

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119417.2 + phase: 0
(898 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
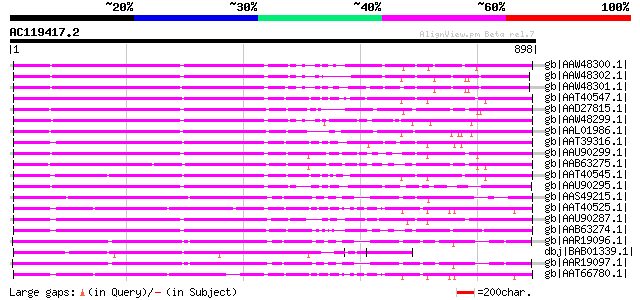
gb|AAW48300.1| potato resistance-like protein I2GA-SH23-1 [Solan... 481 e-134
gb|AAW48302.1| potato resistance-like protein I2GA-SH194-2 [Sola... 457 e-127
gb|AAW48301.1| potato resistance-like protein I2GA-SH23-3 [Solan... 456 e-126
gb|AAT40547.1| putative plant disease resistant protein [Solanum... 453 e-125
gb|AAD27815.1| disease resistance protein I2 [Lycopersicon escul... 452 e-125
gb|AAW48299.1| potato late blight resistance protein R3a [Solanu... 450 e-124
gb|AAL01986.1| I2C-5 [Lycopersicon pimpinellifolium] 447 e-124
gb|AAT39316.1| putative resistance complex protein I2C-2 [Solanu... 446 e-123
gb|AAU90299.1| putative disease resistance protein I2C-5 [Solanu... 436 e-120
gb|AAB63275.1| resistance complex protein I2C-2 [Lycopersicon es... 431 e-119
gb|AAT40545.1| putative plant disease resistant protein [Solanum... 429 e-118
gb|AAU90295.1| putative disease resistance protein I2 [Solanum d... 429 e-118
gb|AAS49215.1| disease resistance protein [Glycine max] 428 e-118
gb|AAT40525.1| putative disease resistance complex protein I2C-1... 426 e-117
gb|AAU90287.1| putative disease resistance protein I2C-5 [Solanu... 424 e-117
gb|AAB63274.1| resistance complex protein I2C-1 [Lycopersicon es... 422 e-116
gb|AAR19096.1| NBS-LRR type disease resistance protein RPG1-B [G... 409 e-112
dbj|BAB01339.1| disease resistance comples protein [Arabidopsis ... 408 e-112
gb|AAR19097.1| NBS-LRR type disease resistance protein RPG1-B [G... 408 e-112
gb|AAT66780.1| putative disease resistance protein [Solanum demi... 405 e-111
>gb|AAW48300.1| potato resistance-like protein I2GA-SH23-1 [Solanum tuberosum]
Length = 1265
Score = 481 bits (1238), Expect = e-134
Identities = 335/932 (35%), Positives = 494/932 (52%), Gaps = 112/932 (12%)
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
IL S+IW +P+++I+P+L L+Y LP+HLKRCF++C+IFPK YPF ++++I LW+A G +
Sbjct: 399 ILRSEIWELPHNDILPALMLSYNDLPAHLKRCFSFCAIFPKDYPFRKEQVIHLWIANGLI 458
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIER---SNDDIVKEKFVMHDVVYDLATIASGKSCC 123
+ +E+ G+ YF EL SRSL ER ++ + F+MHD+V DLA +AS K C
Sbjct: 459 PQE--DEIIEDSGNQYFLELRSRSLFERVPNPSEGNTENLFLMHDLVNDLAQVASSKLCI 516
Query: 124 RF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESY--LSCKV 179
R G + E H++Y+ E F K + + LR+ LPI L + Y LS +V
Sbjct: 517 RLEESQGYHLLEKGRHLSYSMGEDGEFEKLTPLYKLERLRTLLPICIDLTDCYHPLSKRV 576
Query: 180 IDDLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYLQT 238
+++P ++ LR+LSLS+Y I LP+ + KL LR+L++SHT+IK PD+ C LY L+T
Sbjct: 577 QLNILPRLRSLRVLSLSHYRIKDLPDDLFIKLKLLRFLDISHTEIKRFPDSICALYNLET 636
Query: 239 LLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL--TVFLVGKQKV 296
LLLS C L ELP+ + KLINLRHLDIS T + KMP+ + +L++LQ L FLVG
Sbjct: 637 LLLSSCADLEELPLQMEKLINLRHLDISNTCLLKMPLHLSKLKSLQVLVGAKFLVG---- 692
Query: 297 GLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQQT--EESPT 354
GL + +LG+ NL G L + LQN +D EA A ++ K H+++L + W + + + S T
Sbjct: 693 GLRMEDLGEVHNLYGSLSVVELQNVVDSREAVKAKMREKNHVDKLSLEWSESSSADNSQT 752
Query: 355 NEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVP 414
IL+EL+P N+K+L I Y G +FP+WL D F +V LS+++C+ C +LP LGQ+P
Sbjct: 753 ERDILDELRPHKNIKELQIIGYRGTNFPNWLADPLFLKLVQLSLRNCKNCYSLPALGQLP 812
Query: 415 FLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKF 474
FLK L I GM + + EFYG + S +PF LEKLEF MP W++W +F
Sbjct: 813 FLKLLSIGGMPGITEVTEEFYG-----SWSSKKPFNCLEKLEFKDMPEWKQWDQLGSGEF 867
Query: 475 PFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQG 534
