

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121233.5 - phase: 0
(301 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
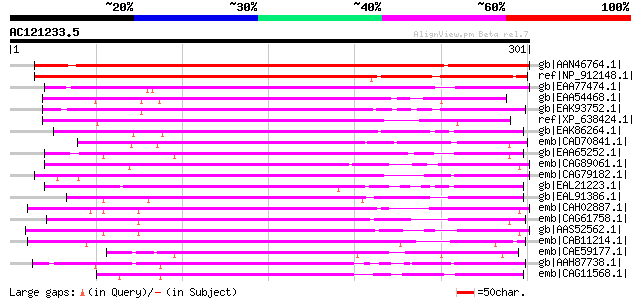
Score E
Sequences producing significant alignments: (bits) Value
gb|AAN46764.1| At3g22660/MWI23_3 [Arabidopsis thaliana] gi|92796... 362 7e-99
ref|NP_912148.1| putative EBNA1-binding protein homolog; Ebp2p [... 350 4e-95
gb|EAA77474.1| hypothetical protein FG07457.1 [Gibberella zeae P... 161 2e-38
gb|EAA54468.1| hypothetical protein MG02453.4 [Magnaporthe grise... 161 2e-38
gb|EAK93752.1| likely EBNA-like pre-rRNA processing factor EBP2p... 159 1e-37
ref|XP_638424.1| hypothetical protein DDB0186212 [Dictyostelium ... 158 2e-37
gb|EAK86264.1| hypothetical protein UM04809.1 [Ustilago maydis 5... 156 8e-37
emb|CAD70841.1| related to rRNA processing protein EBP2 [Neurosp... 154 3e-36
gb|EAA65252.1| hypothetical protein AN0074.2 [Aspergillus nidula... 149 1e-34
emb|CAG89061.1| unnamed protein product [Debaryomyces hansenii C... 148 2e-34
emb|CAG79182.1| unnamed protein product [Yarrowia lipolytica CLI... 145 2e-33
gb|EAL21223.1| hypothetical protein CNBD2780 [Cryptococcus neofo... 143 7e-33
gb|EAL91386.1| rRNA processing protein (Ebp2), putative [Aspergi... 142 2e-32
emb|CAH02887.1| unnamed protein product [Kluyveromyces lactis NR... 141 3e-32
emb|CAG61758.1| unnamed protein product [Candida glabrata CBS138... 141 3e-32
gb|AAS52562.1| AEL123Wp [Ashbya gossypii ATCC 10895] gi|45190484... 137 4e-31
emb|CAB11214.1| SPAC17H9.05 [Schizosaccharomyces pombe] gi|19114... 134 4e-30
emb|CAE59177.1| Hypothetical protein CBG02485 [Caenorhabditis br... 132 9e-30
gb|AAH87738.1| EBNA1 binding protein 2 (predicted) [Rattus norve... 129 1e-28
emb|CAG11568.1| unnamed protein product [Tetraodon nigroviridis] 129 1e-28
>gb|AAN46764.1| At3g22660/MWI23_3 [Arabidopsis thaliana] gi|9279684|dbj|BAB01241.1|
nucleolar protein-like [Arabidopsis thaliana]
gi|16604529|gb|AAL24270.1| AT3g22660/MWI23_3
[Arabidopsis thaliana] gi|15228822|ref|NP_188905.1| rRNA
processing protein-related [Arabidopsis thaliana]
gi|21362538|sp|Q9LUJ5|EBP2_ARATH Probable rRNA
processing protein EBP2 homolog
Length = 293
Score = 362 bits (929), Expect = 7e-99
Identities = 185/288 (64%), Positives = 229/288 (79%), Gaps = 7/288 (2%)
Query: 15 ELPLVNDDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPE 74
E+ ++++D D EAE LS+ S++E+E KLAEP+KTA+ NRD LLDKL DISWPE
Sbjct: 12 EMNMIDEDDATDSEAESLSD----SDTENEITEKLAEPTKTAIYNRDGLLDKLQDISWPE 67
Query: 75 NVEWIHKLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYA 134
+V+W HKL+++IDQ VDVNDDLARE AFYTQALEGTR+AF KL +G+ FLRPA+YYA
Sbjct: 68 DVDWTHKLTVEIDQGGAVDVNDDLARETAFYTQALEGTREAFGKLNEMGVNFLRPANYYA 127
Query: 135 EMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIES 194
EMVK+D HMEKVKSRLL EK+++ E+EERRKAR+ KR++KEVQSQK+KERAK+KKD+IES
Sbjct: 128 EMVKSDVHMEKVKSRLLHEKKQIEESEERRKARDNKRMAKEVQSQKMKERAKEKKDNIES 187
Query: 195 VKKWRKQRQQSGFADDGADKALDFEDGKVFER-SKKKRPGVSPGDRSGGKAKQAFGKGKK 253
VKKWRKQRQQSGF+D + LDFE GK F+R KKRPGVSPGDRSGGK + G
Sbjct: 188 VKKWRKQRQQSGFSDKAGEPELDFESGKSFQRGGGKKRPGVSPGDRSGGKGRPTSRMGN- 246
Query: 254 PKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKKRKR 301
KK + ++SKFG GG+KGL KQNTA+TTNDF G + G G+K++KR
Sbjct: 247 -KKREFRDSKFGHGGRKGLSKQNTAETTNDFKGGFRGGKASGNKRQKR 293
>ref|NP_912148.1| putative EBNA1-binding protein homolog; Ebp2p [Oryza sativa
(japonica cultivar-group)] gi|29027795|dbj|BAC65931.1|
putative EBNA1-binding protein homolog; Ebp2p [Oryza
sativa (japonica cultivar-group)]
