

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC124216.10 - phase: 0
(543 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
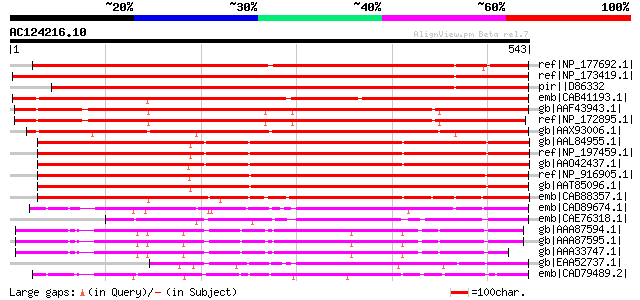
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_177692.1| glyoxal oxidase-related [Arabidopsis thaliana] ... 665 0.0
ref|NP_173419.1| glyoxal oxidase-related [Arabidopsis thaliana] ... 658 0.0
pir||D86332 hypothetical protein F6F9.4 [imported] - Arabidopsis... 638 0.0
emb|CAB41193.1| putative protein [Arabidopsis thaliana] gi|15230... 582 e-164
gb|AAF43943.1| Weak similarity to glyoxal oxidase (glx2) from Ph... 556 e-157
ref|NP_172895.1| glyoxal oxidase-related [Arabidopsis thaliana] 553 e-156
gb|AAX93006.1| probable galactose oxidase (EC 1.1.3.9) F15B8.190... 494 e-138
gb|AAL84955.1| AT5g19580/T20D1_100 [Arabidopsis thaliana] gi|247... 478 e-133
ref|NP_197459.1| glyoxal oxidase-related [Arabidopsis thaliana] 478 e-133
gb|AAO42437.1| putative glyoxal oxidase (glx1) [Arabidopsis thal... 470 e-131
ref|NP_916905.1| putative glyoxal oxidase [Oryza sativa (japonic... 447 e-124
gb|AAT85096.1| putative glyoxal oxidase [Oryza sativa (japonica ... 442 e-122
emb|CAB88357.1| putative protein [Arabidopsis thaliana] gi|26449... 426 e-118
emb|CAD89674.1| glyoxal oxidase [Botryotinia fuckeliana] 213 1e-53
emb|CAE76318.1| related to glyoxal oxidase precursor [Neurospora... 209 2e-52
gb|AAA87594.1| glyoxal oxidase precursor [Phanerochaete chrysosp... 206 2e-51
gb|AAA87595.1| glyoxal oxidase precursor [Phanerochaete chrysosp... 204 8e-51
gb|AAA33747.1| glyoxal oxidase 201 4e-50
gb|EAA52737.1| hypothetical protein MG05865.4 [Magnaporthe grise... 196 2e-48
emb|CAD79489.2| Glyoxaloxidase 2 [Ustilago maydis] gi|46096864|g... 189 3e-46
>ref|NP_177692.1| glyoxal oxidase-related [Arabidopsis thaliana]
gi|9369366|gb|AAF87115.1| F10A5.18 [Arabidopsis
thaliana]
Length = 547
Score = 665 bits (1716), Expect = 0.0
Identities = 328/526 (62%), Positives = 404/526 (76%), Gaps = 16/526 (3%)
Query: 25 VAAGN-GQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGLSKLPLPNGKCRHDPRETTV 83
VA+G+ G W++L ++GI AMH QLLHNDR++++DRT+FG S + LPNG CR P +
Sbjct: 29 VASGDEGTWELLLPNVGISAMHSQLLHNDRVIMYDRTNFGPSNISLPNGACRSSPGDAVS 88
Query: 84 KTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQTGGYNDGDRTIRMFDTC- 142
KTDCTAHSVEY++ N RPL VQ++ WCSSG V P GTL+QTGG DG+R +R+ D C
Sbjct: 89 KTDCTAHSVEYDVALNRIRPLTVQSNTWCSSGGVTPDGTLLQTGGDLDGERKVRLMDPCD 148
Query: 143 -NNCDWQEFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFYPKNNI-GVYRLPFLEQ 200
N+CDW E D GLAARRWYATNHILPDGRQIIIGGR QFNYEF+PK N Y +PFL +
Sbjct: 149 DNSCDWIEVDNGLAARRWYATNHILPDGRQIIIGGRGQFNYEFFPKTNAPNFYSIPFLSE 208
Query: 201 TNDAGAENNLYPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGS 260
TND G ENNLYPFV LN DGNLFIFANNRAIL DY+ N VVRT+P+IPGGDPRSYPS+GS
Sbjct: 209 TNDPGDENNLYPFVFLNTDGNLFIFANNRAILLDYSTNTVVRTYPEIPGGDPRSYPSTGS 268
Query: 261 GVLLPLKNLQSKFIEAEVLICGGAPKGSYQKASKREFLGALNTCARIKITDPNPTWVVET 320
VLLP+KNL EVL+CGGAPKGSY + + F+ AL+TCARI I D NP W+VE
Sbjct: 269 AVLLPIKNLV-----LEVLVCGGAPKGSYNLSWRNTFVKALDTCARININDVNPQWIVEK 323
Query: 321 MPRARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPS 380
MPRARVMGDM++LP+G+VL+ING SGTA WE GR+PVL+P LY + P+G+RFE+QNPS
Sbjct: 324 MPRARVMGDMMLLPDGNVLLINGGSSGTAAWELGREPVLHPDLYHPDKPVGSRFEVQNPS 383
Query: 381 HTPRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPR 440
PRMYHS A L+RDGR+LVGGSNPH YNF VLFPTEL +EAFSPSYL+ +++++RP
Sbjct: 384 TIPRMYHSIATLLRDGRILVGGSNPHAFYNFTGVLFPTELRLEAFSPSYLDTKYSSLRPS 443
Query: 441 IVASTSELQ-KHGQKLGLRFQVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLE---SN 496
IV + +G+ L LRF V + K+ V VTML P F THSFSM+QRLLVL+ S
Sbjct: 444 IVDPRPQTTVNYGRVLRLRFIVSGRV-KSPVKVTMLFPSFTTHSFSMHQRLLVLDHVISF 502
Query: 497 KVNIVEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQI 542
K+ I Y+V+V P S ILAPPG+Y++FVV+++IPSEG+W+++
Sbjct: 503 KLGI--SKIYEVRVRTPSSAILAPPGYYMVFVVNQDIPSEGLWVRL 546
>ref|NP_173419.1| glyoxal oxidase-related [Arabidopsis thaliana]
gi|16604507|gb|AAL24259.1| At1g19900/F6F9_4 [Arabidopsis
thaliana]
Length = 548
Score = 658 bits (1697), Expect = 0.0
Identities = 317/544 (58%), Positives = 400/544 (73%), Gaps = 6/544 (1%)
Query: 4 ATYLLFLLPILITTITTTPYLVAAGNGQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFG 63
AT++ + L + T ++ +A G W+ + ++GI AMHMQLLHNDR+V++DRT+FG
Sbjct: 5 ATFIFYALFLTTLQFLLTHHVSSAARGLWKYIAPNVGISAMHMQLLHNDRVVMYDRTNFG 64
Query: 64 LSKLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTL 123
S + LPNG CR +P++ K DCTAHS+EY++ +NT RPL VQ++ WCSSGSV P G L
Sbjct: 65 PSNISLPNGNCRDNPQDAVSKIDCTAHSIEYDVATNTIRPLTVQSNTWCSSGSVRPDGVL 124
Query: 124 VQTGGYNDGDRTIRMFDTCNN--CDWQEFDGGLAARRWYATNHILPDGRQIIIGGRKQFN 181
VQTGG DG+ R F CNN CDW E + GL RRWYA+NHILPDG+QI++GG+ QFN
Sbjct: 125 VQTGGDRDGELKTRTFSPCNNNQCDWVEMNNGLKKRRWYASNHILPDGKQIVMGGQGQFN 184
Query: 182 YEFYPKN-NIGVYRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNRAILFDYTKNVV 240
YEF+PK N V LPFL +T+D G ENNLYPFV +N DGNLF+FANNRAIL DY KN V
Sbjct: 185 YEFFPKTTNPNVVALPFLAETHDQGQENNLYPFVFMNTDGNLFMFANNRAILLDYVKNTV 244
Query: 241 VRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLICGGAPKGSYQKASKREFLGA 300
V+TFP IPGGDPR+YPS+GS VLLPLKNL++ +E EVL+CGGAPKGSY A K+ F+ A
Sbjct: 245 VKTFPAIPGGDPRNYPSTGSAVLLPLKNLEADNVETEVLVCGGAPKGSYNLARKKTFVKA 304
Query: 301 LNTCARIKITDPNPTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLN 360
L+TCARIKI D P W VE MP ARVMGDM+ LPNGDVL+ING GTA WE GR PVL
Sbjct: 305 LDTCARIKINDAKPEWAVEKMPHARVMGDMIPLPNGDVLLINGGSFGTAAWELGRTPVLA 364
Query: 361 PVLYKTNNPIGARFELQNPSHTPRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTEL 420
P LY NP+G+RFE P+ PRMYHS AIL+RDGRVLVGGSNPH YN+ VLFPTEL