P L+ L++ +C EL +E + I LSS+KS +++ G
Sbjct: 868 PI--LEKLLIENCPEL---------GLETVPI--------------QLSSLKSFEVI--G 900
Query: 535 SPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPT 594
SP + ++ D + + G + ++ L + +SLT+FP + LPT
Sbjct: 901 SPMVGVVFYD-------AQLEGMKQ-------------IEELRISDCNSLTSFPFSILPT 940
Query: 595 SLQSLRIDECQNLAFLRP-ETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCS 653
+L+ + I +CQ L +P S + +TLE +C D ++ L P + L +E C
Sbjct: 941 TLKRIEISDCQKLKLEQPVGEMSMFLEELTLENCDCIDDISPELL---PRARTLFVEDCH 997
Query: 654 SLKSIFISEKNSSLSLST---------------LQSLKVSNCKSLRSLPQRMDTLFVLKS 698
+L I +L + + SL + L+ LP+RM L L S
Sbjct: 998 NLTRFLIPTATETLLIGNCKNVEKLSVACGGPQMTSLSIDGSLKLKWLPERMQEL--LPS 1055
Query: 699 LTLDKLSLCCEVACLP--------PKLQFMHIESLGLATPVTEWGFQSLCFLSDLHIGGD 750
L +LS C E+ P +LQ + E L EW Q L L+DL I D
Sbjct: 1056 LKYLQLSNCPEIESFPEGGLPFNLQQLQICNCEK--LVNGRKEWRLQRLLCLTDLFIDHD 1113
Query: 751 NIVNTLL--KKKLLPPLLVSLTITNLTEMMRLKGNRLQHISTLKNLSF-----KCCSTLE 803
++ + LP +L I+NL L L+ + +L+NL + S LE
Sbjct: 1114 GSDEEIVGGENWELPSSTQTLGISNL---KTLSSQHLKRLISLQNLYIEGNVPQIQSMLE 1170
Query: 804 TCKDFFPSFLKSLVFINCPKLMSLPD-MFPSSLETLEFDDCPRLGLLPRSGFPSSLKLLS 862
+ + L+SL N P L SLP+ PSSL L CP L LP G PSSL L
Sbjct: 1171 QGQFSHLTSLQSLQIENFPNLQSLPESALPSSLSQLRISLCPNLQSLPLKGMPSSLSKLY 1230
Query: 863 ISHCPLLKSRYATKRNDHLSKISHIPVVKING 894
I CPLLK + ++ I+ P +KING
Sbjct: 1231 IRDCPLLKPLLEFDKGEYWPNIAPFPTIKING 1262
>gb|AAW48302.1| potato resistance-like protein I2GA-SH194-2 [Solanum tuberosum]
Length = 1286
Score = 457 bits (1176), Expect = e-127
Identities = 329/942 (34%), Positives = 472/942 (49%), Gaps = 153/942 (16%)
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
IL S+IW +P+++I+P+L L+Y LP+HLKRCF+YCSIFPK YPF ++++I LW+A G +
Sbjct: 399 ILRSEIWELPHNDILPALILSYNDLPAHLKRCFSYCSIFPKDYPFRKEQVIHLWIANGLV 458
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIER---SNDDIVKEKFVMHDVVYDLATIASGKSCC 123
+ +E+ G+ YF EL SRSL +R ++ + F MHD+V DLA IAS K C
Sbjct: 459 PQG--DEIIEDSGNQYFLELRSRSLFQRVPNPSEGNTENLFFMHDLVNDLAQIASSKLCI 516
Query: 124 RF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSCKVID 181
R G + E H++Y++ F K + + LR+ LPI + +LS +V
Sbjct: 517 RLEESQGSHMLEQSRHLSYSKGYGGEFEKLTPLYKLEQLRTLLPICIDINCCFLSKRVQH 576
Query: 182 DLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYLQTLL 240
+++P ++ LR LSLS Y I LPN + KL LR+L+LS I+ LPD+ C LY L TLL
Sbjct: 577 NILPRLRSLRALSLSGYMIKELPNDLFIKLKLLRFLDLSEAWIEKLPDSVCGLYNLDTLL 636
Query: 241 LSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL--TVFLVGKQKVGL 298
LS C+ L ELP+ + KLINLRHLDISYT++ KMP+ + +L +LQ L FLVG GL
Sbjct: 637 LSSCYNLEELPLQMEKLINLRHLDISYTRLLKMPLHLSKLISLQVLVGAKFLVG----GL 692
Query: 299 SIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQQT--EESPTNE 356
+ +LG+ NL G L + LQN +D EA A ++ K H+++L + W + + + S T
Sbjct: 693 RMEDLGEVYNLYGSLSVVELQNVVDSREAVKAKMREKNHVDKLSLEWSESSSADNSQTER 752
Query: 357 VILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFL 416
IL+EL+P N+K+L I Y G FP+WL D F +V LSI +C+ C +LP LGQ+PFL
Sbjct: 753 DILDELRPHKNIKELQIIGYRGTKFPNWLADPLFLKLVQLSIDNCKNCYSLPALGQLPFL 812
Query: 417 KELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFPF 476
K L I GM + + EFYG + S +PF SL +L F MP W++W +FP
Sbjct: 813 KFLSIRGMHGITEVTEEFYG-----SCSSKKPFNSLVELRFEDMPEWKQWDLLGSGEFPI 867
Query: 477 PRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQGSP 536
L+ L++ +C EL S+E + I LSS+KS ++ GSP
Sbjct: 868 --LEKLLIENCPEL---------SLETVPI--------------QLSSLKSFEV--SGSP 900
Query: 537 ELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTSL 596
+ FP + LPT+L
Sbjct: 901 ----------------------------------------------MVINFPFSILPTTL 914
