLVSKVPALLHVMSVRKKIGTMLPY-VRC--PFRRSINKGVGNRRYLLESNDFFALRDL 817
P L + V L+ + +R+ I M PY + C + + +R++ +E++D ++L+DL
Sbjct: 290 PLLFNYVEELVEIRKLRQDILLMKPYFITCKEAMEARLLLQLQDRQHFVENDDMYSLQDL 349
Query: 818 IDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPF--QV 875
+D+S G + T K HI C C G C ++ ++FPF
Sbjct: 350 LDISSGRLGCSLTEIHTTFAK---HIKLDCERCQAKGFVCELCKE----GDILFPFDSHT 402
Query: 876 CICQ 879
+CQ
Sbjct: 403 SVCQ 406
>ref|XP_315962.2| ENSANGP00000013469 [Anopheles gambiae str. PEST]
gi|55238742|gb|EAA11791.2| ENSANGP00000013469 [Anopheles
gambiae str. PEST]
Length = 393
Score = 103 bits (256), Expect = 3e-20
Identities = 64/221 (28%), Positives = 113/221 (50%), Gaps = 24/221 (10%)
Query: 667 LEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCEYTGQLFCSSCHTNETAVLPARV 726
L Q +TCA C G + + + +PRLC+YTG +C +CH N+T+++PAR+
Sbjct: 162 LAFQKYTCAEC------GTQLSYKKLNSI---QPRLCDYTGLYYCPACHWNDTSIIPARI 212
Query: 727 LHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLVSKVPALLHVMSVRKKIGTMLPY 786
++WDF V + ++ ++ + P++ + NP L + +P L V R ++G M Y
Sbjct: 213 TNNWDFVPRKVCRASRQQINLLLAKPVIRLEEKNPRLFTFIPQLAEVKRARVQLGEMKRY 272
Query: 787 -VRCPF---RRSINKGVGNRRYLLESNDFFALRDLIDLSKGVFAALPVMVETVSRKILEH 842
+ C R + K +G RR+L+ES + +++ DL+ + G L + T+ R EH
Sbjct: 273 LIACRLADETRLVVKQIGKRRHLMESIELYSVADLVGVEDG---TLLNYLRTI-RATFEH 328
Query: 843 ITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ---VCICQC 880
C++C C + C++ ++FPF +C QC
Sbjct: 329 HIRNCVICSGKAYIC---EFCNN-DQILFPFDDDAICCTQC 365
>emb|CAB91652.1| unnamed protein product [Homo sapiens]
Length = 926
Score = 102 bits (255), Expect = 4e-20
Identities = 67/232 (28%), Positives = 111/232 (46%), Gaps = 31/232 (13%)
Query: 648 GKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCEYTG 707
G + +V IP K L++Q CAGC R F + +P+LC ++G
Sbjct: 679 GFLLQYLVAIPMEKG----LDSQGCFCAGCSRQIG------------FSFVRPKLCAFSG 722
Query: 708 QLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLVSKV 767
+C CH ++ +V+PAR++H+WD T P+ + A +L I P++ + VN L V
Sbjct: 723 LYYCDICHQDDASVIPARIIHNWDLTKRPICRQALKFLTQIRAQPLINLQMVNASLYEHV 782
Query: 768 PALLHVMSVR----KKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKG 823
+H++ R K +G L R + ++K + +R YLLES F++ DL ++ G
Sbjct: 783 ER-MHLIGRRREQLKLLGDYLGLCRSGALKELSKRLNHRNYLLESPHRFSVADLQQIADG 841
Query: 824 VFAA-LPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
V+ L ++E S+ + C +C G C Q +IFPF+
Sbjct: 842 VYEGFLKALIEFASQHVY-----HCDLCTQRGFICQICQH----HDIIFPFE 884
>ref|NP_001009677.1| pleckstrin homology domain containing, family M (with RUN domain)
member 1 [Rattus norvegicus] gi|56270126|gb|AAH87058.1|
Pleckstrin homology domain containing, family M (with RUN
domain) member 1 (predicted) [Rattus norvegicus]
Length = 1059
Score = 102 bits (255), Expect = 4e-20
Identities = 68/231 (29%), Positives = 112/231 (48%), Gaps = 29/231 (12%)
Query: 648 GKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCEYTG 707
G + +V IP+ K L++Q CAGC R F + +P+LC ++G
Sbjct: 812 GFLLQYLVAIPTEKG----LDSQGCFCAGCSRQIG------------FSFVRPKLCAFSG 855
Query: 708 QLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLVSKV 767
+C CH ++ +V+PAR++H+WD T PV + A +L I P++ + VN L V
Sbjct: 856 LYYCDFCHQDDASVIPARIIHNWDLTKRPVCRQALKFLAQIRAQPLINLQLVNASLYEHV 915
Query: 768 PALLHVMSVR---KKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKGV 824
