

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121245.1 - phase: 0 /pseudo
(660 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
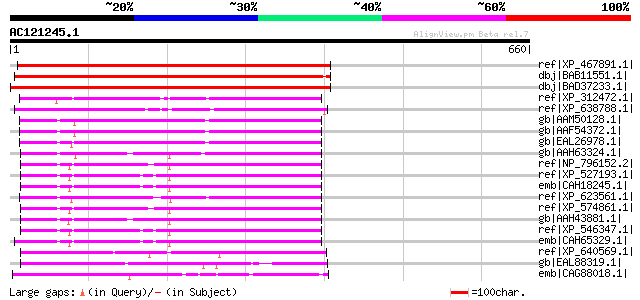
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_467891.1| LMBR1 integral membrane protein-like [Oryza sat... 625 e-177
dbj|BAB11551.1| unnamed protein product [Arabidopsis thaliana] g... 591 e-167
dbj|BAD37233.1| LMBR1 integral membrane protein-like [Oryza sati... 579 e-164
ref|XP_312472.1| ENSANGP00000003057 [Anopheles gambiae str. PEST... 174 6e-42
ref|XP_638788.1| hypothetical protein DDB0185797 [Dictyostelium ... 171 9e-41
gb|AAM50128.1| GH05505p [Drosophila melanogaster] 153 1e-35
gb|AAF54372.1| CG8135-PA [Drosophila melanogaster] gi|24645348|r... 153 1e-35
gb|EAL26978.1| GA20843-PA [Drosophila pseudoobscura] 151 7e-35
gb|AAH63324.1| Unknown (protein for MGC:73387) [Danio rerio] gi|... 149 2e-34
ref|NP_796152.2| hypothetical protein LOC320506 [Mus musculus] g... 148 5e-34
ref|XP_527193.1| PREDICTED: similar to hypothetical protein [Pan... 148 5e-34
emb|CAH18245.1| hypothetical protein [Homo sapiens] gi|56790897|... 148 5e-34
ref|XP_623561.1| PREDICTED: similar to ENSANGP00000003057 [Apis ... 147 1e-33
ref|XP_574861.1| PREDICTED: similar to RIKEN cDNA 9930036E21 gen... 147 1e-33
gb|AAH43881.1| Dkfzp434h2226-prov protein [Xenopus laevis] 145 4e-33
ref|XP_546347.1| PREDICTED: similar to hypothetical protein [Can... 145 4e-33
emb|CAH65329.1| hypothetical protein [Gallus gallus] 145 4e-33
ref|XP_640569.1| hypothetical protein DDB0204597 [Dictyostelium ... 128 5e-28
gb|EAL88319.1| LMBR and BAG domain protein, putative [Aspergillu... 127 1e-27
emb|CAG88018.1| unnamed protein product [Debaryomyces hansenii C... 120 1e-25
>ref|XP_467891.1| LMBR1 integral membrane protein-like [Oryza sativa (japonica
cultivar-group)] gi|46805692|dbj|BAD17093.1| LMBR1
integral membrane protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 734
Score = 625 bits (1611), Expect = e-177
Identities = 296/398 (74%), Positives = 349/398 (87%)
Query: 10 ISFLWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGL 69
I F WS SYWSTF+LTWAVVP +QGYEDAGDFTVK RL+TS+H NL+FY +G++ LFGL
Sbjct: 66 IGFFWSWSYWSTFILTWAVVPTIQGYEDAGDFTVKERLKTSIHMNLLFYSIVGAIGLFGL 125
Query: 70 ILLITLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLSH 129
ILL+ +++ W G + GFAMACSNTFGLVTGAFLLGFG+SEIP+ IW NADW +QK LSH
Sbjct: 126 ILLLVMHRAWDGGIVGFAMACSNTFGLVTGAFLLGFGLSEIPRNIWKNADWTHRQKVLSH 185
Query: 130 KVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKPQ 189
+VAKMAVKLD+AHQ++SNAIV+ QATS QMSKRD LRPYM+IIDKML QML EDPSFKP
Sbjct: 186 RVAKMAVKLDNAHQEYSNAIVVAQATSNQMSKRDLLRPYMDIIDKMLAQMLREDPSFKPS 245
Query: 190 GGRLGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVKNYDRRD 249
GGRLGE+DMDYDTD+K+MA+LRR+LRRA E+YYR +SEY +V+EALELEDT+KNY+RRD
Sbjct: 246 GGRLGENDMDYDTDDKTMATLRRQLRRAHEEYYRCKSEYMTYVMEALELEDTIKNYERRD 305
Query: 250 STGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAILLAEATIL 309
+ GW+++S R R G +G++LDT+EF+WRC+LRKQL+K A++LG MS AILLAEAT+L
Sbjct: 306 ANGWKFVSSFRESRPGTLGSLLDTMEFIWRCVLRKQLQKGFAIVLGCMSAAILLAEATLL 365
Query: 310 PSGVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYSLTPRQTSS 369
PSGVDLSLFSILV + G EVLVQ+AAFVPLMYMC+CTYYSLF++GM MFYSLTPRQTSS
Sbjct: 366 PSGVDLSLFSILVKSVGKQEVLVQVAAFVPLMYMCICTYYSLFQIGMLMFYSLTPRQTSS 425
Query: 370 VSLLMICSMVARYAAPISYNFLNLINIGGDRKTVFEKR 407
VSLLMICSMVARYA PISYNFLNLI +GGD KT FEKR
Sbjct: 426 VSLLMICSMVARYAPPISYNFLNLIRLGGDAKTTFEKR 463
>dbj|BAB11551.1| unnamed protein product [Arabidopsis thaliana]
gi|15238400|ref|NP_201332.1| LMBR1 integral membrane
family protein [Arabidopsis thaliana]
Length = 733
Score = 591 bits (1523), Expect = e-167
Identities = 284/401 (70%), Positives = 342/401 (84%), Gaps = 2/401 (0%)
Query: 7 NVTISFLWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVAL 66
N ISFLWS SYWSTFLLTWAVVPL+QG+EDAGDFTV RL+TS+H NLVFYL LG + L
Sbjct: 67 NGAISFLWSWSYWSTFLLTWAVVPLIQGFEDAGDFTVSERLKTSVHVNLVFYLVLGFIGL 126
Query: 67 FGLILLITLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKF 126
GLILLI +++ W+GS+ G+AMACSNTFGLVTGAFLLGFG+SEIPK +W NADW +QK
Sbjct: 127 LGLILLIMMHRNWTGSILGYAMACSNTFGLVTGAFLLGFGLSEIPKTLWKNADWTTRQKV 186
Query: 127 LSHKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSF 186
LSHK+AK+AVKLD+AHQ+ SNAIV+ QATS QMSKRD +RPYMN+ID ML +M EDPSF