Query: 597 QSLRIDECQNLAFLRP-ETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCSSL 655
+ +RI +CQ L +P S + +TL+ +C D ++ L P + L + C +L
Sbjct: 915 KRIRIIDCQKLKLEQPVGEMSMFLEELTLQNCDCIDDISPELL---PRARHLCVYDCHNL 971
Query: 656 KSIFISEKNSSLSL---------------STLQSLKVSNCKSLRSLPQRMDTLFVLKSLT 700
I + SL + + + SL + C L+ LP+RM LF SL
Sbjct: 972 TRFLIPTASESLYICNCENVEVLSVACGGTQMTSLSIDGCLKLKGLPERMQELF--PSLN 1029
Query: 701 LDKLSLCCEVACLPPKLQFMHIESL------GLATPVTEWGFQSLCFLSDLHIGGD-NIV 753
LS C E+ P +++ L L EW Q L L H G D IV
Sbjct: 1030 TLHLSNCPEIESFPEGGLPFNLQQLIIYNCKKLVNGRKEWHLQRLTELIIYHDGSDEEIV 1089
Query: 754 NTLLKKKLLPPLLVSLTITNLTEM-------------MRLKGN-----------RLQHIS 789
+ LP + +L I NL + + +KGN + H++
Sbjct: 1090 GG--QNWELPSSIQTLRIWNLETLSSQHLKRLISLQNLSIKGNVPQIQSMLEQGQFSHLT 1147
Query: 790 TLKNLSFKCCSTLETCKDFFPSFLKSLVFINCPKLMSLPDM-FPSSLETLEFDDCPRLGL 848
+L++L +L + PS L L +CP L SLP+ PSSL L ++CP L
Sbjct: 1148 SLQSLQISSLQSLP--ESALPSSLSQLTISHCPNLQSLPEFALPSSLSQLTINNCPNLQS 1205
Query: 849 LPRSGFPSSLKLLSISHCPLLKSRYATKRNDHLSK--ISHIP 888
L S PSSL L ISHCP L+S LS+ ISH P
Sbjct: 1206 LSESTLPSSLSQLEISHCPKLQSLPELALPSSLSQLTISHCP 1247
>gb|AAW48301.1| potato resistance-like protein I2GA-SH23-3 [Solanum tuberosum]
Length = 1327
Score = 456 bits (1173), Expect = e-126
Identities = 330/929 (35%), Positives = 489/929 (52%), Gaps = 124/929 (13%)
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
IL S+IW +P+++I+P+L L+Y LP+HLKRCF+YC+IFPK YPF +++ I LW+A G +
Sbjct: 400 ILRSEIWELPHNDILPALMLSYNDLPAHLKRCFSYCAIFPKDYPFRKEQAIHLWIANGLV 459
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIER---SNDDIVKEKFVMHDVVYDLATIASGKSCC 123
+ +E+ G+ YF EL SRSL +R ++ ++ F+MHD+V DLA +AS K C
Sbjct: 460 PQG--DEIIEDSGNQYFLELRSRSLFQRVPNPSELNIENLFLMHDLVNDLAQVASSKLCI 517
Query: 124 RF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSR-LQESYLSCK-V 179
R G + E H++Y+ F K + + LR+ LP + + +Y CK V
Sbjct: 518 RLEESQGYHLLEKGRHLSYSMGYGGEFEKLTPLYKLEQLRTLLPTCNYFMPPNYPLCKRV 577
Query: 180 IDDLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYLQT 238
+ +++P ++ LR LSLS+Y I LP+ + KL LR+L++SHT+IK LPD C LY L+T
Sbjct: 578 LHNILPRLRSLRALSLSHYWIKDLPDDLFIKLKLLRFLDISHTEIKRLPDFICGLYNLET 637
Query: 239 LLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL--TVFLVGKQKV 296
LLLS C L ELP+ + KLINLRHLDIS T KMP+ + +L++LQ L FLVG +
Sbjct: 638 LLLSSCGFLEELPLQMEKLINLRHLDISNTSRLKMPLHLSKLKSLQVLVGARFLVG-DRG 696
Query: 297 GLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYW--DQQTEESPT 354
G + +LG+ NL G + + LQN +D EA A ++ K H++ L + W + S T
Sbjct: 697 GSRMEDLGEVHNLYGSVSVLELQNVVDSREAVKAKMREKNHVDRLSLEWSGSSSADNSQT 756
Query: 355 NEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVP 414
IL+EL+P N+K+L I Y G FP+WL D F +V LS+++C+ C +LP LG++P
Sbjct: 757 ERDILDELRPHKNIKELQIIGYRGTKFPNWLADPLFLKLVKLSLRNCKNCYSLPALGELP 816
Query: 415 FLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREW-ISFRGSK 473
LK L I GM + + EFYG + S +PF LEKLEF MP W++W I G
Sbjct: 817 CLKFLCIRGMHGITEVTEEFYG-----SWSSKKPFNCLEKLEFKDMPEWKQWHIPGNGE- 870
Query: 474 FPFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQ 533
FP L+ L +R+C EL S+E + I LSS+KSL+++
Sbjct: 871 --FPILEDLSIRNCPEL---------SLETVPI--------------QLSSLKSLEVI-- 903
Query: 534 GSPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLP 593
GSP + ++ +D + + G ++ L I ++SLT+FP + LP
Sbjct: 904 GSPMVGVVFDD-------AQLEGMKQIEEL--------------RISVNSLTSFPFSILP 942
Query: 594 TSLQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCS 653
T+L+++ I +CQ S + +TL + N C +LT F + + L I C
Sbjct: 943 TTLKTIEITDCQKCEM------SMFLEELTLNVYN-CHNLTRFLIP--TATESLFILYCE 993