Length = 291
Score = 350 bits (897), Expect = 4e-95
Identities = 185/290 (63%), Positives = 230/290 (78%), Gaps = 11/290 (3%)
Query: 15 ELPLVNDDTMIDGEAEYLSEEMSPSESESEEDVKL-AEPSKTAVNNRDALLDKLGDISWP 73
E PLV D+ ++D + + + + SESE + + AEPSK AV N++ +L+KL DI+WP
Sbjct: 7 EDPLVRDEAILDDDDDDVDTDEEESESEDDSGEEFHAEPSKKAVYNKEGILEKLEDIAWP 66
Query: 74 ENVEWIHKLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYY 133
ENV+W HKL+I+ DQ ++VDVNDDLARELAFYTQAL+GTRQAFEKLQS+ + FLRPADYY
Sbjct: 67 ENVDWRHKLTIEHDQGEKVDVNDDLARELAFYTQALDGTRQAFEKLQSMKVRFLRPADYY 126
Query: 134 AEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIE 193
AEMVKTD+HM K+K RLL+EK+K+ EAEER+KAREAK+ +KEVQ+QK KERAKQKK+ IE
Sbjct: 127 AEMVKTDAHMHKIKGRLLSEKKKIEEAEERKKAREAKKRAKEVQAQKEKERAKQKKEQIE 186
Query: 194 SVKKWRKQRQQSGFA---DDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGK 250
SVKKWRKQRQQ GFA DDG D L+FE + F++SKKKRPGVSPGDRSGG AK K
Sbjct: 187 SVKKWRKQRQQGGFAKGNDDGPD--LNFEGDEGFKQSKKKRPGVSPGDRSGGLAK----K 240
Query: 251 GKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKKRK 300
GK+ K ++SKFG GG+KGLKKQNTA+TTNDF GF++ +K+RK
Sbjct: 241 GKQGKNRKSRDSKFGHGGRKGLKKQNTAETTNDFRGFNQMDK-SQNKRRK 289
>gb|EAA77474.1| hypothetical protein FG07457.1 [Gibberella zeae PH-1]
gi|46126159|ref|XP_387633.1| hypothetical protein
FG07457.1 [Gibberella zeae PH-1]
Length = 418
Score = 161 bits (408), Expect = 2e-38
Identities = 110/289 (38%), Positives = 157/289 (54%), Gaps = 21/289 (7%)
Query: 21 DDTMIDGEAEYLSEEMSPSESESEEDVKLAEP-SKTAVNNRDALLDKLGDISWPENVEW- 78
+D D EA+ + MS E +ED + P + +NN ALL L IS P +
Sbjct: 131 EDEEEDPEADDIP--MSDLEDLDDEDKEDIIPHQRLTINNTTALLAALNRISVPTDKSVP 188
Query: 79 --IHK--LSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYA 134
H+ +S E DV DDL RELAFYTQ+LE TR A + L+ G+PF RP DY+A
Sbjct: 189 FATHQSLVSSSATAESIPDVQDDLQRELAFYTQSLEATRTARKLLRQEGIPFSRPKDYFA 248
Query: 135 EMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIES 194
EM+K D+HMEKVK++L+ E A+E RK R+ K+ K+VQ K++ER K+K++ ++
Sbjct: 249 EMIKEDAHMEKVKAKLVEEASNKKAAQEARKMRDLKKFGKQVQVAKMQERHKEKRETLDK 308
Query: 195 VKKWRKQRQQSGFADDGADKALDFEDGKVFE-RSKKKRPGVSPGDRSGGKAKQAFGKGKK 253
+K +++R ++G A +A F+ G E +S R G G +GG K+A
Sbjct: 309 IKNLKRKRSETGGAGLDTKEADIFDVGVEKEMKSHNPRSGAGRGAGAGGNHKRA------ 362
Query: 254 PKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGF-SKKGAVGGSKKRKR 301
KN K+GFGGKK K A ++ D GF +KK GG + R
Sbjct: 363 -----KKNEKYGFGGKKRHGKSGDAMSSGDLSGFDAKKMKAGGRVSKPR 406
>gb|EAA54468.1| hypothetical protein MG02453.4 [Magnaporthe grisea 70-15]
gi|39968721|ref|XP_365751.1| hypothetical protein
MG02453.4 [Magnaporthe grisea 70-15]
Length = 446
Score = 161 bits (408), Expect = 2e-38
Identities = 109/283 (38%), Positives = 157/283 (54%), Gaps = 24/283 (8%)
Query: 20 NDDTMIDGEAEYLSEEMSPSESES-EEDVK--LAEPSKTAVNNRDALLDKLGDISWPEN- 75
NDD D E + E++ S+ E ED K L + +NN ALL L I+ P +
Sbjct: 143 NDDDEEDEEEDEDDEDIPMSDLEDLPEDDKSDLVPHQRLTINNTTALLAALKRIALPTDG 202
Query: 76 -VEWIHKLSID-----IDQEQEV-DVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLR 128
V + + + E + DV+DDLARELA Y Q+L+ ++A + L++ G+PF R
Sbjct: 203 TVPFATHMCVTPAPGTAPTEASIPDVSDDLARELALYKQSLDAAKRARQLLRAEGVPFTR 262
Query: 129 PADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQK 188
P DY+AEMVK D HMEKVK++L+ + A E RK R+ K+ K+VQ KL+ERAK K
Sbjct: 263 PGDYFAEMVKPDEHMEKVKAKLVEDATAKKAAAEARKLRDLKKFGKQVQVAKLQERAKAK 322
Query: 189 KDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAF 248
KD +E +K +++RQ+ G + G +A F+ G ++ K+P SGGK AF
Sbjct: 323 KDTLEKIKDLKRKRQEGGSSALGEREADIFDVG---VENELKKP-------SGGKRNSAF 372