Sbjct: 187 LSHKIAKIAVKLDNAHQELSNAIVVAQATSTQMSKRDPMRPYMNVIDAMLAKMFREDPSF 246
Query: 187 KPQGGRLGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVKNYD 246
KPQGG+LGE+DMDYDTDEKSMA+LRR LR A+++YYRY+SEY +V EAL LEDT+KNY+
Sbjct: 247 KPQGGQLGENDMDYDTDEKSMATLRRHLRNAKDEYYRYKSEYLTYVTEALVLEDTMKNYE 306
Query: 247 RRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAILLAEA 306
RRD+TGW+YIS R R GK+G +LD++EF+WRCIL+KQ++ LAV++G MS AILLAEA
Sbjct: 307 RRDATGWKYISSFRATRTGKMGNLLDSLEFMWRCILKKQIQMVLAVVMGIMSAAILLAEA 366
Query: 307 TILPSGVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYSLTPRQ 366
T+L S +DLSLFSIL+ + E+LVQ AFVPL+YMCVCTYYSLFK+GM M YSLTPRQ
Sbjct: 367 TLLLSKLDLSLFSILISSVKSDELLVQAFAFVPLVYMCVCTYYSLFKIGMLMIYSLTPRQ 426
Query: 367 TSSVSLLMICSMVARYAAPISYNFLNLINIGGDRKTVFEKR 407
TSSV+LLMICSM+ARYA PISYNF+NLI + + T+FEK+
Sbjct: 427 TSSVNLLMICSMIARYAPPISYNFINLIQLHSE--TIFEKK 465
>dbj|BAD37233.1| LMBR1 integral membrane protein-like [Oryza sativa (japonica
cultivar-group)] gi|51091371|dbj|BAD36105.1| LMBR1
integral membrane protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 715
Score = 579 bits (1493), Expect = e-164
Identities = 275/407 (67%), Positives = 337/407 (82%)
Query: 1 TLNDAVNVTISFLWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLS 60
T+ + + F WS +YWSTF L+W++VP LQGYEDAGDFTVK RL+TS+H NLV+Y
Sbjct: 57 TITGSQEGDVGFFWSWTYWSTFFLSWSIVPTLQGYEDAGDFTVKERLKTSIHKNLVYYKI 116
Query: 61 LGSVALFGLILLITLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADW 120
+GS+ L G+IL+IT+ W+G + GFAMACSNTFGLVTGAFLLGFG+SEIPK IW ADW
Sbjct: 117 IGSIGLVGVILIITMRHDWAGGIMGFAMACSNTFGLVTGAFLLGFGLSEIPKNIWKTADW 176
Query: 121 AIQQKFLSHKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQML 180
+QKFL H++A MA K D+AHQ++ +AI + QATSKQM+KR+ LRP+M+IID ML QML
Sbjct: 177 TRRQKFLYHRIANMAGKFDNAHQEYCHAIAVVQATSKQMTKREPLRPFMDIIDDMLAQML 236
Query: 181 NEDPSFKPQGGRLGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELED 240
+DP FKP GG+LGE DMDYDTDE +MASLRR+LRRA E+YYR +S+YT +V+EALELED
Sbjct: 237 RDDPLFKPSGGKLGEDDMDYDTDENTMASLRRQLRRANEEYYRCKSKYTSYVMEALELED 296
Query: 241 TVKNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFA 300
T+KNY++RD+ W+Y+S LR R +G+ LD IEF+WRCIL+KQL K LAVILG +S A
Sbjct: 297 TIKNYEQRDANEWKYVSGLRESRSCTLGSFLDFIEFIWRCILKKQLLKVLAVILGCISAA 356
Query: 301 ILLAEATILPSGVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFY 360
ILLAEAT+LPS VDLSLFS+L + G EVLVQ+ AF+PLMYMC+CTYYSLF++GM + Y
Sbjct: 357 ILLAEATLLPSDVDLSLFSVLTNVVGKQEVLVQVVAFIPLMYMCICTYYSLFRIGMMVVY 416
Query: 361 SLTPRQTSSVSLLMICSMVARYAAPISYNFLNLINIGGDRKTVFEKR 407
SLTPRQTSSVSLLMICSMVARYAAPISYNFLNLI++GG+ KT FEKR
Sbjct: 417 SLTPRQTSSVSLLMICSMVARYAAPISYNFLNLIHLGGNSKTTFEKR 463
>ref|XP_312472.1| ENSANGP00000003057 [Anopheles gambiae str. PEST]
gi|21296027|gb|EAA08172.1| ENSANGP00000003057 [Anopheles
gambiae str. PEST]
Length = 695
Score = 174 bits (442), Expect = 6e-42
Identities = 108/392 (27%), Positives = 207/392 (52%), Gaps = 17/392 (4%)
Query: 13 LWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVF---YLSLGSVALFGL 69
LW + YWS+ LTW ++PL+Q Y AGDFT+K +LR++L N ++ YL + + L L
Sbjct: 100 LWRIIYWSSQFLTWLIMPLMQSYLKAGDFTIKGKLRSALVDNAIYYGTYLFICGILLIYL 159
Query: 70 ILLITLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLSH 129
L ++ W ++ A + SNT+GL LLG+ + E+P+ +W N+ ++
Sbjct: 160 ALQPGISLDWQ-KLKAIASSASNTWGLFLLVLLLGYALVEVPRSLWNNSKPGFTLQYAYF 218
Query: 130 KVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYM-NIIDKMLVQMLNEDPSFKP 188
K++K++ + +A ++ + + Q+ S+ + R LRP + II K+ +++
Sbjct: 219 KLSKLSSEKAEAEENVDDVLESLQSASRAIPPRHELRPALETIIRKVPTELMERASRISR 278
Query: 189 QGGRLGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVKNYDRR 248
+ G S M + EK++ L R++ ++ + R + ++ V + L LED KN
Sbjct: 279 EDG----SPMAIPS-EKALVRLHRQVIKSLQTLQRTEALWSVQVNKVLHLEDVAKNAVSL 333
Query: 249 DSTGWRYISCLRPERIGKVGAVLD-TIEFLWRCILRKQLEKSLAVILGFMSFAILLAEAT 307
D R+ S R+G A+ T+++ W C+++ K+LA+I F+SF ++ +E T
Sbjct: 334 DH---RFKSEFPKHRVGMARALYSPTVDWYWECVVKAPFLKALAIITAFLSFTVVWSELT 390
Query: 308 ILPSGVDLSLFSILVHAA--GHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMF-MFYSLTP 364
LS+F+ +++ A + V +++ + + L Y+C C Y ++F++ ++Y
Sbjct: 391 FFNRQPVLSIFANVLNVAKGSYDFVTIEIFSMLTLCYLCYCAYSTVFRIKFLNLYYLAAH 450
Query: 365 RQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
QT+ SL+ ++ R P+ NFL +I++
Sbjct: 451 HQTNEYSLIFSGMLLCRLTPPMCLNFLGMIHM 482
>ref|XP_638788.1| hypothetical protein DDB0185797 [Dictyostelium discoideum]
