

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122165.8 - phase: 1 /pseudo
(144 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
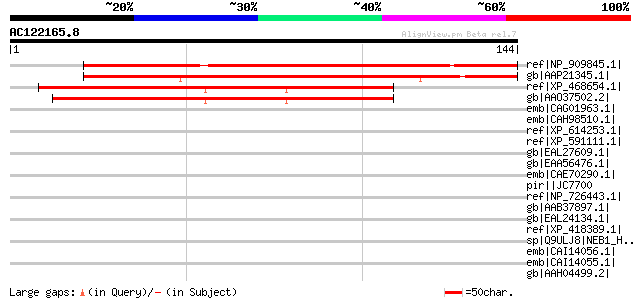
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_909845.1| unknown protein [Oryza sativa (japonica cultiva... 144 3e-34
gb|AAP21345.1| At5g55640 [Arabidopsis thaliana] gi|9758598|dbj|B... 144 3e-34
ref|XP_468654.1| unknown protein [Oryza sativa (japonica cultiva... 95 4e-19
gb|AAO37502.2| expressed protein [Oryza sativa (japonica cultiva... 92 3e-18
emb|CAG01963.1| unnamed protein product [Tetraodon nigroviridis] 37 0.100
emb|CAH98510.1| protein gar2, putative [Plasmodium berghei] 37 0.13
ref|XP_614253.1| PREDICTED: similar to Translocation protein SEC... 36 0.17
ref|XP_591111.1| PREDICTED: similar to Translocation protein SEC... 36 0.17
gb|EAL27609.1| GA20616-PA [Drosophila pseudoobscura] 36 0.17
gb|EAA56476.1| hypothetical protein MG06447.4 [Magnaporthe grise... 36 0.22
emb|CAE70290.1| Hypothetical protein CBG16810 [Caenorhabditis br... 35 0.29
pir||JC7700 38K ribosome-associated protein, Ray38p - yeast (Can... 35 0.29
ref|NP_726443.1| CG30165-PA [Drosophila melanogaster] gi|2162674... 35 0.38
gb|AAB37897.1| Hypothetical protein B0432.11 [Caenorhabditis ele... 35 0.38
gb|EAL24134.1| protein phosphatase 1, regulatory (inhibitor) sub... 35 0.38
ref|XP_418389.1| PREDICTED: similar to hypothetical protein [Gal... 35 0.38
sp|Q9ULJ8|NEB1_HUMAN Neurabin-I (Neural tissue-specific F-actin ... 35 0.38
emb|CAI14056.1| Mov10, Moloney leukemia virus 10, homolog (mouse... 35 0.50
emb|CAI14055.1| Mov10, Moloney leukemia virus 10, homolog (mouse... 35 0.50
gb|AAH04499.2| MOV10 protein [Homo sapiens] 35 0.50
>ref|NP_909845.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|28269467|gb|AAO38010.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 157
Score = 144 bits (364), Expect = 3e-34
Identities = 71/123 (57%), Positives = 89/123 (71%), Gaps = 3/123 (2%)
Query: 22 DTDGFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPS 81
D DGFVIP+L Q+ D +++ A + P + KEEKIYLGPHGAPPSQ KQQE+
Sbjct: 35 DADGFVIPNLTTQEDDVTEHTAPKPKDPEPLKE--KEEKIYLGPHGAPPSQAKQQEINIV 92
Query: 82 NRKQRFKQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDWLDPHCHEAE 141
RKQRF+ KLKEAD + +G +ENKV+ LREL+G S G+ K SSP+DWLDPHCHE+E
Sbjct: 93 GRKQRFRNKLKEADNKFTGNAQENKVETLRELMGARTHSKGVPK-SSPRDWLDPHCHESE 151
Query: 142 FER 144
F+R
Sbjct: 152 FDR 154
>gb|AAP21345.1| At5g55640 [Arabidopsis thaliana] gi|9758598|dbj|BAB09231.1| unnamed
protein product [Arabidopsis thaliana]
gi|18423771|ref|NP_568828.1| expressed protein
[Arabidopsis thaliana] gi|13899123|gb|AAK48983.1|
Unknown protein [Arabidopsis thaliana]
Length = 149
Score = 144 bits (364), Expect = 3e-34
Identities = 76/127 (59%), Positives = 94/127 (73%), Gaps = 5/127 (3%)
Query: 22 DTDGFVIPSLGIQDSDQSKNNAASVE-SPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIP 80