Sbjct: 365 PDLYHPENPVGSRFESLRPTTIPRMYHSAAILLRDGRVLVGGSNPHAFYNYTGVLFPTEL 424
Query: 421 SIEAFSPSYLEPRFANVRPRIVA-STSELQKHGQKLGLRFQVKAALDKNLVYVTMLAPPF 479
S+EAFSP YL+ F+N+RP+I++ + K+G L L+F V + VTM+ P F
Sbjct: 425 SLEAFSPVYLQREFSNLRPKIISPEPQSMIKYGTNLKLKFSVTGEV-TTPAKVTMVFPTF 483
Query: 480 NTHSFSMNQRLLVLESNK-VNIVEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGI 538
THSF+MNQR+LVL++ K + Y+VQV P S +A PG+Y++FVV+++IPSEG+
Sbjct: 484 TTHSFAMNQRVLVLDNVKFTRKGKSPMYEVQVRTPRSANIAWPGYYMIFVVNQDIPSEGV 543
Query: 539 WIQI 542
W+++
Sbjct: 544 WVKL 547
>pir||D86332 hypothetical protein F6F9.4 [imported] - Arabidopsis thaliana
gi|10086483|gb|AAG12543.1| Unknown Protein [Arabidopsis
thaliana]
Length = 504
Score = 638 bits (1645), Expect = 0.0
Identities = 307/504 (60%), Positives = 379/504 (74%), Gaps = 6/504 (1%)
Query: 44 MHMQLLHNDRIVIFDRTDFGLSKLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRP 103
MHMQLLHNDR+V++DRT+FG S + LPNG CR +P++ K DCTAHS+EY++ +NT RP
Sbjct: 1 MHMQLLHNDRVVMYDRTNFGPSNISLPNGNCRDNPQDAVSKIDCTAHSIEYDVATNTIRP 60
Query: 104 LFVQTDVWCSSGSVNPKGTLVQTGGYNDGDRTIRMFDTCNN--CDWQEFDGGLAARRWYA 161
L VQ++ WCSSGSV P G LVQTGG DG+ R F CNN CDW E + GL RRWYA
Sbjct: 61 LTVQSNTWCSSGSVRPDGVLVQTGGDRDGELKTRTFSPCNNNQCDWVEMNNGLKKRRWYA 120
Query: 162 TNHILPDGRQIIIGGRKQFNYEFYPKN-NIGVYRLPFLEQTNDAGAENNLYPFVILNVDG 220
+NHILPDG+QI++GG+ QFNYEF+PK N V LPFL +T+D G ENNLYPFV +N DG
Sbjct: 121 SNHILPDGKQIVMGGQGQFNYEFFPKTTNPNVVALPFLAETHDQGQENNLYPFVFMNTDG 180
Query: 221 NLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLI 280
NLF+FANNRAIL DY KN VV+TFP IPGGDPR+YPS+GS VLLPLKNL++ +E EVL+
Sbjct: 181 NLFMFANNRAILLDYVKNTVVKTFPAIPGGDPRNYPSTGSAVLLPLKNLEADNVETEVLV 240
Query: 281 CGGAPKGSYQKASKREFLGALNTCARIKITDPNPTWVVETMPRARVMGDMVMLPNGDVLI 340
CGGAPKGSY A K+ F+ AL+TCARIKI D P W VE MP ARVMGDM+ LPNGDVL+
Sbjct: 241 CGGAPKGSYNLARKKTFVKALDTCARIKINDAKPEWAVEKMPHARVMGDMIPLPNGDVLL 300
Query: 341 INGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHTPRMYHSTAILVRDGRVLV 400
ING GTA WE GR PVL P LY NP+G+RFE P+ PRMYHS AIL+RDGRVLV
Sbjct: 301 INGGSFGTAAWELGRTPVLAPDLYHPENPVGSRFESLRPTTIPRMYHSAAILLRDGRVLV 360
Query: 401 GGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIVA-STSELQKHGQKLGLRF 459
GGSNPH YN+ VLFPTELS+EAFSP YL+ F+N+RP+I++ + K+G L L+F
Sbjct: 361 GGSNPHAFYNYTGVLFPTELSLEAFSPVYLQREFSNLRPKIISPEPQSMIKYGTNLKLKF 420
Query: 460 QVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNK-VNIVEGTTYDVQVTMPGSPIL 518
V + VTM+ P F THSF+MNQR+LVL++ K + Y+VQV P S +
Sbjct: 421 SVTGEV-TTPAKVTMVFPTFTTHSFAMNQRVLVLDNVKFTRKGKSPMYEVQVRTPRSANI 479
Query: 519 APPGFYLLFVVHKEIPSEGIWIQI 542
A PG+Y++FVV+++IPSEG+W+++
Sbjct: 480 AWPGYYMIFVVNQDIPSEGVWVKL 503
>emb|CAB41193.1| putative protein [Arabidopsis thaliana]
gi|15230360|ref|NP_191321.1| glyoxal oxidase-related
[Arabidopsis thaliana] gi|7485502|pir||T06758 probable
galactose oxidase (EC 1.1.3.9) F15B8.190 [similarity] -
Arabidopsis thaliana
Length = 547
Score = 582 bits (1500), Expect = e-164
Identities = 282/545 (51%), Positives = 384/545 (69%), Gaps = 15/545 (2%)
Query: 4 ATYLLFLLPILITTITTTPYLVAAGNGQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFG 63
AT +L L +++ P+L+ +W++L SIGI AMHMQLLHN +++FDRTDFG
Sbjct: 11 ATTILCLSMAILSEGQANPFLLQLD--RWEMLLPSIGISAMHMQLLHNGMVIMFDRTDFG 68
Query: 64 LSKLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTL 123
S + LP G CR+DP +T K DC+AHSV Y++ SNT+RPL VQTD WCSSG+V P GTL
Sbjct: 69 TSNVSLPGGICRYDPTDTAEKFDCSAHSVLYDVVSNTYRPLNVQTDTWCSSGAVLPNGTL 128
Query: 124 VQTGGYNDGDRTIRMFDTC---NNCDWQEFDGGLAARRWYATNHILPDGRQIIIGGRKQF 180
VQTGGYNDG+R RMF C + CDW EF L+ RRWYATN ILPDGR I++GGR+QF
Sbjct: 129 VQTGGYNDGERAARMFSPCGYSDTCDWIEFPQYLSQRRWYATNQILPDGRIIVVGGRRQF 188
Query: 181 NYEFYPKNN--IGVYRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNRAILFDYTKN 238
NYE +P+++ RL FL +T+D ENNLYPF+ L DGNLF+FAN R+I+FDY KN
Sbjct: 189 NYELFPRHDSRSRSSRLEFLRETSDGSNENNLYPFIHLLPDGNLFVFANTRSIVFDYKKN 248
Query: 239 VVVRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLICGGAPKGSYQKASKREFL 298
+V+ FP+IPGGDPR+YPSSGS +L PL + +E E+++CGG+PKG + R F
Sbjct: 249 RIVKEFPEIPGGDPRNYPSSGSSILFPLDDTNDANVEVEIMVCGGSPKGGF----SRGFT 304
Query: 299 GALNTCARIKITDPNPTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPV 358
A +TC R+K++D +P+W +ETMP RVMGDM++LP GDV+I+NGAG+GTAGWE RDP+
Sbjct: 305 RATSTCGRLKLSDQSPSWEMETMPLPRVMGDMLLLPTGDVIIVNGAGAGTAGWEKARDPI 364
Query: 359 LNPVLYKTNNPIGARFELQNPSHTPRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPT 418
+ PV+Y+ P F + + PRMYHS+AIL+ DGRVLVGGSNPH+ YNF NV +PT
Sbjct: 365 IQPVIYQ---PFDHLFTVMSTPSRPRMYHSSAILLPDGRVLVGGSNPHVYYNFTNVEYPT 421
Query: 419 ELSIEAFSPSYLEPRFANVRPRIVASTSELQKHGQKLGLRFQVKAALDKNLVYVTMLAPP 478
+LS+EA+SP YL +RP+I+ ++ ++ + + + F + L +L+ V ++AP
Sbjct: 422 DLSLEAYSPPYLFFTSDPIRPKILLTSDKVLSYKRLFNVDFSIAQFLTVDLLSVRIVAPS 481
Query: 479 FNTHSFSMNQRLLVLESNKVNIVEGT-TYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEG 537
F THSF+MNQR+++L+ V + T +Y V P + +APPG+Y++F+VH IPS
Sbjct: 482 FTTHSFAMNQRMVILKLLSVTRDQLTNSYRVSALGPSTAEIAPPGYYMIFLVHAGIPSSA 541
Query: 538 IWIQI 542
W+QI
Sbjct: 542 AWVQI 546
>gb|AAF43943.1| Weak similarity to glyoxal oxidase (glx2) from Phanerochaete
chrysosporium gb|L47287. [Arabidopsis thaliana]
gi|25352602|pir||H86278 F14L17.20 protein - Arabidopsis
thaliana
Length = 564
Score = 556 bits (1434), Expect = e-157
Identities = 282/555 (50%), Positives = 380/555 (67%), Gaps = 26/555 (4%)
Query: 6 YLLFLLPILITTITTTPYLVAAGNGQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGLS 65
+ FL + +P + G G+W +LQ S+GI AMHMQLLHN+++VIFDRTD+G S
Sbjct: 18 FFFFLCSTSDLLLPRSPLAILTG-GRWDLLQPSVGISAMHMQLLHNNKVVIFDRTDYGPS 76
Query: 66 KLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQ 125
+ LP+ C++ DC+AHS+ Y++ SNTFRPL ++ D WCSSGS+N G+L+Q
