

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124952.12 + phase: 2 /pseudo
(313 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
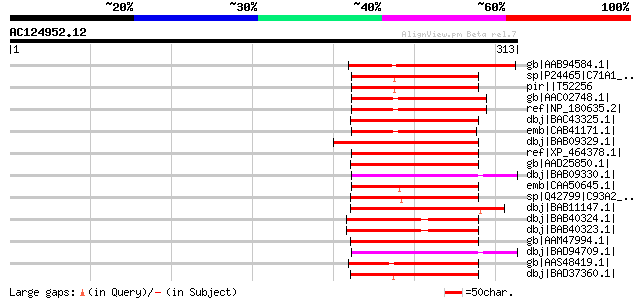
Score E
Sequences producing significant alignments: (bits) Value
gb|AAB94584.1| CYP71A10 [Glycine max] gi|7430655|pir||T05735 cyt... 81 3e-14
sp|P24465|C71A1_PERAE Cytochrome P450 71A1 (CYPLXXIA1) (ARP-2) 75 2e-12
pir||T52256 cytochrome P-450LXXIA1 [similarity] - avocado gi|166... 75 2e-12
gb|AAC02748.1| putative cytochrome P450 [Arabidopsis thaliana] g... 74 5e-12
ref|NP_180635.2| cytochrome P450 71A13, putative (CYP71A13) [Ara... 74 5e-12
dbj|BAC43325.1| putative cytochrome P450 [Arabidopsis thaliana] ... 71 4e-11
emb|CAB41171.1| cytochrome P450-like protein [Arabidopsis thalia... 70 6e-11
dbj|BAB09329.1| cytochrome P450 [Arabidopsis thaliana] gi|152390... 70 6e-11
ref|XP_464378.1| putative cytochrome P450 [Oryza sativa (japonic... 70 8e-11
gb|AAD25850.1| putative cytochrome P450 [Arabidopsis thaliana] g... 70 8e-11
dbj|BAB09330.1| cytochrome P450 [Arabidopsis thaliana] gi|152390... 69 1e-10
emb|CAA50645.1| P450 hydroxylase [Solanum melongena] gi|584861|s... 69 1e-10
sp|Q42799|C93A2_SOYBN Cytochrome P450 93A2 gi|1408322|dbj|BAA130... 69 1e-10
dbj|BAB11147.1| cytochrome P450 [Arabidopsis thaliana] gi|152402... 69 2e-10
dbj|BAB40324.1| cytochrome P450 [Asparagus officinalis] 69 2e-10
dbj|BAB40323.1| cytochrome P450 [Asparagus officinalis] 69 2e-10
gb|AAM47994.1| putative cytochrome P450 [Arabidopsis thaliana] g... 68 3e-10
dbj|BAD94709.1| cytochrome P450 [Arabidopsis thaliana] 68 4e-10
gb|AAS48419.1| flavonoid 3'-hydroxylase [Allium cepa] 68 4e-10
dbj|BAD37360.1| putative cytochrome P450 [Oryza sativa (japonica... 68 4e-10
>gb|AAB94584.1| CYP71A10 [Glycine max] gi|7430655|pir||T05735 cytochrome P450 71A10
- soybean
Length = 513
Score = 81.3 bits (199), Expect = 3e-14
Identities = 44/103 (42%), Positives = 65/103 (62%), Gaps = 2/103 (1%)
Query: 210 LKKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALL 269
+ ++K + +D FLD VI EH+ S + K DFL ILLQLQ+ G +F+L +++LKA+L
Sbjct: 248 IPEMKTTFLAVDAFLDEVIAEHESSNK--KNDDFLGILLQLQECGRLDFQLDRDNLKAIL 305
Query: 270 MNMFLGGSDTTSTTLEWAMAACKKSSHNEESTGGGKENCRGIN 312
++M +GGSDTTSTTLEW A ++ + + GIN
Sbjct: 306 VDMIIGGSDTTSTTLEWTFAEFLRNPNTMKKAQEEVRRVVGIN 348
>sp|P24465|C71A1_PERAE Cytochrome P450 71A1 (CYPLXXIA1) (ARP-2)
Length = 471
Score = 75.1 bits (183), Expect = 2e-12
Identities = 37/83 (44%), Positives = 57/83 (68%), Gaps = 5/83 (6%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRR-----DPKKKDFLDILLQLQDDGLTEFELTQNDLK 266
++K + E+D F+D VI +H +SR+ ++KD +D+LL LQ D L +N+LK
Sbjct: 236 RLKRNHGELDAFVDHVIDDHLLSRKANGSDGVEQKDLVDVLLHLQKDSSLGVHLNRNNLK 295
Query: 267 ALLMNMFLGGSDTTSTTLEWAMA 289
A++++MF GG+DTT+ TLEWAMA
Sbjct: 296 AVILDMFSGGTDTTAVTLEWAMA 318
>pir||T52256 cytochrome P-450LXXIA1 [similarity] - avocado