gi|60467407|gb|EAL65433.1| hypothetical protein
DDB0185797 [Dictyostelium discoideum]
Length = 733
Score = 171 bits (432), Expect = 9e-41
Identities = 112/403 (27%), Positives = 198/403 (48%), Gaps = 18/403 (4%)
Query: 7 NVTISFLWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVAL 66
N + +++ Y+ T LLTW V PL+ + AGDF + R+ S+ N YL G + L
Sbjct: 115 NDKLEYIYQTFYFGTLLLTWLVYPLMGSFVLAGDFHLSGRITRSIKENAYLYLIFGVIGL 174
Query: 67 FGLILLITLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKF 126
+I L+ + + S+ GFAMA +NT+GL L+G+G+ E P+ IW+++ ++ K
Sbjct: 175 VVMIWLLAVKQLDWNSMVGFAMAAANTWGLCLVIILMGYGLVETPRSIWVSSQRSLVLKH 234
Query: 127 LSHKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSF 186
L K ++ A+++ + + + ++ K D Y+ II + Q E +
Sbjct: 235 LQFKAVELLNSKKKANEELIATMKVIRRIQEKTKKYDPYEKYIKII---VDQCPPEQYAL 291
Query: 187 KPQGGRLGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVKNYD 246
+G G+ + Y + +L RL+ A R Y + ++EA EL+D +
Sbjct: 292 VQRGE--GDGEATYSV----LVALNSRLKNAITNSQRAEFLYDQCLVEAFELQDIQNSAI 345
Query: 247 RRD-STGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAILLAE 305
D + W + R +R G+ LD +E++W L + + A++ +S I+ +E
Sbjct: 346 SLDKNVNWSF----REQRQGRFAKKLDIMEWIWYNYLEISVFRVAAIVFAVLSLLIIWSE 401
Query: 306 ATILPSGVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYSLTPR 365
+ + D+S+ S +V + + + VQ F PL Y + Y +LFK+ +F +Y L P
Sbjct: 402 FALAFTSFDISVLSNIVKHSNVSNIFVQFILFFPLGYEALTCYSTLFKIRIFNYYRLIPH 461
Query: 366 QTS-SVSLLMICSMVARYAAPISYNFLNLINIGG---DRKTVF 404
Q S S S++ + + R AP+ YNF+ IN+ D +T F
Sbjct: 462 QHSDSNSIIFSAAYLCRLGAPLCYNFIQFINMNSGIEDNRTSF 504
>gb|AAM50128.1| GH05505p [Drosophila melanogaster]
Length = 694
Score = 153 bits (387), Expect = 1e-35
Identities = 105/395 (26%), Positives = 195/395 (48%), Gaps = 17/395 (4%)
Query: 13 LWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILL 72
LW + YWS+ LTW ++PL+Q Y AGDFTVK +L+++L N ++Y S + + G++L+
Sbjct: 104 LWRIIYWSSQFLTWLIMPLMQSYLKAGDFTVKGKLKSALIENAIYYGSY--LFICGVLLI 161
Query: 73 ITLNKFWS---GSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLSH 129
K S ++ A + SNT+GL LLG+ + E+P+ +W NA ++
Sbjct: 162 YIAVKGESLDWQKLKAIASSASNTWGLFLLILLLGYALVEVPRSLWNNAKPGFALQYAYF 221
Query: 130 KVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPS--FK 187
K AK++ + +A + + + Q S+ + LRP + I + + L E S F
Sbjct: 222 KAAKLSTEKAEAEEHVDDILESLQGLSRVIPNNHELRPCLETILRKVPIELQERASRNFA 281
Query: 188 PQGGR-LGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVKNYD 246
GG +G + EK++ + +++ ++ + R + ++ V L LED KN
Sbjct: 282 RTGGSGMGATSSTILPSEKALVRIHKQVIKSLQTLQRTEALWSVQVQTVLHLEDVAKNIH 341
Query: 247 RRDSTGWRYISCLRPERIGKVGAVL--DTIEFLWRCILRKQLEKSLAVILGFMSFAILLA 304
D R P + ++ + ++++ W C+L+ K++ V+ MS ++ +
Sbjct: 342 SSD----RRFKSEFPRQRTQLERICYSASLQWYWECLLKAPFLKTMCVLTATMSAIVVWS 397
Query: 305 EATILPSGVDLSLFSILVHAA--GHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYSL 362
E T LS+F+ +++ A + +++ + V L Y CTY ++ ++ Y L
Sbjct: 398 ELTFFSRHPVLSIFANVIYVAKESYDFFTIEVFSMVVLCYFFYCTYSTILRIRFLNLYYL 457
Query: 363 TP-RQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
P QT+ SL+ ++ R P+ NFL LI++
Sbjct: 458 APHHQTNEHSLIFSGMLLCRLTPPMCLNFLGLIHM 492
>gb|AAF54372.1| CG8135-PA [Drosophila melanogaster] gi|24645348|ref|NP_649890.1|
CG8135-PA [Drosophila melanogaster]
Length = 694
Score = 153 bits (387), Expect = 1e-35
Identities = 105/395 (26%), Positives = 195/395 (48%), Gaps = 17/395 (4%)
Query: 13 LWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILL 72
LW + YWS+ LTW ++PL+Q Y AGDFTVK +L+++L N ++Y S + + G++L+
Sbjct: 104 LWRIIYWSSQFLTWLIMPLMQSYLKAGDFTVKGKLKSALIENAIYYGSY--LFICGVLLI 161
Query: 73 ITLNKFWS---GSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLSH 129
K S ++ A + SNT+GL LLG+ + E+P+ +W NA ++
Sbjct: 162 YIAVKGESLDWQKLKAIASSASNTWGLFLLILLLGYALVEVPRSLWNNAKPGFALQYAYF 221
Query: 130 KVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPS--FK 187
K AK++ + +A + + + Q S+ + LRP + I + + L E S F
Sbjct: 222 KAAKLSTEKAEAEEHVDDILESLQGLSRVIPNNHELRPCLETILRKVPIELQERASRNFA 281
Query: 188 PQGGR-LGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVKNYD 246
GG +G + EK++ + +++ ++ + R + ++ V L LED KN
Sbjct: 282 RTGGSGMGATSSTILPSEKALVRIHKQVIKSLQTLQRTEALWSVQVQTVLHLEDVAKNIH 341
Query: 247 RRDSTGWRYISCLRPERIGKVGAVL--DTIEFLWRCILRKQLEKSLAVILGFMSFAILLA 304
D R P + ++ + ++++ W C+L+ K++ V+ MS ++ +
Sbjct: 342 SSD----RRFKSEFPRQRTQLERICYSASLQWYWECLLKAPFLKTMCVLTATMSAMVVWS 397