Sbjct: 77 NVSLPSQTCQN-----ATVFDCSAHSILYDVASNTFRPLTLRYDTWCSSGSLNASGSLIQ 131
Query: 126 TGGYNDGDRTIRMFDTCN------NCDWQEFDGGLAARRWYATNHILPDGRQIIIGGRKQ 179
TGGY +G+RT+R+F C+ +CDW E L++RRWY+TN ILPDGR II+GGR+
Sbjct: 132 TGGYGNGERTVRVFTPCDGGVGSVSCDWIENRAYLSSRRWYSTNQILPDGRIIIVGGRRA 191
Query: 180 FNYEFYPKN-NIGVYRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNRAILFDYTKN 238
FNYEFYPK+ V+ L FL +T D ENNLYPF+ L DGNLFIFAN R+ILFD+ +
Sbjct: 192 FNYEFYPKDPGESVFNLRFLAETRDPNEENNLYPFLHLLPDGNLFIFANRRSILFDFVNH 251
Query: 239 VVVRTFPQIPGGDPRSYPSSGSGVLLPL---KNLQSKFIEAEVLICGGAPKGSYQKASK- 294
+++ FPQIPGGD R+YPS+GS VLLPL ++ I AEV++CGGAP G++ KA++
Sbjct: 252 RIIKEFPQIPGGDKRNYPSTGSSVLLPLFLTGDINRTKITAEVMVCGGAPPGAFFKAART 311
Query: 295 --REFLGALNTCARIKITDPNPTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWE 352
+ F+ TC R+K+TDP+P WV+E MP RVM DM++LPNGDVLIINGA +GTAGWE
Sbjct: 312 IPKIFVAGSRTCGRLKVTDPDPKWVMEQMPSPRVMSDMLLLPNGDVLIINGAANGTAGWE 371
Query: 353 YGRDPVLNPVLYKTNNPIGA-RFELQNPSHTPRMYHSTAILVRDGRVLVGGSNPHIGYNF 411
+ VLNP+LY P RFE+ P+ PRMYHS ++L+ DGRVLVGGSNPH YNF
Sbjct: 372 DATNAVLNPILYLPEEPDQTRRFEILTPTRIPRMYHSASLLLSDGRVLVGGSNPHRNYNF 431
Query: 412 NNVLFPTELSIEAFSPSYLEPRFANVRPRIVASTSEL---QKHGQKLGLRFQVKA-ALDK 467
+PTELS+EA+ P YL+P++A VRP I+ T EL +GQ + F + A +
Sbjct: 432 TARPYPTELSLEAYLPRYLDPQYARVRPTII--TVELAGNMLYGQAFAVTFAIPAFGMFD 489
Query: 468 NLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPILAPPGFYLLF 527
V V ++AP F+THS +MNQRLLVL +V+ + Y V P + +APPG+Y++F
Sbjct: 490 GGVSVRLVAPSFSTHSTAMNQRLLVLRVRRVSQLSVFAYKADVDGPTNSYVAPPGYYMMF 549
Query: 528 VVHKEIPSEGIWIQI 542
VVH+ IPS +W++I
Sbjct: 550 VVHRGIPSVAVWVKI 564
>ref|NP_172895.1| glyoxal oxidase-related [Arabidopsis thaliana]
Length = 849
Score = 553 bits (1426), Expect = e-156
Identities = 281/552 (50%), Positives = 377/552 (67%), Gaps = 26/552 (4%)
Query: 6 YLLFLLPILITTITTTPYLVAAGNGQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGLS 65
+ FL + +P + G G+W +LQ S+GI AMHMQLLHN+++VIFDRTD+G S
Sbjct: 18 FFFFLCSTSDLLLPRSPLAILTG-GRWDLLQPSVGISAMHMQLLHNNKVVIFDRTDYGPS 76
Query: 66 KLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQ 125
+ LP+ C++ DC+AHS+ Y++ SNTFRPL ++ D WCSSGS+N G+L+Q
Sbjct: 77 NVSLPSQTCQN-----ATVFDCSAHSILYDVASNTFRPLTLRYDTWCSSGSLNASGSLIQ 131
Query: 126 TGGYNDGDRTIRMFDTCN------NCDWQEFDGGLAARRWYATNHILPDGRQIIIGGRKQ 179
TGGY +G+RT+R+F C+ +CDW E L++RRWY+TN ILPDGR II+GGR+
Sbjct: 132 TGGYGNGERTVRVFTPCDGGVGSVSCDWIENRAYLSSRRWYSTNQILPDGRIIIVGGRRA 191
Query: 180 FNYEFYPKN-NIGVYRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNRAILFDYTKN 238
FNYEFYPK+ V+ L FL +T D ENNLYPF+ L DGNLFIFAN R+ILFD+ +
Sbjct: 192 FNYEFYPKDPGESVFNLRFLAETRDPNEENNLYPFLHLLPDGNLFIFANRRSILFDFVNH 251
Query: 239 VVVRTFPQIPGGDPRSYPSSGSGVLLPL---KNLQSKFIEAEVLICGGAPKGSYQKASK- 294
+++ FPQIPGGD R+YPS+GS VLLPL ++ I AEV++CGGAP G++ KA++
Sbjct: 252 RIIKEFPQIPGGDKRNYPSTGSSVLLPLFLTGDINRTKITAEVMVCGGAPPGAFFKAART 311
Query: 295 --REFLGALNTCARIKITDPNPTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWE 352
+ F+ TC R+K+TDP+P WV+E MP RVM DM++LPNGDVLIINGA +GTAGWE
Sbjct: 312 IPKIFVAGSRTCGRLKVTDPDPKWVMEQMPSPRVMSDMLLLPNGDVLIINGAANGTAGWE 371
Query: 353 YGRDPVLNPVLYKTNNPIGA-RFELQNPSHTPRMYHSTAILVRDGRVLVGGSNPHIGYNF 411
+ VLNP+LY P RFE+ P+ PRMYHS ++L+ DGRVLVGGSNPH YNF
Sbjct: 372 DATNAVLNPILYLPEEPDQTRRFEILTPTRIPRMYHSASLLLSDGRVLVGGSNPHRNYNF 431
Query: 412 NNVLFPTELSIEAFSPSYLEPRFANVRPRIVASTSEL---QKHGQKLGLRFQVKA-ALDK 467
+PTELS+EA+ P YL+P++A VRP I+ T EL +GQ + F + A +
Sbjct: 432 TARPYPTELSLEAYLPRYLDPQYARVRPTII--TVELAGNMLYGQAFAVTFAIPAFGMFD 489
Query: 468 NLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPILAPPGFYLLF 527
V V ++AP F+THS +MNQRLLVL +V+ + Y V P + +APPG+Y++F
Sbjct: 490 GGVSVRLVAPSFSTHSTAMNQRLLVLRVRRVSQLSVFAYKADVDGPTNSYVAPPGYYMMF 549
Query: 528 VVHKEIPSEGIW 539
VVH+ IPS +W
Sbjct: 550 VVHRGIPSVAVW 561
>gb|AAX93006.1| probable galactose oxidase (EC 1.1.3.9) F15B8.190 [similarity] -
Arabidopsis thaliana [Oryza sativa (japonica
cultivar-group)]
Length = 577
Score = 494 bits (1271), Expect = e-138
Identities = 262/538 (48%), Positives = 346/538 (63%), Gaps = 20/538 (3%)
Query: 18 ITTTPYLVAAGNGQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGLSKLPLPNGKCRHD 77
+ P +V AG +WQ+L ++ G+ AMHMQLL D +++FDRTD G S + L
Sbjct: 40 VAQQPAVVLAG--EWQLLHQNTGVSAMHMQLLPGDYVLMFDRTDSGPSNISLDALSPCAA 97
Query: 78 PRETTVKT------DCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQTGGYND 131
T + DCTAHSV +++SN RP + T+ WCSS ++ P GTL+QTGG+++
Sbjct: 98 AATTALAAGGGGAVDCTAHSVLLDLRSNALRPYPLATNPWCSSAALLPNGTLLQTGGFSN 157
Query: 132 GDRTIRMFDTCNNCDWQEFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFYPKNNIG 191
GDR R+F W + LA RRWYAT+ +L DGR +I+GGR+QFN+EF+P ++
Sbjct: 158 GDRIARLFSPSTG--WVDLPSFLAVRRWYATDILLADGRVLILGGRRQFNFEFFPHDDAP 215
Query: 192 VYR---LPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFPQIP 248
+ PFLE+T D AE+NLYPF+ L D +F+FAN+RA++FD +R P IP
Sbjct: 216 APQPTLFPFLEETTDMDAEDNLYPFLHLLPDATVFVFANDRAVVFDPYNRAPLRRLPAIP 275
Query: 249 GGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLICGGAPKGSYQKASKR-EFLGALNTCARI 307
GG PR+YPSSGS VLLPL+ AEVL+CGGAP+G+Y+ A + F A TC RI
Sbjct: 276 GGVPRNYPSSGSSVLLPLRPDSPS--HAEVLVCGGAPRGAYRLALRNGTFAPADRTCGRI 333
Query: 308 KITDPNPTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTN 367
TD NP W +E MP R MGDMV+LP GDVLI+NGA +GTAGWE GR+PV PVLYK +
Sbjct: 334 APTDANPVWAMEEMPLPRAMGDMVLLPTGDVLIVNGAAAGTAGWELGREPVTYPVLYKPD 393
Query: 368 NPIGARFELQNPSHTPRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSP 427
+GARFE+ S PRMYHS+A L GRVLVGGSNPH+GY F+NV +PTELS+EAF P
Sbjct: 394 MQLGARFEVLAASTIPRMYHSSATLDTLGRVLVGGSNPHVGYVFDNVTYPTELSLEAFLP 453
Query: 428 SYLEPRFANVRPRIVASTSELQKHGQKLGLRFQVKAAL---DKNLVYVTMLAPPFNTHSF 484
Y + R VRPR+VA+ SE+ +G+ +RF+V V V +AP F THSF