gi|166949|gb|AAA32913.1| cytochrome P-450LXXIA1
(cyp71A1)
Length = 502
Score = 75.1 bits (183), Expect = 2e-12
Identities = 37/83 (44%), Positives = 57/83 (68%), Gaps = 5/83 (6%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRR-----DPKKKDFLDILLQLQDDGLTEFELTQNDLK 266
++K + E+D F+D VI +H +SR+ ++KD +D+LL LQ D L +N+LK
Sbjct: 236 RLKRNHGELDAFVDHVIDDHLLSRKANGSDGVEQKDLVDVLLHLQKDSSLGVHLNRNNLK 295
Query: 267 ALLMNMFLGGSDTTSTTLEWAMA 289
A++++MF GG+DTT+ TLEWAMA
Sbjct: 296 AVILDMFSGGTDTTAVTLEWAMA 318
>gb|AAC02748.1| putative cytochrome P450 [Arabidopsis thaliana]
gi|13878363|sp|O49342|C71AD_ARATH Cytochrome P450 71A13
Length = 497
Score = 73.9 bits (180), Expect = 5e-12
Identities = 35/83 (42%), Positives = 55/83 (66%), Gaps = 2/83 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
KIK+ S D +D+V+ EH + D K DF+DILL ++ D + F++ +ND+K ++++
Sbjct: 237 KIKEVSRGFSDLMDKVVQEHLEASND--KADFVDILLSIEKDKNSGFQVQRNDIKFMILD 294
Query: 272 MFLGGSDTTSTTLEWAMAACKKS 294
MF+GG+ TTST LEW M +S
Sbjct: 295 MFIGGTSTTSTLLEWTMTELIRS 317
>ref|NP_180635.2| cytochrome P450 71A13, putative (CYP71A13) [Arabidopsis thaliana]
Length = 503
Score = 73.9 bits (180), Expect = 5e-12
Identities = 35/83 (42%), Positives = 55/83 (66%), Gaps = 2/83 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
KIK+ S D +D+V+ EH + D K DF+DILL ++ D + F++ +ND+K ++++
Sbjct: 243 KIKEVSRGFSDLMDKVVQEHLEASND--KADFVDILLSIEKDKNSGFQVQRNDIKFMILD 300
Query: 272 MFLGGSDTTSTTLEWAMAACKKS 294
MF+GG+ TTST LEW M +S
Sbjct: 301 MFIGGTSTTSTLLEWTMTELIRS 323
>dbj|BAC43325.1| putative cytochrome P450 [Arabidopsis thaliana]
gi|28973083|gb|AAO63866.1| putative cytochrome p450
[Arabidopsis thaliana] gi|15223580|ref|NP_175469.1|
cytochrome P450 family protein [Arabidopsis thaliana]
gi|12322341|gb|AAG51197.1| cytochrome P450, putative
[Arabidopsis thaliana] gi|25282575|pir||G96541 probable
cytochrome P450 [imported] - Arabidopsis thaliana
gi|9454558|gb|AAF87881.1| Putative cytochrome P450
[Arabidopsis thaliana]
Length = 533
Score = 71.2 bits (173), Expect = 4e-11
Identities = 29/79 (36%), Positives = 55/79 (68%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
K+I + S+ D+ L+++I EH+ + +D +D+LL++ D E ++T+N +KAL++
Sbjct: 250 KEIMEVSQRYDELLEKIIKEHEEDPNKKEDRDMMDVLLEVCADDKAEVKITRNQIKALIV 309
Query: 271 NMFLGGSDTTSTTLEWAMA 289
+FLGG+DT++ T++W MA
Sbjct: 310 ELFLGGTDTSAQTIQWIMA 328
>emb|CAB41171.1| cytochrome P450-like protein [Arabidopsis thaliana]
gi|13878399|sp|Q9STK7|C71AQ_ARATH Cytochrome P450 71A26
gi|22331672|ref|NP_680106.1| cytochrome P450 71A26,
putative (CYP71A26) [Arabidopsis thaliana]
Length = 489
Score = 70.5 bits (171), Expect = 6e-11
Identities = 31/77 (40%), Positives = 54/77 (69%), Gaps = 2/77 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
+++ ++ ++D F +RV+ +H RD DF+D+LL +Q D FE+ + +KA++MN
Sbjct: 230 QLEKTANDVDKFFERVVQDHVDGNRD--MTDFVDVLLAIQRDKTVGFEINRVSIKAIVMN 287
