

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121241.7 + phase: 0 /pseudo
(69 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
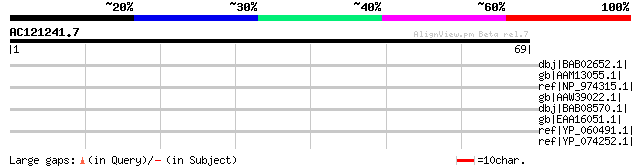
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB02652.1| unnamed protein product [Arabidopsis thaliana] 38 0.079
gb|AAM13055.1| unknown protein [Arabidopsis thaliana] gi|2508351... 38 0.079
ref|NP_974315.1| NHL repeat-containing protein [Arabidopsis thal... 38 0.079
gb|AAW39022.1| At5g55530 [Arabidopsis thaliana] gi|42573680|ref|... 35 0.67
dbj|BAB08570.1| unnamed protein product [Arabidopsis thaliana] 35 0.67
gb|EAA16051.1| putative yir1 protein [Plasmodium yoelii yoelii] 31 7.4
ref|YP_060491.1| LPXTG anchored putative adhesin [Streptococcus ... 31 9.6
ref|YP_074252.1| putative Fe-S oxidoreductase [Symbiobacterium t... 31 9.6
>dbj|BAB02652.1| unnamed protein product [Arabidopsis thaliana]
Length = 511
Score = 37.7 bits (86), Expect = 0.079
Identities = 20/39 (51%), Positives = 28/39 (71%), Gaps = 3/39 (7%)
Query: 26 TKEETGWPSFRQVV-DLLKLSLEEFTSSIL--KFMSKPH 61
TKEE GWPSF Q++ DL KL+LE TS ++ +F + P+
Sbjct: 326 TKEEPGWPSFGQLLTDLCKLALEFITSHLVPARFQTNPN 364
>gb|AAM13055.1| unknown protein [Arabidopsis thaliana] gi|25083516|gb|AAN72090.1|
unknown protein [Arabidopsis thaliana]
gi|22331093|ref|NP_188104.2| NHL repeat-containing
protein [Arabidopsis thaliana]
Length = 492
Score = 37.7 bits (86), Expect = 0.079
Identities = 20/39 (51%), Positives = 28/39 (71%), Gaps = 3/39 (7%)
Query: 26 TKEETGWPSFRQVV-DLLKLSLEEFTSSIL--KFMSKPH 61
TKEE GWPSF Q++ DL KL+LE TS ++ +F + P+
Sbjct: 307 TKEEPGWPSFGQLLTDLCKLALEFITSHLVPARFQTNPN 345
>ref|NP_974315.1| NHL repeat-containing protein [Arabidopsis thaliana]
Length = 493
Score = 37.7 bits (86), Expect = 0.079
Identities = 20/39 (51%), Positives = 28/39 (71%), Gaps = 3/39 (7%)
Query: 26 TKEETGWPSFRQVV-DLLKLSLEEFTSSIL--KFMSKPH 61
TKEE GWPSF Q++ DL KL+LE TS ++ +F + P+
Sbjct: 308 TKEEPGWPSFGQLLTDLCKLALEFITSHLVPARFQTNPN 346
>gb|AAW39022.1| At5g55530 [Arabidopsis thaliana] gi|42573680|ref|NP_974936.1| C2
domain-containing protein [Arabidopsis thaliana]
gi|30696612|ref|NP_200364.2| C2 domain-containing
protein [Arabidopsis thaliana]
gi|42573678|ref|NP_974935.1| C2 domain-containing
protein [Arabidopsis thaliana]
gi|59958302|gb|AAX12861.1| At5g55530 [Arabidopsis
thaliana]
Length = 405
Score = 34.7 bits (78), Expect = 0.67
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 6/72 (8%)
Query: 4 QPSERDFKGEASSDKSKSTLERTKEETGWPSFRQ-VVDLLKLSLEEFTSSILKF-----M 57
Q SE + GE S +K+ ++ K E +Q +VD+ S+++FT S+ K +
Sbjct: 311 QESESEASGETSEEKTVKSVLTVKVEPESKVVQQDIVDMYMKSMQQFTDSLAKMKLPLDI 370
