

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124951.10 - phase: 0
(446 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
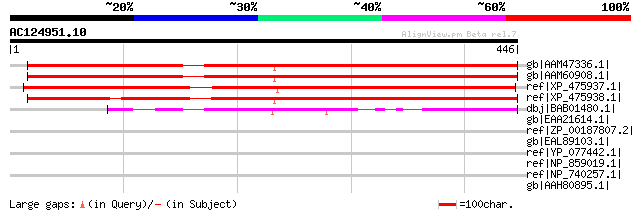
Sequences producing significant alignments: (bits) Value
gb|AAM47336.1| AT3g27930/K24A2_2 [Arabidopsis thaliana] gi|18087... 569 e-161
gb|AAM60908.1| unknown [Arabidopsis thaliana] 566 e-160
ref|XP_475937.1| unknown protein [Oryza sativa (japonica cultiva... 520 e-146
ref|XP_475938.1| unknown protein [Oryza sativa (japonica cultiva... 441 e-122
dbj|BAB01480.1| unnamed protein product [Arabidopsis thaliana] 291 4e-77
gb|EAA21614.1| probable leucyl-tRNA synthetase-related [Plasmodi... 35 4.7
ref|ZP_00187807.2| COG4166: ABC-type oligopeptide transport syst... 35 4.7
gb|EAL89103.1| leucyl-tRNA synthetase [Aspergillus fumigatus Af293] 35 6.2
ref|YP_077442.1| Polysaccharide deacetylase,Carbohydrate Esteras... 35 6.2
ref|NP_859019.1| VP0 [Bovine kobuvirus] 35 6.2
ref|NP_740257.1| unnamed protein product [Bovine kobuvirus] gi|2... 35 6.2
gb|AAH80895.1| Hoxa1-prov protein [Xenopus tropicalis] gi|561185... 35 6.2
>gb|AAM47336.1| AT3g27930/K24A2_2 [Arabidopsis thaliana] gi|18087557|gb|AAL58910.1|
AT3g27930/K24A2_2 [Arabidopsis thaliana]
gi|18405582|ref|NP_566828.1| expressed protein
[Arabidopsis thaliana]
Length = 425
Score = 569 bits (1467), Expect = e-161
Identities = 278/434 (64%), Positives = 337/434 (77%), Gaps = 20/434 (4%)
Query: 16 ESPPPVVLVPPLFDFPPIAARNRMLESSYDVVFGKLALRCLFNDYFQQPKHFVTRIMLKP 75
E PPPVVLVPPLFD+PP++AR RMLESSY+++FGKLALRCLF DYF++ F + +LKP
Sbjct: 9 EPPPPVVLVPPLFDYPPLSARTRMLESSYNLLFGKLALRCLFEDYFEEANRFTGKFLLKP 68
Query: 76 IDDPHVDLIATVSGPLDQKPDENINGNALFRWQSDVNDPHTFMDLYVSTSDPVLQMRSCA 135
DDPHVDL+A+VSG +D + + + GNA FRWQSDV+DPHTF+DL VSTS+PVL MRS A
Sbjct: 69 TDDPHVDLVASVSGAVDGRVEGDFVGNAEFRWQSDVDDPHTFVDLSVSTSNPVLLMRSSA 128
Query: 136 YYPRYGFGAFGVFPLLLKNRTSSQIATFTADKVRQLYRETSQDSGVMGLRYGSGNLSCGV 195
YYP+YG GAF V+PL+ K + ++S++ +MGLRYGS NLS G
Sbjct: 129 YYPKYGIGAFAVYPLISK-----------------ITGKSSEEYRIMGLRYGSTNLSVGA 171
Query: 196 TLMPFAKKDELPKSVWLVSKIGRVTAGVQYEPHHEN---AKLSNLMNWSCAMAYGVGSQS 252
T+ PF+ +ELPK WLVSK+G +T GVQYEP H + AK ++ NWSCA YGVGSQS
Sbjct: 172 TVTPFSANNELPKHAWLVSKMGSLTVGVQYEPLHGSKDLAKYTDPRNWSCAAGYGVGSQS 231
Query: 253 PLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENTVVGITNYIDFGFELQTSVDD 312
