

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123570.11 + phase: 0
(398 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
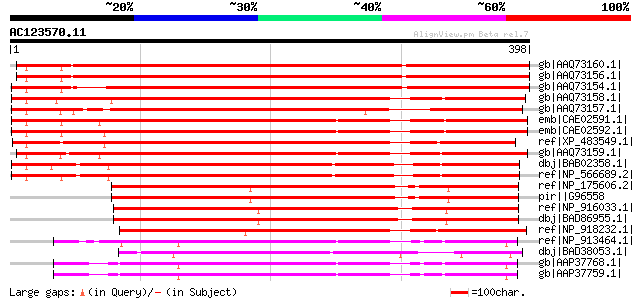
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ73160.1| LysM domain-containing receptor-like kinase 4 [Me... 617 e-175
gb|AAQ73156.1| LysM domain-containing receptor-like kinase 4 [Me... 617 e-175
gb|AAQ73154.1| LysM domain-containing receptor-like kinase 1 [Me... 565 e-160
gb|AAQ73158.1| LysM domain-containing receptor-like kinase 7 [Me... 422 e-117
gb|AAQ73157.1| LysM domain-containing receptor-like kinase 6 [Me... 417 e-115
emb|CAE02591.1| Nod-facor receptor 1a [Lotus corniculatus var. j... 400 e-110
emb|CAE02592.1| Nod-facor receptor 1b [Lotus corniculatus var. j... 399 e-110
ref|XP_483549.1| receptor protein kinase PERK1-like protein [Ory... 392 e-107
gb|AAQ73159.1| LysM domain-containing receptor-like kinase 3 [Me... 388 e-106
dbj|BAB02358.1| unnamed protein product [Arabidopsis thaliana] 387 e-106
ref|NP_566689.2| protein kinase family protein [Arabidopsis thal... 386 e-106
ref|NP_175606.2| protein kinase family protein / peptidoglycan-b... 240 6e-62
pir||G96558 probable protein kinase [imported] - Arabidopsis tha... 240 6e-62
ref|NP_916033.1| putative receptor protein kinase tmk1 precursor... 239 8e-62
dbj|BAD86955.1| putative Nod-factor receptor 1b [Oryza sativa (j... 239 8e-62
ref|NP_918232.1| protein kinase-like [Oryza sativa (japonica cul... 236 9e-61
ref|NP_913464.1| putative receptor protein kinase PERK1 [Oryza s... 214 4e-54
dbj|BAD38053.1| putative light repressible receptor protein kina... 213 8e-54
gb|AAP37768.1| At3g24600 [Arabidopsis thaliana] gi|13877617|gb|A... 213 1e-53
gb|AAP37759.1| At3g24550 [Arabidopsis thaliana] gi|22136228|gb|A... 213 1e-53
>gb|AAQ73160.1| LysM domain-containing receptor-like kinase 4 [Medicago truncatula]
Length = 496
Score = 617 bits (1591), Expect = e-175
Identities = 327/427 (76%), Positives = 354/427 (82%), Gaps = 37/427 (8%)
Query: 6 LLKRYN--VNFSQGSGLVYIRGKDGNGLYVPLPPK------------------------- 38
LL+ +N NFS+GSG+V+I G+D NG+YVPLP +
Sbjct: 62 LLQNFNQDANFSKGSGIVFIPGRDENGVYVPLPSRKAGHLARSLVAAGICIRGVCMVLLL 121
Query: 39 -------YFRKKNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPEFSYEELA 91
YFRKKNGEE SK PPEDSM+PSTKD DKD+ D SKYI VDKSP+FSY+ LA
Sbjct: 122 AICIYVRYFRKKNGEE-SKLPPEDSMSPSTKDGDKDSYSDTRSKYILVDKSPKFSYKVLA 180
Query: 92 NATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELKVLTSVHHWNLVH 151
NAT+NFSLAKKIGQGGFGEVYYG L G+K+AIKKMK QATREFLSELKVLTSV H NLVH
Sbjct: 181 NATENFSLAKKIGQGGFGEVYYGVLGGKKVAIKKMKTQATREFLSELKVLTSVRHLNLVH 240
Query: 152 LIGYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDHSVPV 211
LIGYCVEGFLFLVYEYMENGNLSQHLHNSEKE MTLS RMKIALDVARGLEYIHDHSVPV
Sbjct: 241 LIGYCVEGFLFLVYEYMENGNLSQHLHNSEKELMTLSRRMKIALDVARGLEYIHDHSVPV 300
Query: 212 YIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPENAYGRISR 271
YIHRDIKSDNILLN+NF GK+ADFGLTKLT+ ANS DNT H+AGTFGYMPPENAYGRISR
Sbjct: 301 YIHRDIKSDNILLNKNFNGKIADFGLTKLTNIANSTDNTNHMAGTFGYMPPENAYGRISR 360
Query: 272 KIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVMNQTG 331
K+DVYAFGVVLYELISAKAAVI IDK EFE S EIKTNES DEYKSLVALFDEVM+Q G
Sbjct: 361 KMDVYAFGVVLYELISAKAAVIMIDKNEFE--SHEIKTNESTDEYKSLVALFDEVMDQKG 418
Query: 332 DCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELNSSIDN 391
D I+ LRKLVDPRLG NYSIDSISKMAKLAKACINRDPKQRP MRDVVVSLM+L S+ID+
Sbjct: 419 DPIEGLRKLVDPRLGDNYSIDSISKMAKLAKACINRDPKQRPKMRDVVVSLMKLISTIDD 478
Query: 392 KNSTGSA 398
++ T SA
Sbjct: 479 ESRTDSA 485
>gb|AAQ73156.1| LysM domain-containing receptor-like kinase 4 [Medicago truncatula]
Length = 624
Score = 617 bits (1591), Expect = e-175
Identities = 327/427 (76%), Positives = 354/427 (82%), Gaps = 37/427 (8%)
Query: 6 LLKRYN--VNFSQGSGLVYIRGKDGNGLYVPLPPK------------------------- 38
LL+ +N NFS+GSG+V+I G+D NG+YVPLP +
Sbjct: 190 LLQNFNQDANFSKGSGIVFIPGRDENGVYVPLPSRKAGHLARSLVAAGICIRGVCMVLLL 249
Query: 39 -------YFRKKNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPEFSYEELA 91
YFRKKNGEE SK PPEDSM+PSTKD DKD+ D SKYI VDKSP+FSY+ LA
Sbjct: 250 AICIYVRYFRKKNGEE-SKLPPEDSMSPSTKDGDKDSYSDTRSKYILVDKSPKFSYKVLA 308
Query: 92 NATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELKVLTSVHHWNLVH 151
NAT+NFSLAKKIGQGGFGEVYYG L G+K+AIKKMK QATREFLSELKVLTSV H NLVH
Sbjct: 309 NATENFSLAKKIGQGGFGEVYYGVLGGKKVAIKKMKTQATREFLSELKVLTSVRHLNLVH 368