Query: 272 MFLGGSDTTSTTLEWAM 288
+F+GG+DT+ST +EWAM
Sbjct: 288 VFVGGTDTSSTLMEWAM 304
>dbj|BAB09329.1| cytochrome P450 [Arabidopsis thaliana] gi|15239007|ref|NP_199072.1|
cytochrome P450 family protein [Arabidopsis thaliana]
Length = 499
Score = 70.5 bits (171), Expect = 6e-11
Identities = 32/89 (35%), Positives = 58/89 (64%)
Query: 201 VGLMFSLAKLKKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFEL 260
V L FS K I S E D+FL+R++VEH+ + + +D +D LL+ + E+++
Sbjct: 222 VKLFFSNMFRKDIMGVSREFDEFLERILVEHEENLEGDQDRDMIDHLLEAYRNEEAEYKI 281
Query: 261 TQNDLKALLMNMFLGGSDTTSTTLEWAMA 289
T+ +K+L++ +FLGG+D+++ T++W MA
Sbjct: 282 TRKQIKSLIVEIFLGGTDSSAQTIQWTMA 310
>ref|XP_464378.1| putative cytochrome P450 [Oryza sativa (japonica cultivar-group)]
gi|46390073|dbj|BAD15448.1| putative cytochrome P450
[Oryza sativa (japonica cultivar-group)]
gi|46390042|dbj|BAD15418.1| putative cytochrome P450
[Oryza sativa (japonica cultivar-group)]
Length = 509
Score = 70.1 bits (170), Expect = 8e-11
Identities = 34/78 (43%), Positives = 55/78 (69%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
KI+ M F+D +I EH+ SR + +D LD+LL+LQ D +++ LT ++K++L++
Sbjct: 241 KIERRRRGMMGFIDTIIQEHQESRAAAEDEDLLDVLLRLQKDMDSQYPLTTMNIKSILID 300
Query: 272 MFLGGSDTTSTTLEWAMA 289
MF GS+T++TTL+WAMA
Sbjct: 301 MFGAGSETSATTLQWAMA 318
>gb|AAD25850.1| putative cytochrome P450 [Arabidopsis thaliana]
gi|15225585|ref|NP_179026.1| cytochrome P450 family
protein [Arabidopsis thaliana] gi|25282596|pir||B84514
probable cytochrome P450 [imported] - Arabidopsis
thaliana
Length = 518
Score = 70.1 bits (170), Expect = 8e-11
Identities = 31/79 (39%), Positives = 52/79 (65%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
K+ D S + D+ L+R+IVEH+ D +D+LL + DG E+++T++ LK+L +
Sbjct: 248 KETLDVSRKFDELLERIIVEHEEKTDYDHGMDLMDVLLAVYRDGKAEYKITRDHLKSLFV 307
Query: 271 NMFLGGSDTTSTTLEWAMA 289
+ LGG+DT++ T+EW MA
Sbjct: 308 ELILGGTDTSAQTIEWTMA 326
>dbj|BAB09330.1| cytochrome P450 [Arabidopsis thaliana] gi|15239008|ref|NP_199073.1|
cytochrome P450 71A16, putative (CYP71A16) [Arabidopsis
thaliana] gi|13878374|sp|Q9FH66|C71AG_ARATH Cytochrome
P450 71A16
Length = 497
Score = 69.3 bits (168), Expect = 1e-10
Identities = 33/102 (32%), Positives = 61/102 (59%), Gaps = 3/102 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
K++ +++ DF+++V+ EH+ + D + DF+D+LL +Q D + +L ++DLK ++
Sbjct: 236 KLEKITKQFGDFIEKVLQEHEDTTADKETPDFVDMLLTIQRDETAQCQLDKSDLKVIIFE 295
Query: 272 MFLGGSDTTSTTLEWAMAACKKSSHNEESTGGGKENCRGINK 313
MFLG + TTS +EWAM + N E ++ R ++K
Sbjct: 296 MFLGSTTTTSAVIEWAMT---RLMRNPECLKKLQDEIRSVSK 334
>emb|CAA50645.1| P450 hydroxylase [Solanum melongena]
gi|584861|sp|P37118|C71A2_SOLME Cytochrome P450 71A2
(CYPLXXIA2) (P-450EG4) gi|441185|dbj|BAA03635.1|
Cytochrome P-450EG4 [Solanum melongena]
Length = 505
Score = 69.3 bits (168), Expect = 1e-10