+ + R K +G L R + ++K + +R YLLES F++ DL +++GV
Sbjct: 916 ERMHLIGRSREQLKLLGDYLGLCRSGALKELSKRLSHRNYLLESPHKFSVADLQQIAEGV 975
Query: 825 FAA-LPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
+ L ++E S+ + C +C G C Q C +IFPF+
Sbjct: 976 YEGFLKALIEFASQHVY-----HCDLCTQRGFIC---QICHH-QDIIFPFE 1017
>gb|AAH64361.1| Pleckstrin homology domain containing, family M (with RUN domain)
member 1 [Homo sapiens] gi|40538728|ref|NP_055613.1|
pleckstrin homology domain containing, family M (with RUN
domain) member 1 [Homo sapiens]
Length = 1056
Score = 102 bits (255), Expect = 4e-20
Identities = 67/232 (28%), Positives = 111/232 (46%), Gaps = 31/232 (13%)
Query: 648 GKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCEYTG 707
G + +V IP K L++Q CAGC R F + +P+LC ++G
Sbjct: 809 GFLLQYLVAIPMEKG----LDSQGCFCAGCSRQIG------------FSFVRPKLCAFSG 852
Query: 708 QLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLVSKV 767
+C CH ++ +V+PAR++H+WD T P+ + A +L I P++ + VN L V
Sbjct: 853 LYYCDICHQDDASVIPARIIHNWDLTKRPICRQALKFLTQIRAQPLINLQMVNASLYEHV 912
Query: 768 PALLHVMSVR----KKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKG 823
+H++ R K +G L R + ++K + +R YLLES F++ DL ++ G
Sbjct: 913 ER-MHLIGRRREQLKLLGDYLGLCRSGALKELSKRLNHRNYLLESPHRFSVADLQQIADG 971
Query: 824 VFAA-LPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
V+ L ++E S+ + C +C G C Q +IFPF+
Sbjct: 972 VYEGFLKALIEFASQHVY-----HCDLCTQRGFICQICQH----HDIIFPFE 1014
>dbj|BAA20813.2| KIAA0356 protein [Homo sapiens]
Length = 1058
Score = 102 bits (255), Expect = 4e-20
Identities = 67/232 (28%), Positives = 111/232 (46%), Gaps = 31/232 (13%)
Query: 648 GKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCEYTG 707
G + +V IP K L++Q CAGC R F + +P+LC ++G
Sbjct: 811 GFLLQYLVAIPMEKG----LDSQGCFCAGCSRQIG------------FSFVRPKLCAFSG 854
Query: 708 QLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLVSKV 767
+C CH ++ +V+PAR++H+WD T P+ + A +L I P++ + VN L V
Sbjct: 855 LYYCDICHQDDASVIPARIIHNWDLTKRPICRQALKFLTQIRAQPLINLQMVNASLYEHV 914
Query: 768 PALLHVMSVR----KKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKG 823
+H++ R K +G L R + ++K + +R YLLES F++ DL ++ G
Sbjct: 915 ER-MHLIGRRREQLKLLGDYLGLCRSGALKELSKRLNHRNYLLESPHRFSVADLQQIADG 973
Query: 824 VFAA-LPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQ 874
V+ L ++E S+ + C +C G C Q +IFPF+
Sbjct: 974 VYEGFLKALIEFASQHVY-----HCDLCTQRGFICQICQH----HDIIFPFE 1016
>gb|AAH77567.1| MGC83535 protein [Xenopus laevis]
Length = 443
Score = 102 bits (254), Expect = 6e-20
Identities = 59/184 (32%), Positives = 96/184 (52%), Gaps = 12/184 (6%)
Query: 701 RLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVN 760
R C+YTGQ +C SCH N+ AV+PAR +H+WDF VS+ + YL + P+L + +N
Sbjct: 230 RQCDYTGQYYCISCHWNDLAVIPARAIHNWDFEPRKVSRCSMRYLALMLGRPVLKLREIN 289
Query: 761 PFLVSKVPALLHVMSVRKKIGTMLPY-VRC--PFRRSINKGVGNRRYLLESNDFFALRDL 817
P L + V L+ + +R+ I M PY + C + + +R++ +E++D ++L+DL
Sbjct: 290 PLLFNYVEELVEIRKLRQDILLMKPYFITCKEAMEARLLLQLQDRQHFVENDDMYSLQDL 349
Query: 818 IDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPF--QV 875
+D+S G + T K HI C C G C ++ ++FPF
Sbjct: 350 LDISSGRLGCTLTEIHTTFAK---HIKLDCERCQAKGFVCELCKE----GDILFPFDSHT 402
Query: 876 CICQ 879
+CQ
Sbjct: 403 SVCQ 406
>gb|AAH56097.1| Def8-prov protein [Xenopus laevis]
Length = 443