Query: 152 LIGYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDHSVPV 211
LIGYCVEGFLFLVYEYMENGNLSQHLHNSEKE MTLS RMKIALDVARGLEYIHDHSVPV
Sbjct: 369 LIGYCVEGFLFLVYEYMENGNLSQHLHNSEKELMTLSRRMKIALDVARGLEYIHDHSVPV 428
Query: 212 YIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPENAYGRISR 271
YIHRDIKSDNILLN+NF GK+ADFGLTKLT+ ANS DNT H+AGTFGYMPPENAYGRISR
Sbjct: 429 YIHRDIKSDNILLNKNFNGKIADFGLTKLTNIANSTDNTNHMAGTFGYMPPENAYGRISR 488
Query: 272 KIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVMNQTG 331
K+DVYAFGVVLYELISAKAAVI IDK EFE S EIKTNES DEYKSLVALFDEVM+Q G
Sbjct: 489 KMDVYAFGVVLYELISAKAAVIMIDKNEFE--SHEIKTNESTDEYKSLVALFDEVMDQKG 546
Query: 332 DCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELNSSIDN 391
D I+ LRKLVDPRLG NYSIDSISKMAKLAKACINRDPKQRP MRDVVVSLM+L S+ID+
Sbjct: 547 DPIEGLRKLVDPRLGDNYSIDSISKMAKLAKACINRDPKQRPKMRDVVVSLMKLISTIDD 606
Query: 392 KNSTGSA 398
++ T SA
Sbjct: 607 ESRTDSA 613
>gb|AAQ73154.1| LysM domain-containing receptor-like kinase 1 [Medicago truncatula]
Length = 590
Score = 565 bits (1457), Expect = e-160
Identities = 304/428 (71%), Positives = 338/428 (78%), Gaps = 56/428 (13%)
Query: 2 IEIELLKRYN--VNFSQGSGLVYIRGKDGNGLYVPLPPK--------------------- 38
++ ELL++Y VNFS+GSGLV+I GKD NG+YVPLP
Sbjct: 177 VKPELLQKYTPGVNFSKGSGLVFIPGKDKNGVYVPLPHGKAGHLARSLATAVGGTCTVLL 236
Query: 39 --------YFRKKNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPEFSYEEL 90
YFR KN +E SK P SKYI VDKSP+FSYEEL
Sbjct: 237 LAISIYAIYFRNKNAKE-SKLP---------------------SKYIVVDKSPKFSYEEL 274
Query: 91 ANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELKVLTSVHHWNLV 150
ANATD FSLA KIGQGGFGEVYYGE RG+K AIKKMKMQATREFL+ELK+LT VHH NLV
Sbjct: 275 ANATDKFSLANKIGQGGFGEVYYGEPRGKKTAIKKMKMQATREFLAELKILTRVHHCNLV 334
Query: 151 HLIGYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDHSVP 210
HLIGYCVEG LFLVYEY++NGNLSQ+LH+SE+ PMT STRM+IALDVARGLEYIH+HSVP
Sbjct: 335 HLIGYCVEGSLFLVYEYIDNGNLSQNLHDSERGPMTWSTRMQIALDVARGLEYIHEHSVP 394
Query: 211 VYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPENAYGRIS 270
VYIHRDIKSDNILLNENFTGK+ADFGLT+LTD+ANS DNT+HVAGTFGYMPPEN YGRIS
Sbjct: 395 VYIHRDIKSDNILLNENFTGKIADFGLTRLTDSANSTDNTLHVAGTFGYMPPENVYGRIS 454
Query: 271 RKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVMNQT 330
RKIDVYAFGVVLYELISAK AVIKIDKTEFE EI+TNESIDEYKSLVALFDEV++Q
Sbjct: 455 RKIDVYAFGVVLYELISAKPAVIKIDKTEFE---SEIRTNESIDEYKSLVALFDEVIDQK 511
Query: 331 GDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELNSSID 390
GD I+ LR LVDPRL NYSIDSISKMAKLA+AC+NRDPK+RPTMR VVVSLM LNS+ID
Sbjct: 512 GDPIEGLRNLVDPRLEDNYSIDSISKMAKLARACLNRDPKRRPTMRAVVVSLMTLNSTID 571
Query: 391 NKNSTGSA 398
+ + + SA
Sbjct: 572 DGSRSASA 579
>gb|AAQ73158.1| LysM domain-containing receptor-like kinase 7 [Medicago truncatula]
Length = 620
Score = 422 bits (1085), Expect = e-117
Identities = 243/436 (55%), Positives = 292/436 (66%), Gaps = 57/436 (13%)
Query: 2 IEIELLKRYN--VNFSQGSGLVYIRGKDGNGLYVP------------------------- 34
I+ ELL++YN VNFSQGSGLVYI GKD N YVP
Sbjct: 185 IDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHISTGGLSGGVITGISVGAVAGLI 244
Query: 35 -----LPPKYFRKKNGEEESKFPPEDS-MTPSTKDVDKDTNDDNGSKY---------IWV 79
+ Y+RKK ++ E S + K+ + N G+ I V
Sbjct: 245 LLSFCIYVTYYRKKKIRKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPANMIGIRV 304
Query: 80 DKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELK 139
+KS EFSYEELANAT+NF++A KIGQGGFGEVYY EL G+K AIKKM M+AT+EFL+ELK
Sbjct: 305 EKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKATKEFLAELK 364
Query: 140 VLTSVHHWNLVHLIGYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVAR 199
VLT VHH NLV LIGYCVEG LFLVYEY++NGNL QHL +S+ EP++ S R+KIALD AR
Sbjct: 365 VLTRVHHVNLVRLIGYCVEGSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSAR 424
Query: 200 GLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGY 259
GLEYIH+H+VP YIHRDIKS+NILL++NF KVADFGLTKL DA S+ TV++AGTFGY
Sbjct: 425 GLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGY 484
Query: 260 MPPENAYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSL 319
MPPE AYG +S KIDVYAFGVVLYELISAKAAV I +S + K L
Sbjct: 485 MPPEYAYGSVSSKIDVYAFGVVLYELISAKAAV--------------IMGEDSGADLKGL 530
Query: 320 VALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV 379
V LF+EV +Q I+ L+KLVDPRLG NY ID + KMA+LAK C N DP+QRP M VV
Sbjct: 531 VVLFEEVFDQPHP-IEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVV 589
Query: 380 VSLMELNSSIDNKNST 395
V+L L S+ ++ + T
