

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121233.1 - phase: 0 /pseudo
(1172 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
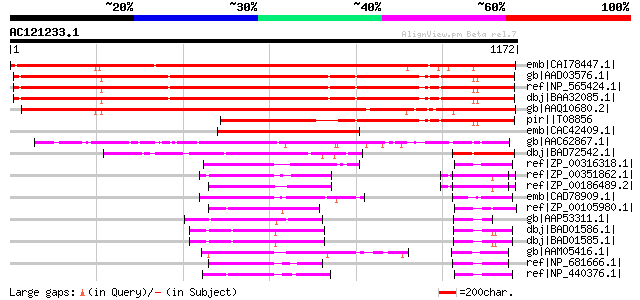
Score E
Sequences producing significant alignments: (bits) Value
emb|CAI78447.1| osmosensor histidine-aspartate kinase [Populus x... 1528 0.0
gb|AAD03576.1| putative histidine kinase [Arabidopsis thaliana] ... 1432 0.0
ref|NP_565424.1| histidine kinase 1 [Arabidopsis thaliana] 1432 0.0
dbj|BAA32085.1| histidine kinase 1 [Arabidopsis thaliana] gi|254... 1430 0.0
gb|AAQ10680.2| cold inducible histidine kinase 1 [Catharanthus r... 1411 0.0
pir||T08856 hypothetical protein A_TM017A05.5 - Arabidopsis thal... 722 0.0
emb|CAC42409.1| putative osmosensor histidine Kinase [Populus eu... 479 e-133
gb|AAC62867.1| putative histidine kinase [Arabidopsis thaliana] ... 280 2e-73
dbj|BAD72542.1| putative histidine kinase 1 [Oryza sativa (japon... 193 3e-47
ref|ZP_00316318.1| COG0642: Signal transduction histidine kinase... 182 8e-44
ref|ZP_00351862.1| COG0642: Signal transduction histidine kinase... 180 2e-43
ref|ZP_00186489.2| COG0642: Signal transduction histidine kinase... 180 2e-43
emb|CAD78909.1| sensory transduction histidine kinase-putative s... 177 1e-42
ref|ZP_00105980.1| COG2202: FOG: PAS/PAC domain [Nostoc punctifo... 177 2e-42
gb|AAP53311.1| putative histidine kinase [Oryza sativa (japonica... 167 2e-39
dbj|BAD01586.1| histidine kinase 3 [Zea mays] 165 1e-38
dbj|BAD01585.1| histidine kinase 3 [Zea mays] 165 1e-38
gb|AAM05416.1| sensory transduction histidine kinase [Methanosar... 160 2e-37
ref|NP_681666.1| two-component hybrid sensor and regulator [Ther... 160 2e-37
ref|NP_440376.1| hybrid sensory kinase [Synechocystis sp. PCC 68... 156 3e-36
>emb|CAI78447.1| osmosensor histidine-aspartate kinase [Populus x canadensis]
Length = 1249
Score = 1528 bits (3956), Expect = 0.0
Identities = 819/1236 (66%), Positives = 952/1236 (76%), Gaps = 77/1236 (6%)
Query: 1 MATKCMHVFDRLLLWFATCWKKNSTLTGRRKFHKDVEKEDFQYSTTQCLSSYYSVFVVRL 60
MAT V +R+L FAT +KN+ GRR FH++VE+ +FQY T CLSSYYSVFVVRL
Sbjct: 23 MATPLRKVCNRIL-GFATSCRKNTAPYGRRIFHREVEQGEFQYGNTHCLSSYYSVFVVRL 81
Query: 61 AIMVMLAILIGLLTILTWHFTKIYTTKSLNSLAYDLRYELLQRPILRMWNILNSTAEITT 120
AIM MLAILIGLLTILTWHFT+ YT KSL++LA LRYELLQRPILRMWNILNSTAEIT
Sbjct: 82 AIMAMLAILIGLLTILTWHFTRSYTKKSLDTLASGLRYELLQRPILRMWNILNSTAEITA 141
Query: 121 AQVKLSEYVIRSHGNLATQAEQVEMYESMRAVTWALFASRKALNSITVKYRNGFVQAFHR 180
AQVKLSEYVI + QAEQVE+YE MR VTWALF+SRKALN+IT+ YRNGFVQAFHR
Sbjct: 142 AQVKLSEYVIGRYSKTTIQAEQVELYEVMRHVTWALFSSRKALNAITINYRNGFVQAFHR 201
Query: 181 DLKDNNIFYIYTDLSY--------NETNSFAAH-----EDTNSNKSAIWYREQLDPVNGE 227
D + NN FYIY+DL ++ N F +H + +SN SAIWYRE LDP +GE
Sbjct: 202 DHRSNNTFYIYSDLRNYSINTKGPSDANMFLSHPAWNDQSIHSNFSAIWYREPLDPTSGE 261
Query: 228 KIGKAMKIAPEDSISIAGLSQVPDGVASWHVSVGKFTDSPLLSAALPVWDSSNKSIVAVV 287
KIGKA I P+D I+IAGLSQVPDGVASWHV+V K+TDSPLLSAALPVWD+ NKSIVAVV
Sbjct: 262 KIGKASPIPPDDLINIAGLSQVPDGVASWHVAVSKYTDSPLLSAALPVWDAYNKSIVAVV 321
Query: 288 GVTTALYSVGQLMKELVDKHSGHMYLTSQEGYLLATSTNDPLLTNSTKKPKLKMAVDCDN 347
GVTTALYSVGQLM+ELV+ H G++YLTSQEGYLLATSTN PLLTNST+ P L MAVD +
Sbjct: 322 GVTTALYSVGQLMRELVEVHKGYIYLTSQEGYLLATSTNAPLLTNSTR-PNLIMAVDTEE 380
Query: 348 EVIREGAMWLKKTYENNFPPSHEVHEENARLGHQQYYIDSFFLILKKLPLVGVIIIPRKH 407
IR GA WL++ Y N FPP H VH ENA+LG QQ YIDSFFL LKKLP+VGVIIIPR++
Sbjct: 381 PTIRMGARWLERVYGNKFPPGHVVHVENAKLGKQQCYIDSFFLNLKKLPIVGVIIIPRRY 440
Query: 408 IMGQADERAFKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEA 467
IMG+ DERAFKTLVILISASLCI+VIGCV ILILTNGVSKEM LRAELISHL+ARR+AEA
Sbjct: 441 IMGKVDERAFKTLVILISASLCILVIGCVFILILTNGVSKEMKLRAELISHLDARRRAEA 500
Query: 468 SSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNN 527
S+NYKSQFLANMSHELRTPMAAVIGLLDIL+ DD LTNEQ A VTQIRKCSTALL LLNN
Sbjct: 501 SNNYKSQFLANMSHELRTPMAAVIGLLDILICDDCLTNEQYANVTQIRKCSTALLRLLNN 560
Query: 528 ILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSA 587
ILD+SKVESGKLVLE+AEFD GRELEGL+DMFSVQCINHNVE +LDLSD+MPKLVRGDSA
Sbjct: 561 ILDLSKVESGKLVLEDAEFDLGRELEGLIDMFSVQCINHNVEAVLDLSDEMPKLVRGDSA 620
Query: 588 RVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQHEN 647
RVVQIFANLI+NSIKFT +GHIILRGWCEN N+ + F L+QK C+ K K++Q N
Sbjct: 621 RVVQIFANLISNSIKFTTTGHIILRGWCENLNNTYNDTQFHLDQKKMRCAIKPKLRQQGN 680
Query: 648 HSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRS 707
H KKA +NKMILWFE+DDTGCGIDPSKW+SVFESFEQAD STTRLHGGTGLGLCIVR+
Sbjct: 681 HLKKACKKENKMILWFEIDDTGCGIDPSKWESVFESFEQADPSTTRLHGGTGLGLCIVRT 740
Query: 708 LVNKMGGEIKIVKKEGPGTLMRLYMQLTAPVDATEQHCQVDFANNNLMVLLALHGNMSRL 767
LVNKMGGEIK+VKK GPGTLMRLY+ L P D + HCQVDF+++N +VL+AL+G+M R+
Sbjct: 741 LVNKMGGEIKVVKKNGPGTLMRLYLLLKTPADGADLHCQVDFSSHNAVVLVALNGSMGRV 800
Query: 768 ITSKWLQKNGVLTMEASEWNGLTQILKELFEAKTSTHNNDFDTHFPAPEGLNSKFISIQE 827
I S+WL++ G+ T+ SEWN LT++L++ F A+ N FD E L S+ ++I++
Sbjct: 801 IMSQWLREIGLTTLGVSEWNELTRVLRKFFHAR--RRENGFDVQCSLNEPLKSEVLNIED 858
Query: 828 LPNPTFVIVVDIDLLDLSTDIWKEQLNFLHKYYARAKFVWLQNHDSSNTVKTELRKKGHI 887
+ + F+IVVD+ LLDLSTDIWKEQ+NFL + +AKF W+ NHD+SN +K ELRKKGH+
Sbjct: 859 MKD-LFIIVVDVGLLDLSTDIWKEQINFLDNFSGKAKFAWMLNHDTSNAIKMELRKKGHL 917
Query: 888 LTVNKPLYKAKMIHILEDVIKERNVEVQKKN-------MKEDDLHESLEIEYTHHCDVAS 940
L VNKPLYKAKMIHILE VIKE+++E QKK+ K+ D+HE LEI+ TH D S
Sbjct: 918 LMVNKPLYKAKMIHILETVIKEKDLEYQKKSSNAARAMAKDGDMHECLEIDSTHF-DTTS 976
Query: 941 SDGSDISEIGSSNLVTA----NGEKQREEVVRINPSSLYQKSNCLLGLSNGYME------ 990
S+ SD +E+G SN + + K+REE+ S Q CL+ L++ E
Sbjct: 977 SEESDTAEMGDSNSPSTFHLRDVRKEREEIA---CQSQCQTFKCLIELADADAEAREDPG 1033
Query: 991 ------------------HKEAFISTRAKGEDSK----GGETSR-----VSGSSKAMNGK 1023
+K+ ST + E SK ETS S S+KA N +
Sbjct: 1034 QNRPNLQGTQYGNDMLLCNKQVPFSTATRNESSKHDERNSETSSHKEQGNSYSNKAGNQQ 1093
Query: 1024 KSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALN----------GMPG 1073
K+L+GLRILLAEDT V+QRVATIMLEKMGA V+ VGDG QAV+ALN PG
Sbjct: 1094 KALDGLRILLAEDTPVLQRVATIMLEKMGAKVITVGDGLQAVEALNCSLSEKDCRRESPG 1153
Query: 1074 VERNTITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAM 1133
+ NT QT+I SYDLILMDCQMPKMDGYEATK IRKSE GT HIPIVALTAHAM
Sbjct: 1154 NDGNT-GLQTDIQESQSYDLILMDCQMPKMDGYEATKAIRKSETGTDLHIPIVALTAHAM 1212
Query: 1134 SCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTKRE 1169
S DEAKCLEVGMDAYLTKPID+KLM STILSLT+R+
Sbjct: 1213 SSDEAKCLEVGMDAYLTKPIDYKLMVSTILSLTRRK 1248
>gb|AAD03576.1| putative histidine kinase [Arabidopsis thaliana]
gi|7486989|pir||T00842 probable histidine kinase
[imported] - Arabidopsis thaliana
Length = 1190
Score = 1432 bits (3706), Expect = 0.0
Identities = 760/1192 (63%), Positives = 917/1192 (76%), Gaps = 56/1192 (4%)
Query: 8 VFDRLLLWFATCWKKNSTLTGRRKF---HKDVEKEDFQYSTTQCLSSYYSVFVVRLAIMV 64
VFD++ W T W+ N L R+ DVE+++FQY+++ CLSSYYSVFVVRLAIMV
Sbjct: 13 VFDKMTEW-VTPWRSN--LESPREMMILRGDVEQDEFQYASSHCLSSYYSVFVVRLAIMV 69
Query: 65 MLAILIGLLTILTWHFTKIYTTKSLNSLAYDLRYELLQRPILRMWNILNSTAEITTAQVK 124
MLAILIGLLT+LTWHFT+IYT +SL +LAY LRYELLQRP+LRMW++LN+T+E+TTAQVK
Sbjct: 70 MLAILIGLLTVLTWHFTRIYTKQSLQTLAYGLRYELLQRPVLRMWSVLNTTSELTTAQVK 129
Query: 125 LSEYVIRSHGNLATQAEQVEMYESMRAVTWALFASRKALNSITVKYRNGFVQAFHRDLKD 184
LSEYVI+ + TQ E VEMY++M+ VTWALFAS KALN+IT+ YRNGFVQAFHRD
Sbjct: 130 LSEYVIKKYDKPTTQEELVEMYQAMKDVTWALFASAKALNAITINYRNGFVQAFHRDPAS 189
Query: 185 NNIFYIYTDLSYNETNSFAAHEDTNS--------NKSAIWYREQLDPVNGEKIGKAMKIA 236
++ FYI++DL N + S ED + N SAIWY++QLDPV GE +GK +KI
Sbjct: 190 SSTFYIFSDLK-NYSISGTGPEDVSGWNNKSIHGNMSAIWYQQQLDPVTGENLGKPLKIP 248
Query: 237 PEDSISIAGLSQVPDGVASWHVSVGKFTDSPLLSAALPVWDSSNKSIVAVVGVTTALYSV 296
P+D I+IAG+SQVPDG ASWHV+V K+ DSPLLSAALPV+D+SNKSIVAVVGVTTALYSV