PL+PSFN +EL +SSQF+ASFYQH+VVQRRV+NP EEN VVGITNYIDFGFELQ+ VDD
Sbjct: 232 PLTPSFNIGIELARSSQFIASFYQHVVVQRRVQNPFEENQVVGITNYIDFGFELQSRVDD 291
Query: 313 AIAANNISDSTFQIAASWQANKNFLVKAKAGPKSSTMALAFKSWWKPSFTFSISATRDRA 372
+ N DS Q+AASWQANKNFL+K K G SST++LAFKSWWKPSF F+ISAT +
Sbjct: 292 SKTPPNAPDSLLQVAASWQANKNFLLKGKVGAHSSTLSLAFKSWWKPSFAFNISATTNHR 351
Query: 373 DGQVQYGFGLQSESLREASYQRADPNFVMLTPSKEHLAEGIVWETGKRPMLQSDIDAGHF 432
G VQ GFGL+ ++LREASYQRADPNFVMLTP+KEHLAEGIVW+ GKRPM Q+D+DA +F
Sbjct: 352 TGNVQCGFGLRVDNLREASYQRADPNFVMLTPNKEHLAEGIVWKMGKRPMYQADVDAENF 411
Query: 433 DGLPRELRPLDKIL 446
LP+ELRP KIL
Sbjct: 412 SELPKELRPSQKIL 425
>gb|AAM60908.1| unknown [Arabidopsis thaliana]
Length = 425
Score = 566 bits (1459), Expect = e-160
Identities = 276/434 (63%), Positives = 336/434 (76%), Gaps = 20/434 (4%)
Query: 16 ESPPPVVLVPPLFDFPPIAARNRMLESSYDVVFGKLALRCLFNDYFQQPKHFVTRIMLKP 75
E PPPVVLVPPLFD+PP++AR RMLESSY+++FGKLALRCLF DYF++ F + +LKP
Sbjct: 9 EPPPPVVLVPPLFDYPPLSARTRMLESSYNLLFGKLALRCLFEDYFEEANRFTAKFLLKP 68
Query: 76 IDDPHVDLIATVSGPLDQKPDENINGNALFRWQSDVNDPHTFMDLYVSTSDPVLQMRSCA 135
DDPHVDL+A+VSG +D + + + GNA FRWQSDV+DPHTF+DL VSTS+PVL MRS A
Sbjct: 69 TDDPHVDLVASVSGAVDGRVEGDFVGNAEFRWQSDVDDPHTFVDLSVSTSNPVLLMRSSA 128
Query: 136 YYPRYGFGAFGVFPLLLKNRTSSQIATFTADKVRQLYRETSQDSGVMGLRYGSGNLSCGV 195
YYP+YG GAF V+PL+ K + ++S++ +MGLRYGS LS G
Sbjct: 129 YYPKYGIGAFAVYPLISK-----------------ITGKSSEEYRIMGLRYGSTXLSVGA 171
Query: 196 TLMPFAKKDELPKSVWLVSKIGRVTAGVQYEPHHEN---AKLSNLMNWSCAMAYGVGSQS 252
T+ PF+ +ELPK WLVSK+G +T GVQYEP H + AK ++ NWSCA YGVGSQS
Sbjct: 172 TVTPFSANNELPKHAWLVSKMGSLTVGVQYEPLHGSKDLAKYTDPRNWSCAAGYGVGSQS 231
Query: 253 PLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENTVVGITNYIDFGFELQTSVDD 312
PL+PSFN +EL +SSQF+ASFYQH+VVQR+V+NP EEN VVGITNYIDFGFELQ+ VDD
Sbjct: 232 PLTPSFNMGIELARSSQFIASFYQHVVVQRKVQNPFEENQVVGITNYIDFGFELQSRVDD 291
Query: 313 AIAANNISDSTFQIAASWQANKNFLVKAKAGPKSSTMALAFKSWWKPSFTFSISATRDRA 372
+ N DS Q+AASWQANKNFL+K K G SST++LAFKSWWKPSF F+ISAT +
Sbjct: 292 SKTPPNAPDSLLQVAASWQANKNFLLKGKVGAHSSTLSLAFKSWWKPSFAFNISATTNHR 351
Query: 373 DGQVQYGFGLQSESLREASYQRADPNFVMLTPSKEHLAEGIVWETGKRPMLQSDIDAGHF 432
G VQ GFGL+ ++LREASYQRADPNFVMLTP+KEHLAEGIVW+ GKRPM Q+D+DA +F
Sbjct: 352 TGNVQCGFGLRVDNLREASYQRADPNFVMLTPNKEHLAEGIVWKMGKRPMYQADVDAENF 411
Query: 433 DGLPRELRPLDKIL 446
LP+ELRP KIL
Sbjct: 412 SELPKELRPSQKIL 425