Sbjct: 590 VALTTLTSTTEDWDIT 605
>gb|AAQ73157.1| LysM domain-containing receptor-like kinase 6 [Medicago truncatula]
Length = 574
Score = 417 bits (1071), Expect = e-115
Identities = 236/426 (55%), Positives = 287/426 (66%), Gaps = 72/426 (16%)
Query: 2 IEIELLKRYN--VNFSQGSGLVYIRGKDGNGLYVPLPP---------------------- 37
IE ELL+RYN VNFS+GSGLV+I GKD NG Y+PL P
Sbjct: 170 IEAELLQRYNPGVNFSKGSGLVFIPGKDQNGSYLPLHPSTVGTVAITGISVGVLAALLLL 229
Query: 38 ------KYFRKKNGEE--ESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPEFSYEE 89
KY+ KK ++ E +DS K TN + I V+KS EFSY+E
Sbjct: 230 LFFVYIKYYLKKKNKKTWEKNLILDDS---KMKSAQIGTNIAS----IMVEKSEEFSYKE 282
Query: 90 LANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELKVLTSVHHWNL 149
L+ AT+NFS+A KIG+GGFGEV+Y ELRGQK AIKKMKM+A++EF +ELKVLT VHH NL
Sbjct: 283 LSIATNNFSMANKIGEGGFGEVFYAELRGQKAAIKKMKMKASKEFCAELKVLTLVHHLNL 342
Query: 150 VHLIGYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDHSV 209
V LIGYCVEGFLFLVYEY++NGNLSQ+LH+SE+EP++ STRM+IALD ARGLEYIH+H+V
Sbjct: 343 VGLIGYCVEGFLFLVYEYIDNGNLSQNLHDSEREPLSWSTRMQIALDSARGLEYIHEHTV 402
Query: 210 PVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPENAYGRI 269
PVYIHRDIKS+NILL+++F KVADFGL+KL D NS +T+ GTFGYMPPE A G +
Sbjct: 403 PVYIHRDIKSENILLDKSFCAKVADFGLSKLADVGNSTSSTIVAEGTFGYMPPEYACGSV 462
Query: 270 SR--KIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVM 327
S K+DVYAFGVVLYELISAKAA FDEV
Sbjct: 463 SSSPKVDVYAFGVVLYELISAKAA-------------------------------FDEVF 491
Query: 328 NQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELNS 387
D + ++ LVDPRLG NYSIDS+ KMA+LAKAC RDP+ RP+MR +VV+LM L S
Sbjct: 492 GYDQDPTEGIKNLVDPRLGDNYSIDSVCKMAQLAKACTMRDPQLRPSMRSIVVALMTLTS 551
Query: 388 SIDNKN 393
+ ++ N
Sbjct: 552 TTEDWN 557
>emb|CAE02591.1| Nod-facor receptor 1a [Lotus corniculatus var. japonicus]
gi|37651058|emb|CAE02589.1| Nod-factor receptor 1a
[Lotus corniculatus var. japonicus]
Length = 621
Score = 400 bits (1029), Expect = e-110
Identities = 232/439 (52%), Positives = 288/439 (64%), Gaps = 59/439 (13%)
Query: 2 IEIELLKRYN--VNFSQGSGLVYIRGKDGNGLYVPLPPK--------------------- 38
++ L++ +N VNFS+ SG+ +I G+ NG+YVPL +
Sbjct: 186 LDAGLIQSFNPSVNFSKDSGIAFIPGRYKNGVYVPLYHRTAGLASGAAVGISIAGTFVLL 245
Query: 39 ------YFR-KKNGEEESKFPPEDSMTPSTKDVDKD------------TNDDNGSKYIWV 79
Y R +K EE++K P + SM ST+D T G I V
Sbjct: 246 LLAFCMYVRYQKKEEEKAKLPTDISMALSTQDASSSAEYETSGSSGPGTASATGLTSIMV 305
Query: 80 DKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELK 139
KS EFSY+ELA AT+NFSL KIGQGGFG VYY ELRG+K AIKKM +QA+ EFL ELK
Sbjct: 306 AKSMEFSYQELAKATNNFSLDNKIGQGGFGAVYYAELRGKKTAIKKMDVQASTEFLCELK 365
Query: 140 VLTSVHHWNLVHLIGYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVAR 199
VLT VHH NLV LIGYCVEG LFLVYE+++NGNL Q+LH S KEP+ S+R++IALD AR
Sbjct: 366 VLTHVHHLNLVRLIGYCVEGSLFLVYEHIDNGNLGQYLHGSGKEPLPWSSRVQIALDAAR 425
Query: 200 GLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGY 259
GLEYIH+H+VPVYIHRD+KS NIL+++N GKVADFGLTKL + NS T + GTFGY
Sbjct: 426 GLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQT-RLVGTFGY 484
Query: 260 MPPENA-YGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKS 318
MPPE A YG IS KIDVYAFGVVL+ELISAK AV +KT E + E K
Sbjct: 485 MPPEYAQYGDISPKIDVYAFGVVLFELISAKNAV--------------LKTGELVAESKG 530
Query: 319 LVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDV 378
LVALF+E +N++ C D LRKLVDPRLG NY IDS+ K+A+L +AC +P RP+MR +
Sbjct: 531 LVALFEEALNKSDPC-DALRKLVDPRLGENYPIDSVLKIAQLGRACTRDNPLLRPSMRSL 589
Query: 379 VVSLMELNSSIDNKNSTGS 397
VV+LM L+S ++ + S
Sbjct: 590 VVALMTLSSLTEDCDDESS 608
>emb|CAE02592.1| Nod-facor receptor 1b [Lotus corniculatus var. japonicus]
gi|37651060|emb|CAE02590.1| Nod-factor receptor 1b
[Lotus corniculatus var. japonicus]
Length = 623
Score = 399 bits (1025), Expect = e-110
Identities = 231/441 (52%), Positives = 291/441 (65%), Gaps = 61/441 (13%)
Query: 2 IEIELLKRYN--VNFSQGSGLVYIRGKDGNGLYVPLPPK--------------------- 38
++ L++ +N VNFS+ SG+ +I G+ NG+YVPL +
Sbjct: 186 LDAGLIQSFNPSVNFSKDSGIAFIPGRYKNGVYVPLYHRTAGLASGAAVGISIAGTFVLL 245
Query: 39 ------YFR-KKNGEEESKFPPEDSMTPSTKDVDKDTNDD--------------NGSKYI 77
Y R +K EE++K P + SM ST+D + ++ + G I
Sbjct: 246 LLAFCMYVRYQKKEEEKAKLPTDISMALSTQDGNASSSAEYETSGSSGPGTASATGLTSI 305
Query: 78 WVDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSE 137
V KS EFSY+ELA AT+NFSL KIGQGGFG VYY ELRG+K AIKKM +QA+ EFL E
Sbjct: 306 MVAKSMEFSYQELAKATNNFSLDNKIGQGGFGAVYYAELRGKKTAIKKMDVQASTEFLCE 365