Sbjct: 454 PYFDARLDGVRPRLVAAPSEV-GYGEAAAVRFEVPGGAVSGGPEEVRVAAVAPAFATHSF 512
Query: 485 SMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQI 542
MNQR++ L V + Y+ QV P SP +APPG+YL FV+H +PS W+++
Sbjct: 513 GMNQRVVSLAVGTVAQLAAGLYEAQVAAPPSPSVAPPGYYLWFVLHAGVPSTAAWVRM 570
>gb|AAL84955.1| AT5g19580/T20D1_100 [Arabidopsis thaliana]
gi|24797058|gb|AAN64541.1| At5g19580/T20D1_100
[Arabidopsis thaliana]
Length = 594
Score = 478 bits (1229), Expect = e-133
Identities = 246/522 (47%), Positives = 332/522 (63%), Gaps = 12/522 (2%)
Query: 30 GQWQVLQKSIGIVAMHMQLLHN-DRIVIFDRTDFGLSKLPLPNGKCRHDPRETTVKTDCT 88
G+W++ ++ G+ MH L+ +++ +D T + +SK+ LP G H T K DC
Sbjct: 77 GKWELFLENSGVSGMHAILMPVINKVQYYDATIWRISKIKLPPGVPCHVVDAKTNKVDCW 136
Query: 89 AHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQTGGYNDGDRTIRMFDTCNNCDWQ 148
AHS+ ++ + +PL + TD WCSSG + GTLV TGGY G T R +C NC W+
Sbjct: 137 AHSILMDVNTGALKPLGLSTDTWCSSGGLTVNGTLVSTGGYGGGANTARYLSSCENCKWE 196
Query: 149 EFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFYPK---NNIGVYRLPFLEQTNDAG 205
E+ LAA+RWY+T LPDG+ +IGGR NYE+ P+ NN ++ L QT+D
Sbjct: 197 EYPQALAAKRWYSTQATLPDGKFFVIGGRDALNYEYIPEEGQNNRKLFDSLLLRQTDDP- 255
Query: 206 AENNLYPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLP 265
ENNLYPFV LN DGNLFIFANNR+IL N V++ FPQ+PGG R+YP SGS LLP
Sbjct: 256 KENNLYPFVWLNTDGNLFIFANNRSILLSPKTNQVIKEFPQLPGG-ARNYPGSGSSALLP 314
Query: 266 LKNL--QSKFIEAEVLICGGAPKGSYQKASKREFLGALNTCARIKITDPNPTWVVETMPR 323
++ K I AEVL+CGG+ + +Y KA K+ + AL CARI+I P W E MP
Sbjct: 315 IQLYVKNPKVIPAEVLVCGGSKQDAYYKAGKKIYEPALQDCARIRINSAKPRWKTEMMPT 374
Query: 324 ARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHTP 383
R+M D V+LPNGD+L++NGA G +GW YG+DP P+LYK + G RF P+ P
Sbjct: 375 PRIMSDTVILPNGDILLVNGAKRGCSGWGYGKDPAFAPLLYKPHAARGKRFRQLKPTTIP 434
Query: 384 RMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIVA 443
RMYHS+AI++ DG+VLVGGSN + GY + NV FPTEL +E FSP YL+P AN+RP+IV
Sbjct: 435 RMYHSSAIILPDGKVLVGGSNTNDGYKY-NVEFPTELRVEKFSPPYLDPALANIRPKIVT 493
Query: 444 S-TSELQKHGQKLGLRFQVK-AALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIV 501
+ T + K+GQ ++ +K K + VTMLAP F THS SMN R+L+L N V
Sbjct: 494 TGTPKQVKYGQFFNVKVDLKEKGATKGNLKVTMLAPAFTTHSISMNMRMLILGVNNVK-P 552
Query: 502 EGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQIL 543
G YD+Q P + +APPG+YL+F ++K +PS G WIQ++
Sbjct: 553 AGAGYDIQAVAPPNGNIAPPGYYLIFAIYKGVPSTGEWIQVV 594
>ref|NP_197459.1| glyoxal oxidase-related [Arabidopsis thaliana]
Length = 594
Score = 478 bits (1229), Expect = e-133
Identities = 246/522 (47%), Positives = 332/522 (63%), Gaps = 12/522 (2%)
Query: 30 GQWQVLQKSIGIVAMHMQLLHN-DRIVIFDRTDFGLSKLPLPNGKCRHDPRETTVKTDCT 88
G+W++ ++ G+ MH L+ +++ +D T + +SK+ LP G H T K DC
Sbjct: 77 GKWELFLENSGVSGMHAILMPVINKVQYYDATIWRISKIKLPPGVPCHVVDAKTNKVDCW 136
Query: 89 AHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQTGGYNDGDRTIRMFDTCNNCDWQ 148
AHS+ ++ + +PL + TD WCSSG + GTLV TGGY G T R +C NC W+
Sbjct: 137 AHSILMDVNTGALKPLGLSTDTWCSSGGLTVNGTLVSTGGYGGGANTARYLSSCENCKWE 196
Query: 149 EFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFYPK---NNIGVYRLPFLEQTNDAG 205
E+ LAA+RWY+T LPDG+ +IGGR NYE+ P+ NN ++ L QT+D
Sbjct: 197 EYPQALAAKRWYSTQATLPDGKFFVIGGRDALNYEYIPEEGQNNRKLFDSLLLRQTDDP- 255
Query: 206 AENNLYPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLP 265
ENNLYPFV LN DGNLFIFANNR+IL N V++ FPQ+PGG R+YP SGS LLP
Sbjct: 256 EENNLYPFVWLNTDGNLFIFANNRSILLSPKTNQVIKEFPQLPGG-ARNYPGSGSSALLP 314
Query: 266 LKNL--QSKFIEAEVLICGGAPKGSYQKASKREFLGALNTCARIKITDPNPTWVVETMPR 323
++ K I AEVL+CGG+ + +Y KA K+ + AL CARI+I P W E MP
Sbjct: 315 IQLYVKNPKVIPAEVLVCGGSKQDAYYKAGKKIYEPALQDCARIRINSAKPRWKTEMMPT 374
Query: 324 ARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHTP 383
R+M D V+LPNGD+L++NGA G +GW YG+DP P+LYK + G RF P+ P
Sbjct: 375 PRIMSDTVILPNGDILLVNGAKRGCSGWGYGKDPAFAPLLYKPHAARGKRFRQLKPTTIP 434
Query: 384 RMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIVA 443
RMYHS+AI++ DG+VLVGGSN + GY + NV FPTEL +E FSP YL+P AN+RP+IV
Sbjct: 435 RMYHSSAIILPDGKVLVGGSNTNDGYKY-NVEFPTELRVEKFSPPYLDPALANIRPKIVT 493
Query: 444 S-TSELQKHGQKLGLRFQVK-AALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIV 501
+ T + K+GQ ++ +K K + VTMLAP F THS SMN R+L+L N V
Sbjct: 494 TGTPKQVKYGQFFNVKVDLKEKGATKGNLKVTMLAPAFTTHSISMNMRMLILGVNNVK-P 552
Query: 502 EGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQIL 543
G YD+Q P + +APPG+YL+F ++K +PS G WIQ++
Sbjct: 553 AGAGYDIQAVAPPNGNIAPPGYYLIFAIYKGVPSTGEWIQVV 594
>gb|AAO42437.1| putative glyoxal oxidase (glx1) [Arabidopsis thaliana]
gi|27754377|gb|AAO22637.1| putative glyoxal oxidase
(glx1) [Arabidopsis thaliana]
gi|15220398|ref|NP_176897.1| glyoxal oxidase-related
[Arabidopsis thaliana] gi|9828629|gb|AAG00252.1|
F1N21.11 [Arabidopsis thaliana]
Length = 615
Score = 470 bits (1210), Expect = e-131
Identities = 254/523 (48%), Positives = 329/523 (62%), Gaps = 12/523 (2%)
Query: 30 GQWQVLQKSIGIVAMHMQLLHN-DRIVIFDRTDFGLSKLPLPNGKCRHDPRETTVKTDCT 88
GQW++ K+ G+ AMH L+ +++ +D T + +S++ LP G H K DC
Sbjct: 96 GQWELFMKNSGVSAMHAILMPLINKVQFYDATIWRISQIKLPPGVPCHVFDAKKNKVDCW 155
Query: 89 AHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQTGGYNDGDRTIRMFDTCNNCDWQ 148
AHSV +I + +PL + TD WCSSG + GTLV TGG+ G T R TC NC W
Sbjct: 156 AHSVLVDINTGDIKPLALTTDTWCSSGGLTVNGTLVSTGGFQGGANTARYLSTCENCVWI 215
Query: 149 EFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFY---PKNNIGVYRLPFLEQTNDAG 205
E+ LAARRWY+T LPDG I++GGR NYE+ +NN +Y L QT+D
Sbjct: 216 EYPKALAARRWYSTQATLPDGTFIVVGGRDALNYEYILPEGQNNKKLYDSQLLRQTDDP- 274
Query: 206 AENNLYPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLP 265
ENNLYPFV LN DGNLFIFANNR+IL N V++ FPQ+PGG R+YP S S LLP