Query: 305 EATILPSGVDLSLFSILVHAA--GHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYSL 362
E T LS+F+ +++ A + +++ + V L Y CTY ++ ++ Y L
Sbjct: 398 ELTFFSRHPVLSIFANVIYVAKESYDFFTIEVFSMVVLCYFFYCTYSTILRIRFLNLYYL 457
Query: 363 TP-RQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
P QT+ SL+ ++ R P+ NFL LI++
Sbjct: 458 APHHQTNEHSLIFSGMLLCRLTPPMCLNFLGLIHM 492
>gb|EAL26978.1| GA20843-PA [Drosophila pseudoobscura]
Length = 699
Score = 151 bits (381), Expect = 7e-35
Identities = 103/396 (26%), Positives = 194/396 (48%), Gaps = 19/396 (4%)
Query: 13 LWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILL 72
LW + YWS+ LTW ++PL+Q Y AGDFT+K +L+++L N ++Y S + + G++L+
Sbjct: 104 LWRIIYWSSQFLTWLIMPLMQSYLKAGDFTIKGKLKSALIENAIYYGSY--LFICGVLLI 161
Query: 73 ITLNK----FWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLS 128
K W ++ A + SNT+GL LLG+ + E+P+ +W NA ++
Sbjct: 162 YIAVKGVPLDWQ-KLKAIASSASNTWGLFLLILLLGYALVEVPRSLWNNAKPGFTLQYAY 220
Query: 129 HKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPS--F 186
K AK++ +A + + + Q S+ + LRP + I + + L E S F
Sbjct: 221 FKAAKLSTDKAEAEEHVDDILESLQCISRVVPNNHELRPCVETIMRKVPIELQERASRNF 280
Query: 187 KPQGGR-LGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVKNY 245
GG +G + EK++ + +++ ++ + R + ++ V L LED +N
Sbjct: 281 SRTGGSGMGATSSTILPSEKALVRIHKQVIKSLQTLQRTEALWSVQVDTVLHLEDVARNI 340
Query: 246 DRRDSTGWRYISCLRPERIGKVGAVL--DTIEFLWRCILRKQLEKSLAVILGFMSFAILL 303
+ R P + ++ + T+++ W C+L+ K+L V+ MS ++
Sbjct: 341 HSAE----RRFKSEFPRQRTQLERIFYSATLQWYWECLLKAPFLKTLCVVTATMSAMVVW 396
Query: 304 AEATILPSGVDLSLFSILVHAA--GHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYS 361
+E T LS+F+ +++ A + +++ + + L Y CTY ++ ++ Y
Sbjct: 397 SEVTFFSRDPVLSIFANVIYLAKESYDFFTIEVFSMMVLCYFFYCTYSTILRIRFLNLYY 456
Query: 362 LTP-RQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
L P QT+ SL+ ++ R P+ NFL LI++
Sbjct: 457 LAPHHQTNEHSLIFSGMLLCRLTPPMCLNFLGLIHM 492
>gb|AAH63324.1| Unknown (protein for MGC:73387) [Danio rerio]
gi|41152363|ref|NP_956238.1| hypothetical protein
LOC335257 [Danio rerio]
Length = 704
Score = 149 bits (377), Expect = 2e-34
Identities = 101/400 (25%), Positives = 206/400 (51%), Gaps = 29/400 (7%)
Query: 14 WSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFG-LILL 72
W + YW++ LTW ++P +Q Y +G FT+ +++T+L N ++Y + + +FG L++
Sbjct: 117 WRVVYWTSQCLTWLLLPFMQSYARSGGFTITGKIKTALIENAIYYGTY--LFIFGSLLIY 174
Query: 73 ITLNKFWSGS---VRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLSH 129
+ ++ W S ++ + +NT+GL LLG+G+ +IP+ W +
Sbjct: 175 VAVHPQWHLSWYELQTIGITAANTWGLFLLVLLLGYGLVDIPRSYWEASRQGHLLIKTYF 234
Query: 130 KVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDP-SFKP 188
K AK+ + D+ ++ + + +++ K + Y + + K + +L + P ++
Sbjct: 235 KAAKLMTEKADSEENLEDVM-------EEVRKINESIKYNHPLRKNVDTILRKCPLEYQE 287
Query: 189 QGGRLGESDMDYD------TDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTV 242
+ GR + D+D E+S++ L +++ A +++ R R ++ + +A LED
Sbjct: 288 KMGRNMDDFEDFDDKQNSYPTERSLSKLHKQVIYAVQRHNRTRVQWQILLEQAFHLEDVA 347
Query: 243 KNYDRRDSTGWRYI-SCLRPERIGKVGAVL--DTIEFLWRCILRKQLEKSLAVILGFMSF 299
KN ST +++ S PE +G + T E+ W C+L++ + LAV+L S
Sbjct: 348 KN---ETSTSRQFVHSFAPPEPVGWFRRYIYTPTAEWYWECLLKQWFYRVLAVVLALFSV 404
Query: 300 AILLAEATILPSGVDLSLFSILVHAA--GHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMF 357
A++ +E T + LSLF++ + A + + +++A F+ + ++C C Y ++F++ +F
Sbjct: 405 AVVWSECTFFSTHPVLSLFAVFIQLAERDYNYLYIEMACFITIFFLCTCVYSTVFRIRVF 464
Query: 358 MFYSL-TPRQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
+Y L + QT + SL + R P+ NFL LI++
Sbjct: 465 NYYYLASHHQTDAYSLQFSGMLFCRLTPPLCLNFLGLIHM 504
>ref|NP_796152.2| hypothetical protein LOC320506 [Mus musculus]
gi|26348001|dbj|BAC37649.1| unnamed protein product [Mus
musculus]
Length = 694
Score = 148 bits (374), Expect = 5e-34
Identities = 98/397 (24%), Positives = 191/397 (47%), Gaps = 23/397 (5%)
Query: 14 WSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILLI 73
W + YW++ LTW ++P +Q Y +G F++ +++T+L N ++Y + + +FG L+
Sbjct: 111 WRVVYWTSQFLTWILLPFMQSYARSGGFSITGKIKTALIENAIYYGTY--LLIFGAFLIY 168
Query: 74 T-----LNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLS 128
L+ W+ ++ +A +NT+GL LLG+G+ EIP+ W A
Sbjct: 169 VAVNPRLHLEWN-QLQTIGIAAANTWGLFLLVLLLGYGLVEIPRSYWNGAKRGYLLMKTY 227
Query: 129 HKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKP 188
K AK+ + DA ++ + + + ++ + LR ++ I K ++