Query: 138 LKVLTSVHHWNLVHLIGYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDV 197
LKVLT VHH NLV LIGYCVEG LFLVYE+++NGNL Q+LH S KEP+ S+R++IALD
Sbjct: 366 LKVLTHVHHLNLVRLIGYCVEGSLFLVYEHIDNGNLGQYLHGSGKEPLPWSSRVQIALDA 425
Query: 198 ARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTF 257
ARGLEYIH+H+VPVYIHRD+KS NIL+++N GKVADFGLTKL + NS T + GTF
Sbjct: 426 ARGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQT-RLVGTF 484
Query: 258 GYMPPENA-YGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEY 316
GYMPPE A YG IS KIDVYAFGVVL+ELISAK AV +KT E + E
Sbjct: 485 GYMPPEYAQYGDISPKIDVYAFGVVLFELISAKNAV--------------LKTGELVAES 530
Query: 317 KSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMR 376
K LVALF+E +N++ C D LRKLVDPRLG NY IDS+ K+A+L +AC +P RP+MR
Sbjct: 531 KGLVALFEEALNKSDPC-DALRKLVDPRLGENYPIDSVLKIAQLGRACTRDNPLLRPSMR 589
Query: 377 DVVVSLMELNSSIDNKNSTGS 397
+VV+LM L+S ++ + S
Sbjct: 590 SLVVALMTLSSLTEDCDDESS 610
>ref|XP_483549.1| receptor protein kinase PERK1-like protein [Oryza sativa (japonica
cultivar-group)] gi|38175551|dbj|BAD01244.1| receptor
protein kinase PERK1-like protein [Oryza sativa
(japonica cultivar-group)] gi|50725672|dbj|BAD33138.1|
receptor protein kinase PERK1-like protein [Oryza sativa
(japonica cultivar-group)]
Length = 594
Score = 392 bits (1007), Expect = e-107
Identities = 219/394 (55%), Positives = 274/394 (68%), Gaps = 24/394 (6%)
Query: 3 EIELLKRYN--VNFSQGSGLVYIRGKDGNGLYVPLPPKYFRKKNGEEESKFPPEDSMTPS 60
++++++RYN + + GSG+VYI KD NG Y+PL R+K + EDS
Sbjct: 195 QLDVVRRYNPGMESATGSGIVYIPVKDPNGSYLPLKSPG-RRKAKQATLLQSSEDSTQLG 253
Query: 61 TKDVDKDTNDD----NGSKYIWVDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGEL 116
T +DK T + I VDKS EFSYEEL+NAT FS+ KIGQGGFG VYY EL
Sbjct: 254 TISMDKVTPSTIVGPSPVAGITVDKSVEFSYEELSNATQGFSIGNKIGQGGFGAVYYAEL 313
Query: 117 RGQKIAIKKMKMQATREFLSELKVLTSVHHWNLVHLIGYCVEGFLFLVYEYMENGNLSQH 176
RG+K AIKKM MQAT EFL+ELKVLT VHH NLV LIGYC+E LFLVYE++ENGNLSQH
Sbjct: 314 RGEKAAIKKMDMQATHEFLAELKVLTHVHHLNLVRLIGYCIESSLFLVYEFIENGNLSQH 373
Query: 177 LHNSEKEPMTLSTRMKIALDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFG 236
L EP++ + R++IALD ARGLEYIH+H+VPVYIHRDIKS NIL+++N+ KVADFG
Sbjct: 374 LRGMGYEPLSWAARIQIALDSARGLEYIHEHTVPVYIHRDIKSANILIDKNYRAKVADFG 433
Query: 237 LTKLTDAANSADNT-VHVAGTFGYMPPENA-YGRISRKIDVYAFGVVLYELISAKAAVIK 294
LTKLT+ ++ T V GTFGYMPPE A YG +S K+DVYAFGVVLYELISAK A+
Sbjct: 434 LTKLTEVGGTSMPTGTRVVGTFGYMPPEYARYGDVSPKVDVYAFGVVLYELISAKEAI-- 491
Query: 295 IDKTEFEFKSLEIKTNESIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSI 354
+++ ES + K LV LF+E +N + D + LR L+DP+LG +Y IDSI
Sbjct: 492 ------------VRSTESSSDSKGLVYLFEEALN-SPDPKEGLRTLIDPKLGEDYPIDSI 538
Query: 355 SKMAKLAKACINRDPKQRPTMRDVVVSLMELNSS 388
K+ +LAK C DPK RP+MR VVV+LM L+S+
Sbjct: 539 LKLTQLAKVCTQEDPKLRPSMRSVVVALMTLSST 572
>gb|AAQ73159.1| LysM domain-containing receptor-like kinase 3 [Medicago truncatula]
gi|34485516|gb|AAQ73155.1| LysM domain-containing
receptor-like kinase 3 [Medicago truncatula]
Length = 620
Score = 388 bits (996), Expect = e-106
Identities = 227/436 (52%), Positives = 281/436 (64%), Gaps = 62/436 (14%)
Query: 6 LLKRYN--VNFSQGSGLVYIRGKDGNGLYVPLPP-------------------------- 37
L++ +N NFS GSG+V+I G+D NG + PL
Sbjct: 190 LIQNFNQDANFSIGSGIVFIPGRDQNGHFFPLYSRTGIAKGSAVGIAMAGIFGLLLFVIY 249
Query: 38 ---KYFRKKNGEEESKFPPEDSMTPSTKDVDKD------------TNDDNGSKYIWVDKS 82
KYF+KK EEE P+ S ST+D T G I V KS
Sbjct: 250 IYAKYFQKK--EEEKTKLPQTSRAFSTQDASGSAEYETSGSSGHATGSAAGLTGIMVAKS 307
Query: 83 PEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELKVLT 142
EF+Y+ELA AT+NFSL KIGQGGFG VYY ELRG+K AIKKM +QA+ EFL ELKVLT
Sbjct: 308 TEFTYQELAKATNNFSLDNKIGQGGFGAVYYAELRGEKTAIKKMDVQASSEFLCELKVLT 367
Query: 143 SVHHWNLVHLIGYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLE 202
VHH NLV LIGYCVEG LFLVYE+++NGNL Q+LH EP+ S+R++IALD ARGLE
Sbjct: 368 HVHHLNLVRLIGYCVEGSLFLVYEHIDNGNLGQYLHGIGTEPLPWSSRVQIALDSARGLE 427
Query: 203 YIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPP 262
YIH+H+VPVYIHRD+KS NIL+++N GKVADFGLTKL + NS +T + GTFGYMPP
Sbjct: 428 YIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLHT-RLVGTFGYMPP 486
Query: 263 ENA-YGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVA 321
E A YG +S KIDVYAFGVVLYELI+AK AV +KT ES+ E K LV
Sbjct: 487 EYAQYGDVSPKIDVYAFGVVLYELITAKNAV--------------LKTGESVAESKGLVQ 532
Query: 322 LFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVS 381