Sbjct: 275 EENNLYPFVWLNTDGNLFIFANNRSILLSPKTNKVLKEFPQLPGG-ARNYPGSASSALLP 333
Query: 266 LKNLQSK--FIEAEVLICGGAPKGSYQKASKREFLG-ALNTCARIKITDPNPTWVVETMP 322
++ I A+VL+CGGA + +Y +A + + AL CAR+ I P W ETMP
Sbjct: 334 IRLYVQNPAIIPADVLVCGGAKQDAYFRAERLKIYDWALKDCARLNINSAKPVWKTETMP 393
Query: 323 RARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHT 382
+RVM D V+LPNG++LIINGA G++GW ++P P+LYK N P+G RF+ PS
Sbjct: 394 TSRVMSDTVILPNGEILIINGAKRGSSGWHLAKEPNFAPLLYKPNKPLGQRFKELAPSTI 453
Query: 383 PRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIV 442
PR+YHS AI + DG+VLVGGSN + GY F NV +PTEL IE FSP YL+P AN+RPRIV
Sbjct: 454 PRVYHSIAIALPDGKVLVGGSNTNNGYQF-NVEYPTELRIEKFSPPYLDPALANMRPRIV 512
Query: 443 ASTSELQ-KHGQKLGLRFQVKAA-LDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNI 500
+ + Q K+GQ ++ ++K + K V VTMLAP F THS SMN RLL+L N V
Sbjct: 513 NTATPKQIKYGQMFDVKIELKQQNVAKENVMVTMLAPSFTTHSVSMNMRLLMLGINNVKN 572
Query: 501 VEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQIL 543
V G + +Q P S LAPPG+YLLF V+ +PS G WIQI+
Sbjct: 573 VGGDNHQIQAVAPPSGKLAPPGYYLLFAVYNGVPSVGEWIQIV 615
>ref|NP_916905.1| putative glyoxal oxidase [Oryza sativa (japonica cultivar-group)]
gi|20161084|dbj|BAB90014.1| glyoxal oxidase
precursor-like [Oryza sativa (japonica cultivar-group)]
Length = 624
Score = 447 bits (1150), Expect = e-124
Identities = 229/520 (44%), Positives = 320/520 (61%), Gaps = 10/520 (1%)
Query: 30 GQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGLSKLPLPNGKCRHDPRETTVKT-DCT 88
G W ++ ++ G+ AMH+ ++ + + ++FD G S + LP G+CR DPR DC
Sbjct: 106 GGWNLVSENSGVSAMHLVVMQHGKAIMFDTCTTGRSLMRLPPGRCRPDPRSKQPGAMDCW 165
Query: 89 AHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQTGGYNDGDRTIRMFDTCNNCDWQ 148
AH+VE++ + R L + TD WCSSG+ + G +VQTGG+ +GD+++R C CDW+
Sbjct: 166 AHAVEFDYNTGALRSLKIVTDTWCSSGAFDADGNMVQTGGFFEGDKSVRYLSACGTCDWK 225
Query: 149 EFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFYP---KNNIGVYRLPFLEQTNDAG 205
EF LA RWY T +LPDG I+IGGR+ F+YEF P + N L L T D
Sbjct: 226 EFPKSLADGRWYGTQLVLPDGSFIVIGGRRAFSYEFVPAAGRANARATPLRLLRDTTD-D 284
Query: 206 AENNLYPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLP 265
ENNLYPFV L DG LFIFAN+R+I+F+Y VVR P +PGG R+YP+S LLP
Sbjct: 285 VENNLYPFVNLLPDGTLFIFANDRSIVFNYRTGQVVRELPILPGGS-RNYPASAMSTLLP 343
Query: 266 LKNLQSKFIEAEVLICGGAPKGSYQKASKREFLGALNTCARIKITDPNPTWVVETMPRAR 325
L + + AEV+ICGGA K +++ F AL CARI + P W ++ MP R
Sbjct: 344 LDLRKGAGLSAEVIICGGATKNAFKLGETSTFPPALRDCARINPSKPGARWALDQMPSGR 403
Query: 326 VMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHTPRM 385
VMGD+++LP GD+L++NGA G +GW +GR +L+PVLY G RF + NPS+ PRM
Sbjct: 404 VMGDVLILPTGDLLMLNGAAKGCSGWGFGRQALLSPVLYSPYLRRGKRFRVLNPSNIPRM 463
Query: 386 YHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIVAST 445
YHST+ L+ D VLV GSN + YNF+ V FPTE+ +E F+P YL P+ + RP I A++
Sbjct: 464 YHSTSALLPDATVLVAGSNTNSAYNFSGVDFPTEVRVERFTPPYLSPQLSPNRPAIDAAS 523
Query: 446 --SELQKHGQKLGLRFQVKA-ALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVE 502
+ ++G + RF A + + VTM APPF TH +SMNQRLL+L +
Sbjct: 524 VPGDGMRYGARFTFRFTTPAQGVGQGDFKVTMYAPPFTTHGYSMNQRLLILPVTAF-AAQ 582
Query: 503 GTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQI 542
G + V V P P LAPPG+Y+++VV K +PS+ W+++
Sbjct: 583 GQRHTVTVDAPPKPELAPPGYYMVYVVAKGVPSKAAWVKM 622
>gb|AAT85096.1| putative glyoxal oxidase [Oryza sativa (japonica cultivar-group)]
Length = 622
Score = 442 bits (1138), Expect = e-122
Identities = 228/523 (43%), Positives = 323/523 (61%), Gaps = 13/523 (2%)
Query: 30 GQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGLSKLPLPNGKCRHDPRETTVKT-DCT 88
G W ++ ++ G+ AMH+ ++ + + ++FD + G S + LP CR DPR T DC
Sbjct: 102 GSWTIVSENSGVSAMHLAVMRHGKAIMFDTSTTGRSLMRLPMNNCRADPRAKREGTMDCW 161
Query: 89 AHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQTGGYNDGDRTIRMFDTCNNCDWQ 148
AH+VE++ + R L TD WCSSG+ + G L+QTGGY +GD+ +R D C+ CDW+
Sbjct: 162 AHAVEFDYSTGALRSLKTATDTWCSSGAFDADGNLIQTGGYFEGDKAVRRLDACDTCDWR 221
Query: 149 EFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFYPK---NNIGVYRLPFLEQTNDAG 205
E+ A RWYAT +LPDGR I+ GGR+ F+YEF P+ N + P L +T D
Sbjct: 222 EYPNSFAEGRWYATQQVLPDGRFIVFGGRRAFSYEFVPQPGMTNGQSIKFPLLRETTD-D 280
Query: 206 AENNLYPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLP 265
ENNLYPFV L DGNLF+FAN+R+++FD+ VVR P++ GG R++P+S +LP
Sbjct: 281 VENNLYPFVNLLPDGNLFVFANDRSVVFDHRTGKVVRELPKLAGGG-RNHPASAMSAMLP 339
Query: 266 L--KNL-QSKFIEAEVLICGGAPKGSYQKASKREFLGALNTCARIKITDPNPTWVVETMP 322
L +NL + E EV++CGGA K +++ + L CARI + + W VE MP
Sbjct: 340 LDLRNLTRGADPEPEVIVCGGALKTAFRLGENNTYQPTLRDCARINLGKIDAVWAVEAMP 399
Query: 323 RARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHT 382
RVMGD+++LP GD+L++NGA G++GW + R P+L+P+LY +P G+RF S
Sbjct: 400 VGRVMGDLLVLPTGDLLMLNGAAKGSSGWGFARQPILSPILYSPRHPEGSRFRPLAASTV 459
Query: 383 PRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRI- 441
RMYHST+ ++ D VLV G N + YNF+ V FPTE+ +E F+P YL R I
Sbjct: 460 ARMYHSTSAVLPDATVLVAGGNTNAAYNFSGVDFPTEVRVERFAPPYLSRELTGNRAVID 519
Query: 442 VAST-SELQKHGQKLGLRFQVK-AALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVN 499
VAS + ++G K RF AA++ V VTM APPF TH +SMNQRLLVL +
Sbjct: 520 VASVPAGGMRYGTKFTFRFHTPVAAVEWGDVRVTMYAPPFTTHGYSMNQRLLVLPVAGFS 579
Query: 500 IVEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQI 542
+G Y++ V P P LAPPG+YL++VV K++PSE W++I
Sbjct: 580 -AQGQMYELTVDTPRKPELAPPGYYLVYVVSKDVPSEAAWVKI 621
>emb|CAB88357.1| putative protein [Arabidopsis thaliana] gi|26449362|dbj|BAC41808.1|
unknown protein [Arabidopsis thaliana]
gi|27363378|gb|AAO11608.1| At3g53950/F5K20_250
[Arabidopsis thaliana] gi|15809876|gb|AAL06866.1|
AT3g53950/F5K20_250 [Arabidopsis thaliana]
gi|15232379|ref|NP_190963.1| glyoxal oxidase-related
[Arabidopsis thaliana] gi|11282416|pir||T45935 probable