>ref|XP_475937.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|47900285|gb|AAT39153.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 424
Score = 520 bits (1340), Expect = e-146
Identities = 251/436 (57%), Positives = 322/436 (73%), Gaps = 21/436 (4%)
Query: 13 NEVESPPPVVLVPPLFDFPPIAARNRMLESSYDVVFGKLALRCLFNDYFQQPKHFVTRIM 72
++ E PPP+VLVPPLFD+PPIAAR RM +Y+++FGKL+L+ LF DYF + +R+M
Sbjct: 6 SKAEPPPPMVLVPPLFDYPPIAARTRMSVPAYELMFGKLSLQNLFEDYFDHAGNMTSRVM 65
Query: 73 LKPIDDPHVDLIATVSGPLDQKPDENINGNALFRWQSDVNDPHTFMDLYVSTSDPVLQMR 132
LKP++DPHVDLIATVS D+ + G+ALFRWQ D++DPHTF+DL VSTS+P+LQ+R
Sbjct: 66 LKPLEDPHVDLIATVSAAADKIDGTQVKGDALFRWQKDLDDPHTFVDLLVSTSNPMLQVR 125
Query: 133 SCAYYPRYGFGAFGVFPLLLKNRTSSQIATFTADKVRQLYRETSQDSGVMGLRYGSGNLS 192
SCAY+P+Y GAFG FPLL+ NR S+ D GVMG+RYGS NLS
Sbjct: 126 SCAYHPKYRVGAFGTFPLLMGNRVRSE------------------DYGVMGVRYGSENLS 167
Query: 193 CGVTLMPFAKKDELPKSVWLVSKIGRVTAGVQYEPHHENAKL---SNLMNWSCAMAYGVG 249
G + +PF ELP WLV + G ++AGVQY+P N L ++ NW+CA++YGVG
Sbjct: 168 FGSSFVPFPGSAELPSGAWLVGRKGSLSAGVQYKPLSGNKHLMPYTDWKNWNCAISYGVG 227
Query: 250 SQSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENTVVGITNYIDFGFELQTS 309
SPLSPSF FSLEL +S++F+ASFYQHMVVQRRVKNP E++ +VGITNYIDFG EL T
Sbjct: 228 LTSPLSPSFIFSLELARSTEFIASFYQHMVVQRRVKNPFEDDQIVGITNYIDFGLELATR 287
Query: 310 VDDAIAANNISDSTFQIAASWQANKNFLVKAKAGPKSSTMALAFKSWWKPSFTFSISATR 369
+D + + ++S FQ AASWQANKNFL K K GP S++ALAFKSWW+PSFTFS++A
Sbjct: 288 IDKDKPSESANNSLFQFAASWQANKNFLFKGKLGPSKSSVALAFKSWWRPSFTFSVTAVN 347
Query: 370 DRADGQVQYGFGLQSESLREASYQRADPNFVMLTPSKEHLAEGIVWETGKRPMLQSDIDA 429
D G YGFG++ E LR+ SYQRADPN+VMLTPSKEHLA G++ E GKRPM Q+++D+
Sbjct: 348 DHLKGTRSYGFGIRVEDLRQPSYQRADPNYVMLTPSKEHLAPGVLREYGKRPMFQAEVDS 407
Query: 430 GHFDGLPRELRPLDKI 445
G++D LP EL+P+ KI
Sbjct: 408 GNYDHLPTELKPISKI 423
>ref|XP_475938.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|47900286|gb|AAT39154.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 414
Score = 441 bits (1135), Expect = e-122
Identities = 225/435 (51%), Positives = 293/435 (66%), Gaps = 30/435 (6%)
Query: 16 ESPPPVVLVPPLFDFPPIAARNRMLESSYDVVFGKLALRCLFNDYFQQPKHFVT-RIMLK 74
E PP +VLVPP F FP AAR RM +Y+V+FGKL R LF+DYF Q + IML+
Sbjct: 6 EMPPAMVLVPPPFTFPAAAARTRMAVPAYEVMFGKLQRRSLFDDYFDQVGSITSGMIMLR 65
Query: 75 PIDDPHVDLIATVSGPLDQKPDENINGNALFRWQSDVNDPHTFMDLYVSTSDPVLQMRSC 134
P+ D HVDL A ++ G ALFRWQ ++DP+TFMDL++ST P++Q+RSC