Sbjct: 249 PDDLINIAGISQVPDGEASWHVTVSKYMDSPLLSAALPVFDASNKSIVAVVGVTTALYSV 308
Query: 297 GQLMKELVDKHSGHMYLTSQEGYLLATSTNDPLLTNSTKKPKLKMAVDCDNEVIREGAMW 356
GQLM++LV+ H GH+YLTSQEGYLLATST+ PLL N++ P+L A D + VI+ GA W
Sbjct: 309 GQLMRDLVEVHGGHIYLTSQEGYLLATSTDGPLLKNTSNGPQLMKATDSEEWVIKSGAQW 368
Query: 357 LKKTYENNFPPSHEVHEENARLGHQQYYIDSFFLILKKLPLVGVIIIPRKHIMGQADERA 416
L+KTY + P H VH EN +LG Q+YYIDSF+L LK+LP+VGV+IIPRK IMG+ DERA
Sbjct: 369 LEKTYGSKRP--HVVHAENVKLGDQRYYIDSFYLNLKRLPIVGVVIIPRKFIMGKVDERA 426
Query: 417 FKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFL 476
FKTL+ILISAS+CI IGCVCILILTNGVSKEM LRAELI L+ARR+AEASSNYKSQFL
Sbjct: 427 FKTLIILISASVCIFFIGCVCILILTNGVSKEMKLRAELIRQLDARRRAEASSNYKSQFL 486
Query: 477 ANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVES 536
ANMSHELRTPMAAVIGLLDIL+SDD L+NEQ ATVTQIRKCSTALL LLNNILD+SKVES
Sbjct: 487 ANMSHELRTPMAAVIGLLDILISDDCLSNEQYATVTQIRKCSTALLRLLNNILDLSKVES 546
Query: 537 GKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANL 596
GKLVLEEAEFD GRELEGLVDMFSVQCINHNVE +LDLSDDMP LVRGDSAR+VQIFANL
Sbjct: 547 GKLVLEEAEFDLGRELEGLVDMFSVQCINHNVETVLDLSDDMPALVRGDSARLVQIFANL 606
Query: 597 INNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNID 656
I+NSIKFT +GHIILRGWCEN NS D + +++++ KTK QH NH +K+
Sbjct: 607 ISNSIKFTTTGHIILRGWCENINSLHDEMSVSVDRRKPWAPMKTKQVQHRNHLQKSCKNA 666
Query: 657 NKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEI 716
NKM+LWFEVDDTGCGIDPSKWDSVFESFEQAD STTR HGGTGLGLCIVR+LVNKMGGEI
Sbjct: 667 NKMVLWFEVDDTGCGIDPSKWDSVFESFEQADPSTTRTHGGTGLGLCIVRNLVNKMGGEI 726
Query: 717 KIVKKEGPGTLMRLYMQLTAPVDATEQHCQVDFANNNLMVLLALHGNMSRLITSKWLQKN 776
K+V+K G GTLMRLY+ L+ P D +Q+ Q DF+ L+V+L+++G+ +R+ITSKWL+K+
Sbjct: 727 KVVQKNGLGTLMRLYLILSTP-DTVDQNIQPDFSKYGLVVMLSMYGSTARMITSKWLRKH 785
Query: 777 GVLTMEASEWNGLTQILKELFEAKTSTHNNDFDTHFPAPEGLNSKFISIQELPNPTFVIV 836
G+ T+EAS+WN LTQI+++L E T + +N FD+ + L ++ +I E+ NP FVIV
Sbjct: 786 GIATVEASDWNELTQIIRDLLE--TGSRDNSFDSQHNISDPLRAELSNIVEIKNPVFVIV 843
Query: 837 VDIDLLDLSTDIWKEQLNFLHKYYARAKFVWLQNHDSSNTVKTELRKKGHILTVNKPLYK 896
VDI +LDL+T+IWKEQLN+L ++ +AKF WL HD+SNTVKTELR+KGH++ VNKPLYK
Sbjct: 844 VDIGVLDLTTNIWKEQLNYLDRFSNKAKFAWLLKHDTSNTVKTELRRKGHVMMVNKPLYK 903
Query: 897 AKMIHILEDVIKERN----VEVQKKNMKEDDLHESLEIEYTHHCDVASSDGSDISEIGSS 952
AKMI ILE VIK R +++ + D+ H+ LEI+ T +S D S+ S
Sbjct: 904 AKMIQILEAVIKNRKRGLCNDLRNRGNGSDESHDCLEIDPTQFDTCSSDDSSETS----- 958
Query: 953 NLVTANGEKQREEVVRINPSSLYQK--SNCLLGLSNGYMEHKEAFISTRAKGEDSKGGET 1010
GEKQ ++ V+ PS+L+ N L+ + + A S K + + +
Sbjct: 959 ------GEKQVDKSVK--PSTLHSPVLKNYLIDATTSNDDSTSA--SMTQKNPEEEDWKD 1008
Query: 1011 SRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALN- 1069
SG + +KSLEG+RILLAEDT V+QRVATIMLEKMGATV AV DGQQAVD+LN
Sbjct: 1009 RLYSGIALDGKNQKSLEGIRILLAEDTPVLQRVATIMLEKMGATVTAVWDGQQAVDSLNY 1068
Query: 1070 -----GMPGVERNTI---------TSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKS 1115
P E + T +T + + YDLILMDCQMPKMDGYEATK IR++
Sbjct: 1069 KSINAQAPTEEHKSFEEETANKVTTRETSLRNSSPYDLILMDCQMPKMDGYEATKAIRRA 1128
Query: 1116 EIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTK 1167
EIGT HIPIVALTAHAMS DEAKCLEVGMDAYLTKPID KLM STILSLTK
Sbjct: 1129 EIGTELHIPIVALTAHAMSSDEAKCLEVGMDAYLTKPIDRKLMVSTILSLTK 1180
>ref|NP_565424.1| histidine kinase 1 [Arabidopsis thaliana]
Length = 1207
Score = 1432 bits (3706), Expect = 0.0
Identities = 760/1192 (63%), Positives = 917/1192 (76%), Gaps = 56/1192 (4%)
Query: 8 VFDRLLLWFATCWKKNSTLTGRRKF---HKDVEKEDFQYSTTQCLSSYYSVFVVRLAIMV 64
VFD++ W T W+ N L R+ DVE+++FQY+++ CLSSYYSVFVVRLAIMV
Sbjct: 30 VFDKMTEW-VTPWRSN--LESPREMMILRGDVEQDEFQYASSHCLSSYYSVFVVRLAIMV 86
Query: 65 MLAILIGLLTILTWHFTKIYTTKSLNSLAYDLRYELLQRPILRMWNILNSTAEITTAQVK 124
MLAILIGLLT+LTWHFT+IYT +SL +LAY LRYELLQRP+LRMW++LN+T+E+TTAQVK
Sbjct: 87 MLAILIGLLTVLTWHFTRIYTKQSLQTLAYGLRYELLQRPVLRMWSVLNTTSELTTAQVK 146
Query: 125 LSEYVIRSHGNLATQAEQVEMYESMRAVTWALFASRKALNSITVKYRNGFVQAFHRDLKD 184
LSEYVI+ + TQ E VEMY++M+ VTWALFAS KALN+IT+ YRNGFVQAFHRD
Sbjct: 147 LSEYVIKKYDKPTTQEELVEMYQAMKDVTWALFASAKALNAITINYRNGFVQAFHRDPAS 206
Query: 185 NNIFYIYTDLSYNETNSFAAHEDTNS--------NKSAIWYREQLDPVNGEKIGKAMKIA 236
++ FYI++DL N + S ED + N SAIWY++QLDPV GE +GK +KI
Sbjct: 207 SSTFYIFSDLK-NYSISGTGPEDVSGWNNKSIHGNMSAIWYQQQLDPVTGENLGKPLKIP 265
Query: 237 PEDSISIAGLSQVPDGVASWHVSVGKFTDSPLLSAALPVWDSSNKSIVAVVGVTTALYSV 296
P+D I+IAG+SQVPDG ASWHV+V K+ DSPLLSAALPV+D+SNKSIVAVVGVTTALYSV
Sbjct: 266 PDDLINIAGISQVPDGEASWHVTVSKYMDSPLLSAALPVFDASNKSIVAVVGVTTALYSV 325
Query: 297 GQLMKELVDKHSGHMYLTSQEGYLLATSTNDPLLTNSTKKPKLKMAVDCDNEVIREGAMW 356
GQLM++LV+ H GH+YLTSQEGYLLATST+ PLL N++ P+L A D + VI+ GA W
Sbjct: 326 GQLMRDLVEVHGGHIYLTSQEGYLLATSTDGPLLKNTSNGPQLMKATDSEEWVIKSGAQW 385
Query: 357 LKKTYENNFPPSHEVHEENARLGHQQYYIDSFFLILKKLPLVGVIIIPRKHIMGQADERA 416
L+KTY + P H VH EN +LG Q+YYIDSF+L LK+LP+VGV+IIPRK IMG+ DERA
Sbjct: 386 LEKTYGSKRP--HVVHAENVKLGDQRYYIDSFYLNLKRLPIVGVVIIPRKFIMGKVDERA 443
Query: 417 FKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFL 476
FKTL+ILISAS+CI IGCVCILILTNGVSKEM LRAELI L+ARR+AEASSNYKSQFL
Sbjct: 444 FKTLIILISASVCIFFIGCVCILILTNGVSKEMKLRAELIRQLDARRRAEASSNYKSQFL 503
Query: 477 ANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVES 536
ANMSHELRTPMAAVIGLLDIL+SDD L+NEQ ATVTQIRKCSTALL LLNNILD+SKVES
Sbjct: 504 ANMSHELRTPMAAVIGLLDILISDDCLSNEQYATVTQIRKCSTALLRLLNNILDLSKVES 563
Query: 537 GKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANL 596
GKLVLEEAEFD GRELEGLVDMFSVQCINHNVE +LDLSDDMP LVRGDSAR+VQIFANL
Sbjct: 564 GKLVLEEAEFDLGRELEGLVDMFSVQCINHNVETVLDLSDDMPALVRGDSARLVQIFANL 623
Query: 597 INNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNID 656
I+NSIKFT +GHIILRGWCEN NS D + +++++ KTK QH NH +K+
Sbjct: 624 ISNSIKFTTTGHIILRGWCENINSLHDEMSVSVDRRKPWAPMKTKQVQHRNHLQKSCKNA 683
Query: 657 NKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEI 716
NKM+LWFEVDDTGCGIDPSKWDSVFESFEQAD STTR HGGTGLGLCIVR+LVNKMGGEI
Sbjct: 684 NKMVLWFEVDDTGCGIDPSKWDSVFESFEQADPSTTRTHGGTGLGLCIVRNLVNKMGGEI 743
Query: 717 KIVKKEGPGTLMRLYMQLTAPVDATEQHCQVDFANNNLMVLLALHGNMSRLITSKWLQKN 776
K+V+K G GTLMRLY+ L+ P D +Q+ Q DF+ L+V+L+++G+ +R+ITSKWL+K+
Sbjct: 744 KVVQKNGLGTLMRLYLILSTP-DTVDQNIQPDFSKYGLVVMLSMYGSTARMITSKWLRKH 802
Query: 777 GVLTMEASEWNGLTQILKELFEAKTSTHNNDFDTHFPAPEGLNSKFISIQELPNPTFVIV 836
G+ T+EAS+WN LTQI+++L E T + +N FD+ + L ++ +I E+ NP FVIV
Sbjct: 803 GIATVEASDWNELTQIIRDLLE--TGSRDNSFDSQHNISDPLRAELSNIVEIKNPVFVIV 860
Query: 837 VDIDLLDLSTDIWKEQLNFLHKYYARAKFVWLQNHDSSNTVKTELRKKGHILTVNKPLYK 896
VDI +LDL+T+IWKEQLN+L ++ +AKF WL HD+SNTVKTELR+KGH++ VNKPLYK
Sbjct: 861 VDIGVLDLTTNIWKEQLNYLDRFSNKAKFAWLLKHDTSNTVKTELRRKGHVMMVNKPLYK 920
Query: 897 AKMIHILEDVIKERN----VEVQKKNMKEDDLHESLEIEYTHHCDVASSDGSDISEIGSS 952
AKMI ILE VIK R +++ + D+ H+ LEI+ T +S D S+ S
Sbjct: 921 AKMIQILEAVIKNRKRGLCNDLRNRGNGSDESHDCLEIDPTQFDTCSSDDSSETS----- 975
Query: 953 NLVTANGEKQREEVVRINPSSLYQK--SNCLLGLSNGYMEHKEAFISTRAKGEDSKGGET 1010
GEKQ ++ V+ PS+L+ N L+ + + A S K + + +
Sbjct: 976 ------GEKQVDKSVK--PSTLHSPVLKNYLIDATTSNDDSTSA--SMTQKNPEEEDWKD 1025
Query: 1011 SRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALN- 1069
SG + +KSLEG+RILLAEDT V+QRVATIMLEKMGATV AV DGQQAVD+LN
Sbjct: 1026 RLYSGIALDGKNQKSLEGIRILLAEDTPVLQRVATIMLEKMGATVTAVWDGQQAVDSLNY 1085
Query: 1070 -----GMPGVERNTI---------TSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKS 1115
P E + T +T + + YDLILMDCQMPKMDGYEATK IR++
Sbjct: 1086 KSINAQAPTEEHKSFEEETANKVTTRETSLRNSSPYDLILMDCQMPKMDGYEATKAIRRA 1145
Query: 1116 EIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTK 1167
EIGT HIPIVALTAHAMS DEAKCLEVGMDAYLTKPID KLM STILSLTK
Sbjct: 1146 EIGTELHIPIVALTAHAMSSDEAKCLEVGMDAYLTKPIDRKLMVSTILSLTK 1197
>dbj|BAA32085.1| histidine kinase 1 [Arabidopsis thaliana] gi|25456590|pir||T52459