Identities = 34/84 (40%), Positives = 56/84 (66%), Gaps = 6/84 (7%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPK------KKDFLDILLQLQDDGLTEFELTQNDL 265
K+K ++E+D FL+ VI EH + ++ + KDF+D+LL++Q+ T+F L ++ L
Sbjct: 238 KVKKVAKELDMFLEIVIEEHIIRKKKEEYTSTGEAKDFVDVLLEIQNGNETDFPLQRDSL 297
Query: 266 KALLMNMFLGGSDTTSTTLEWAMA 289
KA+L++ F G+DTT TL+W MA
Sbjct: 298 KAILLDSFAAGTDTTFATLDWTMA 321
>sp|Q42799|C93A2_SOYBN Cytochrome P450 93A2 gi|1408322|dbj|BAA13076.1| cytochrome P-450
(CYP93A2) [Glycine max]
Length = 502
Score = 69.3 bits (168), Expect = 1e-10
Identities = 33/86 (38%), Positives = 57/86 (65%), Gaps = 7/86 (8%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKK-------KDFLDILLQLQDDGLTEFELTQN 263
K+I+ + D LDR+I + + RR+ K+ KD LD+LL + +D +E +LT+
Sbjct: 228 KRIRKTRIRFDAVLDRIIKQREEERRNNKEIGGTRQFKDILDVLLDIGEDDSSEIKLTKE 287
Query: 264 DLKALLMNMFLGGSDTTSTTLEWAMA 289
++KA +M++F+ G+DT++ T+EWAMA
Sbjct: 288 NIKAFIMDIFVAGTDTSAATMEWAMA 313
>dbj|BAB11147.1| cytochrome P450 [Arabidopsis thaliana] gi|15240211|ref|NP_196307.1|
cytochrome P450 family protein [Arabidopsis thaliana]
Length = 507
Score = 68.9 bits (167), Expect = 2e-10
Identities = 33/103 (32%), Positives = 66/103 (64%), Gaps = 8/103 (7%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPK-KKDFLDILLQLQDDGLTEFELTQNDLKALL 269
K++K++ ++ D ++R++ EH+ S+++ +++ LD+LL + +D E +LT+ ++KA +
Sbjct: 239 KRLKNARDKYDVIIERIMEEHESSKKNATGERNMLDVLLDIYEDKNAEMKLTRENIKAFI 298
Query: 270 MNMFLGGSDTTSTTLEWAMA-------ACKKSSHNEESTGGGK 305
MN++ GG+DT++ T+EWA+A KK+ E G K
Sbjct: 299 MNIYGGGTDTSAITVEWALAELINHPEIMKKAQQEIEQVVGNK 341
>dbj|BAB40324.1| cytochrome P450 [Asparagus officinalis]
Length = 501
Score = 68.6 bits (166), Expect = 2e-10
Identities = 33/82 (40%), Positives = 58/82 (70%), Gaps = 5/82 (6%)
Query: 209 KLKKIKDSSEEMDDFLDRVIVEH-KMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKA 267
KL K+K ++ ++D+F VI H K R++ K+D +D+LL+L+++G +LT + +K
Sbjct: 238 KLGKMKRNARDLDEFYQEVIDAHMKDGRKEDGKEDIVDVLLRLREEG----QLTMDHIKG 293
Query: 268 LLMNMFLGGSDTTSTTLEWAMA 289
LMN+F+GG+DT++ ++ WAMA
Sbjct: 294 ALMNIFVGGTDTSAASIAWAMA 315
>dbj|BAB40323.1| cytochrome P450 [Asparagus officinalis]
Length = 501
Score = 68.6 bits (166), Expect = 2e-10
Identities = 33/82 (40%), Positives = 58/82 (70%), Gaps = 5/82 (6%)
Query: 209 KLKKIKDSSEEMDDFLDRVIVEH-KMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKA 267
KL K+K ++ ++D+F VI H K R++ K+D +D+LL+L+++G +LT + +K
Sbjct: 238 KLGKMKRNARDLDEFYQEVIDAHMKDGRKEDGKEDIVDVLLRLREEG----QLTMDHIKG 293
Query: 268 LLMNMFLGGSDTTSTTLEWAMA 289
LMN+F+GG+DT++ ++ WAMA
Sbjct: 294 ALMNIFVGGTDTSAASIAWAMA 315
>gb|AAM47994.1| putative cytochrome P450 [Arabidopsis thaliana]
gi|15223584|ref|NP_175471.1| cytochrome P450, putative
[Arabidopsis thaliana] gi|17064994|gb|AAL32651.1|
cytochrome P450, putative [Arabidopsis thaliana]
gi|12322346|gb|AAG51202.1| cytochrome P450, putative
[Arabidopsis thaliana] gi|25282576|pir||A96542 probable
