

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121238.3 - phase: 0 /pseudo
(423 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
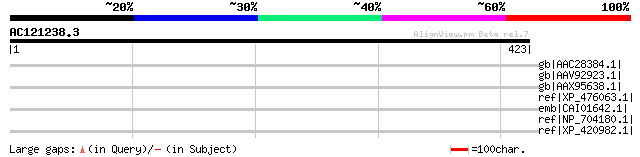
gb|AAC28384.1| mariner transposase [Glycine max] gi|7488706|pir|... 47 0.001
gb|AAV92923.1| transposase [Phytophthora infestans] 40 0.18
gb|AAX95638.1| hypothetical protein [Oryza sativa (japonica cult... 38 0.52
ref|XP_476063.1| hypothetical protein [Oryza sativa (japonica cu... 35 3.4
emb|CAI01642.1| protein kinase, putative [Plasmodium berghei] 35 5.7
ref|NP_704180.1| hypothetical protein [Plasmodium falciparum 3D7... 34 9.8
ref|XP_420982.1| PREDICTED: similar to CEGP1 protein [Gallus gal... 34 9.8
>gb|AAC28384.1| mariner transposase [Glycine max] gi|7488706|pir||T06335 probable
transposase - soybean transposon mariner element
Soymar1
Length = 425
Score = 46.6 bits (109), Expect = 0.001
Identities = 29/74 (39%), Positives = 44/74 (59%), Gaps = 3/74 (4%)
Query: 2 KRKRTFLDNYQRQNICEVLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWR--QKIETG 59
+RK L N +R I ++L+ S K+ +GV + ++SS+ V I IW+ ++ ET
Sbjct: 2 QRKVKMLSNEERITIYQLLLQKSVDGKLPQGVKESVASSFSVCRKTIDRIWKRAKESETH 61
Query: 60 DVCHKKTK-CGRKK 72
DV HKKTK GRK+
Sbjct: 62 DVSHKKTKNSGRKR 75
>gb|AAV92923.1| transposase [Phytophthora infestans]
Length = 200
Score = 39.7 bits (91), Expect = 0.18
Identities = 23/67 (34%), Positives = 38/67 (56%), Gaps = 6/67 (8%)
Query: 12 QRQNICEVLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIET-GD-----VCHKK 65
+RQ + E L+ S G + G + R+++ ++V V+ IWR+ I + GD V +K
Sbjct: 58 ERQAVWETLLLRSVGEVVKHGEIVRVAAYFRVGRQVVERIWRRGINSMGDRVAAVVKSRK 117
Query: 66 TKCGRKK 72
+ CGRKK
Sbjct: 118 SACGRKK 124
>gb|AAX95638.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|50918023|ref|XP_469408.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
gi|28273357|gb|AAO38443.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
Length = 565
Score = 38.1 bits (87), Expect = 0.52
Identities = 21/72 (29%), Positives = 34/72 (47%), Gaps = 7/72 (9%)
Query: 8 LDNYQRQNICEVLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQK-------IETGD 60
L N QRQ I E+L+ S K+ K + ++ + VS+ + IW++ ++
Sbjct: 144 LTNTQRQRIYELLLAKSKDGKLEKDTTRAVAQVFNVSIRTVQRIWKRAKLCIAHGVQVNV 203
Query: 61 VCHKKTKCGRKK 72
K CGRKK
Sbjct: 204 DSRKCYNCGRKK 215
>ref|XP_476063.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|47777448|gb|AAT38081.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 572
Score = 35.4 bits (80), Expect = 3.4
Identities = 15/58 (25%), Positives = 31/58 (52%), Gaps = 4/58 (6%)
Query: 8 LDNYQRQNICEVLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIETGDVCHKK 65
L N QR+ I ++L+ S K+ K + ++ + VS+ + IW++ +CH++
Sbjct: 151 LTNIQRRGIYQLLLQKSKDGKLEKHTTRLVAQEFHVSIRTVQRIWKR----AKICHEQ 204
>emb|CAI01642.1| protein kinase, putative [Plasmodium berghei]
Length = 178
Score = 34.7 bits (78), Expect = 5.7
Identities = 28/110 (25%), Positives = 48/110 (43%), Gaps = 24/110 (21%)
Query: 19 VLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIETGDVCHKKTKCGRKK------ 72
+L++ S KI+ +K + +++D Y ++ + + H K K + K
Sbjct: 74 ILLDGSLNAKISDFGIKEIEQCLDINIDYSYIVFPNNVIKFNNKHFKNKIKKIKIVNKGS 133
Query: 73 ---------KSHRYRKNT*--NTATQT*QSSISSRSIRVYHWTTNEVNKK 111
K+H Y+ NT N ++ T SS V+ WT+ EVNKK
Sbjct: 134 EDMLHVFSSKNHIYKYNTREINVSSNTHNSS-------VFFWTSPEVNKK 176
>ref|NP_704180.1| hypothetical protein [Plasmodium falciparum 3D7]
gi|23498918|emb|CAD50996.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 3351
Score = 33.9 bits (76), Expect = 9.8
Identities = 25/75 (33%), Positives = 35/75 (46%), Gaps = 11/75 (14%)
Query: 7 FLDNYQRQNICEVLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIET-----GDV 61
F D Y R+NI NS + INK ++KR + + ++ + R I DV
Sbjct: 1174 FYDAYDRENI-----NSKYVKVINKYIIKRNKNFVLIKKRIMNLMKRDNIHIRGYLRDDV 1228
Query: 62 CHKKTKCGRKKKSHR 76
K KC +KKK HR
Sbjct: 1229 LRNK-KCHKKKKDHR 1242
>ref|XP_420982.1| PREDICTED: similar to CEGP1 protein [Gallus gallus]
Length = 914
Score = 33.9 bits (76), Expect = 9.8
Identities = 18/61 (29%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 19 VLINSSSGRKINKGVVKRLSSSYQVSVDVIYAIWRQKIETGDVCHKKTKCGRKKKSHRYR 78
V + SSG+K++ G + RL ++ ++ + + + + E D C +C RKK R+R
Sbjct: 503 VNLTCSSGKKVH-GALSRLPATREIFITADFELETNQKEVTDTCD--VRCARKKTEKRFR 559
Query: 79 K 79
K
Sbjct: 560 K 560
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.372 0.162 0.611
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 510,111,154
Number of Sequences: 2540612
Number of extensions: 15669276
Number of successful extensions: 77184
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 77178
Number of HSP's gapped (non-prelim): 10
length of query: 423
length of database: 863,360,394
effective HSP length: 131
effective length of query: 292
effective length of database: 530,540,222
effective search space: 154917744824
effective search space used: 154917744824
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC121238.3