LF+E +++ D ++ LRKLVDPRL NY IDS+ KMA+L +AC +P RP+MR +VV+
Sbjct: 533 LFEEALHRM-DPLEGLRKLVDPRLKENYPIDSVLKMAQLGRACTRDNPLLRPSMRSIVVA 591
Query: 382 LMELNSSIDNKNSTGS 397
LM L+S ++ + S
Sbjct: 592 LMTLSSPTEDCDDDSS 607
>dbj|BAB02358.1| unnamed protein product [Arabidopsis thaliana]
Length = 603
Score = 387 bits (995), Expect = e-106
Identities = 219/418 (52%), Positives = 277/418 (65%), Gaps = 47/418 (11%)
Query: 2 IEIELLKRYN--VNFSQGSGLVYIRGKDGNG---------------LYVPLPPKYFRKKN 44
+ ++L+RYN VNF+ G+G+VY+ G+DG G L + Y +KN
Sbjct: 187 VSADILQRYNPGVNFNSGNGIVYVPGRDGVGAGVIAGIVIGVIVALLLILFIVYYAYRKN 246
Query: 45 GEEESKFPPEDSMTPSTK-DVDKDTNDDNGS----------KYIWVDKSPEFSYEELANA 93
+ F S+ STK D T+ +G I VDKS EFS EELA A
Sbjct: 247 KSKGDSF--SSSIPLSTKADHASSTSLQSGGLGGAGVSPGIAAISVDKSVEFSLEELAKA 304
Query: 94 TDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELKVLTSVHHWNLVHLI 153
TDNF+L+ KIGQGGFG VYY ELRG+K AIKKM M+A+++FL+ELKVLT VHH NLV LI
Sbjct: 305 TDNFNLSFKIGQGGFGAVYYAELRGEKAAIKKMDMEASKQFLAELKVLTRVHHVNLVRLI 364
Query: 154 GYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDHSVPVYI 213
GYCVEG LFLVYEY+ENGNL QHLH S +EP+ + R++IALD ARGLEYIH+H+VPVY+
Sbjct: 365 GYCVEGSLFLVYEYVENGNLGQHLHGSGREPLPWTKRVQIALDSARGLEYIHEHTVPVYV 424
Query: 214 HRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPENAYGRISRKI 273
HRDIKS NIL+++ F KVADFGLTKLT+ SA T GTFGYM PE YG +S K+
Sbjct: 425 HRDIKSANILIDQKFRAKVADFGLTKLTEVGGSA--TRGAMGTFGYMAPETVYGEVSAKV 482
Query: 274 DVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVMNQTGDC 333
DVYAFGVVLYELISAK AV+K+ E++ E++ LV +F+E +T D
Sbjct: 483 DVYAFGVVLYELISAKGAVVKM--------------TEAVGEFRGLVGVFEESFKET-DK 527
Query: 334 IDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVVVSLMELNSSIDN 391
+ LRK++DPRLG +Y DS+ KMA+L KAC + + RP+MR +VV+L L SS N
Sbjct: 528 EEALRKIIDPRLGDSYPFDSVYKMAELGKACTQENAQLRPSMRYIVVALSTLFSSTGN 585
>ref|NP_566689.2| protein kinase family protein [Arabidopsis thaliana]
Length = 617
Score = 386 bits (991), Expect = e-106
Identities = 219/432 (50%), Positives = 277/432 (63%), Gaps = 61/432 (14%)
Query: 2 IEIELLKRYN--VNFSQGSGLVYIRGKDGNGLYVPLPPK--------------------- 38
+ ++L+RYN VNF+ G+G+VY+ G+D NG + P
Sbjct: 187 VSADILQRYNPGVNFNSGNGIVYVPGRDPNGAFPPFKSSKQDGVGAGVIAGIVIGVIVAL 246
Query: 39 --------YFRKKNGEEESKFPPEDSMTPSTK-DVDKDTNDDNGS----------KYIWV 79
Y +KN + F S+ STK D T+ +G I V
Sbjct: 247 LLILFIVYYAYRKNKSKGDSF--SSSIPLSTKADHASSTSLQSGGLGGAGVSPGIAAISV 304
Query: 80 DKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELK 139
DKS EFS EELA ATDNF+L+ KIGQGGFG VYY ELRG+K AIKKM M+A+++FL+ELK
Sbjct: 305 DKSVEFSLEELAKATDNFNLSFKIGQGGFGAVYYAELRGEKAAIKKMDMEASKQFLAELK 364
Query: 140 VLTSVHHWNLVHLIGYCVEGFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVAR 199
VLT VHH NLV LIGYCVEG LFLVYEY+ENGNL QHLH S +EP+ + R++IALD AR
Sbjct: 365 VLTRVHHVNLVRLIGYCVEGSLFLVYEYVENGNLGQHLHGSGREPLPWTKRVQIALDSAR 424
Query: 200 GLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGY 259
GLEYIH+H+VPVY+HRDIKS NIL+++ F KVADFGLTKLT+ SA T GTFGY
Sbjct: 425 GLEYIHEHTVPVYVHRDIKSANILIDQKFRAKVADFGLTKLTEVGGSA--TRGAMGTFGY 482
Query: 260 MPPENAYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSL 319
M PE YG +S K+DVYAFGVVLYELISAK AV+K+ E++ E++ L
Sbjct: 483 MAPETVYGEVSAKVDVYAFGVVLYELISAKGAVVKM--------------TEAVGEFRGL 528
Query: 320 VALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV 379
V +F+E +T D + LRK++DPRLG +Y DS+ KMA+L KAC + + RP+MR +V
Sbjct: 529 VGVFEESFKET-DKEEALRKIIDPRLGDSYPFDSVYKMAELGKACTQENAQLRPSMRYIV 587
Query: 380 VSLMELNSSIDN 391
V+L L SS N
Sbjct: 588 VALSTLFSSTGN 599
>ref|NP_175606.2| protein kinase family protein / peptidoglycan-binding LysM
domain-containing protein [Arabidopsis thaliana]
Length = 651
Score = 240 bits (612), Expect = 6e-62
Identities = 139/321 (43%), Positives = 200/321 (62%), Gaps = 20/321 (6%)
Query: 79 VDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSEL 138
++K F+YEE+ ATD FS + +G G +G VY+G LR Q++A+K+M T+EF +E+
Sbjct: 323 IEKPMVFTYEEIRAATDEFSDSNLLGHGNYGSVYFGLLREQEVAVKRMTATKTKEFAAEM 382
Query: 139 KVLTSVHHWNLVHLIGYCVE-GFLFLVYEYMENGNLSQHLHNSEKE---PMTLSTRMKIA 194
KVL VHH NLV LIGY LF+VYEY+ G L HLH+ + + P++ R +IA
Sbjct: 383 KVLCKVHHSNLVELIGYAATVDELFVVYEYVRKGMLKSHLHDPQSKGNTPLSWIMRNQIA 442
Query: 195 LDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTV-HV 253
LD ARGLEYIH+H+ Y+HRDIK+ NILL+E F K++DFGL KL + + +V V
Sbjct: 443 LDAARGLEYIHEHTKTHYVHRDIKTSNILLDEAFRAKISDFGLAKLVEKTGEGEISVTKV 502