DTDGFVIPSL I++ + NN + VE S S+ K+K EE IYLGPHGAPPSQ +
Sbjct: 20 DTDGFVIPSLEIEEDKVNNNNDSEVETSKPSSPKSKAEENIYLGPHGAPPSQLQDGGSNT 79
Query: 81 SNRKQRFKQKLKEADKRISGTGRENKVDNLRELVG---GEKTSVGMAKGSSPKDWLDPHC 137
S+RKQRFKQ LKEAD+++SG+GRENK+ NLRELVG G + M KG S +DWLDPHC
Sbjct: 80 SSRKQRFKQMLKEADQKMSGSGRENKMANLRELVGGGAGAEKGTNMGKGVS-RDWLDPHC 138
Query: 138 HEAEFER 144
HE++FE+
Sbjct: 139 HESQFEK 145
>ref|XP_468654.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 172
Score = 94.7 bits (234), Expect = 4e-19
Identities = 53/103 (51%), Positives = 66/103 (63%), Gaps = 2/103 (1%)
Query: 9 VLKCMFFGLLLVVDTDGFVIPSLGIQDSDQSKNNAASVESPNSAAK-TKKEEKIYLGPHG 67
VL+ MF + V D DGFVIPSL +++SD AA V P K TK EKIYLGPHG
Sbjct: 50 VLRKMFLHVRNVTDNDGFVIPSLSVEESDLGDWEAAQVSRPQPPPKATKDTEKIYLGPHG 109
Query: 68 APPSQPKQQE-VIPSNRKQRFKQKLKEADKRISGTGRENKVDN 109
APPS+ K+QE + R K K+KEAD+++ GTGR+NK N
Sbjct: 110 APPSRAKKQEDTAAAATGYRDKSKVKEADQKVLGTGRDNKGGN 152
>gb|AAO37502.2| expressed protein [Oryza sativa (japonica cultivar-group)]
Length = 119
Score = 92.0 bits (227), Expect = 3e-18
Identities = 51/99 (51%), Positives = 63/99 (63%), Gaps = 2/99 (2%)
Query: 13 MFFGLLLVVDTDGFVIPSLGIQDSDQSKNNAASVESPNSAAK-TKKEEKIYLGPHGAPPS 71
MF + V D DGFVIPSL +++SD AA V P K TK EKIYLGPHGAPPS
Sbjct: 1 MFLHVRNVTDNDGFVIPSLSVEESDLGDWEAAQVSRPQPPPKATKDTEKIYLGPHGAPPS 60
Query: 72 QPKQQE-VIPSNRKQRFKQKLKEADKRISGTGRENKVDN 109
+ K+QE + R K K+KEAD+++ GTGR+NK N
Sbjct: 61 RAKKQEDTAAAATGYRDKSKVKEADQKVLGTGRDNKGGN 99
>emb|CAG01963.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1372
Score = 37.0 bits (84), Expect = 0.100
Identities = 24/100 (24%), Positives = 42/100 (42%), Gaps = 5/100 (5%)
Query: 31 LGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQK 90
LG D D++ + +++E ++ + K+Y S P Q + S R
Sbjct: 717 LGSDDDDETNDEESALELRQFSSVAHRFSKVYSSTEHLSTSTPSNQSLSSSERSHS---- 772
Query: 91 LKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPK 130
+E ++R G G + D R L GGE+ + G P+
Sbjct: 773 -EEKEERWEGRGTLSPGDGYRHLSGGERRADGHRGSLRPR 811
>emb|CAH98510.1| protein gar2, putative [Plasmodium berghei]
Length = 509
Score = 36.6 bits (83), Expect = 0.13
Identities = 25/86 (29%), Positives = 40/86 (46%), Gaps = 1/86 (1%)
Query: 35 DSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEA 94
DS+ KN V+S KKE I LG Q K++ I ++ + KQKL +
Sbjct: 178 DSENEKNKTKEVKSTFPIENNKKE-SILLGNQNKEDGQGKRKNDISLDKGKNKKQKLGKE 236
Query: 95 DKRISGTGRENKVDNLRELVGGEKTS 120
+K +G + K +N + + +TS
Sbjct: 237 NKNQNGINNKQKKNNGIDSMANNETS 262
>ref|XP_614253.1| PREDICTED: similar to Translocation protein SEC63 homolog, partial
[Bos taurus]
Length = 623
Score = 36.2 bits (82), Expect = 0.17
Identities = 24/97 (24%), Positives = 43/97 (43%)
Query: 47 ESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGRENK 106
+ P AK+KK++ + P P SQPKQQ+ +N + +KE ++ +S G +++
Sbjct: 381 KGPKKTAKSKKKKPLKKKPTPVPLSQPKQQKQKQANGVVGSEAAVKEDEEEVSDKGSDSE 440
Query: 107 VDNLRELVGGEKTSVGMAKGSSPKDWLDPHCHEAEFE 143