Query: 654 SLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLFVLKSLTLDKLSLCCEVACL 713
+++ + ++ + ++ SL + C L+ LP+RM LF SL LS C E+
Sbjct: 994 NVEILLVACGGTQIT-----SLSIDGCLKLKGLPERMQELF--PSLNTLHLSNCPEIESF 1046
Query: 714 PPKLQFMHIESL------GLATPVTEWGFQSLCFLSDLHIGGD-NIVNTLLKKKLLPPLL 766
P +++ L L EW Q L L H G D IV + LP +
Sbjct: 1047 PEGGLPFNLQQLIIYNCKKLVNGRKEWHLQRLTELIIYHDGSDEEIVGG--QNWELPSSI 1104
Query: 767 VSLTITNLTEM-------------MRLKGN-----------RLQHISTLKNLSFKCCSTL 802
+L I NL + + +KGN + H+++L++L +L
Sbjct: 1105 QTLRIWNLETLSSQHLKRLISLQNLSIKGNVPQIQSMLEQGQFSHLTSLQSLQISSLQSL 1164
Query: 803 ETCKDFFPSFLKSLVFINCPKLMSLPDM-FPSSLETLEFDDCPRLGLLPRSGFPSSLKLL 861
+ PS L L +CP L SLP+ PSSL L ++CP L L S PSSL L
Sbjct: 1165 P--ESALPSSLSQLTISHCPNLQSLPEFALPSSLSQLTINNCPNLQSLSESTLPSSLSQL 1222
Query: 862 SISHCPLLKSRYATKRNDHLSK--ISHIP 888
ISHCP L+S LS+ ISH P
Sbjct: 1223 EISHCPKLQSLPELALPSSLSQLTISHCP 1251
>gb|AAT40547.1| putative plant disease resistant protein [Solanum demissum]
Length = 1406
Score = 453 bits (1166), Expect = e-125
Identities = 325/947 (34%), Positives = 493/947 (51%), Gaps = 93/947 (9%)
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
IL S++W +P+++I+P+L L+Y LP+HLK+CF+YC+IFPK YPF ++++I LW+A G L
Sbjct: 491 ILRSEMWELPDNDILPALMLSYNDLPTHLKQCFSYCAIFPKDYPFRKEQVIQLWIANGLL 550
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVK--EKFVMHDVVYDLATIASGKSCCR 124
+ + +E++G+ YF EL SRSL ER + + E+F+MHD++ DLA +AS K C R
Sbjct: 551 KGLQKDETIEDLGNLYFLELRSRSLFERVRESSKRNEEEFLMHDLINDLAQVASSKLCIR 610
Query: 125 F--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESY-LSCKVID 181
G + E +++Y+ + +F K + + K LR+ LPI + S+ LS +V+
Sbjct: 611 LEDNEGSHMLEKCRNLSYSLGD-GVFEKLKPLYKSKQLRTLLPINIQRGYSFPLSKRVLY 669
Query: 182 DLIPSIKRLRMLSLSNYNITVLPNSI-NKLVQLRYLNLSHTDIKCLPDTTCDLYYLQTLL 240
+++P + LR LSLS+Y I LPN + L LR L+LS T I+ LPD+ C LY L+ LL
Sbjct: 670 NILPRLTSLRALSLSHYRIKELPNDLFITLKLLRILDLSQTAIRKLPDSICALYNLEILL 729
Query: 241 LSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLT--VFLVGKQKVGL 298
LS C L ELP H+ KLINLRHLD + T + KMP+ +L+NL L F++G L
Sbjct: 730 LSSCIYLEELPPHMEKLINLRHLDTTGTSLLKMPLHPSKLKNLHVLVGFKFILGGCN-DL 788
Query: 299 SIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQQ-TEESPTNEV 357
+ +LG+ NL G + + LQN +D EA +AN+ K H+E L + W + + S T
Sbjct: 789 RMVDLGELHNLHGSISVLELQNVVDRREALNANMMKKEHVEMLSLEWSESIADSSQTEGD 848
Query: 358 ILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFLK 417
IL++LQP+ N+K+L I Y G FP+W+ D SF +V +S+ +C C +LP LGQ+P LK
Sbjct: 849 ILDKLQPNTNIKELEIAGYRGTKFPNWMADHSFLKLVGVSLSNCNNCASLPALGQLPSLK 908
Query: 418 ELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFPFP 477
L + GM R+ + EFYG T S +PF SLEKLEF MP W++W K FP
Sbjct: 909 FLTVRGMHRITEVSEEFYG-----TLSSKKPFNSLEKLEFAEMPEWKQWHVL--GKGEFP 961
Query: 478 RLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQGSPE 537
L ++ DC +L G LP L S+ + I C + T LS++K ++ SP+
Sbjct: 962 ALHDFLIEDCPKLIGKLPEKLCSLRGLRISKCPEL--SPETPIQLSNLKEFKVV--ASPK 1017
Query: 538 LSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTSLQ 597
+ +L +D+ L S + G +++ L C+ SLT P + LP++L+
Sbjct: 1018 VGVLFDDA--QLFTSQLQGMKQIVEL--------CIHD-----CHSLTFLPISILPSTLK 1062
Query: 598 SLRIDECQNLAFLRPETWSNYTSLVTLE--LKNCCDSLTSFQLNGFPVLQILSIEGCSSL 655
+ I C+ L L S + LE + CDS+ P LS+ C +L
Sbjct: 1063 KIEIYHCRKLK-LEASMISRGDCNMFLENLVIYGCDSIDDISPELVPRSHYLSVNSCPNL 1121
Query: 656 KSIFISEKNSSLSL----------------STLQSLKVSNCKSLRSLPQRMDTLFVLKSL 699
+ I + L + + L++L + +C+ L+ LP+ M L + SL
Sbjct: 1122 TRLLIPTETEKLYIWHCKNLEILSVASGTQTMLRNLSIRDCEKLKWLPECMQEL--IPSL 1179
Query: 700 TLDKLSLCCEVAC-----LPPKLQFMHIE-SLGLATPVTEWGFQSLCFLSDLHIGGDNIV 753
+L C E+ LP LQ + I L EW Q L L +L I D