Query: 249 G---KGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFS 288
G + + +K KN K+GFGGKK K A ++ D GFS
Sbjct: 373 GDRNRDEPNRKRQKKNDKYGFGGKKKYSKSGDAMSSGDLSGFS 415
>gb|EAK93752.1| likely EBNA-like pre-rRNA processing factor EBP2p [Candida albicans
SC5314] gi|46434305|gb|EAK93718.1| likely EBNA-like
pre-rRNA processing factor EBP2p [Candida albicans
SC5314]
Length = 426
Score = 159 bits (401), Expect = 1e-37
Identities = 101/284 (35%), Positives = 157/284 (54%), Gaps = 17/284 (5%)
Query: 20 NDDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPEN---- 75
+D+ D EAE +++ S+ E + D + +K +NN AL + L I P +
Sbjct: 146 DDEVENDDEAE---DDIPLSDVEVDSDADIVPHTKLTINNMAALRESLARIELPWSKHSF 202
Query: 76 VEWIHKLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAE 135
+E S D + + D+ DD RELAFY Q L+ +Q+ + L + +PF RP DY+AE
Sbjct: 203 IEHQSITSADKTESEIKDIYDDTERELAFYKQGLDAVKQSRKTLLKLKIPFSRPMDYFAE 262
Query: 136 MVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESV 195
MVK+D HM+K+K++LL E +EE ++ R+ K+ K+VQ L+ERAKQKK+ +E +
Sbjct: 263 MVKSDEHMDKLKNKLLTEAANKKASEEAKRQRQLKKFGKQVQHATLQERAKQKKETLEKI 322
Query: 196 KKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPK 255
K +K+R + ++D D + E+ E ++K G G SGG K + K
Sbjct: 323 KSLKKKRGANEISNDD-DFQIALEEAT--ENNQKYGHG---GSGSGGDNK----RRKPNS 372
Query: 256 KGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKKR 299
K K++K+GFGGKK K++N A ++ D GFS + G S R
Sbjct: 373 KRLAKDAKYGFGGKKRGKRENDAASSADISGFSTRKMKGKSTSR 416
>ref|XP_638424.1| hypothetical protein DDB0186212 [Dictyostelium discoideum]
gi|60467019|gb|EAL65061.1| hypothetical protein
DDB0186212 [Dictyostelium discoideum]
Length = 546
Score = 158 bits (400), Expect = 2e-37
Identities = 95/280 (33%), Positives = 147/280 (51%), Gaps = 29/280 (10%)
Query: 20 NDDTMIDGEAEYLSEEMSPSESESEEDVKL----AEPSKTAVNNRDALLDKLGDISWPEN 75
+D+ +ID + E L E + E ++ ++ + K VN+ + + KL D
Sbjct: 227 SDEELIDEDGELLDSENEMDKEEKKQRLEQQHLKSNRQKYVVNDIEGMKSKLRDFKLSSG 286
Query: 76 -VEWIHKLSIDIDQEQEV-DVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYY 133
V W+H L++ ++ D++DD RE+AFY Q L+ + + GL R D++
Sbjct: 287 KVPWVHTLAVTSQTPLDIEDIHDDFHREIAFYKQTLQSVTECEKLCAQNGLTVRRKPDFF 346
Query: 134 AEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIE 193
AEM+K+D M K+K+ + +EK+++ +E RK RE K+ K+VQ+QKL+ER KQK D IE
Sbjct: 347 AEMIKSDQQMHKIKTNIQSEKKRVETSEMIRKKREIKKFGKQVQTQKLQERQKQKSDAIE 406
Query: 194 SVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKK 253
SVKKWRK R++ +D+ +D D + K ++ K K
Sbjct: 407 SVKKWRKNREKGNVSDEFNIDLID--------------------DIAAKKDRKKVEKSKL 446
Query: 254 PKKGD---VKNSKFGFGGKKGLKKQNTADTTNDFGGFSKK 290
P KGD K++K+GFGGKK K N + ND GFS K
Sbjct: 447 PVKGDKRKKKDAKYGFGGKKRYAKTNDKGSVNDMSGFSVK 486
>gb|EAK86264.1| hypothetical protein UM04809.1 [Ustilago maydis 521]
gi|49077026|ref|XP_402424.1| hypothetical protein
UM04809.1 [Ustilago maydis 521]
Length = 608
Score = 156 bits (394), Expect = 8e-37
Identities = 99/281 (35%), Positives = 145/281 (51%), Gaps = 13/281 (4%)
Query: 26 DGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDI-----SWPENVEWIH 80
D + E ++ E P + E E K + + +NN AL DI S + WI
Sbjct: 304 DEDEEPIAYEDLPDDVELSEQAKASRVQRIKINNESALRRIYEDIRLDSPSSSSKLAWIE 363
Query: 81 KLSIDID---QEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMV 137
+ I Q D +DL RELAFY QAL + + +Q+ G PF RPADY+AEMV
Sbjct: 364 TMRITYPISVSSQVTDAENDLERELAFYKQALHAAVEGKKLIQASGTPFTRPADYFAEMV 423
Query: 138 KTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKK 197
KTD HME+++ +LL E + +E +K RE K+ K+VQ +KL+ER + KK+ ++