Sbjct: 228 FKAAKLMTEKADAEENLEDVMEEVRKVNESIKYNHPLRKCVDTILKKC------PTDYQE 281
Query: 189 QGGRLGESDMDYD------TDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTV 242
+ GR + D+D EKS+ L +++ + +++ R + ++ + +A LED
Sbjct: 282 KMGRNMDDYEDFDEKRNTYPSEKSLVKLHKQVIYSVQRHRRTQVQWQILLEQAFYLEDVA 341
Query: 243 KNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAIL 302
KN PE T+E+ W C+LR ++LAV+L S ++
Sbjct: 342 KNETSATHQFVHTFQSPEPENRFIQYFYNPTVEWYWECLLRPWFHRTLAVVLSIFSVIVV 401
Query: 303 LAEATILPSGVDLSLFSILVHAAGHT--EVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFY 360
+E T + LSLF++ + A T + +++A F+ + ++ +C Y ++F++ +F +Y
Sbjct: 402 WSECTFFSTTPVLSLFAVFIQLAERTYNYIYIEIACFLSIFFLSICVYSTVFRIRVFNYY 461
Query: 361 SL-TPRQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
L + QT + SLL + R P+ NFL L ++
Sbjct: 462 YLASHHQTDAYSLLFSGMLFCRLTPPLCLNFLGLTHM 498
>ref|XP_527193.1| PREDICTED: similar to hypothetical protein [Pan troglodytes]
Length = 652
Score = 148 bits (374), Expect = 5e-34
Identities = 101/397 (25%), Positives = 192/397 (47%), Gaps = 23/397 (5%)
Query: 14 WSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILLI 73
W + YW++ LTW ++P +Q Y +G F++ +++T+L N ++Y + + +FG L+
Sbjct: 111 WRVVYWTSQFLTWILLPFMQSYARSGGFSITGKIKTALIENAIYYGTY--LLIFGAFLIY 168
Query: 74 T-----LNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLS 128
L+ W+ ++ +A +NT+GL LLG+G+ EIP+ W A
Sbjct: 169 VAVNPHLHLEWN-QLQTIGIAAANTWGLFLLVLLLGYGLVEIPRSYWNGAKRGYLLMKTY 227
Query: 129 HKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKP 188
K AK+ + DA ++ +A+ + ++ + LR +D +L + E ++
Sbjct: 228 FKAAKLMTEKADAEENLEDAMEEVRKVNESIKYNHPLR---KCVDTILKKCPTE---YQE 281
Query: 189 QGGRLGESDMDYD------TDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTV 242
+ GR + D+D EKS+ L +++ + +++ R + ++ + +A LED
Sbjct: 282 KMGRNMDDYEDFDEKHNIYPSEKSLVKLHKQVIYSVQRHRRTQVQWQILLEQAFYLEDVA 341
Query: 243 KNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAIL 302
KN PE T E+ W C+LR K LAV+L S ++
Sbjct: 342 KNETSATHQFVHTFQSPEPENRFIQYFYNPTFEWYWECLLRPWFYKILAVVLSIFSVIVV 401
Query: 303 LAEATILPSGVDLSLFSILVHAAGHT--EVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFY 360
+E T + LSLF++ + A T + +++A F+ + ++ +C Y ++F++ +F +Y
Sbjct: 402 WSECTFFSTTPVLSLFAVFIQLAEKTYNYIYIEIACFLSIFFLSICVYSTVFRIRVFNYY 461
Query: 361 SL-TPRQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
L + QT + SLL + R P+ NFL L ++
Sbjct: 462 YLASHHQTDAYSLLFSGMLFCRLTPPLCLNFLGLTHM 498
>emb|CAH18245.1| hypothetical protein [Homo sapiens] gi|56790897|ref|NP_001007528.1|
hypothetical protein LOC92255 [Homo sapiens]
Length = 695
Score = 148 bits (374), Expect = 5e-34
Identities = 101/397 (25%), Positives = 192/397 (47%), Gaps = 23/397 (5%)
Query: 14 WSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILLI 73
W + YW++ LTW ++P +Q Y +G F++ +++T+L N ++Y + + +FG L+
Sbjct: 111 WRVVYWTSQFLTWILLPFMQSYARSGGFSITGKIKTALIENAIYYGTY--LLIFGAFLIY 168
Query: 74 T-----LNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLS 128
L+ W+ ++ +A +NT+GL LLG+G+ EIP+ W A
Sbjct: 169 VAVNPHLHLEWN-QLQTIGIAAANTWGLFLLVLLLGYGLVEIPRSYWNGAKRGYLLMKTY 227
Query: 129 HKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKP 188
K AK+ + DA ++ +A+ + ++ + LR +D +L + E ++
Sbjct: 228 FKAAKLMTEKADAEENLEDAMEEVRKVNESIKYNHPLR---KCVDTILKKCPTE---YQE 281
Query: 189 QGGRLGESDMDYD------TDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTV 242
+ GR + D+D EKS+ L +++ + +++ R + ++ + +A LED
Sbjct: 282 KMGRNMDDYEDFDEKHSIYPSEKSLVKLHKQVIYSVQRHRRTQVQWQILLEQAFYLEDVA 341
Query: 243 KNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAIL 302
KN PE T E+ W C+LR K LAV+L S ++
Sbjct: 342 KNETSATHQFVHTFQSPEPENRFIQYFYNPTFEWYWECLLRPWFYKILAVVLSIFSVIVV 401
Query: 303 LAEATILPSGVDLSLFSILVHAAGHT--EVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFY 360
+E T + LSLF++ + A T + +++A F+ + ++ +C Y ++F++ +F +Y
Sbjct: 402 WSECTFFSTTPVLSLFAVFIQLAEKTYNYIYIEIACFLSIFFLSICVYSTVFRIRVFNYY 461
Query: 361 SL-TPRQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
L + QT + SLL + R P+ NFL L ++
Sbjct: 462 YLASHHQTDAYSLLFSGMLFCRLTPPLCLNFLGLTHM 498
>ref|XP_623561.1| PREDICTED: similar to ENSANGP00000003057 [Apis mellifera]
Length = 688
Score = 147 bits (371), Expect = 1e-33
Identities = 101/397 (25%), Positives = 193/397 (48%), Gaps = 27/397 (6%)
Query: 13 LWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILL 72
LW + YW++ LTW ++PL+Q Y AGDFTV+ +L+++L N ++Y S + + G++L+