Sbjct: 66 PLVDSHVDLTAKMT---------TTGGEALFRWQRYLDDPNTFMDLHLSTPKPMVQLRSC 116
Query: 135 AYYPRYGFGAFGVFPLLLKNRTSSQIATFTADKVRQLYRETSQDSGVMGLRYGSGNLSCG 194
AYYP+Y GAFG FPLL NR S+ D G+MGLRYGS NLS G
Sbjct: 117 AYYPKYRIGAFGTFPLLKANRDCSE-----------------GDYGIMGLRYGSENLSIG 159
Query: 195 VTLMPFAKKDELPKSVWLVSKIGRVTAGVQYEPHHEN---AKLSNLMNWSCAMAYGVGSQ 251
+ +PFA ++P WLV + G ++AG+QY+P E+ L++L NW+CA++YG+GS
Sbjct: 160 ASFLPFALSGQVPYGAWLVGRKGNISAGIQYKPLCESMHPVPLTDLKNWNCAISYGMGST 219
Query: 252 SPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENTVVGITNYIDFGFELQTSVD 311
SPLSPSFNFSLELV+++Q VASFYQH VVQRRV NP EE ++G TN++DFG EL TS+D
Sbjct: 220 SPLSPSFNFSLELVRNTQLVASFYQHFVVQRRVMNPREEEHIIGTTNFVDFGLELATSLD 279
Query: 312 DAIAANNISDSTFQIAASWQANKNFLVKAKAGPKSSTMALAFKSWWKPSFTFSISATRDR 371
A N S+ +FQ+AASWQA+KNFLVK K GP S+MALA KSWW+P FTFS +A D
Sbjct: 280 KDKAKENASNPSFQVAASWQASKNFLVKGKLGPSKSSMALAMKSWWRPFFTFSFTAMYDH 339
Query: 372 ADGQVQYGFGLQSESLREASYQRADPNFVMLTPSKEHLAEGIVWETGKRPMLQSDIDAGH 431
G YGFG+ E L+E SYQ A N+V++T +KE + + + GK+ + Q DID+G+
Sbjct: 340 LKGTGSYGFGISIEDLKEPSYQMAYSNYVIVTQNKEDVEPRFLKKLGKKYIFQPDIDSGN 399
Query: 432 FDGLPRELRPLDKIL 446
+D LP L+P+DKIL
Sbjct: 400 YDNLPTGLKPIDKIL 414
>dbj|BAB01480.1| unnamed protein product [Arabidopsis thaliana]
Length = 351
Score = 291 bits (744), Expect = 4e-77
Identities = 173/366 (47%), Positives = 213/366 (57%), Gaps = 80/366 (21%)
Query: 87 VSGPLDQKPDENINGNALFRWQSDVNDPHTFMDLYVSTSDPVLQMRSCAYYPRYGFGAFG 146
VSG +D + + + GNA FRWQ VL MRS AYYP+YG GAF
Sbjct: 60 VSGAVDGRVEGDFVGNAEFRWQR------------------VLLMRSSAYYPKYGIGAFA 101
Query: 147 VFPLLLKNRTSSQIATFTADKVRQLYRETSQDSGVMGLRYGSGNLSCGVTLMPFAKKDEL 206
V+PL+ K + ++S++ +MGLRYGS NLS G T+ PF+ +EL
Sbjct: 102 VYPLISK-----------------ITGKSSEEYRIMGLRYGSTNLSVGATVTPFSANNEL 144
Query: 207 PKSVWLVSKIGRVTAGVQYEPHH---ENAKLSNLMNWSCAMAYGVGSQSPLSPSFNFSLE 263
PK WLVSK+G +T GVQYEP H + AK ++ NWSCA YGVGSQSPL+PSFN +E
Sbjct: 145 PKHAWLVSKMGSLTVGVQYEPLHGSKDLAKYTDPRNWSCAAGYGVGSQSPLTPSFNIGIE 204
Query: 264 LVKSSQFVASFYQH---MVVQRRVKNPLEENTVVGITNYIDFGFELQTSVDDAIAANNIS 320
L +SSQF S Q+ ++ +V+NP EEN VVGITNYIDFGFELQ
Sbjct: 205 LARSSQFSTSLAQYEIDLMDLEQVQNPFEENQVVGITNYIDFGFELQ------------- 251
Query: 321 DSTFQIAASWQANKNFLVKAKAGPKSSTMALAFKSWWKPSFTFSISATRDRADGQVQYGF 380
S F IA AG + AT + G VQ GF
Sbjct: 252 -SRFFIAGGC---------VLAGQQE----------------LFAEATTNHRTGNVQCGF 285