cytochrome P450 [imported] - Arabidopsis thaliana
gi|9454560|gb|AAF87883.1| Putative cytochrome P450
[Arabidopsis thaliana]
Length = 519
Score = 68.2 bits (165), Expect = 3e-10
Identities = 27/79 (34%), Positives = 56/79 (70%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLM 270
K+I + S+ D+ L+++I EH+ + + + +D +D+LL++ D EF++++N +KAL +
Sbjct: 251 KEIMEVSQRYDELLEKIIKEHEENPNNGEDRDMMDVLLEVCADDNAEFKISRNQIKALFV 310
Query: 271 NMFLGGSDTTSTTLEWAMA 289
+FL G+DT++ T++W +A
Sbjct: 311 EIFLAGTDTSAQTIQWILA 329
>dbj|BAD94709.1| cytochrome P450 [Arabidopsis thaliana]
Length = 497
Score = 67.8 bits (164), Expect = 4e-10
Identities = 32/102 (31%), Positives = 61/102 (59%), Gaps = 3/102 (2%)
Query: 212 KIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTEFELTQNDLKALLMN 271
K++ +++ DF+++V+ EH+ + D + DF+D+LL +Q D + +L ++DL+ ++
Sbjct: 236 KLEKITKQFGDFIEKVLQEHEDTTADKETPDFVDMLLTIQRDETAQCQLDKSDLEVIIFE 295
Query: 272 MFLGGSDTTSTTLEWAMAACKKSSHNEESTGGGKENCRGINK 313
MFLG + TTS +EWAM + N E ++ R ++K
Sbjct: 296 MFLGSTTTTSAVIEWAMT---RLMRNPECLKKLQDEIRSVSK 334
>gb|AAS48419.1| flavonoid 3'-hydroxylase [Allium cepa]
Length = 510
Score = 67.8 bits (164), Expect = 4e-10
Identities = 34/81 (41%), Positives = 54/81 (65%), Gaps = 4/81 (4%)
Query: 210 LKKIKDSSEEMDDFLDRVIVEHKMSRRDPKKKDFLDILLQLQDDGLTE-FELTQNDLKAL 268
++K+KD D FLD+VI EH+M D L +L++ +DD + +LT D+KAL
Sbjct: 237 VRKMKDVHARFDSFLDKVIAEHRMV---DNAHDLLSLLIRSKDDVDNDGVKLTNTDIKAL 293
Query: 269 LMNMFLGGSDTTSTTLEWAMA 289
L+N+F G+DT+S+T+EWA++
Sbjct: 294 LLNLFTAGTDTSSSTVEWALS 314
>dbj|BAD37360.1| putative cytochrome P450 [Oryza sativa (japonica cultivar-group)]
Length = 504
Score = 67.8 bits (164), Expect = 4e-10
Identities = 33/84 (39%), Positives = 53/84 (62%), Gaps = 5/84 (5%)
Query: 211 KKIKDSSEEMDDFLDRVIVEHKMSR-----RDPKKKDFLDILLQLQDDGLTEFELTQNDL 265
++ + S EM +D VI EH+ R RD + +D LD+LL++Q DG + L +
Sbjct: 228 RRAEAHSREMTRLMDGVIEEHRQRRAATGWRDEEDEDLLDVLLRIQKDGGLQIPLDMGTI 287
Query: 266 KALLMNMFLGGSDTTSTTLEWAMA 289
+A+++++F GS+TT TTL+WAMA
Sbjct: 288 RAIIIDLFSAGSETTGTTLQWAMA 311
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.353 0.156 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 449,777,644
Number of Sequences: 2540612
Number of extensions: 15178875
Number of successful extensions: 85560
Number of sequences better than 10.0: 1467
Number of HSP's better than 10.0 without gapping: 516
Number of HSP's successfully gapped in prelim test: 951
Number of HSP's that attempted gapping in prelim test: 84236
Number of HSP's gapped (non-prelim): 1578
length of query: 313
length of database: 863,360,394
effective HSP length: 128
effective length of query: 185
effective length of database: 538,162,058
effective search space: 99559980730
effective search space used: 99559980730
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 75 (33.5 bits)
Medicago: description of AC124952.12