Query: 254 AGTFGYMPPEN-AYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNES 312
GT+GY+ PE + G + K D+YAFGVVL+E+IS + AVI+ + I T
Sbjct: 503 VGTYGYLAPEYLSDGLATSKSDIYAFGVVLFEIISGREAVIRTE---------AIGTKN- 552
Query: 313 IDEYKSLVALFDEVMNQTGDCID--DLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPK 370
E + L ++ V+ + D ++ L++ VDP + Y D + K+A LAK C++ DP
Sbjct: 553 -PERRPLASIMLAVLKNSPDSMNMSSLKEFVDPNMMDLYPHDCLFKIATLAKQCVDDDPI 611
Query: 371 QRPTMRDVVVSLME-LNSSID 390
RP M+ VV+SL + L SSI+
Sbjct: 612 LRPNMKQVVISLSQILLSSIE 632
>pir||G96558 probable protein kinase [imported] - Arabidopsis thaliana
gi|9802793|gb|AAF99862.1| Putative protein kinase
[Arabidopsis thaliana]
Length = 601
Score = 240 bits (612), Expect = 6e-62
Identities = 139/321 (43%), Positives = 200/321 (62%), Gaps = 20/321 (6%)
Query: 79 VDKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSEL 138
++K F+YEE+ ATD FS + +G G +G VY+G LR Q++A+K+M T+EF +E+
Sbjct: 273 IEKPMVFTYEEIRAATDEFSDSNLLGHGNYGSVYFGLLREQEVAVKRMTATKTKEFAAEM 332
Query: 139 KVLTSVHHWNLVHLIGYCVE-GFLFLVYEYMENGNLSQHLHNSEKE---PMTLSTRMKIA 194
KVL VHH NLV LIGY LF+VYEY+ G L HLH+ + + P++ R +IA
Sbjct: 333 KVLCKVHHSNLVELIGYAATVDELFVVYEYVRKGMLKSHLHDPQSKGNTPLSWIMRNQIA 392
Query: 195 LDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTV-HV 253
LD ARGLEYIH+H+ Y+HRDIK+ NILL+E F K++DFGL KL + + +V V
Sbjct: 393 LDAARGLEYIHEHTKTHYVHRDIKTSNILLDEAFRAKISDFGLAKLVEKTGEGEISVTKV 452
Query: 254 AGTFGYMPPEN-AYGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNES 312
GT+GY+ PE + G + K D+YAFGVVL+E+IS + AVI+ + I T
Sbjct: 453 VGTYGYLAPEYLSDGLATSKSDIYAFGVVLFEIISGREAVIRTE---------AIGTKN- 502
Query: 313 IDEYKSLVALFDEVMNQTGDCID--DLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPK 370
E + L ++ V+ + D ++ L++ VDP + Y D + K+A LAK C++ DP
Sbjct: 503 -PERRPLASIMLAVLKNSPDSMNMSSLKEFVDPNMMDLYPHDCLFKIATLAKQCVDDDPI 561
Query: 371 QRPTMRDVVVSLME-LNSSID 390
RP M+ VV+SL + L SSI+
Sbjct: 562 LRPNMKQVVISLSQILLSSIE 582
>ref|NP_916033.1| putative receptor protein kinase tmk1 precursor [Oryza sativa
(japonica cultivar-group)]
Length = 612
Score = 239 bits (611), Expect = 8e-62
Identities = 140/320 (43%), Positives = 196/320 (60%), Gaps = 19/320 (5%)
Query: 80 DKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELK 139
+K F+Y+E+ +TD+FS A +G G +G VYYG LR Q++AIK+M T+EF+ E+K
Sbjct: 284 EKPIVFTYQEILASTDSFSDANLLGHGTYGSVYYGVLRDQEVAIKRMTATKTKEFIVEMK 343
Query: 140 VLTSVHHWNLVHLIGYCV-EGFLFLVYEYMENGNLSQHLHNSEKEPMTLST---RMKIAL 195
VL VHH +LV LIGY + L+L+YEY + G+L HLH+ + + T + R++IAL
Sbjct: 344 VLCKVHHASLVELIGYAASKDELYLIYEYSQKGSLKNHLHDPQSKGYTSLSWIYRVQIAL 403
Query: 196 DVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTV-HVA 254
D ARGLEYIH+H+ Y+HRDIKS NILL+E+F K++DFGL KL + A+ +V V
Sbjct: 404 DAARGLEYIHEHTKDHYVHRDIKSSNILLDESFRAKISDFGLAKLVVKSTDAEASVTKVV 463
Query: 255 GTFGYMPPENAY-GRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESI 313
GTFGY+ PE G + K DVYAFGVVL+ELIS K A+ + D S
Sbjct: 464 GTFGYLAPEYLRDGLATTKNDVYAFGVVLFELISGKEAITRTDGL----------NEGSN 513
Query: 314 DEYKSLVALFDEVMNQTGDC--IDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQ 371
E +SL ++ + + + L+ +DP L Y D + KMA LAK C+ DP
Sbjct: 514 SERRSLASVMLSALKNCRNSMYMGSLKDCIDPNLMDLYPHDCVYKMAMLAKQCVEEDPVL 573
Query: 372 RPTMRDVVVSLME-LNSSID 390
RP M+ V++L + L SSI+
Sbjct: 574 RPDMKQAVITLSQILLSSIE 593
>dbj|BAD86955.1| putative Nod-factor receptor 1b [Oryza sativa (japonica
cultivar-group)]
Length = 420
Score = 239 bits (611), Expect = 8e-62
Identities = 140/320 (43%), Positives = 196/320 (60%), Gaps = 19/320 (5%)
Query: 80 DKSPEFSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELK 139
+K F+Y+E+ +TD+FS A +G G +G VYYG LR Q++AIK+M T+EF+ E+K
Sbjct: 92 EKPIVFTYQEILASTDSFSDANLLGHGTYGSVYYGVLRDQEVAIKRMTATKTKEFIVEMK 151
Query: 140 VLTSVHHWNLVHLIGYCV-EGFLFLVYEYMENGNLSQHLHNSEKEPMTLST---RMKIAL 195
VL VHH +LV LIGY + L+L+YEY + G+L HLH+ + + T + R++IAL
Sbjct: 152 VLCKVHHASLVELIGYAASKDELYLIYEYSQKGSLKNHLHDPQSKGYTSLSWIYRVQIAL 211
Query: 196 DVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTV-HVA 254
D ARGLEYIH+H+ Y+HRDIKS NILL+E+F K++DFGL KL + A+ +V V
Sbjct: 212 DAARGLEYIHEHTKDHYVHRDIKSSNILLDESFRAKISDFGLAKLVVKSTDAEASVTKVV 271
Query: 255 GTFGYMPPENAY-GRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESI 313
GTFGY+ PE G + K DVYAFGVVL+ELIS K A+ + D S
Sbjct: 272 GTFGYLAPEYLRDGLATTKNDVYAFGVVLFELISGKEAITRTDGL----------NEGSN 321
Query: 314 DEYKSLVALFDEVMNQTGDC--IDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQ 371