Query: 381 GLQSESLREASYQRADPNFVMLTPSKEHLAEGIVWETGKRPMLQSDIDAGHFDGLPRELR 440
GL+ ++LREASYQRADPNFVMLTP+KEHLAEGIVW+ GKRPM Q+D+DA +F LP+ELR
Sbjct: 286 GLRVDNLREASYQRADPNFVMLTPNKEHLAEGIVWKMGKRPMYQADVDAENFSELPKELR 345
Query: 441 PLDKIL 446
P KIL
Sbjct: 346 PSQKIL 351
>gb|EAA21614.1| probable leucyl-tRNA synthetase-related [Plasmodium yoelii yoelii]
Length = 1278
Score = 35.0 bits (79), Expect = 4.7
Identities = 33/129 (25%), Positives = 58/129 (44%), Gaps = 10/129 (7%)
Query: 237 LMNWSCAMAYGVGSQSPLSPSFNFS-LELVKSSQ-FVASFYQHMVVQRRVKNPLEENTVV 294
L +WSC+ +YG+G+ P N S E +K+ + +V ++ V + + L ++T+
Sbjct: 670 LDDWSCSRSYGLGTHMPNFNEKNKSENESIKTDENYVRKTDENYVRKTELIESLSDSTI- 728
Query: 295 GITNYIDFGFELQTSVDDA------IAANNISDSTFQIAASWQANKNFLVKAKAGPKSST 348
Y LQ +VD I N+I+D+ F N + + K K +
Sbjct: 729 -YMAYYTISHYLQGNVDGKEKGILNIDHNDINDAFFDYVFDISNNTSNISKNITIEKLNI 787
Query: 349 MALAFKSWW 357
M +FK W+
Sbjct: 788 MRKSFKYWY 796
>ref|ZP_00187807.2| COG4166: ABC-type oligopeptide transport system, periplasmic
component [Rubrobacter xylanophilus DSM 9941]
Length = 541
Score = 35.0 bits (79), Expect = 4.7
Identities = 17/39 (43%), Positives = 23/39 (58%), Gaps = 3/39 (7%)
Query: 90 PLDQKPDENINGNALFRWQ---SDVNDPHTFMDLYVSTS 125
P D+K + NG WQ +D NDP TF+DL++S S
Sbjct: 420 PFDRKLELEANGEFELSWQGWIADYNDPMTFLDLFLSDS 458
>gb|EAL89103.1| leucyl-tRNA synthetase [Aspergillus fumigatus Af293]
Length = 1127
Score = 34.7 bits (78), Expect = 6.2
Identities = 25/96 (26%), Positives = 38/96 (39%), Gaps = 3/96 (3%)
Query: 228 HHENAKLSNLMNWSCAMAYGVGSQSPLSPSF---NFSLELVKSSQFVASFYQHMVVQRRV 284
H L+ L W+CA YG+GS+ P P F + S V + + + Y H
Sbjct: 605 HGFEKNLTWLNRWACARTYGLGSKLPWDPKFLVESLSDSTVYMAYYTVAHYLHGDRYGNT 664
Query: 285 KNPLEENTVVGITNYIDFGFELQTSVDDAIAANNIS 320
PL D+ F + D+ I+ + IS
Sbjct: 665 PGPLNIKPEQMTDEVWDYIFTRRELSDELISKSGIS 700
>ref|YP_077442.1| Polysaccharide deacetylase,Carbohydrate Esterase Family 4 [Bacillus
licheniformis ATCC 14580] gi|52001862|gb|AAU21804.1|
Polysaccharide deacetylase,Carbohydrate Esterase Family
4 [Bacillus licheniformis ATCC 14580]
gi|52784013|ref|YP_089842.1| YbaN [Bacillus
licheniformis ATCC 14580] gi|52346515|gb|AAU39149.1|
YbaN [Bacillus licheniformis DSM 13]
Length = 254
Score = 34.7 bits (78), Expect = 6.2
Identities = 20/65 (30%), Positives = 35/65 (53%), Gaps = 2/65 (3%)
Query: 294 VGITNYIDFGFELQTSVDDAIAANNISDSTFQIAASWQANKNFLVK--AKAGPKSSTMAL 351