galactose oxidase (EC 1.1.3.9) F5K20.250 [similarity] -
Arabidopsis thaliana
Length = 545
Score = 426 bits (1096), Expect = e-118
Identities = 231/522 (44%), Positives = 318/522 (60%), Gaps = 19/522 (3%)
Query: 30 GQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGLSKLPLPNGKCRHDPRETTVKTDCTA 89
G W+++ + GI +MH + + +++ DRT+ G S+ L +CR DP++ +K DC A
Sbjct: 34 GSWELIVQDAGIASMHTAVTRFNTVILLDRTNIGPSRKALDRHRCRRDPKDAALKRDCYA 93
Query: 90 HSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQTGGYNDGDRTIRMFDTCN---NCD 146
HSV +++ +N RPL +QTD WCSSG G+L+QTGG DG + IR F+ C+ CD
Sbjct: 94 HSVLFDLGTNQIRPLMIQTDTWCSSGQFLSDGSLLQTGGDKDGFKKIRKFEPCDPNETCD 153
Query: 147 WQEF-DGGLAARRWYATNHILPDGRQIIIGGRKQFNYEFYPKNNIGVYRLPFLEQTNDAG 205
W E D L RWYA+N ILPDG II+GGR E+YP G FL D
Sbjct: 154 WVELQDTELITGRWYASNQILPDGSVIIVGGRGTNTVEYYPPRENGAVPFQFLADVEDKQ 213
Query: 206 AENNLYPFVILNVD---GNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGV 262
+N LYP+V L D GNLFIFAN+RA+ +D+ N VV+ +P + GG PR+YPS GS
Sbjct: 214 MDN-LYPYVHLLPDDDGGNLFIFANSRAVKYDHRINAVVKEYPPLDGG-PRNYPSGGSSA 271
Query: 263 LLPLKNLQSKFIEAEVLICGGAPKGSYQKASKREFLGALNTCARIKITDPNPTWVVETMP 322
+L + Q F AE+LICGGA G++ ++ A TC RI T +P WV E MP
Sbjct: 272 MLAI---QGDFTTAEILICGGAQSGAF--TARAIDAPAHGTCGRIVATAADPVWVTEEMP 326
Query: 323 RARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHT 382
R+MGDMV LP G++LIINGA +G+ G+E G DP L P+LY+ + PIG RF NP
Sbjct: 327 FGRIMGDMVNLPTGEILIINGAQAGSQGFEMGSDPCLYPLLYRPDQPIGLRFMTLNPGTV 386
Query: 383 PRMYHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIV 442
PRMYHSTA L+ DGR+L+ GSNPH Y F N FPTEL IEAFSP YL P AN+RP I
Sbjct: 387 PRMYHSTANLLPDGRILLAGSNPHYFYKF-NAEFPTELRIEAFSPEYLSPDRANLRPEI- 444
Query: 443 ASTSELQKHGQKLGLRFQVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVE 502
++ ++G+ + V + ++ + + PF THSFS QRL+ L + ++ +
Sbjct: 445 QEIPQIIRYGEVFDVFVTVPLPV-VGIIQMNWGSAPFATHSFSQGQRLVKL-TVAPSVPD 502
Query: 503 GT-TYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQIL 543
G Y +Q T P + ++PPG+Y+ F V++ +PS WI+I+
Sbjct: 503 GVGRYRIQCTAPPNGAVSPPGYYMAFAVNQGVPSIARWIRIV 544
>emb|CAD89674.1| glyoxal oxidase [Botryotinia fuckeliana]
Length = 656
Score = 213 bits (543), Expect = 1e-53
Identities = 168/554 (30%), Positives = 253/554 (45%), Gaps = 62/554 (11%)
Query: 21 TPYLVAAGNGQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGLSKLPLPNGKCRHDPRE 80
+P A G + ++ S G+ AMH L+ N R++ D+ + ++L LPNG
Sbjct: 133 SPAPAAPVGGAFNIVGSS-GVPAMHAALMPNGRVMFLDKLE-NYTQLKLPNGYY------ 184
Query: 81 TTVKTDCTAHSVEYNIKSNTFR-PLFVQTDVWCSSGSVNPKGTLVQTGG----------Y 129
A S EY+ +N PL +T+ +CS G+ G +V GG
Sbjct: 185 --------AMSSEYDPATNAVATPLAYKTNAFCSGGTFLADGRVVSLGGNAPLDWLDPNI 236
Query: 130 NDGDRTIRMFD------TCNNCDWQEFDGGLAARRWYATNHILPDGRQIIIGGRKQFNYE 183
DG IR + + N DW E LA+ RWYAT + DG + G
Sbjct: 237 GDGFDAIRYLERSSTDASLNGKDWSEPGNKLASARWYATAQTMGDGTIFVAFGSLNGLDP 296
Query: 184 FYPKNNIGVYRLPFLEQTNDAGA-------ENN----LYPFVILNVDGNLFIFANNRAIL 232
NN Y + F G E N +YPFV L DGNLF+F + + +
Sbjct: 297 TVKTNNNPTYEI-FSATAVSQGKNIDMEILEKNQPYYMYPFVHLLNDGNLFVFVSKSSQV 355
Query: 233 FDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLICGGAPKGSYQKA 292
+ N +V+ P++ GD R+YP++G VLLPL + +++ICGG G+YQ
Sbjct: 356 LNVGTNTIVKELPEL-AGDYRTYPNTGGSVLLPLSSANKW--NPDIIICGG---GAYQDI 409
Query: 293 SKREFLGALNTCARIKITDPNPTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWE 352
+ +C RI+ NPTW ++ MP R M + +LP+G V+ +NG G G+
Sbjct: 410 TSP----TEPSCGRIQPLSANPTWELDAMPEGRGMVEGTLLPDGTVVWLNGGNLGAQGFG 465
Query: 353 YGRDPVLNPVLYKTNNPIGARFELQNPSHTPRMYHSTAILVRDGRVLVGGSNPHIGYNFN 412
+DP L +LY G RF S PR+YHS ++L+ DG ++V GSNP
Sbjct: 466 LAKDPTLEALLYDPTKAKGQRFSTLATSTIPRLYHSVSLLLLDGTLMVAGSNPVEMPKLQ 525
Query: 413 NVL---FPTELSIEAFSPSYLEPRFANVRP-RIVASTSELQKHGQKLGLRFQVKAALDKN 468
+ TE +E + P YL A RP + S+ + G L + F A
Sbjct: 526 PDAADPYVTEFRVENYVPPYLSGDNAKKRPTNVKLSSGSFKADGSTLDVTFDCPAG--AK 583
Query: 469 LVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPILAPPGFYLLFV 528
V VT+ F THS M R+L L+ N + T + VT P + +APPG Y++++
Sbjct: 584 AVTVTLYHGGFVTHSVHMGHRMLHLD-NTGFVAGATQQKLTVTRPPNNNVAPPGPYVVYI 642
Query: 529 VHKEIPSEGIWIQI 542
+ IP+ G ++ +
Sbjct: 643 LVDGIPAMGQFVTV 656
>emb|CAE76318.1| related to glyoxal oxidase precursor [Neurospora crassa]
gi|32422431|ref|XP_331659.1| hypothetical protein
[Neurospora crassa] gi|28926493|gb|EAA35466.1|
hypothetical protein [Neurospora crassa]
Length = 1105
Score = 209 bits (531), Expect = 2e-52
Identities = 152/457 (33%), Positives = 220/457 (47%), Gaps = 47/457 (10%)
Query: 101 FRPLFVQTDVWCSSGSVNPK--GTLVQTGGYNDGDRTIRMFDTCNNCDWQEF--DGGLAA 156
FR L ++TDV+C+ G P G + GG++ GD T DW+E + L A
Sbjct: 660 FRELHLKTDVFCAGGVTLPDKVGRQLTVGGWS-GDSTYGTRLYWPGHDWEENVNELSLQA 718
Query: 157 RRWYATNHILPDGRQIIIGGRKQFNYEFYPKNNIGVYR------LPFLEQTNDAGAENNL 210
RWY + I+ +G +IGG N P + Y + +LE+T+ NNL
Sbjct: 719 GRWYPSAMIMANGSIFVIGGETGSNAAAVPSIEVLPYTGTKPLFMEWLERTDP----NNL 774
Query: 211 YPFVILNVDGNLFIFANNRAILFDYTKNVVVRTFPQIPGG--DP---RSYPSSGSGVLLP 265
YPFV + G +F+ N A + D ++ P++PG DP R+YP G+ VLLP
Sbjct: 775 YPFVAVLPSGGIFVQYWNEARILDEKTFATIKVLPKVPGAVNDPTSGRTYPLEGAAVLLP 834
Query: 266 LKNLQSKFIEAEVLICGGAPKGSYQKASKREFLGALNTCARIKITDPNPTWVVETMPRAR 325
+ S+ + VLICGG+ G AL+ C + D NPTWV+E MP R
Sbjct: 835 QRYPYSENLG--VLICGGSNVGPGY---------ALDNCVSTRPDDANPTWVIERMPSFR 883
Query: 326 VMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNNPIGARFELQNPSHTPRM 385
VM M LP+G LI NGA G AG+ +P LN +LY P G+R + + RM
Sbjct: 884 VMPCMAPLPDGTYLIANGAHHGVAGFGLANNPNLNALLYDPTKPYGSRITVMANTTIARM 943
Query: 386 YHSTAILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIVAST 445
YHS AI + DGRV++ GS+P N P E +E F P YL N +PR +
Sbjct: 944 YHSEAITLLDGRVMISGSDPQDAVN------PEEYRVEVFVPPYL----LNGKPRPTFTL 993