E +SL ++ + + + L+ +DP L Y D + KMA LAK C+ DP
Sbjct: 322 SERRSLASVMLSALKNCRNSMYMGSLKDCIDPNLMDLYPHDCVYKMAMLAKQCVEEDPVL 381
Query: 372 RPTMRDVVVSLME-LNSSID 390
RP M+ V++L + L SSI+
Sbjct: 382 RPDMKQAVITLSQILLSSIE 401
>ref|NP_918232.1| protein kinase-like [Oryza sativa (japonica cultivar-group)]
Length = 599
Score = 236 bits (602), Expect = 9e-61
Identities = 131/319 (41%), Positives = 198/319 (62%), Gaps = 22/319 (6%)
Query: 85 FSYEELANATDNFSLAKKIGQGGFGEVYYGELRGQKIAIKKMKMQATREFLSELKVLTSV 144
FS + +AT NF +KIG+GG+G VY G + +IA+KKMK ++EF +ELKVL +
Sbjct: 283 FSLRAIEDATSNFDEKRKIGEGGYGSVYLGFIGTHEIAVKKMKASKSKEFFAELKVLCKI 342
Query: 145 HHWNLVHLIGYCV-EGFLFLVYEYMENGNLSQHLHN---SEKEPMTLSTRMKIALDVARG 200
HH N+V LIGY + L+LVYEY++NG+LS+HLH+ +P++ + R +IA+D ARG
Sbjct: 343 HHINVVELIGYAAGDDHLYLVYEYVQNGSLSEHLHDPLLKGHQPLSWTARTQIAMDSARG 402
Query: 201 LEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSAD-NTVHVAGTFGY 259
+EYIHDH+ Y+HRDIK+ NILL+ KVADFGL KL ++ + + GT GY
Sbjct: 403 IEYIHDHTKTCYVHRDIKTSNILLDNGLRAKVADFGLVKLVQRSDEDECLATRLVGTPGY 462
Query: 260 MPPENAYG-RISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKS 318
+PPE+ ++ K DVYAFGVVL ELI+ A+ ++ N+ ++ KS
Sbjct: 463 LPPESVLELHMTTKSDVYAFGVVLAELITGLRAL--------------VRDNKEANKTKS 508
Query: 319 LVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDV 378
L+++ + + D L +VDP L NY I+ + K+A ++ C++ DP RP MR+V
Sbjct: 509 LISIMRKAF-KPEDLESSLETIVDPYLKDNYPIEEVCKLANISMWCLSEDPLHRPEMREV 567
Query: 379 VVSLMELN-SSIDNKNSTG 396
+ L +++ +SI+ + S G
Sbjct: 568 MPILAQIHMASIEWEASLG 586
>ref|NP_913464.1| putative receptor protein kinase PERK1 [Oryza sativa (japonica
cultivar-group)] gi|17385728|dbj|BAB78668.1| putative
brassinosteroid insensitive 1-associated receptor kinase
1 [Oryza sativa (japonica cultivar-group)]
Length = 597
Score = 214 bits (545), Expect = 4e-54
Identities = 149/383 (38%), Positives = 211/383 (54%), Gaps = 54/383 (14%)
Query: 34 PLPPKYFRKKNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPE--------- 84
P PP + KK S PP P+ +V + +GS Y D S
Sbjct: 154 PPPPDHVLKKVPSHPSPPPP-----PAPLNVH---SGGSGSNYSGGDNSQPLVSPGAALG 205
Query: 85 -----FSYEELANATDNFSLAKKIGQGGFGEVYYGEL-RGQKIAIKKMKM---QATREFL 135
F+YE+L+ ATD FS A +GQGGFG V+ G L G ++A+K+++ Q REF
Sbjct: 206 FSRCTFTYEDLSAATDGFSDANLLGQGGFGYVHKGVLPNGTEVAVKQLRDGSGQGEREFQ 265
Query: 136 SELKVLTSVHHWNLVHLIGYCVEGFL-FLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIA 194
+E+++++ VHH +LV L+GYC+ G LVYEY+ N L HLH + M TR++IA
Sbjct: 266 AEVEIISRVHHKHLVTLVGYCISGGKRLLVYEYVPNNTLELHLHGRGRPTMEWPTRLRIA 325
Query: 195 LDVARGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVA 254
L A+GL Y+H+ P IHRDIKS NILL+ F KVADFGL KLT N+ +T V
Sbjct: 326 LGAAKGLAYLHEDCHPKIIHRDIKSANILLDARFEAKVADFGLAKLTSDNNTHVST-RVM 384
Query: 255 GTFGYMPPENA-YGRISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNES- 312
GTFGY+ PE A G+++ K DV++FGV+L ELI+ + V ++N+S
Sbjct: 385 GTFGYLAPEYASSGQLTEKSDVFSFGVMLLELITGRRPV---------------RSNQSQ 429
Query: 313 IDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQR 372
+D+ SLV +M + D + LVDPRLG Y+ + +++M A AC+ ++R
Sbjct: 430 MDD--SLVDWARPLMMRASD-DGNYDALVDPRLGQEYNGNEMARMIACAAACVRHSARRR 486
Query: 373 PTMRDVV------VSLMELNSSI 389
P M VV VSL +LN +
Sbjct: 487 PRMSQVVRALEGDVSLDDLNEGV 509
>dbj|BAD38053.1| putative light repressible receptor protein kinase [Oryza sativa
(japonica cultivar-group)]
Length = 899
Score = 213 bits (542), Expect = 8e-54
Identities = 141/327 (43%), Positives = 185/327 (56%), Gaps = 47/327 (14%)
Query: 84 EFSYEELANATDNFSLAKKIGQGGFGEVYYGELR-GQKIAIK---KMKMQATREFLSELK 139
EFSY EL + T+NFS +++G+GGFG V+ G L G +A+K + Q +EFL+E +
Sbjct: 565 EFSYRELKHITNNFS--QQVGKGGFGAVFLGYLENGNPVAVKVRSESSSQGGKEFLAEAQ 622
Query: 140 VLTSVHHWNLVHLIGYCVE-GFLFLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVA 198
LT +HH NLV LIGYC + L LVYEYM GNL HL + +P+T R+ IALD A
Sbjct: 623 HLTRIHHKNLVSLIGYCKDKNHLALVYEYMPEGNLQDHLRATTNKPLTWEQRLHIALDAA 682
Query: 199 RGLEYIHDHSVPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFG 258
+GLEY+H P IHRD+KS NILL N K+ADFGLTK+ + + T AGTFG
Sbjct: 683 QGLEYLHVACKPALIHRDVKSRNILLTTNLGAKIADFGLTKVFSESRT-HMTTEPAGTFG 741
Query: 259 YMPPENAYG-RISRKIDVYAFGVVLYELISAKAAVIKIDKT------EFEFKSLEIKTNE 311
Y+ PE IS K DVY+FGVVL ELI+ + VI ID++ EF +SL+ + E
Sbjct: 742 YLDPEYYRNYHISEKSDVYSFGVVLLELITGRPPVIPIDESVSIHIGEFVHQSLDHGSIE 801
Query: 312 SIDEYKSLVALFDEVMNQTGDCIDDLRKLVDPRL--GYNYSIDSISKMAKLAKACINRDP 369