+ EK +D EAE++
Sbjct: 441 EEETNRDSQSEKDDGSDRDSDREQDEKQNKDDEAEWQ 477
>ref|XP_591111.1| PREDICTED: similar to Translocation protein SEC63 homolog, partial
[Bos taurus]
Length = 357
Score = 36.2 bits (82), Expect = 0.17
Identities = 24/97 (24%), Positives = 43/97 (43%)
Query: 47 ESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGRENK 106
+ P AK+KK++ + P P SQPKQQ+ +N + +KE ++ +S G +++
Sbjct: 115 KGPKKTAKSKKKKPLKKKPTPVPLSQPKQQKQKQANGVVGSEAAVKEDEEEVSDKGSDSE 174
Query: 107 VDNLRELVGGEKTSVGMAKGSSPKDWLDPHCHEAEFE 143
+ EK +D EAE++
Sbjct: 175 EEETNRDSQSEKDDGSDRDSDREQDEKQNKDDEAEWQ 211
>gb|EAL27609.1| GA20616-PA [Drosophila pseudoobscura]
Length = 1045
Score = 36.2 bits (82), Expect = 0.17
Identities = 29/82 (35%), Positives = 41/82 (49%), Gaps = 7/82 (8%)
Query: 32 GIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKL 91
G + ++ NNAASV NSA+K + L P+ P P + P+ R++R
Sbjct: 458 GRRGNNNKTNNAASVAKKNSASKPPPPQD--LKPNPLLPPIPIADKTEPAVRQRR----R 511
Query: 92 KEADKRISGTGRENK-VDNLRE 112
K A KR S T EN+ DN R+
Sbjct: 512 KAASKRYSATINENETTDNARD 533
>gb|EAA56476.1| hypothetical protein MG06447.4 [Magnaporthe grisea 70-15]
gi|39977089|ref|XP_369932.1| hypothetical protein
MG06447.4 [Magnaporthe grisea 70-15]
Length = 1592
Score = 35.8 bits (81), Expect = 0.22
Identities = 27/87 (31%), Positives = 41/87 (47%), Gaps = 10/87 (11%)
Query: 45 SVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEADKRISGTGRE 104
S +P S+ K + + L PH PSQ QE++ K++ K+KLK + T RE
Sbjct: 1495 SQSNPGSSTKQSRPNRTGLNPHAQTPSQ-ATQEILDRKAKKKKKKKLKLKTE----TERE 1549
Query: 105 NKVDNLRELVGGEKTSVGMAKGSSPKD 131
++ V G GMA S+ +D
Sbjct: 1550 PRLPEWANKVSG-----GMASESTNRD 1571
>emb|CAE70290.1| Hypothetical protein CBG16810 [Caenorhabditis briggsae]
Length = 823
Score = 35.4 bits (80), Expect = 0.29
Identities = 28/116 (24%), Positives = 52/116 (44%), Gaps = 15/116 (12%)
Query: 40 KNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKLKEA--DKR 97
K++ A +E +S++ + + + +PK++ +P R + FK + +E+ D
Sbjct: 45 KDHLAGIEEMSSSSSDEASDSDDV----MDSEEPKKELKLPKKRSKAFKIESEESSSDSY 100
Query: 98 ISGTGRENKVDNLREL---------VGGEKTSVGMAKGSSPKDWLDPHCHEAEFER 144
SG E++ D+ ++ GG K +KGS KD P C +A R
Sbjct: 101 ESGESEESENDDSEDVGGPGPSSKAFGGSKGKSSKSKGSDQKDAHWPKCSKASIAR 156
>pir||JC7700 38K ribosome-associated protein, Ray38p - yeast (Candida maltosa)
gi|13928591|dbj|BAB47177.1| Ray38p [Candida maltosa]
Length = 269
Score = 35.4 bits (80), Expect = 0.29
Identities = 32/115 (27%), Positives = 46/115 (39%), Gaps = 6/115 (5%)
Query: 35 DSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQ-----QEVIPSNRKQRFKQ 89
D ++KNN V +A K K K P + K+ ++ K++FKQ
Sbjct: 47 DPAKAKNNKKKVTGNEAALKNKTNNKEVAPPQSSAAKHTKKPFDRHSRTGKTDSKKKFKQ 106
Query: 90 KLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDWLDPHCHEAEFER 144
E+DKR E D EL E + A S+PK L + E E +R
Sbjct: 107 GWGESDKRELEGEVEGAEDAEAEL-SAEASENADAADSTPKKTLQEYLAEMELKR 160