Sbjct: 424 KTDEHMERIRQKLLDESAAIKASEAAKKQRELKKFGKKVQVEKLQERIRNKKELNARIQG 483
Query: 198 WRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKG 257
+++ D GAD+ D + +R P G+A A GK K P+
Sbjct: 484 LKRKHGDIEGEDGGADE-FDVRLQEAIGGGDAERSAKKPALGKAGRA--ASGKSKMPR-- 538
Query: 258 DVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKK 298
+++K+GFGGKK K NTA++TNDF ++GA G K
Sbjct: 539 SARDAKYGFGGKKKYSKSNTAESTNDFTVGPERGARGSKVK 579
>emb|CAD70841.1| related to rRNA processing protein EBP2 [Neurospora crassa]
gi|32412540|ref|XP_326750.1| hypothetical protein
[Neurospora crassa] gi|28922297|gb|EAA31538.1|
hypothetical protein [Neurospora crassa]
Length = 436
Score = 154 bits (389), Expect = 3e-36
Identities = 99/270 (36%), Positives = 145/270 (53%), Gaps = 14/270 (5%)
Query: 40 ESESEEDVKLAEPSKTAVNNRDALLDKLGD--ISWPENVEWIHKLSI---DIDQEQEVDV 94
+ E E+VK ++ +NN+ LL L + + V + SI + +E +
Sbjct: 153 DGEEGEEVKAISRARQTINNKQGLLTALKRFALDTTDKVPFAFHQSIVAKKVTEESIPSI 212
Query: 95 NDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRLLAEK 154
+DDLARELAF QALE R L+ G+PF RP DY+AE +++D MEKVK++L+ E
Sbjct: 213 DDDLARELAFMNQALEAARVGRALLRKEGVPFTRPTDYFAETLRSDETMEKVKAKLIEEA 272
Query: 155 QKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADK 214
+ E RK R+ K+ K+VQ K +ERAKQK+D +E + +K+R+ G GA++
Sbjct: 273 TAKKASAEARKQRDLKKFGKQVQVAKQQERAKQKRDTLEKINLLKKKRKDGG-GPTGANE 331
Query: 215 ALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKK 274
A D D V + K PG + + G K+ G K ++ K++KFGFGGKK K
Sbjct: 332 AEDLFDVAV-DNELKGGPGGNNSKKRGRDGKEGGGPNAKRQR---KDAKFGFGGKKKYAK 387
Query: 275 QNTADTTNDFGGFS----KKGAVGGSKKRK 300
N A ++ D GFS K G G K +K
Sbjct: 388 SNDALSSGDVSGFSARKMKSGGPAGGKPKK 417
>gb|EAA65252.1| hypothetical protein AN0074.2 [Aspergillus nidulans FGSC A4]
gi|67515585|ref|XP_657678.1| hypothetical protein
AN0074_2 [Aspergillus nidulans FGSC A4]
gi|49083944|ref|XP_404211.1| hypothetical protein
AN0074.2 [Aspergillus nidulans FGSC A4]
Length = 378
Score = 149 bits (375), Expect = 1e-34
Identities = 108/297 (36%), Positives = 150/297 (50%), Gaps = 38/297 (12%)
Query: 21 DDTMIDGEAEYLSEEMSPSESESEEDVKLAEPS-----------KTAVNNRDALLDKLGD 69
DD D EAE EE E EED+ L++ S + +NN A+L
Sbjct: 86 DDAEDDEEAEDEDEE-----EEEEEDIPLSDLSDDEREDVIPHQRLTINNSTAILASTKR 140
Query: 70 ISWPENVE-WIHKLSIDIDQEQEVDV---NDDLARELAFYTQALEGTRQAFEKLQSIGLP 125
IS+ ++ + S+ E E+D+ NDDL REL+FY A A + L+ G+P
Sbjct: 141 ISFISHLTPFSEHNSLISKAEDEMDIPDANDDLNRELSFYKTAQTAAYTARKLLKKEGVP 200
Query: 126 FLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERA 185
F RP DY+AEMVK+D HM K+K +L E A E RK R+ K+ K+VQ KL++RA
Sbjct: 201 FTRPGDYFAEMVKSDEHMGKIKKKLYDEAASKKAAAEARKQRDLKKFGKQVQVAKLQQRA 260
Query: 186 KQKKDDIESVKKWRKQRQ-QSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKA 244
K+KK+ IE + +K+R+ + D GAD D + +KR + GD SG
Sbjct: 261 KEKKEAIERINDLKKKRKNDTSGQDGGADDLFDVAVEDAVSENPRKR---ARGDTSGPNL 317
Query: 245 KQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFS---KKGAVGGSKK 298
K+ KN K+GFGGKK K A ++ D GFS KGA GG+K+
Sbjct: 318 KR-----------QKKNEKYGFGGKKRHAKSGDAISSGDLRGFSVKKMKGASGGAKR 363
>emb|CAG89061.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50424269|ref|XP_460721.1| unnamed protein product
[Debaryomyces hansenii]
Length = 410
Score = 148 bits (374), Expect = 2e-34
Identities = 96/290 (33%), Positives = 152/290 (52%), Gaps = 29/290 (10%)
Query: 21 DDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLG--DISWPENVEW 78
D+ D E E E++ S+ + ++D + +K +NN AL D L ++ W ++
Sbjct: 140 DEDEKDDEEEEEEEDIPLSDVDIDDDADIVPHTKLTINNMAALKDSLARIELPWSKHSFQ 199