sensory transduction histidine kinase 1 [validated] -
Arabidopsis thaliana
Length = 1207
Score = 1430 bits (3702), Expect = 0.0
Identities = 759/1192 (63%), Positives = 916/1192 (76%), Gaps = 56/1192 (4%)
Query: 8 VFDRLLLWFATCWKKNSTLTGRRKF---HKDVEKEDFQYSTTQCLSSYYSVFVVRLAIMV 64
VFD++ W T W+ N L R+ DVE+++FQY+++ CLSSYYSVFVVRLAIMV
Sbjct: 30 VFDKMTEW-VTPWRSN--LESPREMMILRGDVEQDEFQYASSHCLSSYYSVFVVRLAIMV 86
Query: 65 MLAILIGLLTILTWHFTKIYTTKSLNSLAYDLRYELLQRPILRMWNILNSTAEITTAQVK 124
MLAILIGLLT+LTWHFT+IYT +SL +LAY LRYELLQRP+LRMW++LN+T+E+TTAQVK
Sbjct: 87 MLAILIGLLTVLTWHFTRIYTKQSLQTLAYGLRYELLQRPVLRMWSVLNTTSELTTAQVK 146
Query: 125 LSEYVIRSHGNLATQAEQVEMYESMRAVTWALFASRKALNSITVKYRNGFVQAFHRDLKD 184
LSEYVI+ + TQ E VEMY++M+ VTWALFAS KALN+IT+ YRNGFVQAFHRD
Sbjct: 147 LSEYVIKKYDKPTTQEELVEMYQAMKDVTWALFASAKALNAITINYRNGFVQAFHRDPAS 206
Query: 185 NNIFYIYTDLSYNETNSFAAHEDTNS--------NKSAIWYREQLDPVNGEKIGKAMKIA 236
++ FYI++DL N + S ED + N SAIWY++QLDPV GE +GK +KI
Sbjct: 207 SSTFYIFSDLK-NYSISGTGPEDVSGWNNKSIHGNMSAIWYQQQLDPVTGENLGKPLKIP 265
Query: 237 PEDSISIAGLSQVPDGVASWHVSVGKFTDSPLLSAALPVWDSSNKSIVAVVGVTTALYSV 296
P+D I+IAG+SQVPDG ASWHV+V K+ DSPLLSAALPV+D+SNKSIVAVVGVTTALYSV
Sbjct: 266 PDDLINIAGISQVPDGEASWHVTVSKYMDSPLLSAALPVFDASNKSIVAVVGVTTALYSV 325
Query: 297 GQLMKELVDKHSGHMYLTSQEGYLLATSTNDPLLTNSTKKPKLKMAVDCDNEVIREGAMW 356
GQLM++LV+ H GH+YLTSQEGYLLATST+ PLL N++ P+L A D + VI+ GA W
Sbjct: 326 GQLMRDLVEVHGGHIYLTSQEGYLLATSTDGPLLKNTSNGPQLMKATDSEEWVIKSGAQW 385
Query: 357 LKKTYENNFPPSHEVHEENARLGHQQYYIDSFFLILKKLPLVGVIIIPRKHIMGQADERA 416
L+KTY + P H VH EN +LG Q+YYIDSF+L LK+LP+VGV+IIPRK IMG+ DERA
Sbjct: 386 LEKTYGSKRP--HVVHAENVKLGDQRYYIDSFYLNLKRLPIVGVVIIPRKFIMGKVDERA 443
Query: 417 FKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFL 476
FKTL+ILISAS+CI IGCVCILILTNGVSKEM LRAELI L+ARR+AEASSNYKSQFL
Sbjct: 444 FKTLIILISASVCIFFIGCVCILILTNGVSKEMKLRAELIRQLDARRRAEASSNYKSQFL 503
Query: 477 ANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVES 536
ANMSHELRTPMAAVIGLLDIL+SDD L+NEQ ATVTQIRKCSTALL LLNNILD+SKVES
Sbjct: 504 ANMSHELRTPMAAVIGLLDILISDDCLSNEQYATVTQIRKCSTALLRLLNNILDLSKVES 563
Query: 537 GKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANL 596
GKLVLEEAEFD GRELEGLVDMFSVQCINHNVE +LDLSDDMP LVRGDSAR+VQIFANL
Sbjct: 564 GKLVLEEAEFDLGRELEGLVDMFSVQCINHNVETVLDLSDDMPALVRGDSARLVQIFANL 623
Query: 597 INNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNID 656
I+NSIKFT +GHIILRGWCEN NS D + +++++ KTK QH NH +K+
Sbjct: 624 ISNSIKFTTTGHIILRGWCENINSLHDEMSVSVDRRKPWAPMKTKQVQHRNHLQKSCKNA 683
Query: 657 NKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEI 716
NKM+LWFEVDDTGCGIDPSKWDSVFESFEQAD STTR HGGTGLGLCIVR+LVNKMGGEI
Sbjct: 684 NKMVLWFEVDDTGCGIDPSKWDSVFESFEQADPSTTRTHGGTGLGLCIVRNLVNKMGGEI 743
Query: 717 KIVKKEGPGTLMRLYMQLTAPVDATEQHCQVDFANNNLMVLLALHGNMSRLITSKWLQKN 776
K+V+K G GTLMRLY+ L+ P D +Q+ Q DF+ L+V+L+++G+ +R+ITSKWL+K+
Sbjct: 744 KVVQKNGLGTLMRLYLILSTP-DTVDQNIQPDFSKYGLVVMLSMYGSTARMITSKWLRKH 802
Query: 777 GVLTMEASEWNGLTQILKELFEAKTSTHNNDFDTHFPAPEGLNSKFISIQELPNPTFVIV 836
G+ T+EAS+WN LTQI+++L E T + +N FD+ + L ++ +I E+ NP FVIV
Sbjct: 803 GIATVEASDWNELTQIIRDLLE--TGSRDNSFDSQHNISDPLRAELSNIVEIKNPVFVIV 860
Query: 837 VDIDLLDLSTDIWKEQLNFLHKYYARAKFVWLQNHDSSNTVKTELRKKGHILTVNKPLYK 896
VDI +LDL+T+IWKEQLN+L ++ +AKF WL HD+SNTVKTELR+KGH++ VNKPLYK
Sbjct: 861 VDIGVLDLTTNIWKEQLNYLDRFSNKAKFAWLLKHDTSNTVKTELRRKGHVMMVNKPLYK 920
Query: 897 AKMIHILEDVIKERN----VEVQKKNMKEDDLHESLEIEYTHHCDVASSDGSDISEIGSS 952
AKMI ILE VIK R +++ + D+ H+ LEI+ T +S D S+ S
Sbjct: 921 AKMIQILEAVIKNRKRGLCNDLRNRGNGSDESHDCLEIDPTQFDTCSSDDSSETS----- 975
Query: 953 NLVTANGEKQREEVVRINPSSLYQK--SNCLLGLSNGYMEHKEAFISTRAKGEDSKGGET 1010
GEKQ ++ V+ PS+L+ N L+ + + A S K + + +
Sbjct: 976 ------GEKQVDKSVK--PSTLHSPVLKNYLIDATTSNDDSTSA--SMTQKNPEEEDWKD 1025
Query: 1011 SRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALN- 1069
SG + +KSLEG+R LLAEDT V+QRVATIMLEKMGATV AV DGQQAVD+LN
Sbjct: 1026 RLYSGIALDGKNQKSLEGIRFLLAEDTPVLQRVATIMLEKMGATVTAVWDGQQAVDSLNY 1085
Query: 1070 -----GMPGVERNTI---------TSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKS 1115
P E + T +T + + YDLILMDCQMPKMDGYEATK IR++
Sbjct: 1086 KSINAQAPTEEHKSFEEETANKVTTRETSLRNSSPYDLILMDCQMPKMDGYEATKAIRRA 1145
Query: 1116 EIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTK 1167
EIGT HIPIVALTAHAMS DEAKCLEVGMDAYLTKPID KLM STILSLTK
Sbjct: 1146 EIGTELHIPIVALTAHAMSSDEAKCLEVGMDAYLTKPIDRKLMVSTILSLTK 1197
>gb|AAQ10680.2| cold inducible histidine kinase 1 [Catharanthus roseus]
Length = 1205
Score = 1411 bits (3652), Expect = 0.0
Identities = 763/1183 (64%), Positives = 902/1183 (75%), Gaps = 58/1183 (4%)
Query: 28 GRRKFHKDVEK-EDFQ-YSTTQCLSSYYSVFVVRLAIMVMLAILIGLLTILTWHFTKIYT 85
GR DVE E+FQ ++T CLSSYYSVFVVRLAIMVMLAILIG+LT+LTWHFTK+YT
Sbjct: 35 GRNFNRDDVESAEEFQDAASTLCLSSYYSVFVVRLAIMVMLAILIGMLTLLTWHFTKVYT 94
Query: 86 TKSLNSLAYDLRYELLQRPILRMWNILNSTAEITTAQVKLSEYVIRSHGNLATQAEQVEM 145
T+SL +LAY LRYEL QRPILRMWNILNST EITTAQVKLSEYVIR + QA+QVE+
Sbjct: 95 TRSLKTLAYGLRYELPQRPILRMWNILNSTVEITTAQVKLSEYVIRKYSKPVNQAQQVEL 154
Query: 146 YESMRAVTWALFASRKALNSITVKYRNGFVQAFHRDLKDNNIFYIYTDLS-YN-----ET 199
YE MR VTWALFASR+AL++IT+ YRNGFVQAFHRD + NN +YIY+DL+ Y+ +T
Sbjct: 155 YEVMRDVTWALFASRRALSAITINYRNGFVQAFHRDHRSNNTYYIYSDLANYSISGSYDT 214
Query: 200 NSFAAHEDTNS-----NKSAIWYREQLDPVNGEKIGKAMKIAPEDSISIAGLSQVPDGVA 254
+ A+ + N N SAIWYRE LDP GEKIGK+ I+P++ I+IAG+SQVPDG A
Sbjct: 215 SLLASRQGWNDQSIQGNLSAIWYREPLDPATGEKIGKSSPISPDELINIAGISQVPDGQA 274
Query: 255 SWHVSVGKFTDSPLLSAALPVWDSSNKSIVAVVGVTTALYSVGQLMKELVDKHSGHMYLT 314
SWHV+V K+TDSPLLSAALPVWDS+ + IVAVVGVTTALYSVGQLMKE HSGH+YLT
Sbjct: 275 SWHVAVSKYTDSPLLSAALPVWDSTKEIIVAVVGVTTALYSVGQLMKETCRVHSGHIYLT 334
Query: 315 SQEGYLLATSTNDPLLTNSTKKPKLKMAVDCDNEVIREGAMWLKKTYENNFPPSHEVHEE 374
SQEG+ LATSTN PLL NS+ PKL MAVD VIR GA L+KTY N FPP+HEVH E
Sbjct: 335 SQEGWWLATSTNTPLLMNSSTGPKLMMAVDSQEFVIRSGAECLQKTYGNKFPPNHEVHIE 394
Query: 375 NARLGHQQYYIDSFFLILKKLPLVGVIIIPRKHIMGQADERAFKTLVILISASLCIIVIG 434
NA+LG+Q YYIDSFFL LK+LP+VGVII+PRK++MG+ DERAFKTL+ LISASLCI+VIG
Sbjct: 395 NAKLGNQLYYIDSFFLNLKRLPMVGVIILPRKYVMGKVDERAFKTLLALISASLCILVIG 454
Query: 435 CVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLL 494
CVCI ILTNGVS+EM LRAE ISHL+ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLL
Sbjct: 455 CVCIFILTNGVSREMKLRAEFISHLDARRKAEASSNYKSQFLANMSHELRTPMAAVIGLL 514
Query: 495 DILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEG 554
DIL+SDD LTNEQ A VTQIRKCSTALL LLNNILD+SKVESGKLVLEE EFD GRELEG
Sbjct: 515 DILMSDDCLTNEQFANVTQIRKCSTALLRLLNNILDLSKVESGKLVLEETEFDLGRELEG 574
Query: 555 LVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGW 614
LVDMFSVQCINHNVE ILDLS+DMPK VRGDSARVVQIFANLI+NS+KFT SGHII+RGW
Sbjct: 575 LVDMFSVQCINHNVETILDLSEDMPKFVRGDSARVVQIFANLISNSMKFTTSGHIIVRGW 634
Query: 615 CENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDP 674
CEN N + ++ QK ++ K+K +++ +K+ +NK++LWFEV+DTGCGIDP
Sbjct: 635 CENQNPSISIRSSSINQKECWRLQRLKLKHQDSNDQKSFRKENKVVLWFEVEDTGCGIDP 694
Query: 675 SKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQL 734
SKW+SVFESFEQAD STTRLHGGTGLGLCIVR+LVNKMGGEIK+VKK G GTLMRL + L
Sbjct: 695 SKWESVFESFEQADPSTTRLHGGTGLGLCIVRTLVNKMGGEIKVVKKNGAGTLMRLCLLL 754
Query: 735 TAPV---DATEQHCQVDFANNNLMVLLALHGNMSRLITSKWLQKNGVLTMEASEWNGLTQ 791
P +HCQ F + L VLLAL+G M RLI S+WL+KNGV T EASEWN LTQ
Sbjct: 755 NTPAADPHGIVKHCQFKFEQHKLTVLLALNGRMGRLILSQWLKKNGVNTCEASEWNELTQ 814
Query: 792 ILKELFEAKTSTHNNDFDTHFPAPEGLNSKFISIQELPNPTFVIVVDIDLLDLSTDIWKE 851
IL E F+ K+ H++ + P + + SI F+IV+D+ LLDL TDIWKE
Sbjct: 815 ILLENFQCKSYIHSSPSEISEPEEVDIQASSFSI-------FIIVIDVGLLDLRTDIWKE 867
Query: 852 QLNFLHKYYARAKFVWLQNHDSSNTVKTELRKKGHILTVNKPLYKAKMIHILEDVIKERN 911