SI VD R+ G Y I+S+ K+A LA C
Sbjct: 802 SI---------------------------VDARMGGGGGYDINSVWKVADLALHCKREVS 834
Query: 370 KQRPTMRDVVVSL---MELNSSIDNKN 393
++RPTM +VV L +EL S D K+
Sbjct: 835 RERPTMTEVVAQLKESLELESHGDRKH 861
>gb|AAP37768.1| At3g24600 [Arabidopsis thaliana] gi|13877617|gb|AAK43886.1| protein
kinase-like protein [Arabidopsis thaliana]
Length = 652
Score = 213 bits (541), Expect = 1e-53
Identities = 138/368 (37%), Positives = 200/368 (53%), Gaps = 45/368 (12%)
Query: 34 PLPPKYFRKKNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPEFSYEELANA 93
P PP + G + S P +P + KS F+YEEL+ A
Sbjct: 232 PPPPAFMSSSGGSDYSDLPVLPPPSPGL--------------VLGFSKST-FTYEELSRA 276
Query: 94 TDNFSLAKKIGQGGFGEVYYGEL-RGQKIAIKKMKM---QATREFLSELKVLTSVHHWNL 149
T+ FS A +GQGGFG V+ G L G+++A+K++K Q REF +E+++++ VHH +L
Sbjct: 277 TNGFSEANLLGQGGFGYVHKGILPSGKEVAVKQLKAGSGQGEREFQAEVEIISRVHHRHL 336
Query: 150 VHLIGYCVEGFL-FLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDHS 208
V LIGYC+ G LVYE++ N NL HLH + M STR+KIAL A+GL Y+H+
Sbjct: 337 VSLIGYCMAGVQRLLVYEFVPNNNLEFHLHGKGRPTMEWSTRLKIALGSAKGLSYLHEDC 396
Query: 209 VPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPE-NAYG 267
P IHRDIK+ NIL++ F KVADFGL K+ N+ +T V GTFGY+ PE A G
Sbjct: 397 NPKIIHRDIKASNILIDFKFEAKVADFGLAKIASDTNTHVST-RVMGTFGYLAPEYAASG 455
Query: 268 RISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVM 327
+++ K DV++FGVVL ELI+ + V N +D+ SLV ++
Sbjct: 456 KLTEKSDVFSFGVVLLELITGRRPV--------------DANNVYVDD--SLVDWARPLL 499
Query: 328 NQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV------VS 381
N+ + D L D ++G Y + +++M A AC+ ++RP M +V VS
Sbjct: 500 NRASE-EGDFEGLADSKMGNEYDREEMARMVACAAACVRHSARRRPRMSQIVRALEGNVS 558
Query: 382 LMELNSSI 389
L +LN +
Sbjct: 559 LSDLNEGM 566
>gb|AAP37759.1| At3g24550 [Arabidopsis thaliana] gi|22136228|gb|AAM91192.1| protein
kinase-like protein [Arabidopsis thaliana]
gi|9294050|dbj|BAB02007.1| protein kinase-like protein
[Arabidopsis thaliana] gi|20260332|gb|AAM13064.1|
unknown protein [Arabidopsis thaliana]
gi|16649063|gb|AAL24383.1| protein kinase-like protein
[Arabidopsis thaliana] gi|15983765|gb|AAL10479.1|
AT3g24550/MOB24_8 [Arabidopsis thaliana]
gi|15230130|ref|NP_189098.1| protein kinase family
protein [Arabidopsis thaliana]
Length = 652
Score = 213 bits (541), Expect = 1e-53
Identities = 138/368 (37%), Positives = 200/368 (53%), Gaps = 45/368 (12%)
Query: 34 PLPPKYFRKKNGEEESKFPPEDSMTPSTKDVDKDTNDDNGSKYIWVDKSPEFSYEELANA 93
P PP + G + S P +P + KS F+YEEL+ A
Sbjct: 232 PPPPAFMSSSGGSDYSDLPVLPPPSPGL--------------VLGFSKST-FTYEELSRA 276
Query: 94 TDNFSLAKKIGQGGFGEVYYGEL-RGQKIAIKKMKM---QATREFLSELKVLTSVHHWNL 149
T+ FS A +GQGGFG V+ G L G+++A+K++K Q REF +E+++++ VHH +L
Sbjct: 277 TNGFSEANLLGQGGFGYVHKGILPSGKEVAVKQLKAGSGQGEREFQAEVEIISRVHHRHL 336
Query: 150 VHLIGYCVEGFL-FLVYEYMENGNLSQHLHNSEKEPMTLSTRMKIALDVARGLEYIHDHS 208
V LIGYC+ G LVYE++ N NL HLH + M STR+KIAL A+GL Y+H+
Sbjct: 337 VSLIGYCMAGVQRLLVYEFVPNNNLEFHLHGKGRPTMEWSTRLKIALGSAKGLSYLHEDC 396
Query: 209 VPVYIHRDIKSDNILLNENFTGKVADFGLTKLTDAANSADNTVHVAGTFGYMPPE-NAYG 267
P IHRDIK+ NIL++ F KVADFGL K+ N+ +T V GTFGY+ PE A G
Sbjct: 397 NPKIIHRDIKASNILIDFKFEAKVADFGLAKIASDTNTHVST-RVMGTFGYLAPEYAASG 455
Query: 268 RISRKIDVYAFGVVLYELISAKAAVIKIDKTEFEFKSLEIKTNESIDEYKSLVALFDEVM 327
+++ K DV++FGVVL ELI+ + V N +D+ SLV ++
Sbjct: 456 KLTEKSDVFSFGVVLLELITGRRPV--------------DANNVYVDD--SLVDWARPLL 499
Query: 328 NQTGDCIDDLRKLVDPRLGYNYSIDSISKMAKLAKACINRDPKQRPTMRDVV------VS 381
N+ + D L D ++G Y + +++M A AC+ ++RP M +V VS
Sbjct: 500 NRASE-EGDFEGLADSKMGNEYDREEMARMVACAAACVRHSARRRPRMSQIVRALEGNVS 558
Query: 382 LMELNSSI 389
L +LN +
Sbjct: 559 LSDLNEGM 566
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.135 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 695,036,779
Number of Sequences: 2540612
Number of extensions: 30927964
Number of successful extensions: 141781
Number of sequences better than 10.0: 20404
Number of HSP's better than 10.0 without gapping: 7841
Number of HSP's successfully gapped in prelim test: 12567
Number of HSP's that attempted gapping in prelim test: 94400
Number of HSP's gapped (non-prelim): 25662
length of query: 398
length of database: 863,360,394
effective HSP length: 130
effective length of query: 268
effective length of database: 533,080,834
effective search space: 142865663512
effective search space used: 142865663512
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC123570.11