Query: 79 IHKLSIDIDQ-EQEV-DVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEM 136
H+ ++ E E+ D+ DD RELAFY Q L+ +Q L + +PF RP DY+AEM
Sbjct: 200 EHQSITSAEKTETEIKDIYDDTERELAFYKQGLDAVKQGRAALTKLKVPFSRPMDYFAEM 259
Query: 137 VKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVK 196
+K+D HM+K+KS+LL E +E+ +K R K+ K+VQ+ L+ERAKQKK+ ++ +K
Sbjct: 260 IKSDEHMDKLKSKLLTEAANKKASEDAKKQRHLKKFGKQVQNSTLQERAKQKKETLDKIK 319
Query: 197 KWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKK 256
K++ S +G D + ++ + DR G K +KP
Sbjct: 320 S-LKRKGASNELSNGDDFQIALDE-------------ATSEDRGGD------NKRRKPNS 359
Query: 257 GDV-KNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVG----GSKKRKR 301
+ K+SK+GFGGKK + N A+++ D GFS K G G KR +
Sbjct: 360 KRLGKDSKYGFGGKKRGSRTNDAESSADISGFSSKKMKGKPRPGKSKRSK 409
>emb|CAG79182.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50552382|ref|XP_503601.1| hypothetical protein
[Yarrowia lipolytica]
Length = 388
Score = 145 bits (365), Expect = 2e-33
Identities = 95/296 (32%), Positives = 149/296 (50%), Gaps = 33/296 (11%)
Query: 15 ELPLVNDDTMID-----GEAEYLSEEMSP---SESESEEDVKLAEPSKTAVNNRDALLDK 66
E+P + D T +D E E + E+ S+ E + D K NN+ AL+
Sbjct: 106 EVPQLYDTTRLDLSESESEIENVENEVEDIPLSDVELDSDNDAIPYQKVTKNNKPALMQA 165
Query: 67 LGDISWP-ENVEWIHKLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLP 125
IS P + ++ L + D+ DD+ +ELAFY Q L+ + A ++L +P
Sbjct: 166 KARISLPYAKMTFVDTLVHTSGAIELKDIYDDMEKELAFYKQGLDAAKVARKELLKAKVP 225
Query: 126 FLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERA 185
F RP DY+AEMVK+D HMEK+K +L+ E +EE R+ R+ K+ K+VQ KL+ERA
Sbjct: 226 FTRPLDYFAEMVKSDEHMEKLKQKLIEEATSKKASEEARRQRDLKKFGKKVQHNKLQERA 285
Query: 186 KQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAK 245
K KK+ +E +K +++R + D A++ A
Sbjct: 286 KDKKETLEKIKSLKRKRAGQEMTSEEFDIAVE-----------------------EAAAD 322
Query: 246 QAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKKRKR 301
+GK K+ K++KFGFGGKK +K NT++++ D GF++K KKR++
Sbjct: 323 NHNIEGKNHKR-QAKDAKFGFGGKKRFRKANTSESSGDLSGFNQKRNNAPVKKRQQ 377
>gb|EAL21223.1| hypothetical protein CNBD2780 [Cryptococcus neoformans var.
neoformans B-3501A] gi|57226759|gb|AAW43219.1| rRNA
processing-related protein, putative [Cryptococcus
neoformans var. neoformans JEC21]
gi|58266740|ref|XP_570526.1| rRNA processing-related
protein, putative [Cryptococcus neoformans var.
neoformans JEC21]
Length = 395
Score = 143 bits (360), Expect = 7e-33
Identities = 94/281 (33%), Positives = 152/281 (53%), Gaps = 22/281 (7%)
Query: 21 DDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPENVEWIH 80
D+T+I+ + + + S+ D K +NN+ AL L D N+ W
Sbjct: 114 DNTIINEKPDDDVVSLDGLGSDVSVDEDAVPMQKVTINNKPALR-ALTDSIRVTNMSWPE 172
Query: 81 KLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTD 140
L +D + +VD +DDL RE FY AL QA + +PF RP DYYAEMVK+D
Sbjct: 173 HLVLDSKETADVDPSDDLQRETVFYKIALGCIPQARKLASKHDIPFTRPGDYYAEMVKSD 232
Query: 141 SHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKK---DDIESVKK 197
HME+V+++L+ E Q + ++E+ +K R+ K+ K++Q +KL++R + KK D ++ +K+
Sbjct: 233 EHMERVRTKLVEEAQGIKKSEDAKKQRQLKKFGKQIQHEKLRQREQDKKSFEDRVQGLKR 292
Query: 198 WRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKG 257
RK+ + G D+G + + ED + +++P + GG+A A GK K P+
Sbjct: 293 KRKEGMELG--DEGEEFDIAVED------AMEEQP------QKGGRA--APGKSKMPR-- 334
Query: 258 DVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKK 298
+++KF GG KQNT ++T DFGG +G G + K
Sbjct: 335 HARDAKFSLGGGGRRSKQNTRESTMDFGGGMSRGKGGKADK 375
>gb|EAL91386.1| rRNA processing protein (Ebp2), putative [Aspergillus fumigatus
Af293]
Length = 384