QL++L +Y+ RAKF W+ NHD+SNT+K+ELR++GH+L +N+P+Y AK+IHI E V+K++N
Sbjct: 868 QLSYLDEYFGRAKFAWILNHDTSNTIKSELRQRGHLLMINRPIYMAKLIHIFEAVMKDQN 927
Query: 912 VEVQ------KKNMKEDDLHESLEIEYTHHCDVASSDGSDISEIGSSNLVT--ANGEKQR 963
VE+Q K E L E EI+ C ASSD SD SE +S+ V+ GEK
Sbjct: 928 VELQKYMNTLKATAAEGYLSECHEIDVIPFC-AASSDDSDKSERENSSSVSPVQAGEKTY 986
Query: 964 EEVVRINPSSLYQKSNCLLGLSNGYMEHKEAFISTRAKGEDSKGGETSRVSGSSKAMNGK 1023
E + + +S Q L+N ++E ++ + + ++G + R + ++N K
Sbjct: 987 E---KFHIASACQNRT----LNNYFVEFPQSNLEENSFDMRNRGVKGPRERHKNSSVNSK 1039
Query: 1024 --------------KSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALN 1069
KSL+GLRILLAEDT V+QRVATIMLEK+GA VVAVGDG QAVDAL
Sbjct: 1040 DRDTQSNGTLWNEQKSLDGLRILLAEDTPVLQRVATIMLEKLGAKVVAVGDGLQAVDALR 1099
Query: 1070 GMPGVERNTITSQTEIL----SFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPI 1125
M ++ + S E + S P Y LILMDCQMPKMDGY ATK IRKSE+GT HIPI
Sbjct: 1100 FMFDSNQSNLESPEEDVINSTSLP-YHLILMDCQMPKMDGYAATKAIRKSEMGTGLHIPI 1158
Query: 1126 VALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTKR 1168
VALTAHAMS DEAKCLEVGMDAYLTKPID KLM STILSLT+R
Sbjct: 1159 VALTAHAMSSDEAKCLEVGMDAYLTKPIDRKLMVSTILSLTRR 1201
>pir||T08856 hypothetical protein A_TM017A05.5 - Arabidopsis thaliana
Length = 648
Score = 722 bits (1863), Expect = 0.0
Identities = 403/702 (57%), Positives = 485/702 (68%), Gaps = 85/702 (12%)
Query: 487 MAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEF 546
MAAVIGLLDIL+SDD L+NEQ ATVTQIRKCSTALL LLNNILD+SKVESGKLVLEEAEF
Sbjct: 1 MAAVIGLLDILISDDCLSNEQYATVTQIRKCSTALLRLLNNILDLSKVESGKLVLEEAEF 60
Query: 547 DFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLS 606
D GRELEGLVDMFSVQCINHNVE +LDLSDDMP LVRGDSAR+VQIFANLI+NSIKFT +
Sbjct: 61 DLGRELEGLVDMFSVQCINHNVETVLDLSDDMPALVRGDSARLVQIFANLISNSIKFTTT 120
Query: 607 GHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVD 666
GHIILRGWCEN NS D + +++++ KTK QH NH +K+ NKM+LWFEVD
Sbjct: 121 GHIILRGWCENINSLHDEMSVSVDRRKPWAPMKTKQVQHRNHLQKSCKNANKMVLWFEVD 180
Query: 667 DTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGT 726
DTGCGIDPSKWDSVFESFEQAD STTR HGGTGLGLCIVR+LV
Sbjct: 181 DTGCGIDPSKWDSVFESFEQADPSTTRTHGGTGLGLCIVRNLV----------------- 223
Query: 727 LMRLYMQLTAPVDATEQHCQVDFANNNLMVLLALHGNMSRLITSKWLQKNGVLTMEASEW 786
+L+++G+ +R+ITSKWL+K+G+ T+EAS+W
Sbjct: 224 ------------------------------MLSMYGSTARMITSKWLRKHGIATVEASDW 253
Query: 787 NGLTQILKELFEAKTSTHNNDFDTHFPAPEGLNSKFISIQELPNPTFVIVVDIDLLDLST 846
N LTQI+++L E T + +N FD+ + L ++ +I E+ NP FVIVVDI +LDL+T
Sbjct: 254 NELTQIIRDLLE--TGSRDNSFDSQHNISDPLRAELSNIVEIKNPVFVIVVDIGVLDLTT 311
Query: 847 DIWKEQLNFLHKYYARAKFVWLQNHDSSNTVKTELRKKGHILTVNKPLYKAKMIHILEDV 906
+IWKEQLN+L ++ +AKF WL HD+SNTVKTELR+KGH++ VNKPLYKAKMI ILE V
Sbjct: 312 NIWKEQLNYLDRFSNKAKFAWLLKHDTSNTVKTELRRKGHVMMVNKPLYKAKMIQILEAV 371
Query: 907 IKERNV----EVQKKNMKEDDLHESLEIEYTHHCDVASSDGSDISEIGSSNLVTANGEKQ 962
IK R +++ + D+ H+ LEI+ T +S D S+ S GEKQ
Sbjct: 372 IKNRKRGLCNDLRNRGNGSDESHDCLEIDPTQFDTCSSDDSSETS-----------GEKQ 420
Query: 963 REEVVRINPSSLYQK--SNCLLGLSNGYMEHKEAFISTRAKGEDSKGGETSRVSGSSKAM 1020
++ V+ PS+L+ N L+ + + A S K + + + SG +
Sbjct: 421 VDKSVK--PSTLHSPVLKNYLIDATTSNDDSTSA--SMTQKNPEEEDWKDRLYSGIALDG 476
Query: 1021 NGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALN------GMPGV 1074
+KSLEG+RILLAEDT V+QRVATIMLEKMGATV AV DGQQAVD+LN P
Sbjct: 477 KNQKSLEGIRILLAEDTPVLQRVATIMLEKMGATVTAVWDGQQAVDSLNYKSINAQAPTE 536
Query: 1075 ERNTI---------TSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPI 1125
E + T +T + + YDLILMDCQMPKMDGYEATK IR++EIGT HIPI
Sbjct: 537 EHKSFEEETANKVTTRETSLRNSSPYDLILMDCQMPKMDGYEATKAIRRAEIGTELHIPI 596
Query: 1126 VALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTK 1167
VALTAHAMS DEAKCLEVGMDAYLTKPID KLM STILSLTK
Sbjct: 597 VALTAHAMSSDEAKCLEVGMDAYLTKPIDRKLMVSTILSLTK 638
>emb|CAC42409.1| putative osmosensor histidine Kinase [Populus euramericana]
Length = 331
Score = 479 bits (1232), Expect = e-133
Identities = 236/329 (71%), Positives = 275/329 (82%), Gaps = 2/329 (0%)
Query: 481 HELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLV 540
HELRTPMAAVIGLLDIL+ DD LTNEQ A VTQIRKCSTALL LLNNILD+SKVESGKLV
Sbjct: 1 HELRTPMAAVIGLLDILICDDCLTNEQYANVTQIRKCSTALLRLLNNILDLSKVESGKLV 60
Query: 541 LEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNS 600
LE+AEFD GRELEGL+DMFSVQC NHNVE +LDLSD+MPKL+RGDSARVVQIFANLI+NS
Sbjct: 61 LEDAEFDLGRELEGLIDMFSVQCTNHNVEAVLDLSDEMPKLLRGDSARVVQIFANLISNS 120
Query: 601 IKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMI 660
IKFT +GHIILRGWCEN N+ + F L+QK C+ K K++Q NH KKA +NKMI
Sbjct: 121 IKFTTTGHIILRGWCENLNNTYNDTQFHLDQKKMRCAIKPKLRQQGNHLKKACKKENKMI 180
Query: 661 LWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVK 720
LWFE+DDTGCGIDP KW+SVFESFEQAD STTRLHGGTGLGLCIVR+LVNKMGGEIK+VK
Sbjct: 181 LWFEIDDTGCGIDPRKWESVFESFEQADPSTTRLHGGTGLGLCIVRTLVNKMGGEIKVVK 240
Query: 721 KEGPGTLMRLYMQLTAPVDATEQHCQVDFANNNLMVLLALHGNMSRLITSKWLQKNGVLT 780
K GPGTLMRLY+ L P D + CQVDF+++N +VL+AL+G+M R+I S WL++ G+ T
Sbjct: 241 KNGPGTLMRLYLLLKTPADGADLPCQVDFSSHNAVVLVALYGSMGRVIMSLWLREIGLTT 300
Query: 781 MEASEWNGLTQILKELFEAKTSTHNNDFD 809
+ SEWN LT++L++LF A+ N FD
Sbjct: 301 LGVSEWNELTRVLRKLFHAR--RRENGFD 327
>gb|AAC62867.1| putative histidine kinase [Arabidopsis thaliana]
gi|7487571|pir||T00441 probable histidine kinase
[imported] - Arabidopsis thaliana
gi|15226594|ref|NP_182265.1| cytokinin-responsive
histidine kinase (CKI1) [Arabidopsis thaliana]
gi|1679803|dbj|BAA13416.1| histidine kinase homolog
[Arabidopsis thaliana]
Length = 1122
Score = 280 bits (717), Expect = 2e-73
Identities = 297/1192 (24%), Positives = 498/1192 (40%), Gaps = 191/1192 (16%)
Query: 57 VVRLAIMVMLAILIGLLTILTWHFTKIYTTKSLNSLAYDLRYELLQRPILRMWNILNSTA 116
+V ++ L ++ + I W T K + S DLR L+ + +
Sbjct: 14 IVVFCVLAFLVVVFECIWISNWRTTTENLVKEVASFTEDLRTSLVSE--------IENIG 65
Query: 117 EITTAQVKLSEYVIRSHGNLATQAEQVEMYESMRAVTWALFASRKALNSITVKYRNGFVQ 176
+ T A+ LS + + E + LF + + ++ V
Sbjct: 66 KFTYAKTNLSTIGLARVIDSYITNNDTGFTEIQTQIAPLLFVAYSTILQVSQ------VS 119
Query: 177 AFHRDLKDNNIFYIYTDLSYNETNSFAAHEDTNSNKSAIWYREQLDPVNGEKIGKAMKIA 236
RD + + Y S FA +S WY + +D + G G + K
Sbjct: 120 YISRD----GLMFSYIAESNTSVAVFANSSSNSSRGDYTWYTQTVDQLTGRLNGNSTKSQ 175
Query: 237 PEDSISIAGLSQVPDG---VASWHVSVGKFTDSPLLSAALPVWDSSNKSIVAVVGVTTAL 293
D A S+G + L+ + + ++ S K +V++ L
Sbjct: 176 SLDVTHTDWFQAAQSNNYTTAFVGTSLGGEDNETLIQSVVSLY--SKKGLVSLGFPVKTL 233
Query: 294 YSVGQLMKELVDKHSGHMYLTSQEGYLLAT--STNDPLLTNSTKKPKLKMAVDCDNEVIR 351
V ++ H +Y+ +++G +L S ND ++ C
Sbjct: 234 TEV----LNSLNLHGEELYMWTKDGTVLVREGSLNDSFFISNGSI--------CFGR--E 279
Query: 352 EGAMWLKKTYENNFPPSHEVHEENARLGHQQYYIDSFFLILKKLPLVGVIIIPRK----H 407
++W + EN +EV E RL +Q + + + +PL ++ P K
Sbjct: 280 SNSLWSQCIPENCSSSGYEV--EIKRLRYQAF---CSVIEVSGVPLRYTLMFPNKGGATR 334
Query: 408 IMGQADERAFKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEA 467
I QA++ ++ +V++I V + + +EM++RA LI+ +EA ++AE
Sbjct: 335 IKHQAEKAKYQLIVVMIFLGFGWPVW---FVWFMMQATRREMHMRATLINQMEATQQAER 391
Query: 468 SSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNN 527
S KSQ AN SH++R +A + GL+DI + ++ T+ Q+ C+ L+ LLN+
Sbjct: 392 KSMNKSQAFANASHDIRGALAGMKGLIDICRDGVKPGSDVDTTLNQVNVCAKDLVALLNS 451
Query: 528 ILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMP---KLVRG 584
+LD+SK+ESGK+ L E +F+ + LE ++D + + V+++LD D VRG
Sbjct: 452 VLDMSKIESGKMQLVEEDFNLSKLLEDVIDFYHPVAMKKGVDVVLDPHDGSVFKFSNVRG 511
Query: 585 DSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFG--------- 635
DS R+ QI NL++N++KFT+ GHI +R W + P G + + L P G
Sbjct: 512 DSGRLKQILNNLVSNAVKFTVDGHIAVRAWAQRP---GSNSSVVLASYPKGVSKFVKSMF 568