Score = 101 bits (252), Expect = 9e-20
Identities = 59/184 (32%), Positives = 96/184 (52%), Gaps = 12/184 (6%)
Query: 701 RLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVN 760
R C+YTGQ +C SCH N+ AV+PAR +H+WDF VS+ + YL + P+L + +N
Sbjct: 230 RQCDYTGQYYCISCHWNDLAVIPARAIHNWDFEPCKVSRYSMRYLALMLGRPVLKLREIN 289
Query: 761 PFLVSKVPALLHVMSVRKKIGTMLPY-VRC--PFRRSINKGVGNRRYLLESNDFFALRDL 817
P L + V L+ + +R+ I M PY + C + + +R++ +E++D ++L+DL
Sbjct: 290 PLLFNYVEELVEIRKLRQDILLMKPYFITCKEAMEDRLLLQLQDRQHFVENDDMYSLQDL 349
Query: 818 IDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPF--QV 875
+D+S G + T K HI C C G C ++ ++FPF
Sbjct: 350 LDISSGRLGCSLTEIHTTFAK---HIKLDCERCQAKGFMCELCKE----GDILFPFDSHT 402
Query: 876 CICQ 879
+CQ
Sbjct: 403 SVCQ 406
>emb|CAG31192.1| hypothetical protein [Gallus gallus]
Length = 1005
Score = 101 bits (252), Expect = 9e-20
Identities = 68/231 (29%), Positives = 107/231 (45%), Gaps = 27/231 (11%)
Query: 648 GKTISLIVEIPSNKSTRQLLEAQHHTCAGCHRHFDDGNSSIWDFVQTFGWGKPRLCEYTG 707
G + +V IP K L++Q CAGC R F + KP+LC Y+G
Sbjct: 744 GFLLQYLVAIPVEKG----LDSQSFICAGCSRQIG------------FSFAKPKLCAYSG 787
Query: 708 QLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVNPFLVSKV 767
+C SCH ++ V+P+R++H+WD + V + A +L I + P++ + VN L V
Sbjct: 788 LYYCDSCHRDDETVIPSRLIHNWDLSKRGVCRQALKFLTQICKQPLIDLKLVNESLYDHV 847
Query: 768 PALLHVMSVR---KKIGTMLPYVRCPFRRSINKGVGNRRYLLESNDFFALRDLIDLSKGV 824
+ + R K +G L R + ++K + +R YLLE +++ DL +S G
Sbjct: 848 ERMRRIRRSREQLKLLGDYLSVCRSGALKELSKRLDHRNYLLECPHKYSVADLQQISDGA 907
Query: 825 FAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPFQV 875
F P ++ H C +C G C Q C S +IFPFQ+
Sbjct: 908 FE--PFLLSLT--HFASHHVYSCDLCTQRGFIC---QICGG-SDIIFPFQL 950
>emb|CAF99530.1| unnamed protein product [Tetraodon nigroviridis]
Length = 414
Score = 101 bits (252), Expect = 9e-20
Identities = 57/176 (32%), Positives = 96/176 (54%), Gaps = 10/176 (5%)
Query: 701 RLCEYTGQLFCSSCHTNETAVLPARVLHHWDFTHYPVSQLAKSYLDSIHEHPMLCVTAVN 760
R C+YTGQ +CS+CH N+TA++PARV+H+W+F V + + YL + P+L + +N
Sbjct: 200 RQCDYTGQYYCSTCHWNDTAIVPARVIHNWEFEPRKVCRSSLRYLALMMSRPVLKLKEIN 259
Query: 761 PFLVSKVPALLHVMSVRKKIGTMLPY-VRC--PFRRSINKGVGNRRYLLESNDFFALRDL 817
P L + V L+ + +R+ I M PY + C + + +R++ +E++D ++L+DL
Sbjct: 260 PLLFNFVEELVEIRKLRQDILLMKPYFITCKEAMEARLLLQLQDRQHFVENDDMYSLQDL 319
Query: 818 IDLSKGVFAALPVMVETVSRKILEHITDQCLVCCDVGIPCSARQDCSDPSSLIFPF 873
ID+S G + + T K HI C C G C ++ ++FPF
Sbjct: 320 IDISCGRLSCSLTEIHTTFAK---HIKLDCERCQAKGFVCELCKE----GDILFPF 368
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.315 0.133 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,562,181,321
Number of Sequences: 2540612
Number of extensions: 71252167
Number of successful extensions: 302568
Number of sequences better than 10.0: 1988
Number of HSP's better than 10.0 without gapping: 342
Number of HSP's successfully gapped in prelim test: 1784
Number of HSP's that attempted gapping in prelim test: 273215
Number of HSP's gapped (non-prelim): 10076
length of query: 880
length of database: 863,360,394
effective HSP length: 137
effective length of query: 743
effective length of database: 515,296,550
effective search space: 382865336650
effective search space used: 382865336650
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 80 (35.4 bits)
Medicago: description of AC124952.2