Score = 142 bits (357), Expect = 2e-32
Identities = 97/280 (34%), Positives = 138/280 (48%), Gaps = 29/280 (10%)
Query: 34 EEMSPSESESEEDVKLAEPS-----------KTAVNNRDALLDKLGDISWPENVEWI--H 80
EE E E EED+ L++ S + +NN A+ + IS+ + H
Sbjct: 106 EEGEEEEEEEEEDIPLSDLSEDEREDVVPHQRLTINNSAAINTSIKRISFITSQTPFSEH 165
Query: 81 KLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTD 140
+ + + D NDDL RELAFY +A L+ G+PF RP DY+AEMVKTD
Sbjct: 166 NSLVSKEPIEVPDPNDDLNRELAFYKVCQAAASEARRLLKKEGVPFTRPGDYFAEMVKTD 225
Query: 141 SHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRK 200
HM K+K +L E A E RK R+ K+ K+VQ KL++R K+K++ +E + + +K
Sbjct: 226 EHMSKIKKKLYDEAAAKKAAAEARKQRDLKKFGKQVQVAKLQQRQKEKRETLEKINELKK 285
Query: 201 QRQ--QSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGD 258
+R+ +SG DK D D + + K S G+ SGG + K
Sbjct: 286 KRKADKSGI----TDKDNDLFDIAIEDSGKSNSRKRSRGNESGGPS----------LKRQ 331
Query: 259 VKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKK 298
KN K+GFGGKK K A ++ D FS K GGSK+
Sbjct: 332 KKNEKYGFGGKKRHAKSGDAISSGDLRNFSVKKMKGGSKR 371
>emb|CAH02887.1| unnamed protein product [Kluyveromyces lactis NRRL Y-1140]
gi|50302725|ref|XP_451299.1| unnamed protein product
[Kluyveromyces lactis]
Length = 423
Score = 141 bits (355), Expect = 3e-32
Identities = 99/315 (31%), Positives = 158/315 (49%), Gaps = 49/315 (15%)
Query: 11 LDLMELPLVNDDTMIDGEAEYLSEEMSPSESESEE---------DVKLAEPS------KT 55
LDL +L + D+ D E E EE E E EE DV++ E + KT
Sbjct: 132 LDLEKLANSDSDSDSDEEEEEEEEEEEEEEEEKEEEEEEDVPLSDVEIDEDADVVPYQKT 191
Query: 56 AVNNRDALLDKLGDISWP---ENVEWIHKLSIDIDQEQEV-DVNDDLARELAFYTQALEG 111
+NN A+ D L I P + + ++ ++ ++++ D+ DD RELAFY Q+L+
Sbjct: 192 TINNVKAMKDALERIQKPWEKHSFQEHQSVTSQLNTDEKIKDIYDDTERELAFYKQSLDA 251
Query: 112 TRQAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKR 171
A EKL+ + +PF RP DY+AEMVK+D HM+++K +L+ E + +E R+ R+ K+
Sbjct: 252 VTIAREKLKKLKVPFKRPLDYFAEMVKSDEHMDRLKGKLIKEASEKKARQEARRQRDLKK 311
Query: 172 LSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKR 231
K+VQ L++R K K++ ++ +K +K+R+ + D D + ED
Sbjct: 312 FGKQVQVATLQQRQKDKRETLDKIKDLKKKRKHNEIGGDDFD--IGVED----------- 358
Query: 232 PGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKG 291
A Q+ K K K + KNSK+G GG K K++N A ++ + FS++
Sbjct: 359 ------------AAQSEKKHKPNFKREAKNSKYGSGGLKRYKRKNDAQSSAEMPEFSQRK 406
Query: 292 AVG-----GSKKRKR 301
G G KR R
Sbjct: 407 MKGKAPRPGKSKRSR 421
>emb|CAG61758.1| unnamed protein product [Candida glabrata CBS138]
gi|50292711|ref|XP_448788.1| unnamed protein product
[Candida glabrata]
Length = 387
Score = 141 bits (355), Expect = 3e-32
Identities = 89/284 (31%), Positives = 147/284 (51%), Gaps = 29/284 (10%)
Query: 22 DTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDALLDKLGDISWP---ENVEW 78
D+ D + E E++ S+ E + D + K +NN AL L I P + E
Sbjct: 127 DSENDEDKEDEEEDVPLSDVEFDSDADVVPHHKLTINNTKALKHSLERIQLPWAKHSFEE 186
Query: 79 IHKLSIDIDQEQEV-DVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMV 137
++ + + ++ + D+ DD RELAFY Q+L+ A EKL+ + +PF RP DY+AEMV
Sbjct: 187 HQSVTSETNTDEGMKDIYDDTERELAFYKQSLDAVNIAKEKLKKLKVPFKRPLDYFAEMV 246
Query: 138 KTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKK 197
K D HM+K+K +L+ E + E+ RK R+ K+ K+VQ L++R +K++ ++ +K
Sbjct: 247 KNDEHMDKIKHKLVQEASEKKAREDARKQRQLKKFGKQVQVATLQKRQLEKRETLDRIKN 306
Query: 198 WRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKG 257
+K+R+ + D D + E+ E +K+++P K
Sbjct: 307 LKKKRKHNEIGD--MDFNVGIEEATESEDTKRRKPN---------------------SKR 343
Query: 258 DVKNSKFGFGGKKGLKKQNTADTTNDFGGFS--KKGAVGGSKKR 299
KNSK+G GG K K++N A ++ D GFS KK G S+++