Query: 636 CSRKTKMKQHENH-SKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRL 694
C K + +E S N N M FEVDDTG GI SVFE++ Q T +
Sbjct: 569 CKNKEESSTYETEISNSIRNNANTMEFVFEVDDTGKGIPMEMRKSVFENYVQV-RETAQG 627
Query: 695 HGGTGLGLCIVRSLVNKMGGEIKIVKKE--GPGTLMRLYMQLTA----PVDATEQHCQVD 748
H GTGLGL IV+SLV MGGEI+I K GT + + LT PV + +++
Sbjct: 628 HQGTGLGLGIVQSLVRLMGGEIRITDKAMGEKGTCFQFNVLLTTLESPPVSDMKVRQEIE 687
Query: 749 FAN-----------------------------NNLM----------VLLALHGNMSRLIT 769
NN + V+L L R +T
Sbjct: 688 AGGDYVSTPNLGLTINTSLGGSMNIRNLSPRFNNCLSSSPKQEGSRVVLLLKNEERRRVT 747
Query: 770 SKWLQKNGVLTMEASEWNGLTQILKELFEAKTSTHNNDFDTHFPAPEGLNSKFI------ 823
K+++ G+ +W L+ L+ LF + + P FI
Sbjct: 748 EKYIKNLGIKVTVVEKWEHLSYALERLFGFSPQSSMGRAECSLSCPSSRELPFIGMDGID 807
Query: 824 SIQELPNPTFVIVVDIDLL--DLSTDIWKEQLNFLHKYY------ARAKFVWLQNHDSSN 875
S +LP + + LL D T + E + + ++ K VWL N
Sbjct: 808 SRSQLPKRRSISFSAVVLLVIDAKTGPFFELCDIVKQFRRGLPHGISCKVVWL------N 861
Query: 876 TVKTELRKKGHILTVNKPLYKAKMIHILE-------DVIKERNVEVQKKNMKEDDLHESL 928
T + ++G I + ++PL+ ++++ +L+ V+KE E+Q++++
Sbjct: 862 ESSTRVSERGDI-SCSRPLHGSRLMEVLKMLPEFGGTVLKEPPTELQRESL--------- 911
Query: 929 EIEYTHHCDVASSDGSDISEIGSSNLVTANGEKQREEVVRINPSSLYQK--SNCLLGLSN 986
H VA + +V PSS++ K ++ ++
Sbjct: 912 ----LRHSFVAE-------------------RSPKHKVQEEGPSSMFNKKLGKRIMASTD 948
Query: 987 GYMEHKEAFISTRAK--GEDSKGGETSRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVA 1044
E + + T K G ETS+ S + L G R+L+ +D + ++VA
Sbjct: 949 SESETRVKSVRTGRKPIGNPEDEQETSKPSDD-------EFLRGKRVLVVDDNFISRKVA 1001
Query: 1045 TIMLEKMGATVVAVGD-GQQAVDALNGMPGVERNTITSQTEILSFPSYDLILMDCQMPKM 1103
T L+KMG + V D G++A+ + G+ + + L F D I MDCQMP+M
Sbjct: 1002 TGKLKKMGVSEVEQCDSGKEALRLVT--EGLTQREEQGSVDKLPF---DYIFMDCQMPEM 1056
Query: 1104 DGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAK-CLEVGMDAYLTKPID 1154
DGYEAT+EIRK E PI+A++ H +EA+ ++ GMDA+L K ++
Sbjct: 1057 DGYEATREIRKVEKSYGVRTPIIAVSGHDPGSEEARETIQAGMDAFLDKSLN 1108
>dbj|BAD72542.1| putative histidine kinase 1 [Oryza sativa (japonica
cultivar-group)]
Length = 1048
Score = 193 bits (491), Expect = 3e-47
Identities = 191/645 (29%), Positives = 282/645 (43%), Gaps = 77/645 (11%)
Query: 216 WYREQLDPVNGEKIGKAMKIAPEDSISIAGLSQVPDGVASWHVSVGKFTDSP---LLSAA 272
W+ + +DP G +G A AP + + DG S+ P +L +
Sbjct: 146 WFTQAVDPATGRPVGNATAAAPHQQLPPNVTRLLLDG-GGGGASLADGWARPGVRMLFLS 204
Query: 273 LPVWDSSNKSIVAVVGVTTALYSVGQLMKELVDKHSGHMYLTSQEGYLLATSTNDPLLTN 332
PV ++ A V V + +++L D G Y + G A
Sbjct: 205 APVGGGGG-AVSAAVAVDDVVLRGAAGLRQLRDL--GMYYAVAGNGGATAAPP------- 254
Query: 333 STKKPKLKMAVDCDNEVIREGAMW--LKKTYEN-NFPPSHEVH---EENARLGHQQYYID 386
+P ++ D E A++ +K T + PP +VH + R + I
Sbjct: 255 -APEPAAYRSLLGDGAAAEEMALFSSVKCTASAIDAPPKLDVHGVKSDKYRFACTNFDIS 313
Query: 387 S----FFLILKKLPLVGVIIIPRKHIMGQADERAFKTLVILISASLCIIVIGCVCIL-IL 441
F ++L+K +VGV R T+V + A+ + CV + L
Sbjct: 314 GVQMGFRVVLRKSAMVGVF------------RRGGVTMVAVACAAAAAATVACVLMARAL 361
Query: 442 TNGVSKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDD 501
V++E L A+L H +A R+AE S KS A+ SH++R+ +AAV GL+++ S
Sbjct: 362 RRAVAREAALGADLARHRDALRQAERKSMNKSNAFASASHDIRSALAAVAGLVEV--SRP 419
Query: 502 RLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSV 561
+ Q+ C+ LL +LN+ILD +KVESGK+ LEE EF+ LE VDM +V
Sbjct: 420 EANPNIVDNLNQMELCTNKLLDILNSILDTTKVESGKVQLEEVEFNMADVLEESVDMANV 479
Query: 562 QCINHNVEIILDLSD-DMPKL--VRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENP 618
I +E+I D D + K + GDS R QI NL+ N++KFT GH+ILR W P
Sbjct: 480 VGITKGIEVIWDPCDFSVMKCDNIIGDSKRFKQILDNLLGNAMKFTQEGHVILRAWANRP 539
Query: 619 ---NSYGDSENF---TLEQKPFGCSRKTKM-KQHENHSKKASNIDNKMILWFEVDDTGCG 671
S G F +LE F K + +N N N + +FEV DTG G
Sbjct: 540 IARGSIGAPSRFAYRSLENNFFSFFFGAKEDRVSQNSFNPLQNDPNSVEFYFEVVDTGIG 599
Query: 672 IDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKE--------G 723
I K +SVFE++ Q HGGTGLGL IV+S V MGGEI I +KE G
Sbjct: 600 IPKEKRESVFENYVQVKEG----HGGTGLGLGIVQSFVRLMGGEISIKEKEPGERGTCFG 655
Query: 724 PGTLMRLYMQLTAPVDATEQHCQVD--------FANNNLM----VLLALHGNMSRLITSK 771
L++ A D E V F N +L +HG+ +R +
Sbjct: 656 FNVLLKTSGNQAAEEDIEEGPSTVSELDIRASVFRETNCFKGWHCILFVHGDETRRVLQA 715
Query: 772 WLQKNGVLTMEASEWNGLTQILKELFEAKTSTHNNDFDTHFPAPE 816
W++ G M+ G+ I L +A++S + D D F + E
Sbjct: 716 WMESIG---MKVWMVPGVESISSTLEKARSSRDDCDVDRCFSSKE 757
Score = 118 bits (295), Expect = 1e-24
Identities = 61/144 (42%), Positives = 91/144 (62%), Gaps = 4/144 (2%)
Query: 1024 KSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQT 1083
+ L+G+ +LL EDT V+Q + ML ++GA V GDG +AVD +ER +++ +
Sbjct: 906 RPLDGMHVLLVEDTLVLQTIQRKMLNQLGAIVELAGDGAKAVDMFRD--AIERASVSEEH 963
Query: 1084 EILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEV 1143
+ P YD+I MDCQMP+MDGYEAT+ IR+ E PI+ALTAH+M D K ++V
Sbjct: 964 SV-PLP-YDVIFMDCQMPRMDGYEATRRIREEESRYGIRTPIIALTAHSMEDDLQKAIDV 1021
Query: 1144 GMDAYLTKPIDFKLMESTILSLTK 1167
GMD ++TKPI+ + + + + K
Sbjct: 1022 GMDLHMTKPIERRRIVEAVHGVCK 1045
>ref|ZP_00316318.1| COG0642: Signal transduction histidine kinase [Microbulbifer
degradans 2-40]
Length = 765
Score = 182 bits (461), Expect = 8e-44
Identities = 123/367 (33%), Positives = 185/367 (49%), Gaps = 54/367 (14%)
Query: 448 EMNLRAELISHLEARR---KAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLT 504
E+ L E +E RR +AEA++ KSQF+A MSHE+RTPM VIG+++ +L D L
Sbjct: 226 ELALMREKEEKIETRRLVQEAEAANRAKSQFIATMSHEIRTPMNGVIGMVE-MLRDTTLD 284
Query: 505 NEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCI 564
+ Q + I + +L++++N+ILD SK+E+GK+ LE +FD LE V MFS
Sbjct: 285 DSQRHYLGIIERSGDSLMNIINDILDYSKIEAGKMSLEHMQFDLEELLEDCVQMFSASTD 344
Query: 565 NHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDS 624
N+E+I +S + PK + GD R+ Q+ NLI N+ KFT GHI +
Sbjct: 345 KRNIELICSISPNTPKQLMGDPTRLKQVLVNLIGNAFKFTSEGHIYV------------- 391
Query: 625 ENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESF 684
E AS+ D MI F V+D+G GI+ ++ + +F++F
Sbjct: 392 ---------------------EARQVNASDADLPMI-HFSVEDSGIGIESNQQEKLFDAF 429
Query: 685 EQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQLTAPVDATEQH 744
QAD STTR GGTGLGL I + L MGGEI + +E G+ T A +
Sbjct: 430 CQADGSTTRKFGGTGLGLTICKQLAEMMGGEIGVYSREKAGSTFWFSAVFTHVEKAINEQ 489
Query: 745 CQVDFANNNLMVLLALHGNMSRLITSKWLQKNGVLTMEASEWNGLTQIL---KELFEAKT 801
A L+V+ N S+++ V+ A++WN +I+ ++ E +
Sbjct: 490 PPQSLAGKKLLVV-----NQSKIVEK-------VIASHAADWNIACRIVHSGEKALEVIS 537
Query: 802 STHNNDF 808
H+ DF
Sbjct: 538 GQHHYDF 544
Score = 72.0 bits (175), Expect = 1e-10
Identities = 45/135 (33%), Positives = 68/135 (50%), Gaps = 17/135 (12%)
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSF 1088
L +L+AED V + V +L K DG QA++A+ P
Sbjct: 636 LHVLVAEDNPVNRMVIEGLLAKFDIAPKFAEDGLQALNAVTESP---------------- 679
Query: 1089 PSYDLILMDCQMPKMDGYEATKEIRKSEIGTSF-HIPIVALTAHAMSCDEAKCLEVGMDA 1147
YDLI+MDC+MP+MDG+EAT+ IR E + I+ALTAH + + + GM+
Sbjct: 680 TPYDLIIMDCEMPEMDGFEATRSIRTWESNNNQPATTIIALTAHVEAEHRQRVFDSGMNY 739
Query: 1148 YLTKPIDFKLMESTI 1162
YL+KP+ + + +
Sbjct: 740 YLSKPVTMEKLSEAL 754
>ref|ZP_00351862.1| COG0642: Signal transduction histidine kinase [Rubrobacter
xylanophilus DSM 9941]
Length = 1063
Score = 180 bits (457), Expect = 2e-43
Identities = 113/306 (36%), Positives = 165/306 (52%), Gaps = 32/306 (10%)
Query: 438 ILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDIL 497
IL +++ + EL EA+ +AEA+S KS+F+ANMSHE+RTPM VIG+ ++L
Sbjct: 298 ILAAARDITERKRIEEEL---REAKEQAEAASRAKSEFVANMSHEIRTPMNGVIGMAELL 354
Query: 498 LSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVD 557
L D LT EQ IR ALL +LN+ILD SK+E+G+L LE F+ +E+E +V
Sbjct: 355 L-DTPLTPEQREYARTIRSSGEALLAILNDILDFSKIEAGRLSLERIPFEIHKEVEEVVS 413
Query: 558 MFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCEN 617