Sbjct: 1180 KELELWFCTEIVSFPEGGLPFNLQVLRIHYCKKLVNARKEWHLQRLPCLRELTILHDG-S 1238
Query: 754 NTLLKKKLLPPLLVSLTITN---LTEMMRLKGNRLQHISTLKNLSFKCCSTLETCKDFFP 810
+ + LP + LT++N L+ + L+++ST +L + S LE
Sbjct: 1239 DLAGENWELPCSIRRLTVSNLKTLSSQLFKSLTSLEYLSTGNSLQIQ--SLLEEGLPISL 1296
Query: 811 S----------------------FLKSLVFINCPKLMSLPD-MFPSSLETLEFDDCPRLG 847
S L+ L +C +L S+P+ PSSL L +C +L
Sbjct: 1297 SRLTLFGNHELHSLPIEGLRQLTSLRDLFISSCDQLQSVPESALPSSLSELTIQNCHKLQ 1356
Query: 848 LLPRSGFPSSLKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKING 894
LP G P+S+ LSI CPLLK + ++ KI+HI + I+G
Sbjct: 1357 YLPVKGMPTSISSLSIYDCPLLKPLLEFDKGEYWPKIAHISTINIDG 1403
>gb|AAD27815.1| disease resistance protein I2 [Lycopersicon esculentum]
Length = 1266
Score = 452 bits (1164), Expect = e-125
Identities = 315/949 (33%), Positives = 473/949 (49%), Gaps = 141/949 (14%)
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
IL S+IW +P+++I+P+L L+Y LP+HLKRCF++C+IFPK YPF ++++I LW+A G +
Sbjct: 393 ILRSEIWELPHNDILPALMLSYNDLPAHLKRCFSFCAIFPKDYPFRKEQVIHLWIANGLV 452
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIERSNDDI---VKEKFVMHDVVYDLATIASGKSCC 123
+ + +++G+ YF EL SRSL E+ + ++E F+MHD+V DLA +AS K C
Sbjct: 453 P--VKDEINQDLGNQYFLELRSRSLFEKVPNPSKRNIEELFLMHDLVNDLAQLASSKLCI 510
Query: 124 RF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSCKVID 181
R G + E H++Y+ F K + + LR+ LPI + LS +V+
Sbjct: 511 RLEESQGSHMLEQCRHLSYSIGFNGEFKKLTPLYKLEQLRTLLPIRIEFRLHNLSKRVLH 570
Query: 182 DLIPSIKRLRMLSLSNYNITVLPNSI-NKLVQLRYLNLSHTDIKCLPDTTCDLYYLQTLL 240
+++P+++ LR LS S Y I LPN + KL LR+L++S T I LPD+ C LY L+TLL
Sbjct: 571 NILPTLRSLRALSFSQYKIKELPNDLFTKLKLLRFLDISRTWITKLPDSICGLYNLETLL 630
Query: 241 LSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLT--VFLVGKQKVGL 298
LS C L ELP+ + KLINLRHLD+S T+ KMP+ + RL++LQ L F V G
Sbjct: 631 LSSCADLEELPLQMEKLINLRHLDVSNTRRLKMPLHLSRLKSLQVLVGPKFFVD----GW 686
Query: 299 SIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQQT--EESPTNE 356
+ +LG+ NL G L + L+N +D EA A ++ K H+E+L + W + + + S T
Sbjct: 687 RMEDLGEAQNLHGSLSVVKLENVVDRREAVKAKMREKNHVEQLSLEWSESSIADNSQTES 746
Query: 357 VILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFL 416
IL+EL P N+KK+ I Y G +FP+W+ D F +V LS+++C+ C +LP LGQ+P L
Sbjct: 747 DILDELCPHKNIKKVEISGYRGTNFPNWVADPLFLKLVNLSLRNCKDCYSLPALGQLPCL 806
Query: 417 KELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFPF 476
K L + GM + + EFYG S +PF SLEKLEF M W++W + + F
Sbjct: 807 KFLSVKGMHGIRVVTEEFYGRL-----SSKKPFNSLEKLEFEDMTEWKQWHALGIGE--F 859
Query: 477 PRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQGSP 536
P L+ L +++C EL +P S++++ + C P S L+ M Q
Sbjct: 860 PTLENLSIKNCPELSLEIPIQFSSLKRLEVSDC---PVVFDDAQLFRS--QLEAMKQ--- 911
Query: 537 ELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTSL 596
++ +D+ +S+T+FP + LPT+L
Sbjct: 912 ------------------------------------IEEIDICDCNSVTSFPFSILPTTL 935
Query: 597 QSLRIDECQNLAFLRP--ETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCSS 654
+ ++I C L P E + Y + N C + P + LSIE C +
Sbjct: 936 KRIQISRCPKLKLEAPVGEMFVEYLRV------NDCGCVDDISPEFLPTARQLSIENCQN 989
Query: 655 LKSIFISEKNSSLSLST----------------LQSLKVSNCKSLRSLPQRMDTLFVLKS 698
+ I +L +S + SL + CK L+ LP+ +L S
Sbjct: 990 VTRFLIPTATETLRISNCENVEKLSVACGGAAQMTSLNIWGCKKLKCLPE------LLPS 1043
Query: 699 LTLDKLSLCCEV-ACLPPKLQFMH-IESLGLATPVTEWGFQSLCFLSDLHIGGDNIVNTL 756
L +LS C E+ LP L+ + I L EW Q L L H G D +
Sbjct: 1044 LKELRLSDCPEIEGELPFNLEILRIIYCKKLVNGRKEWHLQRLTELWIDHDGSDEDI--- 1100
Query: 757 LKKKLLPPLLVSLTITNLTEMMRLKGNRLQHISTLKNLSFKC------------------ 798
+ LP + LTI NL + QH+ +L +L + C
Sbjct: 1101 -EHWELPCSIQRLTIKNLKTLSS------QHLKSLTSLQYLCIEGYLSQIQSQGQLSSFS 1153