Query: 58 SKPHTTERSSSD 69
P +E SSSD
Sbjct: 371 DSPTKSENSSSD 382
>dbj|BAB08570.1| unnamed protein product [Arabidopsis thaliana]
Length = 439
Score = 34.7 bits (78), Expect = 0.67
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 6/72 (8%)
Query: 4 QPSERDFKGEASSDKSKSTLERTKEETGWPSFRQ-VVDLLKLSLEEFTSSILKF-----M 57
Q SE + GE S +K+ ++ K E +Q +VD+ S+++FT S+ K +
Sbjct: 345 QESESEASGETSEEKTVKSVLTVKVEPESKVVQQDIVDMYMKSMQQFTDSLAKMKLPLDI 404
Query: 58 SKPHTTERSSSD 69
P +E SSSD
Sbjct: 405 DSPTKSENSSSD 416
>gb|EAA16051.1| putative yir1 protein [Plasmodium yoelii yoelii]
Length = 313
Score = 31.2 bits (69), Expect = 7.4
Identities = 16/52 (30%), Positives = 28/52 (53%), Gaps = 6/52 (11%)
Query: 17 DKSKSTLERTKEETGWPSFRQVV----DLLKLSLEEFTS--SILKFMSKPHT 62
D K + KE+TG+ +F++++ DLL + E+ + KF+ K HT
Sbjct: 123 DNGKDCNSKFKEKTGYNNFKEIIDKRKDLLNIKFEDISKLYDAFKFLCKMHT 174
>ref|YP_060491.1| LPXTG anchored putative adhesin [Streptococcus pyogenes MGAS10394]
gi|50903593|gb|AAT87308.1| LPXTG anchored putative
adhesin [Streptococcus pyogenes MGAS10394]
gi|40218526|gb|AAR83180.1| LPXTG anchored putative
adhesin [Streptococcus pyogenes]
Length = 1123
Score = 30.8 bits (68), Expect = 9.6
Identities = 20/67 (29%), Positives = 35/67 (51%), Gaps = 2/67 (2%)
Query: 3 QQPSERDFKGEASSDKSKSTLERTKEETGWPSFRQVVDLLKLSLEEFTSSILKFMSKPHT 62
++ SE+ + +A+ + +K LE K+E + +V+ K LE+F I K K T
Sbjct: 157 KESSEKVTELKANLESAKKDLE--KKEADYVKENALVERDKKDLEKFEKEIAKAREKKQT 214
Query: 63 TERSSSD 69
TE++ D
Sbjct: 215 TEKAIKD 221
>ref|YP_074252.1| putative Fe-S oxidoreductase [Symbiobacterium thermophilum IAM
14863] gi|51855250|dbj|BAD39408.1| putative Fe-S
oxidoreductase [Symbiobacterium thermophilum IAM 14863]
Length = 636
Score = 30.8 bits (68), Expect = 9.6
Identities = 21/62 (33%), Positives = 30/62 (47%), Gaps = 9/62 (14%)
Query: 1 MLQQPSERDFKGEASSDKSKSTLERTKEETGWPSFRQVVDLLKLSLEEFTSSILKFMSKP 60
+L P+E D EA ++K + LE +EE G R+V T S+ F+ KP
Sbjct: 413 ILGLPTETDADLEAIAEKGRWALEIAEEELGRQEARKV---------RVTISVSVFVPKP 463
Query: 61 HT 62
HT
Sbjct: 464 HT 465
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.306 0.122 0.322
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 94,279,816
Number of Sequences: 2540612
Number of extensions: 2595359
Number of successful extensions: 7103
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 7103
Number of HSP's gapped (non-prelim): 8
length of query: 69
length of database: 863,360,394
effective HSP length: 45
effective length of query: 24
effective length of database: 749,032,854
effective search space: 17976788496
effective search space used: 17976788496
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC121241.7