Sbjct: 103 LWRIVYWTSQCLTWLILPLMQSYIKAGDFTVQGKLKSALIDNAIYYGSY--LFICGVLLI 160
Query: 73 ITLNK----FWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLS 128
K ++ A + SNT+GL LLG+ + E+P+G+W + +
Sbjct: 161 YIALKPGFDLDGQKLKAIASSASNTWGLFLLVLLLGYALVEVPRGLWNASKPGYTLNYSY 220
Query: 129 HKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKP 188
KVAK+++ +A + + + QA + + + I + + L +
Sbjct: 221 FKVAKLSLDKCEAEETVDDILESLQAATISIGPGHPFHCNLETIFQKIPAELKD------ 274
Query: 189 QGGRLGESDMDYDT-----DEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVK 243
R+ + DT EK++ L R+ +A + R +++ V + ++LED +
Sbjct: 275 ---RMNRRQLPDDTPTDTPSEKALIRLHRQTIKALQTLQRIETQWGILVDKIIDLEDIAR 331
Query: 244 NYDRRDSTGWRYISCLRPERIGKVGAVLD-TIEFLWRCILRKQLEKSLAVILGFMSFAIL 302
N D R+ R + + IE+ W+CI++ ++K +A+ G +S A++
Sbjct: 332 NQLNDDR---RFKPTFPTHRSFPFCIIYNPIIEWYWKCIIKSYVQKIVAICTGCLSIAVV 388
Query: 303 LAEATILPSGVDLSLFSILVHAA--GHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFY 360
+E T LSLF+ ++ A + +++ + + + Y+C C Y ++ K+ + Y
Sbjct: 389 WSEVTFFNKSPVLSLFAQFLNLAKENYDYFTIEVLSMLIIAYLCYCAYSTVLKIRVLNLY 448
Query: 361 SLTP-RQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
L P QT+ SL+ M+ R P+ NFL LI++
Sbjct: 449 YLAPHHQTNEYSLIFSGMMLCRLTPPMCLNFLGLIHM 485
>ref|XP_574861.1| PREDICTED: similar to RIKEN cDNA 9930036E21 gene [Rattus
norvegicus]
Length = 694
Score = 147 bits (370), Expect = 1e-33
Identities = 98/397 (24%), Positives = 190/397 (47%), Gaps = 23/397 (5%)
Query: 14 WSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILLI 73
W + YW++ LTW ++P +Q Y +G F++ +++T+L N ++Y + + +FG L+
Sbjct: 111 WRVVYWTSQFLTWILLPFMQSYARSGGFSITGKIKTALIENAIYYGTY--LLIFGAFLIY 168
Query: 74 T-----LNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLS 128
L+ W+ ++ +A +NT+GL LLG+G+ EIP+ W A
Sbjct: 169 VAVNPRLHLEWN-QLQTIGIAAANTWGLFLLVLLLGYGLVEIPRSYWNGAKRGYLLMKTY 227
Query: 129 HKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKP 188
K AK+ + DA ++ + + + ++ + LR ++ I K ++
Sbjct: 228 FKAAKLMTEKADAEENLEDVMEEVRKVNESIKYNHPLRKCVDTILKKC------PTDYQE 281
Query: 189 QGGRLGESDMDYD------TDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTV 242
+ GR + D+D EKS+ L +++ + +++ R + ++ + +A LED
Sbjct: 282 KMGRNMDDYEDFDEKRNTYPSEKSLVKLHKQVIYSVQRHRRTQVQWQILLEQAFYLEDVA 341
Query: 243 KNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAIL 302
KN PE T+E+ W C+LR + LAV+L S ++
Sbjct: 342 KNETSATHQFVHTFQSPEPENRFIQYFYNPTVEWYWECLLRPWFHRILAVVLSIFSVIVV 401
Query: 303 LAEATILPSGVDLSLFSILVHAAGHT--EVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFY 360
+E T + LSLF++ + A T + +++A F+ + ++ +C Y ++F++ +F +Y
Sbjct: 402 WSECTFFSTTPVLSLFAVFIQLAEKTYNYIYIEIACFLSIFFLSICVYSTVFRIRVFNYY 461
Query: 361 SL-TPRQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
L + QT + SLL + R P+ NFL L ++
Sbjct: 462 YLASHHQTDAYSLLFSGMLFCRLTPPLCLNFLGLTHM 498
>gb|AAH43881.1| Dkfzp434h2226-prov protein [Xenopus laevis]
Length = 713
Score = 145 bits (366), Expect = 4e-33
Identities = 99/398 (24%), Positives = 194/398 (47%), Gaps = 25/398 (6%)
Query: 14 WSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILLI 73
W + YW++ LTW ++P +Q Y +G F++ +++T+L N ++Y + + +FG +L+
Sbjct: 128 WRVVYWTSQFLTWILMPFMQSYARSGGFSITGKIKTALIENAIYYGTY--LLIFGALLIY 185
Query: 74 -----TLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLS 128
L+ W ++ +A +NT+GL L+G+G+ EIP+ W A
Sbjct: 186 VAVNPNLHLEWY-QLQTIGIAAANTWGLFLLVLLMGYGLVEIPRSQWNGAKKGYLLMKTY 244
Query: 129 HKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPS-FK 187
K AK+ + DA + + + +++ K + Y + + K + +L + P+ ++
Sbjct: 245 FKAAKLMTEKADAEETLEDVM-------EEVRKVNECIKYNHPLRKCVDTILKKCPAEYQ 297
Query: 188 PQGGRLGESDMDYD------TDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDT 241
+ GR + D++ EK++ L +++ A +++ R + +++ + +A LED
Sbjct: 298 EKMGRNMDDYEDFEEKNISYPSEKTLVKLHKQVIYAVQRHRRTQVQWSILLEQAFHLEDV 357
Query: 242 VKNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAI 301
KN PE TIE+ W C+LR + LAVIL S +
Sbjct: 358 AKNETSAAKQFVHTFPHQEPESWIMRRLYTPTIEWYWECLLRPWCSRILAVILALFSTVV 417
Query: 302 LLAEATILPSGVDLSLFSILVHAA--GHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMF 359
+ +E T + LSLF++ + A H + V++ F+ + ++ +C Y ++F++ +F +
Sbjct: 418 VWSECTFFSAKPVLSLFAVFIQQAEQTHNYIYVEVVCFLSIFFLSICVYSTVFRIRVFNY 477
Query: 360 YSL-TPRQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
Y L + QT + SLL + R P+ NFL L ++
Sbjct: 478 YYLASHHQTDAYSLLFSGMLFCRLTPPLCLNFLGLTHM 515