+ + + +E+I + +P L++GD R+ Q+ NLI N+IKFT G +++
Sbjct: 414 LLAPRAHAKGLELICFIEPGVPPLIKGDPFRLRQVLTNLIGNAIKFTEEGEVVV------ 467
Query: 618 PNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKW 677
S + + E+K S A ++ L FEV D+G GI +
Sbjct: 468 ------SVSLSKEEK----------------SAPAGERGGEIELRFEVKDSGIGIQEEQQ 505
Query: 678 DSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQLTAP 737
+FE+F QAD STTR +GGTGLGL I R LV MGGEI + G+ + P
Sbjct: 506 KHLFEAFSQADASTTRRYGGTGLGLAISRQLVEMMGGEIGLKSSPKEGSTFHFSARFVLP 565
Query: 738 VDATEQ 743
+ E+
Sbjct: 566 EEEEEK 571
Score = 85.9 bits (211), Expect = 7e-15
Identities = 55/136 (40%), Positives = 78/136 (56%), Gaps = 6/136 (4%)
Query: 1023 KKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQ 1082
K+ E ILLAED V Q+VA ML+K+G V +G++A+ L P ++ + +
Sbjct: 759 KEGKERPLILLAEDNPVNQQVAASMLKKLGYRVELAQNGEEALKKLFPPPSSPPSS-SGR 817
Query: 1083 TEILSFPS--YDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIP---IVALTAHAMSCDE 1137
E P Y +LMD QMPKMDG EAT+ IR E +P I+A++AHAM D
Sbjct: 818 EEGGGSPKKKYAAVLMDIQMPKMDGLEATRRIRSKEEEGVLRLPRTPIIAMSAHAMQEDR 877
Query: 1138 AKCLEVGMDAYLTKPI 1153
+ L+ GMD Y++KP+
Sbjct: 878 ERALKAGMDHYISKPV 893
Score = 50.8 bits (120), Expect = 3e-04
Identities = 46/181 (25%), Positives = 82/181 (44%), Gaps = 19/181 (10%)
Query: 997 STRAKGEDSKGGETSRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVV 1056
S + E+ +GG R + ++SL GLR+L+ +D A + + LE
Sbjct: 572 SPKTPPEEGEGGGARRTR-EEEERGRRESLAGLRVLVVDDNATNRTILCRQLEAWRMEPK 630
Query: 1057 AVGDGQQAVDALNGMPGVERNTITSQTEILSFPS-YDLILMDCQMPKMDGYEATKEIR-- 1113
+VG Q+A+ L S T + S S YDL+++D QMP+ DG E + I+
Sbjct: 631 SVGGAQEAIRELEA----------SGTFLASSSSGYDLVILDMQMPETDGLELARRIKSR 680
Query: 1114 ----KSEIGTSFH-IPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTKR 1168
+ + G +F + ++ LT+ E ++A L KP+ + I +L ++
Sbjct: 681 LQEEEEKRGGAFSALRLMLLTSMDPQTKEKIKASGTVEACLQKPVRQSQLYDAIATLFQK 740
Query: 1169 E 1169
+
Sbjct: 741 K 741
>ref|ZP_00186489.2| COG0642: Signal transduction histidine kinase [Rubrobacter
xylanophilus DSM 9941]
Length = 1375
Score = 180 bits (457), Expect = 2e-43
Identities = 107/284 (37%), Positives = 155/284 (53%), Gaps = 27/284 (9%)
Query: 460 EARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCST 519
EA+ +AEA+S KS+F+ANMSHE+RTPM VIG+ ++LL D LT EQ IR
Sbjct: 621 EAKEQAEAASRAKSEFVANMSHEIRTPMNGVIGMAELLL-DTPLTPEQREYARTIRSSGE 679
Query: 520 ALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMP 579
ALL +LN+ILD SK+E+G+L LE F+ +E+E +V + + + +E+I + +P
Sbjct: 680 ALLAILNDILDFSKIEAGRLSLERIPFEIHKEVEEVVSLLAPRAHAKGLELICFIEPGVP 739
Query: 580 KLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRK 639
L++GD R+ Q+ NLI N+IKFT G +++
Sbjct: 740 PLIKGDPFRLRQVLTNLIGNAIKFTEEGEVVVSA-------------------------- 773
Query: 640 TKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTG 699
+ K+ + S A ++ L FEV D+G GI + +FE+F QAD STTR +GGTG
Sbjct: 774 SLSKEEKKKSAPAGERGGEIELRFEVKDSGIGIQEEQQKHLFEAFSQADASTTRRYGGTG 833
Query: 700 LGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQLTAPVDATEQ 743
LGL I R LV MGGEI + G+ + P + E+
Sbjct: 834 LGLAISRQLVEMMGGEIGLKSSPKEGSTFHFSARFVLPEEEEEK 877
Score = 85.9 bits (211), Expect = 7e-15
Identities = 55/136 (40%), Positives = 78/136 (56%), Gaps = 6/136 (4%)
Query: 1023 KKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQ 1082
K+ E ILLAED V Q+VA ML+K+G V +G++A+ L P ++ + +
Sbjct: 1065 KEGKERPLILLAEDNPVNQQVAASMLKKLGYRVELAQNGEEALKKLFPPPSSPPSS-SGR 1123
Query: 1083 TEILSFPS--YDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIP---IVALTAHAMSCDE 1137
E P Y +LMD QMPKMDG EAT+ IR E +P I+A++AHAM D
Sbjct: 1124 EEGGGSPKKKYAAVLMDIQMPKMDGLEATRRIRSKEEEGVLRLPRTPIIAMSAHAMQEDR 1183
Query: 1138 AKCLEVGMDAYLTKPI 1153
+ L+ GMD Y++KP+
Sbjct: 1184 ERALKAGMDHYISKPV 1199
Score = 50.8 bits (120), Expect = 3e-04
Identities = 46/181 (25%), Positives = 82/181 (44%), Gaps = 19/181 (10%)
Query: 997 STRAKGEDSKGGETSRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVV 1056
S + E+ +GG R + ++SL GLR+L+ +D A + + LE
Sbjct: 878 SPKTPPEEGEGGGARRTR-EEEERGRRESLAGLRVLVVDDNATNRTILCRQLEAWRMEPK 936
Query: 1057 AVGDGQQAVDALNGMPGVERNTITSQTEILSFPS-YDLILMDCQMPKMDGYEATKEIR-- 1113
+VG Q+A+ L S T + S S YDL+++D QMP+ DG E + I+
Sbjct: 937 SVGGAQEAIRELEA----------SGTFLASSSSGYDLVILDMQMPETDGLELARRIKSR 986
Query: 1114 ----KSEIGTSFH-IPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTKR 1168
+ + G +F + ++ LT+ E ++A L KP+ + I +L ++
Sbjct: 987 LQEEEEKRGGAFSALRLMLLTSMDPQTKEKIKASGTVEACLQKPVRQSQLYDAIATLFQK 1046
Query: 1169 E 1169
+
Sbjct: 1047 K 1047
>emb|CAD78909.1| sensory transduction histidine kinase-putative sensory histidine
kinase / response regulator hybrid [Rhodopirellula
baltica SH 1] gi|32476458|ref|NP_869452.1| sensory
transduction histidine kinase-putative sensory histidine
kinase / response regulator hybrid [Rhodopirellula
baltica SH 1]
Length = 871
Score = 177 bits (450), Expect = 1e-42
Identities = 137/396 (34%), Positives = 202/396 (50%), Gaps = 41/396 (10%)
Query: 439 LILTNGVSKEMN--LRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDI 496
+I T G+S+++ R+E H EA R A+A++ KS+FLANMSHE+RTPM A+IG+ D
Sbjct: 268 IIGTFGISRDITDLKRSEAALH-EAVRMADAANRSKSEFLANMSHEIRTPMNALIGMAD- 325
Query: 497 LLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLV 556
LLS +L +Q V I++ S LL L+N+ILD SK+E+ +L LE F G+ +E +
Sbjct: 326 LLSQTKLDPDQQDYVQIIQESSNGLLRLINDILDFSKIEARRLELESVPFSLGKTIESTL 385
Query: 557 DMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCE 616
FS + +E+ D+S ++P + GD RV Q+ NLI N+IKFT G + +R
Sbjct: 386 RSFSFKASEKGLELQRDISPELPDRLIGDPGRVRQVLTNLIGNAIKFTDHGGVRVR---- 441
Query: 617 NPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSK 676
E E+ R + + +++ D+ + L V D+G GI P++
Sbjct: 442 -------VEVLRREE------RADRDLAATHSGRESPERDDMVELRLSVCDSGIGIPPAQ 488
Query: 677 WDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQLTA 736
D+V F QAD STTR GGTGLGL I R LV MGGE+ + + G GT QL
Sbjct: 489 QDAVLNPFTQADASTTRRFGGTGLGLSISRQLVELMGGELHLQSEVGVGTTFS--FQLKM 546
Query: 737 PVDATEQHCQVDFANNN-----------LMVLLALHGNMSRLITSKWLQKNGVLTMEASE 785
PV +VD + L VL+A G ++ + + L+ G AS+
Sbjct: 547 PVCPASMPNEVDDEEEDLGGPARQRLAPLHVLVAEDGVTNQHVIAGLLRSLGHKCSIASD 606
Query: 786 W-NGLTQILKELFEAKTSTHNNDFDTHFPAPEGLNS 820
LT+ E ++ D H P +GL +
Sbjct: 607 GRETLTKWRSESYDVVL------MDMHMPVMDGLEA 636
Score = 94.7 bits (234), Expect = 2e-17
Identities = 58/140 (41%), Positives = 73/140 (51%), Gaps = 18/140 (12%)
Query: 1023 KKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQ 1082
++ L L +L+AED Q V +L +G DG R T+T
Sbjct: 569 RQRLAPLHVLVAEDGVTNQHVIAGLLRSLGHKCSIASDG--------------RETLTKW 614
Query: 1083 TEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLE 1142
SYD++LMD MP MDG EAT+ IR+ E GT H PIVALTA AMS D A C E
Sbjct: 615 RS----ESYDVVLMDMHMPVMDGLEATRAIRQEEWGTERHTPIVALTAAAMSEDAAACRE 670
Query: 1143 VGMDAYLTKPIDFKLMESTI 1162
GMD+YLTKPI + + +
Sbjct: 671 AGMDSYLTKPIHSRKLRDAL 690
>ref|ZP_00105980.1| COG2202: FOG: PAS/PAC domain [Nostoc punctiforme PCC 73102]
Length = 1403
Score = 177 bits (448), Expect = 2e-42
Identities = 101/265 (38%), Positives = 154/265 (58%), Gaps = 9/265 (3%)
Query: 460 EARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCST 519
+A+R+AEA++ KS+FLA MSHE+RTPM AVIG+ D+LL D LT +Q V +R
Sbjct: 665 QAKRQAEAANRAKSEFLAMMSHEIRTPMNAVIGMTDLLLDTD-LTPQQQDFVETVRSSGD 723
Query: 520 ALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMP 579
ALL ++N+ILD SK+ESGKL LEE FD +E + + + + N+E++ + +P
Sbjct: 724 ALLTIINDILDFSKIESGKLELEEQPFDLRACVEQAISLLAPKAAQKNIELVYLIQPQVP 783
Query: 580 KLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFT--------LEQ 631