Query: 799 -CSTLET------------CKDFFPSFLKSLVFINCPKLMSL-PDMFPSSLETLEFDDCP 844
++L+T + PS L L +CP L SL PSSL L DCP
Sbjct: 1154 HLTSLQTLQIWNFLNLQSLAESALPSSLSHLEIDDCPNLQSLFESALPSSLSQLFIQDCP 1213
Query: 845 RLGLLPRSGFPSSLKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKIN 893
L LP G PSSL LSI +CPLL + ++ +I+HIP++ I+
Sbjct: 1214 NLQSLPFKGMPSSLSKLSIFNCPLLTPLLEFDKGEYWPQIAHIPIINID 1262
>gb|AAW48299.1| potato late blight resistance protein R3a [Solanum tuberosum]
Length = 1282
Score = 450 bits (1157), Expect = e-124
Identities = 329/932 (35%), Positives = 482/932 (51%), Gaps = 97/932 (10%)
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
IL S+IW +P+++I+P+L L+Y LP+HLKRCF++C+IFPK YPF ++++I LW+A G +
Sbjct: 399 ILRSEIWELPHNDILPALMLSYNDLPAHLKRCFSFCAIFPKDYPFRKEQVIHLWIANGLV 458
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIER---SNDDIVKEKFVMHDVVYDLATIASGKSCC 123
V +E+ G+ YF EL SRSL ER + + F+MHD+V DLA IAS K C
Sbjct: 459 PQEDV--IIEDSGNQYFLELRSRSLFERVPNPSQGNTENLFLMHDLVNDLAQIASSKLCI 516
Query: 124 RF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQES--YLSCKV 179
R G + E H++Y+ F K + + LR+ LP L + +LS +V
Sbjct: 517 RLEESQGSHMLEQSQHLSYSMGYGGEFEKLTPLYKLEQLRTLLPTCIDLPDCCHHLSKRV 576
Query: 180 IDDLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYLQT 238
+ +++P + LR LSLS Y I LPN + KL LR+L++S T+IK LPD+ C LY L+T
Sbjct: 577 LHNILPRLTSLRALSLSCYEIVELPNDLFIKLKLLRFLDISRTEIKRLPDSICALYNLET 636
Query: 239 LLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL--TVFLVGKQKV 296
LLLS C+ L ELP+ + KLINLRHLDIS T++ KMP+ + +L++LQ L FL+G
Sbjct: 637 LLLSSCYDLEELPLQMEKLINLRHLDISNTRLLKMPLHLSKLKSLQVLVGAKFLIG---- 692
Query: 297 GLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYW--DQQTEESPT 354
GL + +LG+ NL G L + LQN +D EA A ++ K H++ L + W + S T
Sbjct: 693 GLRMEDLGEVHNLYGSLSVVELQNVVDRREAVKAKMREKNHVDRLYLEWSGSSSADNSQT 752
Query: 355 NEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVP 414
IL+EL+P N+K + I Y G +FP+WL D F +V LS+++C+ C +LP LGQ+P
Sbjct: 753 ERDILDELRPHKNIKVVKITGYRGTNFPNWLADPLFLKLVKLSLRNCKNCYSLPALGQLP 812
Query: 415 FLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKF 474
FLK L I M + + EFYG + S +PF LEKLEF MP W++W +F
Sbjct: 813 FLKFLSIREMHGITEVTEEFYG-----SWSSKKPFNCLEKLEFKDMPEWKQWDLLGSGEF 867
Query: 475 PFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQG 534
P L+ L++ +C EL S+E + I LSS+KS D++ G
Sbjct: 868 PI--LEKLLIENCPEL---------SLETVPI--------------QLSSLKSFDVI--G 900
Query: 535 SP-----ELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPA 589
SP LS+L P L+ I KL S L+ L LI +
Sbjct: 901 SPLVINFPLSIL----PTTLKRIKISDCQKLKLEQPTGEISMFLEELTLIKCDCIDDISP 956
Query: 590 NGLPTSLQSLRIDECQNLA-FLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILS 648
LP + + L + + NL FL P T+ TL++ NC + G + L+
Sbjct: 957 ELLPRA-RKLWVQDWHNLTRFLIP------TATETLDIWNCENVEILSVACGGTQMTSLT 1009
Query: 649 IEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLP--------QRMDTLFVLKSLT 700
I C LK ++ E+ L L +L+ L +SNC + S P Q++ + K +
Sbjct: 1010 IAYCKKLK--WLPERMQEL-LPSLKELHLSNCPEIESFPEGGLPFNLQQLAIRYCKKLVN 1066
Query: 701 LDKLSLCCEVACLPPKL--------QFMHIESLGLATPVTEWGFQSLCFLSDLHIGGDNI 752
K CL + + + E+ L + + +L LS H+
Sbjct: 1067 GRKEWHLQRRLCLTALIIYHDGSDEEIVGGENWELPSSIQRLTIVNLKTLSSQHLKNLTS 1126
Query: 753 VNTLLKKKLLP---PLLVSLTITNLTEMMRLKGNRLQHI------STLKNLSFKCCSTLE 803
+ L + LP P+L ++LT + L+ + LQ + S+L +L C L+
Sbjct: 1127 LQYLFIRGNLPQIQPMLEQGQCSHLTSLQSLQISSLQSLPESALPSSLSHLEISHCPNLQ 1186
Query: 804 TC-KDFFPSFLKSLVFINCPKLMSLPD-MFPSSLETLEFDDCPRLGLLPRSGFPSSLKLL 861
+ + PS L L NCP L SL + PSSL LE CP L LP G PSSL L
Sbjct: 1187 SLPESALPSSLSQLTINNCPNLQSLSESTLPSSLSQLEISFCPNLQYLPLKGMPSSLSEL 1246