>ref|NP_726443.1| CG30165-PA [Drosophila melanogaster] gi|21626743|gb|AAM68312.1|
CG30165-PA [Drosophila melanogaster]
Length = 469
Score = 35.0 bits (79), Expect = 0.38
Identities = 15/66 (22%), Positives = 36/66 (53%)
Query: 32 GIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRFKQKL 91
G +D + ++ +++ +T EE + L + PS+ +QQ+ P R++ ++++
Sbjct: 131 GKRDQSRERSTSSTARPKPPLRRTATEETLDLADEESTPSRQQQQQPPPQRRQESDRERV 190
Query: 92 KEADKR 97
+E D+R
Sbjct: 191 RERDQR 196
>gb|AAB37897.1| Hypothetical protein B0432.11 [Caenorhabditis elegans]
gi|17531335|ref|NP_493697.1| putative nuclear protein
family member (2A582) [Caenorhabditis elegans]
gi|7494942|pir||T25462 hypothetical protein B0432.11 -
Caenorhabditis elegans
Length = 190
Score = 35.0 bits (79), Expect = 0.38
Identities = 26/88 (29%), Positives = 39/88 (43%), Gaps = 4/88 (4%)
Query: 47 ESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSN----RKQRFKQKLKEADKRISGTG 102
++ N+ ++TKK P AP P + I + + + K+ +KE + SGTG
Sbjct: 40 KTKNAGSRTKKSATSTSIPAPAPAPAPTEATTIAKDSLMGKLEPPKEPVKEPESDKSGTG 99
Query: 103 RENKVDNLRELVGGEKTSVGMAKGSSPK 130
N V LR K+ G KG PK
Sbjct: 100 PVNSVKWLRGRKKAWKSRKGSKKGKKPK 127
>gb|EAL24134.1| protein phosphatase 1, regulatory (inhibitor) subunit 9A [Homo
sapiens] gi|41147635|ref|XP_371933.1| PREDICTED: protein
phosphatase 1, regulatory (inhibitor) subunit 9A [Homo
sapiens] gi|41148006|ref|XP_374491.1| PREDICTED: similar
to Ppp1r9a protein [Homo sapiens]
Length = 1322
Score = 35.0 bits (79), Expect = 0.38
Identities = 30/104 (28%), Positives = 50/104 (47%), Gaps = 13/104 (12%)
Query: 28 IPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRF 87
IP IQ S + +++ ++ ++P+S K EE P + PK + IPS +
Sbjct: 292 IPGEEIQQSKEPEDSTSNQQTPDSIDKDGPEE-----PCAESKAMPKSE--IPSPQ---- 340
Query: 88 KQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKD 131
Q L++A+ + GRE +EL GG+ TS + S K+
Sbjct: 341 SQLLEDAEANL--VGREAAKQQRKELAGGDFTSPDASASSCGKE 382
>ref|XP_418389.1| PREDICTED: similar to hypothetical protein [Gallus gallus]
Length = 618
Score = 35.0 bits (79), Expect = 0.38
Identities = 27/112 (24%), Positives = 52/112 (46%), Gaps = 20/112 (17%)
Query: 36 SDQSKNNAASVESPNSAAKTKKEEKIY--LGPHGAPPSQPKQQEVIPSNRKQ----RFKQ 89
S S +N+ S + +++ ++ + + H A P P+Q + P +K+ K
Sbjct: 105 SSPSSSNSGSYKGSDTSPIMRRSGRYLTCVDHHSAKPQNPEQY-LTPLQQKEVTVRHLKT 163
Query: 90 KLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKDWLDPHCHEAE 141
KLKE+ ++ RE +++ L KT +G + +DW++ CH E
Sbjct: 164 KLKESQSKLKE--RETEIEEL-------KTQLGRMR----EDWIEEECHRVE 202
>sp|Q9ULJ8|NEB1_HUMAN Neurabin-I (Neural tissue-specific F-actin binding protein I)
(Protein phosphatase 1 regulatory subunit 9A)
Length = 1098
Score = 35.0 bits (79), Expect = 0.38
Identities = 30/104 (28%), Positives = 50/104 (47%), Gaps = 13/104 (12%)
Query: 28 IPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGPHGAPPSQPKQQEVIPSNRKQRF 87
IP IQ S + +++ ++ ++P+S K EE P + PK + IPS +
Sbjct: 292 IPGEEIQQSKEPEDSTSNQQTPDSIDKDGPEE-----PCAESKAMPKSE--IPSPQ---- 340