+ GD R+ Q+ NL+NN+IKFT G ++L +S EN + L
Sbjct: 784 TQIVGDMTRLRQVLMNLLNNAIKFTERGEVVLSVELGAGDSVKQGENSSPLHPSLVRLRP 843
Query: 632 KPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLST 691
+ T ++ ++ +++ ++ + + F + DTG GI P K + +F F QAD+S
Sbjct: 844 SLREAAPTTTLRTSSRQARPSASSESPIQIQFAIKDTGIGITPEKIERLFHPFTQADVSM 903
Query: 692 TRLHGGTGLGLCIVRSLVNKMGGEI 716
TR +GGTGLGL I + L MGG +
Sbjct: 904 TRRYGGTGLGLVISKRLSKMMGGTL 928
Score = 89.4 bits (220), Expect = 7e-16
Identities = 54/143 (37%), Positives = 77/143 (53%), Gaps = 19/143 (13%)
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSF 1088
LRILLAEDT V Q+VA +ML+K+G + V +G + + AL P
Sbjct: 1130 LRILLAEDTIVNQKVALLMLQKIGYSADVVTNGVEVLKALQKQP---------------- 1173
Query: 1089 PSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAY 1148
YDL+LMD MP+MDG E + I + E F I+A+TA+AM D CL GMD Y
Sbjct: 1174 --YDLVLMDVHMPEMDGLETARRICQ-EWKVGFRPHIIAITANAMRGDREVCLAAGMDDY 1230
Query: 1149 LTKPIDFKLMESTILSLTKRETL 1171
++KPI + + + ++ +L
Sbjct: 1231 ISKPIQLQKLAQALSKCPRQRSL 1253
Score = 38.1 bits (87), Expect = 1.8
Identities = 30/133 (22%), Positives = 58/133 (43%), Gaps = 19/133 (14%)
Query: 1021 NGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTIT 1080
N L+G R+L+ +D ++ ++ E A ++A+ L G++
Sbjct: 979 NSPMQLQGKRLLIVDDNLTNCQIISLQAESWEMETYAAKSAKEALAQL--AQGIQ----- 1031
Query: 1081 SQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKC 1140
+D+ ++D +MP MDG ++IR+ ++P+V LT+ +
Sbjct: 1032 ----------FDIAILDMEMPAMDGITLARQIRRQ--SGCQNLPLVILTSLGREETFSDF 1079
Query: 1141 LEVGMDAYLTKPI 1153
+V A L+KPI
Sbjct: 1080 GDVEFAASLSKPI 1092
>gb|AAP53311.1| putative histidine kinase [Oryza sativa (japonica cultivar-group)]
gi|37533444|ref|NP_921024.1| putative histidine kinase
[Oryza sativa (japonica cultivar-group)]
gi|20279446|gb|AAM18726.1| putative histidine kinase
[Oryza sativa (japonica cultivar-group)]
Length = 925
Score = 167 bits (423), Expect = 2e-39
Identities = 121/341 (35%), Positives = 178/341 (51%), Gaps = 27/341 (7%)
Query: 405 RKHIMGQADERA--FKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEAR 462
RKH+M + A I+IS+++ IIV+ I+ T +E + L+ R
Sbjct: 269 RKHVMHCRFKHAPSLPWSAIMISSAVAIIVLLVGYIIYATLNSLEEAEDNYTTMRDLKGR 328
Query: 463 RKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALL 522
AEA+ KSQFLA +SHE+RTPM V+G+L +L+ D L Q V ++ +L+
Sbjct: 329 --AEAADVAKSQFLATVSHEIRTPMNGVLGMLQMLM-DTELDTTQRDFVVTAQESGKSLI 385
Query: 523 HLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLV 582
+L+N +LD++K+ESGK+ LE FD L+ +V +FS + +E+ + +SD +P ++
Sbjct: 386 NLINEVLDLAKIESGKIELEAVRFDVRDILDNVVSLFSEKSWAKGIELAVLVSDQVPDVL 445
Query: 583 RGDSARVVQIFANLINNSIKFTLSGHIILRGWC----------------ENPNSYGDSEN 626
GD R QI NL+ NS+KFT GHI +R EN +S+N
Sbjct: 446 IGDPWRFRQIITNLVGNSMKFTEQGHIFIRVHLIEEVKRKMEALDDTSPENIEVTANSKN 505
Query: 627 F----TLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFE 682
TL +RKT + K +S+ + + L V+DTG GI +F
Sbjct: 506 TMPYNTLSGLEVANNRKT--LESFRMFKDSSDAIDSVNLLVTVEDTGIGITKDAQTRIFT 563
Query: 683 SFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEG 723
F QAD ST+R +GGTG+GL I + LV MGGEI V K G
Sbjct: 564 PFMQADGSTSRTYGGTGIGLSITKRLVELMGGEIGFVSKPG 604
Score = 50.4 bits (119), Expect = 3e-04
Identities = 32/91 (35%), Positives = 44/91 (48%), Gaps = 17/91 (18%)
Query: 1026 LEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEI 1085
L G IL+ +D AV + VA L+K GA V V G++A+ L
Sbjct: 792 LTGKNILVVDDNAVNRIVAAGALKKYGAIVTCVDSGKEAISRLQPPH------------- 838
Query: 1086 LSFPSYDLILMDCQMPKMDGYEATKEIRKSE 1116
+D MD QMP+MDG+EAT+ +R E
Sbjct: 839 ----KFDACFMDVQMPEMDGFEATRLVRSVE 865
>dbj|BAD01586.1| histidine kinase 3 [Zea mays]
Length = 1049
Score = 165 bits (417), Expect = 1e-38
Identities = 106/328 (32%), Positives = 173/328 (52%), Gaps = 23/328 (7%)
Query: 417 FKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFL 476
+ ++I + ++ ++++G + L + E + R E + +AEA+ KSQFL
Sbjct: 394 WSAIIISTAVAIIVLLVGHIIYATLNSLEKAEHDYRVMR----ELKGQAEAADVAKSQFL 449
Query: 477 ANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVES 536
A +SHE+RTPM V+G+L +L+ + T +Q VT ++ L++L+N +LD++K+ES
Sbjct: 450 ATVSHEIRTPMNGVLGMLQMLMDTELDTTQQDFVVTA-QESGKVLINLINEVLDLAKIES 508
Query: 537 GKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANL 596
G++ LE FD L+ ++ +F + +E+ + +SD +P ++ GD R QI NL
Sbjct: 509 GRIELEAVPFDVRDILDNVISLFYEKSQAKGIELAVLVSDQVPDVLIGDPWRFRQIITNL 568
Query: 597 INNSIKFTLSGHIILR------------GWCE----NPNSYGDSENFTLEQKPFGCSRKT 640
+ NS+KFT GHI ++ +C+ N DS+N L G
Sbjct: 569 VGNSMKFTEQGHIFVQVHLVKELSRKGNNFCDVSAHNREILYDSDNSMLWNTLSGLEVAD 628
Query: 641 KMKQHENHS--KKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGT 698
+ EN + K ++ + + L V+DTG GI +F F QAD ST+R +GGT
Sbjct: 629 SWRSLENFTMFKNSNGETDTIRLAVRVEDTGIGITKDAQMRIFTPFMQADSSTSRTYGGT 688
Query: 699 GLGLCIVRSLVNKMGGEIKIVKKEGPGT 726
G+GL I + LV MGGEI K G G+
Sbjct: 689 GIGLSITKRLVELMGGEIGFTSKSGVGS 716
Score = 79.3 bits (194), Expect = 7e-13
Identities = 50/155 (32%), Positives = 77/155 (49%), Gaps = 35/155 (22%)
Query: 1026 LEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEI 1085
L G +IL+ +D AV ++VA L+K G+TV V G A+D L
Sbjct: 902 LTGKQILVVDDNAVNRKVAAGSLKKYGSTVTCVDSGSDAIDMLKPPH------------- 948
Query: 1086 LSFPSYDLILMDCQMPKMDGYEATKEIRK-----------SEIGTS-------FHIPIVA 1127
++D MD QMP+MDG+EAT+ IR EI + +H+PI+A
Sbjct: 949 ----TFDACFMDVQMPEMDGFEATRLIRSIEKKINDMIMMGEISANNYGNKAHWHVPILA 1004
Query: 1128 LTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTI 1162
+TA + KC++ GMD Y++KP + + + S +
Sbjct: 1005 MTADVIQATFEKCMQCGMDGYVSKPFEEQQLYSAV 1039
>dbj|BAD01585.1| histidine kinase 3 [Zea mays]
Length = 1201
Score = 165 bits (417), Expect = 1e-38
Identities = 106/328 (32%), Positives = 173/328 (52%), Gaps = 23/328 (7%)
Query: 417 FKTLVILISASLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFL 476
+ ++I + ++ ++++G + L + E + R E + +AEA+ KSQFL
Sbjct: 546 WSAIIISTAVAIIVLLVGHIIYATLNSLEKAEHDYRVMR----ELKGQAEAADVAKSQFL 601
Query: 477 ANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVES 536
A +SHE+RTPM V+G+L +L+ + T +Q VT ++ L++L+N +LD++K+ES
Sbjct: 602 ATVSHEIRTPMNGVLGMLQMLMDTELDTTQQDFVVTA-QESGKVLINLINEVLDLAKIES 660
Query: 537 GKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANL 596
G++ LE FD L+ ++ +F + +E+ + +SD +P ++ GD R QI NL
Sbjct: 661 GRIELEAVPFDVRDILDNVISLFYEKSQAKGIELAVLVSDQVPDVLIGDPWRFRQIITNL 720
Query: 597 INNSIKFTLSGHIILR------------GWCE----NPNSYGDSENFTLEQKPFGCSRKT 640
+ NS+KFT GHI ++ +C+ N DS+N L G
Sbjct: 721 VGNSMKFTEQGHIFVQVHLVKELSRKGNNFCDVSAHNREILYDSDNSMLWNTLSGLEVAD 780
Query: 641 KMKQHENHS--KKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGT 698
+ EN + K ++ + + L V+DTG GI +F F QAD ST+R +GGT
Sbjct: 781 SWRSLENFTMFKNSNGETDTIRLAVRVEDTGIGITKDAQMRIFTPFMQADSSTSRTYGGT 840
Query: 699 GLGLCIVRSLVNKMGGEIKIVKKEGPGT 726
G+GL I + LV MGGEI K G G+
Sbjct: 841 GIGLSITKRLVELMGGEIGFTSKSGVGS 868
Score = 79.3 bits (194), Expect = 7e-13
Identities = 50/155 (32%), Positives = 77/155 (49%), Gaps = 35/155 (22%)
Query: 1026 LEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEI 1085
L G +IL+ +D AV ++VA L+K G+TV V G A+D L
Sbjct: 1054 LTGKQILVVDDNAVNRKVAAGSLKKYGSTVTCVDSGSDAIDMLKPPH------------- 1100
Query: 1086 LSFPSYDLILMDCQMPKMDGYEATKEIRK-----------SEIGTS-------FHIPIVA 1127
++D MD QMP+MDG+EAT+ IR EI + +H+PI+A
Sbjct: 1101 ----TFDACFMDVQMPEMDGFEATRLIRSIEKKINDMIMMGEISANNYGNKAHWHVPILA 1156
Query: 1128 LTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTI 1162
+TA + KC++ GMD Y++KP + + + S +
Sbjct: 1157 MTADVIQATFEKCMQCGMDGYVSKPFEEQQLYSAV 1191
>gb|AAM05416.1| sensory transduction histidine kinase [Methanosarcina acetivorans
str. C2A] gi|20090861|ref|NP_616936.1| sensory