Query: 446 SELQKHGQKLGLRFQVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVEGTT 505
+ + + F + AA + VT+L +TH SM R ++ V+ T
Sbjct: 994 ANRDWDWNQKTIPFTLGAAARNGAITVTLLGSVSSTHGNSMGARTIMPN------VQCTG 1047
Query: 506 YDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQI 542
V P + APPG+Y FV+ +P+ G++++I
Sbjct: 1048 TSCTVDAPPNAHTAPPGWYQFFVLDGGVPAVGVYVRI 1084
>gb|AAA87594.1| glyoxal oxidase precursor [Phanerochaete chrysosporium]
gi|542394|pir||A48296 glyoxal oxidase (EC 1.2.3.-)
precursor - basidiomycete (Phanerochaete chrysosporium)
Length = 559
Score = 206 bits (524), Expect = 2e-51
Identities = 176/577 (30%), Positives = 269/577 (46%), Gaps = 79/577 (13%)
Query: 7 LLFLLPILITTITTTPYLVAAGNGQWQVLQKS--IGIVAMHMQLLHNDRIVIFDRTDFGL 64
+L LL ++ T A+ W+ K GIVA+ ++++ +VIFDR G
Sbjct: 1 MLSLLAVVSLAAATLAAPAASDAPGWRFDLKPNLSGIVALEAIVVNSSLVVIFDRAT-GD 59
Query: 65 SKLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLV 124
L + NG+ + +++ ++T RPL V TD +C+SG++ GT+V
Sbjct: 60 QPLKI-NGE--------------STWGALWDLDTSTVRPLSVLTDSFCASGALLSNGTMV 104
Query: 125 QTGGYNDG----------DRTIRMFDTC-----NNCDWQEFDGG--LAARRWYATNHILP 167
GG G ++ IR+F+ C + C E L RWY ++ +
Sbjct: 105 SMGGTPGGTGGDVAAPPGNQAIRIFEPCASPSGDGCTLFEDPATVHLLEERWYPSSVRIF 164
Query: 168 DGRQIIIGGRKQF----------NYEFYPKNNIGVYRLPFLEQTNDAGAENNLYPFVILN 217
DG +IIGG ++EF+P FLE++ A NL+P
Sbjct: 165 DGSLMIIGGSHVLTPFYNVDPANSFEFFPSKEQTPRPSAFLERSLPA----NLFPRAFAL 220
Query: 218 VDGNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAE 277
DG +FI ANN++I++D KN P IP G + P GS +LLPL FI E
Sbjct: 221 PDGTVFIVANNQSIIYDIEKNTET-ILPDIPNGVRVTNPIDGSAILLPLS--PPDFIP-E 276
Query: 278 VLICGGAPKG-SYQKASKREFLGALNTCARIKITDPN--PTWVVETMPRARVMGDMVMLP 334
VL+CGG+ S S A + C+RIK+T W VE M AR+M ++V +P
Sbjct: 277 VLVCGGSTADTSLPSTSLSSQHPATSQCSRIKLTPEGIKAGWQVEHMLEARMMPELVHVP 336
Query: 335 NGDVLIINGAGSGTAGWEYGRD---------PVLNPVLYKTNNPIGARFELQNPSHT--P 383
NG +LI NGAG+G A D PVL P LY + P+G R T P
Sbjct: 337 NGQILITNGAGTGFAALSAVADPVGNSNADHPVLTPSLYTPDAPLGKRISNAGMPTTTIP 396
Query: 384 RMYHSTAILVRDGRVLVGGSNPHIGY---NFNNVLFPTELSIEAFSPSYLEPRFANVRPR 440
RMYHST L + G +GG+NP++ + + FP+EL IE P P RP
Sbjct: 397 RMYHSTVTLTQQGNFFIGGNNPNMNFTPPGTPGIKFPSELRIETLDP----PFMFRSRPA 452
Query: 441 IVASTSELQKHGQKLGLRFQVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNI 500
++ +L K GQK+ + + + L + V V ++ F++H+F + RL+ +ES+
Sbjct: 453 LLTMPEKL-KFGQKVTVPITIPSDLKASKVQVALMDLGFSSHAFHSSARLVFMESS---- 507
Query: 501 VEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEG 537
+ + T P + + PPG ++F+ ++ S G
Sbjct: 508 ISADRKSLTFTAPPNGRVFPPGPAVVFLTIDDVTSPG 544
>gb|AAA87595.1| glyoxal oxidase precursor [Phanerochaete chrysosporium]
Length = 559
Score = 204 bits (518), Expect = 8e-51
Identities = 175/577 (30%), Positives = 268/577 (46%), Gaps = 79/577 (13%)
Query: 7 LLFLLPILITTITTTPYLVAAGNGQWQVLQKS--IGIVAMHMQLLHNDRIVIFDRTDFGL 64
+L LL ++ T A+ W+ K GIVA+ ++++ +VIFDR G
Sbjct: 1 MLSLLAVVSLAAATLAAPAASDAPGWRFDLKPNLSGIVALEAIVVNSSLVVIFDRAT-GD 59
Query: 65 SKLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLV 124
L + NG+ + +++ ++T RPL V TD +C+SG++ GT+V
Sbjct: 60 QPLKI-NGE--------------STWGALWDLDTSTVRPLSVLTDSFCASGALLSNGTMV 104
Query: 125 QTGGYNDG----------DRTIRMFDTC-----NNCDWQEFDGG--LAARRWYATNHILP 167
GG G ++ IR+F+ C + C E L RWY ++ +
Sbjct: 105 SMGGTPGGTGGDVAAPPGNQAIRIFEPCASPSGDGCTLFEDPATVHLLEERWYPSSVRIF 164
Query: 168 DGRQIIIGGRKQF----------NYEFYPKNNIGVYRLPFLEQTNDAGAENNLYPFVILN 217
DG +IIGG ++EF+P FLE++ A NL+P
Sbjct: 165 DGSLMIIGGSHVLTPFYNVDPANSFEFFPSKEQTPRPSAFLERSLPA----NLFPRAFAL 220
Query: 218 VDGNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAE 277
DG +FI ANN++I++D KN P IP G + P GS +LLPL FI E
Sbjct: 221 PDGTVFIVANNQSIIYDIEKNTET-ILPDIPNGVRVTNPIDGSAILLPLS--PPDFIP-E 276
Query: 278 VLICGGAPKG-SYQKASKREFLGALNTCARIKITDPN--PTWVVETMPRARVMGDMVMLP 334
VL+CGG+ S S A + C+RI +T W VE M AR+M ++V +P
Sbjct: 277 VLVCGGSTADTSLPSTSLSSQHPATSQCSRITLTPEGIKAGWQVEHMLEARMMPELVHVP 336
Query: 335 NGDVLIINGAGSGTAGWEYGRD---------PVLNPVLYKTNNPIGARFELQNPSHT--P 383
NG +LI NGAG+G A D PVL P LY + P+G R T P
Sbjct: 337 NGQILITNGAGTGFAALSAVADPVGNSNADHPVLTPSLYTPDAPLGKRISNAGMPTTTIP 396
Query: 384 RMYHSTAILVRDGRVLVGGSNPHIGY---NFNNVLFPTELSIEAFSPSYLEPRFANVRPR 440
RMYHST L + G +GG+NP++ + + FP+EL IE P P RP
Sbjct: 397 RMYHSTVTLTQQGNFFIGGNNPNMNFTPPGTPGIKFPSELRIETLDP----PFMFRSRPA 452
Query: 441 IVASTSELQKHGQKLGLRFQVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNI 500
++ +L K GQK+ + + + L + V V ++ F++H+F + RL+ +ES+
Sbjct: 453 LLTMPEKL-KFGQKVTVPITIPSDLKASKVQVALMDLGFSSHAFHSSARLVFMESS---- 507
Query: 501 VEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEG 537
+ + T P + + PPG ++F+ ++ S G
Sbjct: 508 ISADRKSLTFTAPPNGRVFPPGPAVVFLTIDDVTSPG 544
>gb|AAA33747.1| glyoxal oxidase
Length = 529
Score = 201 bits (512), Expect = 4e-50
Identities = 173/562 (30%), Positives = 261/562 (45%), Gaps = 79/562 (14%)
Query: 7 LLFLLPILITTITTTPYLVAAGNGQWQVLQKS--IGIVAMHMQLLHNDRIVIFDRTDFGL 64
+L LL ++ T A+ W+ K GIVA+ ++++ +VIFDR G
Sbjct: 1 MLSLLAVVSLAAATLAAPAASDAPGWRFDLKPNLSGIVALEAIVVNSSLVVIFDRAT-GD 59
Query: 65 SKLPLPNGKCRHDPRETTVKTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLV 124
L + NG+ + +++ ++T RPL V TD +C+SG++ GT+V
Sbjct: 60 QPLKI-NGE--------------STWGALWDLDTSTVRPLSVLTDSFCASGALLSNGTMV 104
Query: 125 QTGGYNDG----------DRTIRMFDTC-----NNCDWQEFDGG--LAARRWYATNHILP 167
GG G ++ IR+F+ C + C E L RWY ++ +
Sbjct: 105 SMGGTPGGTGGDVAAPPGNQAIRIFEPCASPSGDGCTLFEDPATVHLLEERWYPSSVRIF 164
Query: 168 DGRQIIIGGRKQF----------NYEFYPKNNIGVYRLPFLEQTNDAGAENNLYPFVILN 217
DG +IIGG ++EF+P FLE++ A NL+P
Sbjct: 165 DGSLMIIGGSHVLTPFYNVDPANSFEFFPSKEQTPRPSAFLERSLPA----NLFPRAFAL 220
Query: 218 VDGNLFIFANNRAILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAE 277