Sbjct: 344 QAKNSKYGQGGMKRFKRKNDAQSSADITGFSNRKKSRPGKSRRK 387
>gb|AAS52562.1| AEL123Wp [Ashbya gossypii ATCC 10895] gi|45190484|ref|NP_984738.1|
AEL123Wp [Eremothecium gossypii]
Length = 395
Score = 137 bits (345), Expect = 4e-31
Identities = 101/313 (32%), Positives = 155/313 (49%), Gaps = 47/313 (15%)
Query: 10 SLDLMELPLVNDDTMIDGE-AEYLSEEMSPSESESEEDVKLAEPS-----------KTAV 57
SLDL +L + ++ + E AE S E +E SEEDV L++ K +
Sbjct: 107 SLDLDKLARSDSESESEDEFAEARSVEEHEAEDGSEEDVPLSDVEFDSDADVVPHQKVTI 166
Query: 58 NNRDALLDKLGDISWP---ENVEWIHKLSIDIDQEQEV-DVNDDLARELAFYTQALEGTR 113
NN A+ D L I P + + ++ + + + D+ DD RELAFY Q+L+
Sbjct: 167 NNTKAISDSLARIQLPWQKHSFQEHQTVTAAANADAGITDIYDDTERELAFYKQSLDAVV 226
Query: 114 QAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLS 173
QA + L ++ +PF RP DY+AEMVKTD HM+K+KS+L+ E EE RK R+ K+
Sbjct: 227 QARDALTALRVPFKRPLDYFAEMVKTDEHMDKLKSKLVKEASDKKAREEARKQRQLKKFG 286
Query: 174 KEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPG 233
K+VQ L++R K+K++ ++ +K R +R+ + DD A+ ED + K
Sbjct: 287 KQVQVATLQQRQKEKRETLDKIKSLRTKRKHNDIDDDAF--AVGVEDAAAEKPHK----- 339
Query: 234 VSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAV 293
G +QA KN+K+G GG K K++N A ++ + FS +
Sbjct: 340 -------AGAKRQA------------KNAKYGQGGLKRFKRKNDAQSSAEMTDFSTRKMK 380
Query: 294 G-----GSKKRKR 301
G G KR R
Sbjct: 381 GKAPRPGKSKRSR 393
>emb|CAB11214.1| SPAC17H9.05 [Schizosaccharomyces pombe]
gi|19114487|ref|NP_593575.1| hypothetical protein
SPAC17H9.05 [Schizosaccharomyces pombe 972h-]
gi|7490725|pir||T37871 hypothetical nucleolar protein -
fission yeast (Schizosaccharomyces pombe)
gi|3183355|sp|O13802|EBP2_SCHPO Probable rRNA processing
protein ebp2
Length = 333
Score = 134 bits (336), Expect = 4e-30
Identities = 95/300 (31%), Positives = 147/300 (48%), Gaps = 32/300 (10%)
Query: 11 LDLMELPLVNDDTMIDGEAEYL-SEEMSPSESES---EEDVKLAEPSKTAVNNRDALLDK 66
++ ++ P++ D E E S+E+ S+ E EED L K A+NN AL +
Sbjct: 44 INTIKSPIIETADTADQENESEGSDEVELSDLEGIELEEDADLIRKRKLAINNTVALENI 103
Query: 67 LGDISWPENVEWIHKLSIDIDQEQEVD-VNDDLARELAFYTQALEGTRQAFEKLQSIGLP 125
I +P+++ ++ ++ + ++ V DDLARELAFY Q + + AF KL+ +
Sbjct: 104 YERIKYPDDISFVENQAVTTKEPIIIENVEDDLARELAFYKQGVSSVKAAFAKLREANVL 163
Query: 126 FLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERA 185
RP DY+AEM+K+D HMEKV+ L+ E +++ +K RE K+ K+VQ K +ER
Sbjct: 164 ISRPHDYFAEMLKSDDHMEKVRQELIKEATAKKLSQQAKKQRELKKFGKQVQLAKQEERQ 223
Query: 186 KQKKDDIESVKKWRKQRQQSGF-ADDGADKALDFEDGKVFE---RSKKKRPGVSPGDRSG 241
++KK+ +E + +++ +D D AL F+ RS K RP +P
Sbjct: 224 REKKETLEKINLLKRKHTGGDLTTEDDFDIALSSASADTFKKGSRSTKSRPQPNP----- 278
Query: 242 GKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADT--TNDFGGFSKKGAVGGSKKR 299
K KN K+GFGG K K N D+ +FG K SKKR
Sbjct: 279 --------------KRQKKNEKYGFGGPKHRSKSNDLDSLAATEFGRKGLKNI--KSKKR 322
>emb|CAE59177.1| Hypothetical protein CBG02485 [Caenorhabditis briggsae]
Length = 323
Score = 132 bits (333), Expect = 9e-30
Identities = 84/252 (33%), Positives = 136/252 (53%), Gaps = 25/252 (9%)
Query: 57 VNNRDALLDKLGDISWPENVEWIHKLSIDIDQEQEVDV---NDDLARELAFYTQALEGTR 113
+N + +KL +I+ +++ W+ L + + E+D+ NDD REL FY QA + +
Sbjct: 76 INKSAEMKEKLAEIT--KDLPWVETLEV-VTPHAEMDMKVENDDFQRELNFYKQAEKAVQ 132
Query: 114 QAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLS 173
A+ +L ++G+ LRP+DYYAEM K+DSHM+KV+ RLL ++ E R+ RE K+ +
Sbjct: 133 TAYPRLLNLGIKVLRPSDYYAEMAKSDSHMQKVRKRLLGIQEMKERQEAFRRIREEKKFA 192
Query: 174 KEVQSQKLKERAKQKKDDIESVKKWRK--QRQQSGFADDGADKALDFEDGKVFERSKKKR 231
+VQ + + + +KK E+VKK +K ++Q ++ LD +D