transduction histidine kinase [Methanosarcina
acetivorans C2A]
Length = 1000
Score = 160 bits (406), Expect = 2e-37
Identities = 141/521 (27%), Positives = 233/521 (44%), Gaps = 102/521 (19%)
Query: 443 NGVSKEMNLRAELISH--------LEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLL 494
NGV M + I+H L+A+ AE+++ K FLANMSHE+RTPM AV+G+L
Sbjct: 408 NGVPVRMFGTLQDITHRKLMEEQLLKAKEAAESANKAKGDFLANMSHEIRTPMNAVLGML 467
Query: 495 DILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEG 554
++LL + LT+EQ + + +LL +++++LD SK+E KL LE+ F+ +
Sbjct: 468 EMLL-ETSLTDEQREYLQLAHASAESLLSIIDDVLDFSKIEQNKLELEQISFELESLISH 526
Query: 555 LVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGW 614
++++ S + + +++ + D+P GD R+ Q+ N++ N+IKFT G I L
Sbjct: 527 IINLLSGKAGSKGLKLAFHIEKDLPSTFIGDPVRLKQVLFNILGNAIKFTKKGEIAL--- 583
Query: 615 CENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDP 674
S + M N S+ ++ L F+V DTG GI P
Sbjct: 584 ----------------------SVEVYMPSERNSGLSNSDFPEEVALLFKVKDTGIGIPP 621
Query: 675 SKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQ- 733
K +F+ F QAD S TR +GGTGLGL I LV M G I + + G G ++
Sbjct: 622 EKLSQIFDPFIQADASVTREYGGTGLGLAISSQLVELMEGRIWVESEVGKGCTFYFTVRV 681
Query: 734 -------------------LTAPVDATEQHCQVDFANNNLMVLLALHGNMSRLITSKWLQ 774
L ++A E + + +L VL A +++ + L+
Sbjct: 682 KRGESSENYGQGNNIPEESLLVCMEAKESYGNGITSGESLNVLFAEDHPINQKLILGLLE 741
Query: 775 KNG-VLTMEASEWNGLTQILKELFEAKTSTHNNDFDTHFPAPEGLNSKFISIQELPNPTF 833
K G LT+ S + L + + F+A D P +GL +
Sbjct: 742 KKGHKLTLVTSGKDALDALSRRDFDAVL------MDIQMPGMDGLEA------------- 782
Query: 834 VIVVDIDLLDLSTDIWKEQLNFLHKYYARAKFVWLQNHDSSNTVKTELRKKGHILTVNKP 893
+ D S+ + + + + + ARA + + + G ++KP
Sbjct: 783 ----TRRIRDPSSGVRRHNIPII-AFTARA----------LKEDREKCFEAGMNYYISKP 827
Query: 894 LYKAKMIHILEDV----------IKERNVE---VQKKNMKE 921
L K K+++ILED+ +ER V+ VQ+KN++E
Sbjct: 828 LKKEKLLNILEDIRVGKNTREKTARERTVQDRIVQEKNIQE 868
Score = 89.4 bits (220), Expect = 7e-16
Identities = 54/134 (40%), Positives = 71/134 (52%), Gaps = 19/134 (14%)
Query: 1021 NGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTIT 1080
NG S E L +L AED + Q++ +LEK G + V G+ A+DAL
Sbjct: 713 NGITSGESLNVLFAEDHPINQKLILGLLEKKGHKLTLVTSGKDALDAL------------ 760
Query: 1081 SQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFH-IPIVALTAHAMSCDEAK 1139
S +D +LMD QMP MDG EAT+ IR G H IPI+A TA A+ D K
Sbjct: 761 ------SRRDFDAVLMDIQMPGMDGLEATRRIRDPSSGVRRHNIPIIAFTARALKEDREK 814
Query: 1140 CLEVGMDAYLTKPI 1153
C E GM+ Y++KP+
Sbjct: 815 CFEAGMNYYISKPL 828
>ref|NP_681666.1| two-component hybrid sensor and regulator [Thermosynechococcus
elongatus BP-1] gi|22294598|dbj|BAC08428.1|
two-component hybrid sensor and regulator
[Thermosynechococcus elongatus BP-1]
Length = 1035
Score = 160 bits (405), Expect = 2e-37
Identities = 100/262 (38%), Positives = 143/262 (54%), Gaps = 30/262 (11%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
AR+ AE +S KS+FLA MSHE+RTPM A+IGL +LL D L +Q V IR A
Sbjct: 483 ARQAAEEASRAKSEFLATMSHEIRTPMNAIIGLTGLLL-DTPLNPQQQEFVNTIRLSGDA 541
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
LL ++N+ILD SK+ESGKL LE F+ +E ++D+ + + VE+ + +P
Sbjct: 542 LLGVINDILDFSKIESGKLELESYPFNLRTCVEEVLDLLASRAAEREVELAAHIDSGVPC 601
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
V GD R+ Q+ NLI N+IKFT G++ L ++ KP S+
Sbjct: 602 AVIGDMGRLRQVLVNLIGNAIKFTQKGNVTLY----------------VKAKPMPMSQYE 645
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
M + ++ F + DTG GI + + +F+ F Q D STTR +GGTGL
Sbjct: 646 FMPSYYSYL-------------FAIQDTGIGISAAGMNRLFKPFSQVDASTTRQYGGTGL 692
Query: 701 GLCIVRSLVNKMGGEIKIVKKE 722
GL I + LV +MGG+I + K+
Sbjct: 693 GLVICQRLVERMGGQIWVESKK 714
Score = 85.9 bits (211), Expect = 7e-15
Identities = 54/132 (40%), Positives = 72/132 (53%), Gaps = 19/132 (14%)
Query: 1023 KKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQ 1082
K L LRIL+AED V Q VA +LEK+G +G + +DA+ P
Sbjct: 902 KAELPPLRILVAEDNKVNQMVALRILEKLGYRGDIAANGLEVLDAVQRQP---------- 951
Query: 1083 TEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIP-IVALTAHAMSCDEAKCL 1141
YD+ILMD QMP+MDG AT+E+ + G P I+A+TA+AM D CL
Sbjct: 952 --------YDVILMDMQMPEMDGITATREVTQLFQGLKKPRPRIIAMTANAMESDRQLCL 1003
Query: 1142 EVGMDAYLTKPI 1153
+ GMD Y++KPI
Sbjct: 1004 DAGMDDYVSKPI 1015
Score = 38.9 bits (89), Expect = 1.0
Identities = 30/140 (21%), Positives = 61/140 (43%), Gaps = 23/140 (16%)
Query: 1026 LEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEI 1085
LE R+L+ +D A +++ + ++K + G A+ L V
Sbjct: 764 LENRRVLIVDDNATNRQILALQIKKWKMQPLVAESGPSALALLQQQGWV----------- 812
Query: 1086 LSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGM 1145
DL ++D QMP+MDG ++IR +P++ LT+ + E++ +
Sbjct: 813 ------DLAILDLQMPEMDGVTLARQIR----AAYGELPLILLTSLGNTLTESE--KALF 860
Query: 1146 DAYLTKPIDFKLMESTILSL 1165
+ ++KP+ + + + SL
Sbjct: 861 SSLISKPVKQSTLYNALNSL 880
>ref|NP_440376.1| hybrid sensory kinase [Synechocystis sp. PCC 6803]
gi|1652132|dbj|BAA17056.1| hybrid sensory kinase
[Synechocystis sp. PCC 6803] gi|7470852|pir||S75142
sensory transduction histidine kinase slr1759 -
Synechocystis sp. (strain PCC 6803)
Length = 1462
Score = 156 bits (395), Expect = 3e-36
Identities = 104/300 (34%), Positives = 155/300 (51%), Gaps = 29/300 (9%)
Query: 445 VSKEMNLRAELIS-HLEARRK---AEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSD 500
+ KE R EL +LE + AEA++ K +FLA MSHE+RTPM VIG+ ++L+
Sbjct: 747 LEKERERRRELAQKNLELEKATWAAEAANRAKGEFLAMMSHEIRTPMNGVIGMTELLIMT 806
Query: 501 DRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFS 560
D L +Q V IR+ LL ++N+ILD SK+E+ KLVLE F+ +E +++MF
Sbjct: 807 D-LNLQQLDYVQTIRQSGETLLTIINDILDFSKIEADKLVLETQAFELRPLIETVLEMFG 865
Query: 561 VQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNS 620
++E+ + P + GD R+ QI +NLI N++KFT G ++L
Sbjct: 866 PIARAKHLELTYGIDPQTPARILGDQVRLRQILSNLIGNALKFTEKGEVVL--------- 916
Query: 621 YGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSV 680
T++ +PF + + H + F + DTG GI + D +
Sbjct: 917 -------TVKGEPFDPAESYHTILNLPHPSHR--------ICFNLRDTGIGIPLDRQDRL 961
Query: 681 FESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQLTAPVDA 740
F+SF Q D STTR +GGTGLGL I + L MGG + + + G G+ R + TA A
Sbjct: 962 FKSFSQVDSSTTRKYGGTGLGLVISQRLTQMMGGVLTVTSEPGVGSNFRFCILTTAQAPA 1021
Score = 85.5 bits (210), Expect = 1e-14
Identities = 52/134 (38%), Positives = 71/134 (52%), Gaps = 19/134 (14%)
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSF 1088
L+ILLAED V Q+VA ML +G V +GQ+ +DAL
Sbjct: 1190 LQILLAEDNLVNQKVAHQMLNNLGYPVAIANNGQEVIDALEKK----------------- 1232
Query: 1089 PSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAY 1148
YDL+LMD QMP MDG A + IR++ + IVA+TA+AM D +CL+ GMD Y
Sbjct: 1233 -FYDLVLMDMQMPVMDGITACRHIRQT-LPLERQPRIVAMTANAMPGDRQECLDAGMDGY 1290
Query: 1149 LTKPIDFKLMESTI 1162
++KPI + +
Sbjct: 1291 ISKPISINQLRKVL 1304
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.133 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,885,929,651
Number of Sequences: 2540612
Number of extensions: 78510705
Number of successful extensions: 262654
Number of sequences better than 10.0: 10173
Number of HSP's better than 10.0 without gapping: 3430
Number of HSP's successfully gapped in prelim test: 6743
Number of HSP's that attempted gapping in prelim test: 239206
Number of HSP's gapped (non-prelim): 18079
length of query: 1172
length of database: 863,360,394
effective HSP length: 139
effective length of query: 1033
effective length of database: 510,215,326
effective search space: 527052431758
effective search space used: 527052431758
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 81 (35.8 bits)
Medicago: description of AC121233.1