Query: 88 KQKLKEADKRISGTGRENKVDNLRELVGGEKTSVGMAKGSSPKD 131
Q L++A+ + GRE +EL GG+ TS + S K+
Sbjct: 341 SQLLEDAEANL--VGREAAKQQRKELAGGDFTSPDASASSCGKE 382
>emb|CAI14056.1| Mov10, Moloney leukemia virus 10, homolog (mouse) [Homo sapiens]
Length = 947
Score = 34.7 bits (78), Expect = 0.50
Identities = 27/104 (25%), Positives = 47/104 (44%), Gaps = 7/104 (6%)
Query: 25 GFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGP---HGAPPSQPKQQEVIPS 81
GF I G+ D+ + N+ S +P AA K+ L P G P+ VI
Sbjct: 717 GFPIIFHGVMGKDEREGNSPSFFNPEEAATVTSYLKLLLAPSSKKGKARLSPRSVGVISP 776
Query: 82 NRKQ--RFKQKLKEADKRISGTG--RENKVDNLRELVGGEKTSV 121
RKQ + + + + D+ + G ++ KV ++ E G E++ +
Sbjct: 777 YRKQVEKIRYCITKLDRELRGLDDIKDLKVGSVEEFQGQERSVI 820
>emb|CAI14055.1| Mov10, Moloney leukemia virus 10, homolog (mouse) [Homo sapiens]
gi|14424568|gb|AAH09312.1| Mov10, Moloney leukemia virus
10, homolog [Homo sapiens] gi|12803447|gb|AAH02548.1|
Mov10, Moloney leukemia virus 10, homolog [Homo sapiens]
gi|14211540|ref|NP_066014.1| Mov10, Moloney leukemia
virus 10, homolog [Homo sapiens]
gi|24638063|sp|Q9HCE1|MOV10_HUMAN Potential helicase
MOV-10 (Moloney leukemia virus 10 protein)
Length = 1003
Score = 34.7 bits (78), Expect = 0.50
Identities = 27/104 (25%), Positives = 47/104 (44%), Gaps = 7/104 (6%)
Query: 25 GFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGP---HGAPPSQPKQQEVIPS 81
GF I G+ D+ + N+ S +P AA K+ L P G P+ VI
Sbjct: 773 GFPIIFHGVMGKDEREGNSPSFFNPEEAATVTSYLKLLLAPSSKKGKARLSPRSVGVISP 832
Query: 82 NRKQ--RFKQKLKEADKRISGTG--RENKVDNLRELVGGEKTSV 121
RKQ + + + + D+ + G ++ KV ++ E G E++ +
Sbjct: 833 YRKQVEKIRYCITKLDRELRGLDDIKDLKVGSVEEFQGQERSVI 876
>gb|AAH04499.2| MOV10 protein [Homo sapiens]
Length = 601
Score = 34.7 bits (78), Expect = 0.50
Identities = 27/104 (25%), Positives = 47/104 (44%), Gaps = 7/104 (6%)
Query: 25 GFVIPSLGIQDSDQSKNNAASVESPNSAAKTKKEEKIYLGP---HGAPPSQPKQQEVIPS 81
GF I G+ D+ + N+ S +P AA K+ L P G P+ VI
Sbjct: 371 GFPIIFHGVMGKDEREGNSPSFFNPEEAATVTSYLKLLLAPSSKKGKARLSPRSVGVISP 430
Query: 82 NRKQ--RFKQKLKEADKRISGTG--RENKVDNLRELVGGEKTSV 121
RKQ + + + + D+ + G ++ KV ++ E G E++ +
Sbjct: 431 YRKQVEKIRYCITKLDRELRGLDDIKDLKVGSVEEFQGQERSVI 474
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 257,206,944
Number of Sequences: 2540612
Number of extensions: 10660816
Number of successful extensions: 41032
Number of sequences better than 10.0: 202
Number of HSP's better than 10.0 without gapping: 33
Number of HSP's successfully gapped in prelim test: 170
Number of HSP's that attempted gapping in prelim test: 40939
Number of HSP's gapped (non-prelim): 244
length of query: 144
length of database: 863,360,394
effective HSP length: 120
effective length of query: 24
effective length of database: 558,486,954
effective search space: 13403686896
effective search space used: 13403686896
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC122165.8