DG +FI ANN++I++D KN P IP G + P GS +LLPL FI E
Sbjct: 221 PDGTVFIVANNQSIIYDIEKNTET-ILPDIPNGVRVTNPIDGSAILLPLS--PPDFIP-E 276
Query: 278 VLICGGAPKG-SYQKASKREFLGALNTCARIKITDPN--PTWVVETMPRARVMGDMVMLP 334
VL+CGG+ S S A + C+RIK+T W VE M AR+M ++V +P
Sbjct: 277 VLVCGGSTADTSLPSTSLSSQHPATSQCSRIKLTPEGIKAGWQVEHMLEARMMPELVHVP 336
Query: 335 NGDVLIINGAGSGTAGWEYGRD---------PVLNPVLYKTNNPIGARFELQNPSHT--P 383
NG +LI NGAG+G A D PVL P LY + P+G R T P
Sbjct: 337 NGQILITNGAGTGFAALSAVADPVGNSNADHPVLTPSLYTPDAPLGKRISNAGMPTTTIP 396
Query: 384 RMYHSTAILVRDGRVLVGGSNPHIGY---NFNNVLFPTELSIEAFSPSYLEPRFANVRPR 440
RMYHST L + G +GG+NP++ + + FP+EL IE P P RP
Sbjct: 397 RMYHSTVTLTQQGNFFIGGNNPNMNFTPPGTPGIKFPSELRIETLDP----PFMFRSRPA 452
Query: 441 IVASTSELQKHGQKLGLRFQVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNI 500
++ +L K GQK+ + + + L + V V ++ F++H+F + RL+ +ES+
Sbjct: 453 LLTMPEKL-KFGQKVTVPITIPSDLKASKVQVALMDLGFSSHAFHSSARLVFMESS---- 507
Query: 501 VEGTTYDVQVTMPGSPILAPPG 522
+ + T P + + PPG
Sbjct: 508 ISADRKSLTFTAPPNGRVFPPG 529
>gb|EAA52737.1| hypothetical protein MG05865.4 [Magnaporthe grisea 70-15]
gi|39976417|ref|XP_369599.1| hypothetical protein
MG05865.4 [Magnaporthe grisea 70-15]
Length = 445
Score = 196 bits (498), Expect = 2e-48
Identities = 140/421 (33%), Positives = 199/421 (47%), Gaps = 41/421 (9%)
Query: 147 WQEFDGGLAARRWYATNHILPDGRQIIIGG-----------RKQFNYEFYPKNNIG---V 192
W E L++ RWY + LPDGR + G YE +N +
Sbjct: 40 WDEPGNKLSSPRWYPSAQTLPDGRVFVASGSLNGLAPENASNNNPTYEMLDRNGVSSGQF 99
Query: 193 YRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNRAILFDY----TKNVVVRTFPQIP 248
++ LE+ +YPF+ L DGNLFIFA + +F T VV+ P++P
Sbjct: 100 VKMDLLERAQPY----YMYPFIHLLNDGNLFIFAAQISQIFSVGSGATTGTVVKEMPELP 155
Query: 249 GGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLICGGAPKGSYQKASKREFLGALNTCARIK 308
G D R+YP++G V+LPL ++LICGG P YQ + +C RIK
Sbjct: 156 G-DYRTYPNTGGSVMLPLSKANG--YTPDILICGGGP---YQDVTAP----TEPSCGRIK 205
Query: 309 ITDPNPTWVVETMPRARVMGDMVMLPNGDVLIINGAGSGTAGWEYGRDPVLNPVLYKTNN 368
D NP W ++ MP RVM + V++ +G V +NGA G G+ P +LY
Sbjct: 206 PLDANPKWEMDAMPDGRVMTEGVLMLDGTVFFVNGAHQGAQGFGVADKPAFTSLLYDPAQ 265
Query: 369 PIGARFELQNPSHTPRMYHSTAILVRDGRVLVGGSNP---HIGYNFNNVLFPTELSIEAF 425
P+G RF S PRMYHS +I++ D VL+ GSNP I + F TE +E +
Sbjct: 266 PLGQRFTTAATSTIPRMYHSVSIMLEDATVLIAGSNPVQQPILEVSADTPFATEFRVERY 325
Query: 426 SPSYLEPRFANVRPRIVASTSELQKHG----QKLGLRFQVKAALDKNLVYVTMLAPPFNT 481
+P YL N+RP + + G L +RF + +A K+ V V + + T
Sbjct: 326 TPPYLSNGKQNLRPLNMTLSGTNMTPGPAGSSVLNVRFGLPSATVKD-VKVALYYNGYVT 384
Query: 482 HSFSMNQRLLVLESNKVNIVEGTTYDVQVTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQ 541
HS M R++ LE V T ++ V P S + PPG+Y+LFV+ IPS G I
Sbjct: 385 HSVHMGHRMVYLEHTGF-AVGKTAQNLMVQPPPSNNITPPGYYILFVIADGIPSVGQQIM 443
Query: 542 I 542
I
Sbjct: 444 I 444
>emb|CAD79489.2| Glyoxaloxidase 2 [Ustilago maydis] gi|46096864|gb|EAK82097.1|
hypothetical protein UM00913.1 [Ustilago maydis 521]
gi|49068478|ref|XP_398528.1| hypothetical protein
UM00913.1 [Ustilago maydis 521]
Length = 625
Score = 189 bits (479), Expect = 3e-46
Identities = 166/573 (28%), Positives = 251/573 (42%), Gaps = 94/573 (16%)
Query: 25 VAAGN-GQWQVLQKSIGIVAMHMQLLHNDRIVIFDRTDFGLSKLPLPNGKCRHDPRETTV 83
V+AG G QV+ S G+ A M L ++ I D+T+ + NG
Sbjct: 27 VSAGKAGTMQVVGNS-GVSAQMMFLGTEQKVYILDKTENNPVSV---NGH---------- 72
Query: 84 KTDCTAHSVEYNIKSNTFRPLFVQTDVWCSSGSVNPKGTLVQTGG--------------- 128
A +VEY+I SN++RP+ V+++ +C+ G G+ + TGG
Sbjct: 73 ----PAWAVEYDINSNSYRPMEVRSNTFCAGGMTLGDGSWLVTGGNKAVTTNGATAKAGA 128
Query: 129 ----YNDGDRTIRMFDTCNN--CDWQEFDGG-LAARRWYATNHILPDGRQIIIGGRKQFN 181
YN G + +R C+N C W + + L RWY T L DG II+GG +
Sbjct: 129 GYGAYNGG-KALRFLSPCDNMQCQWNDQNSNQLNMERWYPTVEPLADGSNIILGGMRDGG 187
Query: 182 -----------YEFYPKNNIGV-YRLPFLEQTNDAGAENNLYPFVILNVDGNLFIFANNR 229
YEFYP + G LP L++T +LYP L G +FI A
Sbjct: 188 FVPSQGSNVPTYEFYPPKSGGASINLPILQRT----VPLSLYPIAYLMSSGEVFIQAGRE 243
Query: 230 AILFDYTKNVVVRTFPQIPGGDPRSYPSSGSGVLLPLKNLQSKFIEAEVLICGGAPKGSY 289
AIL++Y + R F +IPG PR YP+SG +LPL + +L CGG G
Sbjct: 244 AILWNYDQQSE-RAFAKIPGA-PRVYPASGGSAMLPLTPADD--YKETILFCGGTSLGKV 299
Query: 290 QKASKR-------EFLGALNTCARIKITDPNPTWVVETMPRARVMGDMVMLPNGDVLIIN 342
+ A +C +I V+ +P R MG + LP+G + N
Sbjct: 300 SNWGNEGGPSIPISQVPASTSCEQISPFQGGNWESVDDLPERRSMGQFINLPDGTLWFGN 359
Query: 343 GAGSGTAGWE-------------YGRDPVLNPVLYKTNNPIGARFELQNPSHTPRMYHST 389
G +G AG+ YG +P P++Y G R++ ++ R+YHS+
Sbjct: 360 GVTTGVAGYSTDPNSVGKPVGESYGDNPSYQPLVYDPKASRGNRWKRVGSTNIGRLYHSS 419
Query: 390 AILVRDGRVLVGGSNPHIGYNFNNVLFPTELSIEAFSPSYLEPRFANVRPRIVASTSELQ 449
A L+ D +LV GSNP+ N ++V + TE IE + P + + RP S
Sbjct: 420 ATLLPDSSILVAGSNPNADVN-HHVKWKTEYRIERWYPDF----YDQPRPSNDGLPSSFS 474
Query: 450 KHGQKLGLRFQVKAALDKNLVYVTMLAPPFNTHSFSMNQRLLVLESNKVNIVEGTTYDVQ 509
GQ G ++ +A V ++ F+TH +M QR++ L+S G+ V
Sbjct: 475 YGGQ--GFTIRLSSAAQAQKAKVVLIRTGFSTHGMNMGQRMIELKSTH----RGSKLYV- 527
Query: 510 VTMPGSPILAPPGFYLLFVVHKEIPSEGIWIQI 542
+P +P L PG L FVV +PS+G + +
Sbjct: 528 AQLPPNPNLFAPGPALAFVVVDGVPSQGKMVMV 560
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,022,536,974
Number of Sequences: 2540612
Number of extensions: 47506721
Number of successful extensions: 86931
Number of sequences better than 10.0: 79
Number of HSP's better than 10.0 without gapping: 49
Number of HSP's successfully gapped in prelim test: 30
Number of HSP's that attempted gapping in prelim test: 86437
Number of HSP's gapped (non-prelim): 147
length of query: 543
length of database: 863,360,394
effective HSP length: 133
effective length of query: 410
effective length of database: 525,458,998
effective search space: 215438189180
effective search space used: 215438189180
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC124216.10