>ref|XP_546347.1| PREDICTED: similar to hypothetical protein [Canis familiaris]
Length = 668
Score = 145 bits (366), Expect = 4e-33
Identities = 99/397 (24%), Positives = 190/397 (46%), Gaps = 23/397 (5%)
Query: 14 WSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILLI 73
W + YW++ LTW ++P +Q Y +G F++ +++T+L N ++Y + + +FG L+
Sbjct: 111 WRVVYWTSQFLTWILLPFMQSYARSGGFSITGKIKTALIENAIYYGTY--LLIFGAFLIY 168
Query: 74 T-----LNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLS 128
L+ W+ ++ +A +NT+GL LLG+G+ EIP+ W A
Sbjct: 169 VAVNPHLHLEWN-QLQTIGIAAANTWGLFLLVLLLGYGLVEIPRSYWNGAKRGYLLMKTY 227
Query: 129 HKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKP 188
K AK+ + DA + + + + + + LR +D +L + E ++
Sbjct: 228 FKAAKLMTEKADAEETLEDVMEEVRKVNDSIKYNHPLR---KCVDTILKKCPTE---YQE 281
Query: 189 QGGRLGESDMDYD------TDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTV 242
+ GR + D+D EKS+ L +++ + +++ R + ++ + +A LED
Sbjct: 282 KMGRNMDDYEDFDEKHNTYPSEKSLVKLHKQVIYSVQRHRRTQVQWQILLEQAFYLEDVA 341
Query: 243 KNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAIL 302
KN PE T+E+ W C+LR + LAV+L S ++
Sbjct: 342 KNETSATHQFVHTFQSPEPENKFIQYFYNPTVEWYWECLLRPWFYRILAVVLSIFSVIVV 401
Query: 303 LAEATILPSGVDLSLFSILVHAAGHT--EVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFY 360
+E T + LSLF++ + A T + +++A F+ + ++ +C Y ++F++ +F +Y
Sbjct: 402 WSECTFFSTTPVLSLFAVFIQLAEKTYNYIYIEIACFLSIFFLSICVYSTVFRIRVFNYY 461
Query: 361 SL-TPRQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
L + QT + SLL + R P+ NFL L ++
Sbjct: 462 YLASHHQTDAYSLLFSGMLFCRLTPPLCLNFLGLTHM 498
>emb|CAH65329.1| hypothetical protein [Gallus gallus]
Length = 688
Score = 145 bits (366), Expect = 4e-33
Identities = 100/404 (24%), Positives = 192/404 (46%), Gaps = 23/404 (5%)
Query: 7 NVTISFLWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVAL 66
N + W + YW++ LTW ++P +Q Y +G F++ +++T+L N ++Y + + +
Sbjct: 98 NGIMPIFWRVVYWTSQFLTWILLPFMQSYARSGGFSITGKIKTALIENAIYYGTY--LLI 155
Query: 67 FGLILLIT-----LNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWA 121
FG L+ N W+ ++ +A +NT+GL LLG+G+ EIP+ W A
Sbjct: 156 FGAFLIYVAVNPKFNLQWN-QLQTIGIAAANTWGLFLLVLLLGYGLVEIPRSHWNGAKRG 214
Query: 122 IQQKFLSHKVAKMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLN 181
K AK+ + DA ++ + + + S+ + LR +D +L +
Sbjct: 215 YLLMKTYFKAAKLMTEKADAEENLEDIMEEVRKVSESIKYNHPLR---KCVDTILKKCPT 271
Query: 182 EDPSFKPQGGRLGESDMDYD------TDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEA 235
E ++ + GR + D+D EKS+ L +++ + +++ R + ++ + +A
Sbjct: 272 E---YQERMGRNMDDYEDFDERQNSYPSEKSLVKLHKQVIYSVQRHRRTQVQWQILLEQA 328
Query: 236 LELEDTVKNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILG 295
LED KN PE T+E+ W C+LR + LAV+L
Sbjct: 329 FYLEDVAKNETSATRQFVHTFQSQEPENKIIQYFYTPTVEWYWECLLRPWFYRVLAVVLA 388
Query: 296 FMSFAILLAEATILPSGVDLSLFSILVHAAGHT--EVLVQLAAFVPLMYMCVCTYYSLFK 353
S ++ +E T + LSL ++ + A T + +++A F+ + ++ +C Y ++F+
Sbjct: 389 AFSVIVVWSECTFFSTRPVLSLVAVFIQLAEKTYNYIYIEMACFLTIFFLSICVYSTVFR 448
Query: 354 MGMFMFYSL-TPRQTSSVSLLMICSMVARYAAPISYNFLNLINI 396
+ +F +Y L + QT + SLL + R P+ NFL L ++
Sbjct: 449 IRVFNYYYLASHHQTDAYSLLFSGMLFCRLTPPLCLNFLGLTHM 492
>ref|XP_640569.1| hypothetical protein DDB0204597 [Dictyostelium discoideum]
gi|60468593|gb|EAL66596.1| hypothetical protein
DDB0204597 [Dictyostelium discoideum]
Length = 790
Score = 128 bits (322), Expect = 5e-28
Identities = 86/398 (21%), Positives = 183/398 (45%), Gaps = 14/398 (3%)
Query: 14 WSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILLI 73
W Y+ + +L W + P+LQ + AGDF R++ + N++ Y + L G+I+++
Sbjct: 126 WRFLYFGSLILCWVIFPVLQSFSTAGDFRFWERVKRATKENVILYTFMFIAGLIGIIVIL 185
Query: 74 TLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNAD-WAIQQKFLSHKVA 132
++ + S F M +N +G++ +G+G+ ++P+ + +AI + + V
Sbjct: 186 SVKEMDPSSFLSFVMLLANVYGVILITITMGYGLIDVPRNLLRKGSHYAILRNYRVEAVV 245
Query: 133 KMAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKML---VQMLNEDPSFKPQ 189
+ +L+D + + + + + S + + D R Y+++I +L ++ +P
Sbjct: 246 -LKTELEDVKRQLIDHLKLIKTISDRAGQYDPFRIYLDVIISKCPTEYDVLIQEYHAEPL 304
Query: 190 GGRLGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVKNYDRRD 249
+ + Y K + + L ++ + Y + + +A +ED ++ +R
Sbjct: 305 PAGMEAEILSY----KYLVGIHSTLLDLVDRNHSAEVLYERLLGKAFAIEDIIETRERNK 360
Query: 250 STGWRYISCLRPERIG---KVGAVLDTIEFLWRCILRKQLEKSLAVILGFMSFAILLAEA 306
T I K E+LW + ++ +S IL +E