Query: 862 SISHCPLLKSRYATKRNDHLSKISHIPVVKIN 893
SI CPLLK + + ++ I+ P +KI+
Sbjct: 1247 SIYKCPLLKPQLEFDKGEYWPNIAQFPTIKID 1278
>gb|AAL01986.1| I2C-5 [Lycopersicon pimpinellifolium]
Length = 1297
Score = 447 bits (1149), Expect = e-124
Identities = 321/959 (33%), Positives = 485/959 (50%), Gaps = 137/959 (14%)
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
IL S+IW + +++I+P+L L+Y LP+HLKRCF++C+IFPK YPF ++++I LW+A G +
Sbjct: 400 ILRSEIWELRDNDILPALMLSYNDLPAHLKRCFSFCAIFPKDYPFRKEQVIHLWIANGLV 459
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIERSNDDI---VKEKFVMHDVVYDLATIASGKSCC 123
+ + +++G+ YF EL SRSL E+ + ++E F+MHD+V DLA IAS K C
Sbjct: 460 P--VKDEINQDLGNQYFLELRSRSLFEKVPNPSKRNIEELFLMHDLVNDLAQIASSKLCI 517
Query: 124 RFGS--GGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSCKVID 181
R G + E HV+Y+ F K + + LR+ LPI + YLS +V+
Sbjct: 518 RLEERKGSFMLEKSWHVSYSMGRDGEFEKLTPLYKLEQLRTLLPIRIEFRSHYLSKRVLH 577
Query: 182 DLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYLQTLL 240
+++P+++ LR+LSLS+Y LPN + KL LR+L+LS T I LPD+ C LY L+TLL
Sbjct: 578 NILPTLRSLRVLSLSHYKNKELPNDLFIKLKLLRFLDLSCTWITKLPDSICGLYNLETLL 637
Query: 241 LSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL--TVFLVGKQKVGL 298
LS C+KL ELP+ + KLINLRHLD+S T+ KMP+ + RL++LQ L FLV VG
Sbjct: 638 LSSCYKLEELPLQMEKLINLRHLDVSNTRRLKMPLHLSRLKSLQVLVGAEFLV----VGW 693
Query: 299 SIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQQT--EESPTNE 356
+ LG+ NL G L + L+N ++ EA A ++ K H+E+L + W + + + S T
Sbjct: 694 RMEYLGEAQNLYGSLSVVKLENVVNRREAVKAKMREKNHVEQLSLEWSKSSIADNSQTER 753
Query: 357 VILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFL 416
IL+EL P N+K++ I Y G +FP+W+ D F +V LS+ C+ C +LP LGQ+P L
Sbjct: 754 DILDELHPHKNIKEVVISGYRGTNFPNWVADPLFVKLVKLSLSYCKDCYSLPALGQLPCL 813
Query: 417 KELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFPF 476
K L + GM + + EFYG S +PF LEKL+F M W++W + + F
Sbjct: 814 KFLSVKGMHGIRVVTEEFYGRL-----SSKKPFNCLEKLKFEDMTEWKQWHALGIGE--F 866
Query: 477 PRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQGSP 536
P L+ L +++C EL P S++++ ++ C
Sbjct: 867 PTLEKLSIKNCPELSLERPIQFSSLKRLEVVGC--------------------------- 899
Query: 537 ELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTSL 596
P + +F F + ++ L++ +S+T+FP + LPT+L
Sbjct: 900 ---------PVVFDDAQLFRF--------QLEAMKQIEALNISDCNSVTSFPFSILPTTL 942
Query: 597 QSLRIDECQNLAFLRP--ETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCSS 654
+ ++I C L F P E + Y + + CD + P + LSIE C +
Sbjct: 943 KRIQISGCPKLKFEVPVCEMFVEYLGV------SNCDCVDDMSPEFIPTARKLSIESCHN 996
Query: 655 LKSIFISEKNSSLSL----------------STLQSLKVSNCKSLRSLPQRMDTLFVLKS 698
+ I +L + + L SL +S C+ L+ LP+ M L +L S
Sbjct: 997 VTRFLIPTATETLCIFNCENVEKLSVACGGAAQLTSLNISACEKLKCLPENM--LELLPS 1054
Query: 699 LTLDKLSLCCEV-ACLPPKLQFMHIE-SLGLATPVTEWGFQSLCFLSDLHIGGDNIVN-- 754
L +L+ C E+ LP LQ + I L EW Q L L H G D +
Sbjct: 1055 LKELRLTNCPEIEGELPFNLQKLDIRYCKKLLNGRKEWHLQRLTELVIHHDGSDEDIEHW 1114
Query: 755 ------TLLKKKLLPPL----LVSLT----------------------ITNLTEMMRLKG 782
T L+ L L L SLT ++LT + L+
Sbjct: 1115 ELPCSITRLEVSNLITLSSQHLKSLTSLQFLRIVGNLSQIQSQGQLSSFSHLTSLQTLRI 1174
Query: 783 NRLQHI------STLKNLSFKCCSTLETCKD-FFPSFLKSLVFINCPKLMSLPD-MFPSS 834
LQ + S+L +L+ C L++ + PS L L NCP L SL + PSS
Sbjct: 1175 RNLQSLAESALPSSLSHLNIYNCPNLQSLSESALPSSLSHLTIYNCPNLQSLSESALPSS 1234
Query: 835 LETLEFDDCPRLGLLPRSGFPSSLKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKIN 893
L L +CP L L S PSSL L I CPLL+S + ++ +I+HIP ++I+
Sbjct: 1235 LSHLTIYNCPNLQSLSESALPSSLSKLWIFKCPLLRSLLEFVKGEYWPQIAHIPTIQID 1293
>gb|AAT39316.1| putative resistance complex protein I2C-2 [Solanum demissum]
Length = 1284
Score = 446 bits (1148), Expect = e-123