Sbjct: 193 VKVQKEAIANKNNEKKKLAEAVKKHKKGMKQQLEDMLNNVKRHGLDQDD---------DG 243
Query: 232 PGVSPGDRSGGKAKQAFG---KGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDF---- 284
P + GDR GG+ G + K +K+ KFG+GGKK K+N ++ ND
Sbjct: 244 PSSAYGDRRGGRGGSGRGGSMRNAGELKRKLKSDKFGYGGKKKGMKRNNKESFNDLFGAP 303
Query: 285 -GGFSKKGAVGG 295
GGF ++G GG
Sbjct: 304 RGGFGRRGRGGG 315
>gb|AAH87738.1| EBNA1 binding protein 2 (predicted) [Rattus norvegicus]
gi|56790289|ref|NP_001008721.1| EBNA1 binding protein 2
(predicted) [Rattus norvegicus]
Length = 307
Score = 129 bits (323), Expect = 1e-28
Identities = 90/307 (29%), Positives = 154/307 (49%), Gaps = 44/307 (14%)
Query: 14 MELPLVNDDTMIDGEAEYLSEEMSPSESESEEDVK-----LAEPSKTAVNNRDALLDKLG 68
M+ P ++D G E L+ + ++ S +K + E K AVN+ + L L
Sbjct: 1 MDTPPLSDSD--SGSDECLASDQELQDAFSRGLLKPGLNVVLEKPKKAVNDENGLKQCLA 58
Query: 69 DISWPENVEWIHKLSIDI----------------DQEQEVDVNDDLARELAFYTQALEGT 112
+ ++EW+ +L + + DQ++ V+ DD RE++FY QA
Sbjct: 59 EFK--RDLEWVERLDVTLGPVPEASETQSTPQNKDQKKGVNPEDDFQREMSFYRQAQAAV 116
Query: 113 RQAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRL 172
+L + +P RP DY+AEM K+D M+K++ +L ++ M ++E+ ++ R ++
Sbjct: 117 LAVLPRLHQLQVPTKRPTDYFAEMAKSDQQMQKIRQKLQTKQAAMEKSEKAKQLRALRKY 176
Query: 173 SKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRP 232
K+VQ++ L++R ++K + ++KK++K GF+D LDF +G ++P
Sbjct: 177 GKKVQTEVLQKRQQEKAHMMNAIKKYQK-----GFSD-----KLDFLEG-------DQKP 219
Query: 233 GVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGA 292
V G ++GG Q KG K+ KN KFGFGGKK K NT ++ +D F K A
Sbjct: 220 -VERGAKAGGAKGQQISKGPNAKR-RYKNQKFGFGGKKKGSKWNTRESYDDVSSFRGKVA 277
Query: 293 VGGSKKR 299
G +R
Sbjct: 278 HGKGPRR 284
>emb|CAG11568.1| unnamed protein product [Tetraodon nigroviridis]
Length = 285
Score = 129 bits (323), Expect = 1e-28
Identities = 86/262 (32%), Positives = 132/262 (49%), Gaps = 36/262 (13%)
Query: 51 EPSKTAVNNRDA----LLDKLGDISWPENVEWIHKLSIDI----------DQEQEVDVND 96
+ SK VNN +A L D D+ W E ++ + + D+ +++V V+D
Sbjct: 20 DKSKKYVNNVEAMKQCLADFRRDLPWVERLDMTNLPAEDVLSKVEEKFLEKTDEDVTVDD 79
Query: 97 DLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRLLAEKQK 156
D RE+ FY QA QA L G+ RP DY+AEM K+D HM+K++ +L++++
Sbjct: 80 DFQREMLFYRQAQATVLQALPLLSKHGIATKRPEDYFAEMAKSDLHMQKIRKKLISKQMI 139
Query: 157 MVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKAL 216
M +E+ +K RE ++ K+VQ + +++R K+KK + +VKK++K G L
Sbjct: 140 MERSEKAKKLREQRKFGKKVQIEVIQKRQKEKKAMMSAVKKYQK----------GMTDKL 189
Query: 217 DFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQN 276
DF +G K+ + G + K + GK K K+ KFGFGGKK K N
Sbjct: 190 DFLEG------DKREKNTTQGSKKVLNKKGSSGKRK------YKDQKFGFGGKKSGMKWN 237
Query: 277 TADTTNDFGGFSKKGAVGGSKK 298
T D+ ND F K A G K
Sbjct: 238 TKDSYNDVSSFRAKVAHGRGNK 259
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.310 0.130 0.355
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 512,765,564
Number of Sequences: 2540612
Number of extensions: 22543452
Number of successful extensions: 142024
Number of sequences better than 10.0: 3501
Number of HSP's better than 10.0 without gapping: 672
Number of HSP's successfully gapped in prelim test: 3068
Number of HSP's that attempted gapping in prelim test: 122409
Number of HSP's gapped (non-prelim): 12955
length of query: 301
length of database: 863,360,394
effective HSP length: 127
effective length of query: 174
effective length of database: 540,702,670
effective search space: 94082264580
effective search space used: 94082264580
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 75 (33.5 bits)
Medicago: description of AC121233.5