Sbjct: 361 QTAGNNQIAGEERSIQWSFKSTKSNGKFEYLWHMYIHPWYFIIAGLVCVCLSGIILWSEI 420
Query: 307 TI-LPSGVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYSLTPR 365
+ L S D S F + + +Q+ F+P++YMCVC+Y +LFK+ + +Y L P+
Sbjct: 421 VLALVSNPDYSPFYRAI-VRMEPGIGLQIFCFIPMIYMCVCSYSTLFKLRISNYYRLVPQ 479
Query: 366 QTSSVSLLMICSMVARYAAPISYNFLNLINIGGDRKTV 403
Q+++ S++ + + R AAP++YNF+ + ++ D V
Sbjct: 480 QSNTFSIMFSANYLCRLAAPLAYNFIQICHVNQDNSIV 517
>gb|EAL88319.1| LMBR and BAG domain protein, putative [Aspergillus fumigatus Af293]
Length = 1087
Score = 127 bits (319), Expect = 1e-27
Identities = 110/409 (26%), Positives = 180/409 (43%), Gaps = 37/409 (9%)
Query: 14 WSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALFGLILLI 73
W ++YW F+LTWA++PLL Y D+G K+R+ SL N + L + A GLI +
Sbjct: 91 WRITYWLIFVLTWAILPLLGEYIDSGYRDTKSRILYSLRSNARYQLIVLCCAFVGLIYIS 150
Query: 74 TLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQKFLSHKVAK 133
N F S++ MA + +GLV + +G G+ IP+ ++ NA+ + + + + +
Sbjct: 151 IQNGFRFASIKALVMALAYVWGLVLAIYAMGHGLVSIPRTLFRNANVSGRLRRVQAHAPR 210
Query: 134 MAVKLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKPQGGRL 193
+ +L DA D + + +Q T Q K + R + + I+++ +
Sbjct: 211 LHDRLMDAINDLES--LESQVTQLQQRKTGTARDFQDWIEELAETSSPIESRSAILEPSN 268
Query: 194 GESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEALELEDTVKN--------- 244
+ + E+ MA L RRL+RAR Q R+ E+ + VL A +L+ + +
Sbjct: 269 AQGTVPPVITERYMADLTRRLQRARHQKARFVDEWDRLVLLAADLQAIIDSSASRKLDFG 328
Query: 245 YDRRDSTGWRYISCLRP--------ERIGKVGAVLDTI-EFLWRCILRKQLEKSLAVILG 295
R ST +I L P I + VL + CI+ +L KSLA L
Sbjct: 329 QSPRRSTSLPHIKFLTPYMRYHLYVNIIPSLRLVLGALCAIASACIIWSELVKSLAPRLS 388
Query: 296 FMSFAILLAEATILPSGVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTYYSLFKMG 355
++ +I+ P VD QLAA L+YMC +
Sbjct: 389 VVTLSIVSYHKD--PKPVDFGR---------------QLAASAWLLYMCAAALGGVNDAK 431
Query: 356 MFMFYSLTPRQTSSVSLLMICSMVARYAAPISYNFLNLINIGGDRKTVF 404
++ +L R T S S+VAR PI+YNFL + + + T+F
Sbjct: 432 VWGNRALVRRNTYGESACWYASLVARLTVPIAYNFLTFLPLSVRQNTIF 480
>emb|CAG88018.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50422427|ref|XP_459779.1| unnamed protein product
[Debaryomyces hansenii]
Length = 667
Score = 120 bits (302), Expect = 1e-25
Identities = 96/412 (23%), Positives = 178/412 (42%), Gaps = 23/412 (5%)
Query: 4 DAVNVTISFLWSLSYWSTFLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGS 63
D + I +LW +YW TFLLTW ++PLLQ + +G F + ++ + ++ N+ F L +
Sbjct: 68 DLPDKVILYLWKFNYWITFLLTWLILPLLQEFFRSGYFNILSKFKDAVKRNIKFQLIILG 127
Query: 64 VALFGLILLITLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQ 123
V+ GL+ LI ++ +A S+ + LV +L+ G+ IP+ W++
Sbjct: 128 VSTAGLVYLILEVGLSLSHLKLMIIALSHIYSLVLALWLMAHGLISIPRNKWISGSLLQD 187
Query: 124 QKFLSHKVAKMAVKLDDAHQDFSNAIV--------ITQATSKQMSKRDSLRPYMNIIDKM 175
KV K+ L+D F ++ T + + RD + I
Sbjct: 188 LNHHYLKVPKLVDNLEDIKITFKEDVLQVLVLTKNFTSTSGEDFRFRDWILQLHKKIPLD 247
Query: 176 LVQMLNEDPSFKPQGGRLGESDMDYDTDEKSMASLRRRLRRAREQYYRYRSEYTKFVLEA 235
+ + + + + G + + K A+ L + Y +E+ + +
Sbjct: 248 IKEFMEQQYTHDDPGNTISRDQVTTHFMTKLTANFNLNLNKLNS----YEAEFNSVLKKI 303
Query: 236 LELEDTVKNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWRCILRKQLEKSLAVILG 295
+ LED V N D+ R R + + +L +F+ +R + + L+++L
Sbjct: 304 VSLED-VLNCSANDNLQQRSRLVYRID--NHLTLLLPKNKFMLEYYIRPVINRLLSIVLF 360
Query: 296 FMSFAILLAEATILPSGVDLSLFSILVHAAG--HTEVLVQLAAFVPLMYMCVCTYYSLFK 353
SF IL +E +SL +IL+++ G + +L + + YM + SL +
Sbjct: 361 VSSFIILESE---FFHSTPISLMNILIYSTGINNHNLLQLIVCCITFSYMLFASLNSLTR 417
Query: 354 MGMFMFYSLTPRQTSSVSLLMICSMVARYAAPISYNFLNLINIGGDRKTVFE 405
+ +F Y L P + VS + +AR P+SYNF+ L R ++FE
Sbjct: 418 LKIFNMYHLVPHNSDPVSACFYTTYIARLTIPLSYNFITLF---VSRNSIFE 466
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.348 0.152 0.521
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 990,410,332
Number of Sequences: 2540612
Number of extensions: 38412202
Number of successful extensions: 167912
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 43
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 167729
Number of HSP's gapped (non-prelim): 83
length of query: 660
length of database: 863,360,394
effective HSP length: 135
effective length of query: 525
effective length of database: 520,377,774
effective search space: 273198331350
effective search space used: 273198331350
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 79 (35.0 bits)
Medicago: description of AC121245.1