V +T I +G + + D + AN I D+TF ++ASW +V+ K G + +M
Sbjct: 59 VALTFNISWGDQKAMPILDTLKANGIKDATFFLSASWAERHPDVVERIRKDGHQIGSMGY 118
Query: 352 AFKSW 356
A+K++
Sbjct: 119 AYKNY 123
>ref|NP_859019.1| VP0 [Bovine kobuvirus]
Length = 367
Score = 34.7 bits (78), Expect = 6.2
Identities = 19/81 (23%), Positives = 38/81 (46%), Gaps = 3/81 (3%)
Query: 346 SSTMALAFKSWWKPSFTFSISATRDRADGQVQYGFGLQSESLR---EASYQRADPNFVML 402
S+T A FK WW+P+ + ++S D+A ++ L S++++ + PN + L
Sbjct: 62 STTHAQNFKHWWEPAASKALSRGVDKAPDGIEGAGKLASDAIKSKLSGARPGPSPNLIAL 121
Query: 403 TPSKEHLAEGIVWETGKRPML 423
PS ++ P++
Sbjct: 122 NPSSTQHGNAMITTGSTAPVV 142
>ref|NP_740257.1| unnamed protein product [Bovine kobuvirus]
gi|24817743|dbj|BAC23066.1| unnamed protein product
[Bovine kobuvirus]
Length = 2463
Score = 34.7 bits (78), Expect = 6.2
Identities = 19/81 (23%), Positives = 38/81 (46%), Gaps = 3/81 (3%)
Query: 346 SSTMALAFKSWWKPSFTFSISATRDRADGQVQYGFGLQSESLR---EASYQRADPNFVML 402
S+T A FK WW+P+ + ++S D+A ++ L S++++ + PN + L
Sbjct: 249 STTHAQNFKHWWEPAASKALSRGVDKAPDGIEGAGKLASDAIKSKLSGARPGPSPNLIAL 308
Query: 403 TPSKEHLAEGIVWETGKRPML 423
PS ++ P++
Sbjct: 309 NPSSTQHGNAMITTGSTAPVV 329
>gb|AAH80895.1| Hoxa1-prov protein [Xenopus tropicalis]
gi|56118550|ref|NP_001008017.1| hoxa1-prov protein
[Xenopus tropicalis]
Length = 323
Score = 34.7 bits (78), Expect = 6.2
Identities = 24/93 (25%), Positives = 41/93 (43%), Gaps = 1/93 (1%)
Query: 228 HHENAKLSNLMNWSCAMAYGVGSQSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNP 287
HH N +S + SC YG+ + SP F + E S+ F S Y + V++
Sbjct: 77 HHNNLGMS-YTHTSCGSNYGMQNFSPGYSHFPINQEADVSAGFPQSVYSGNIASSVVQHH 135
Query: 288 LEENTVVGITNYIDFGFELQTSVDDAIAANNIS 320
++ + G +YI + ++ A NN+S
Sbjct: 136 QHQSYIEGSAHYIHHSYGPDQNISVANYNNNVS 168
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.135 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 780,508,556
Number of Sequences: 2540612
Number of extensions: 31747710
Number of successful extensions: 75443
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 75421
Number of HSP's gapped (non-prelim): 14
length of query: 446
length of database: 863,360,394
effective HSP length: 131
effective length of query: 315
effective length of database: 530,540,222
effective search space: 167120169930
effective search space used: 167120169930
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Medicago: description of AC124951.10