Identities = 329/936 (35%), Positives = 494/936 (52%), Gaps = 98/936 (10%)
Query: 7 ILNSDIWNIP--NDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEG 64
IL S+IW +P ++ I+P+L L+Y L HLK+CFA+C+I+PK + F+++++I LW+A G
Sbjct: 394 ILRSEIWELPRHSNGILPALMLSYNDLRPHLKQCFAFCAIYPKDHLFSKEQVIHLWIANG 453
Query: 65 FLE--HSMVGKAVEEVGDDYFNELLSRSLIE--RSNDDIVKEKFVMHDVVYDLATIASGK 120
++ HS + YF EL SRSL E R + + +F+MHD+V DLA IAS
Sbjct: 454 LVQQLHS---------ANQYFLELRSRSLFEKVRESSKWNQGEFLMHDLVNDLAQIASSN 504
Query: 121 SCCRF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSCK 178
C R G + E H++Y+ + D F K +T + LR+ LPI +L+ +LS +
Sbjct: 505 LCIRLEENQGSHMLEQTRHLSYSMGDGD-FGKLKTLNKLEQLRTLLPINIQLRWCHLSKR 563
Query: 179 VIDDLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYLQ 237
V+ D++P + LR LSLS+Y PN + KL LR+L+ S T+IK LPD+ C LY L+
Sbjct: 564 VLHDILPRLTSLRALSLSHYKNEEFPNDLFIKLKHLRFLDFSWTNIKNLPDSICVLYNLE 623
Query: 238 TLLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLT--VFLVGKQK 295
TLLLS C L+ELP+H+ KLINLRHLDIS + P+ + +L++L L FL+ +
Sbjct: 624 TLLLSYCSNLMELPLHMEKLINLRHLDISEAYLTT-PLHLSKLKSLDVLVGAKFLLSGRS 682
Query: 296 VGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWD-QQTEESPT 354
G + +LGK NL G L I LQ+ +D E+ AN++ K H+E L + W + S T
Sbjct: 683 -GSRMEDLGKLHNLYGSLSILGLQHVVDRRESLKANMREKKHVERLSLEWSGSNADNSQT 741
Query: 355 NEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVP 414
IL+ELQP+ N+K++ I Y G FP+WL D SF + +S++ C+ C +LP LGQ+P
Sbjct: 742 ERDILDELQPNTNIKEVEINGYRGTKFPNWLADHSFHKLTKVSLRYCKDCDSLPALGQLP 801
Query: 415 FLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKF 474
LK L I GM ++ + EFYG ++S +PF SLE+LEF MP W++W K
Sbjct: 802 CLKFLTIRGMHQITEVTEEFYG-----SSSFTKPFNSLEELEFGEMPEWKQWHVL--GKG 854
Query: 475 PFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQG 534
FP L+ L + DC +L G LP +L S+ ++ I C +L T LS++K ++
Sbjct: 855 EFPVLEELSIEDCPKLIGKLPENLSSLTRLRISKCPEL--SLETPIQLSNLKEFEV--AN 910
Query: 535 SPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPT 594
SP++ ++ +D+ L S + G +++ LD+ SLT+ P + LP+
Sbjct: 911 SPKVGVVFDDA--QLFTSQLEGMKQIVK-------------LDITDCKSLTSLPISILPS 955
Query: 595 SLQSLRIDECQNLAFLRP-----ETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSI 649
+L+ +RI C+ L P ++L +++ C++LT + + +SI
Sbjct: 956 TLKRIRISGCRELKLEAPINAICRVPEFLPRALSLSVRS-CNNLTRLLIP--TATETVSI 1012
Query: 650 EGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLFVLKSLTLDKLSLCCE 709
C +L+ + ++ + + SL + +C+ L+SLP+ M L L SL KL C +
Sbjct: 1013 RDCDNLEILSVA------CGTQMTSLHIYHCEKLKSLPEHMQQL--LPSLKELKLVNCSQ 1064
Query: 710 VAC-----LPPKLQFMHIESL-GLATPVTEWGFQSLCFLSDL---HIGGDNIVNTLLK-- 758
+ LP LQ + I L EW Q L L DL H G D +V K
Sbjct: 1065 IESFPEGGLPFNLQQLWISCCKKLVNGRKEWHLQRLPCLRDLTIHHDGSDEVVLADEKWE 1124
Query: 759 -------------KKLLPPLLVSLT------ITNLTEMMRLKGNRL-QHISTLKNLSFKC 798
K L LL SLT NL +M L L +S +K S
Sbjct: 1125 LPCSIRRLSIWNLKTLSSQLLKSLTSLEYLFANNLPQMQSLLEEGLPSSLSEVKLFSNHD 1184
Query: 799 CSTLETCKDFFPSFLKSLVFINCPKLMSLPDM-FPSSLETLEFDDCPRLGLLPRSGFPSS 857
+L T ++L+ L +C L SLP+ PSSL L +C + LP SG P S
Sbjct: 1185 LHSLPTEGLQRLTWLQRLEIRDCHSLQSLPESGLPSSLSELRIWNCSNVQSLPESGMPPS 1244
Query: 858 LKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKIN 893
+ L IS CPLLK + D+ KI+HIP + I+
Sbjct: 1245 ISNLYISKCPLLKPLLEFNKGDYWPKIAHIPTIYID 1280
>gb|AAU90299.1| putative disease resistance protein I2C-5 [Solanum demissum]
Length = 1266
Score = 436 bits (1120), Expect = e-120
Identities = 327/926 (35%), Positives = 476/926 (51%), Gaps = 102/926 (11%)
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
IL S+IW +P+++I+P+L L+Y LP HLKRCF+YC+IFPK YPF ++++I LW+A G +
Sbjct: 400 ILRSEIWELPDNDILPALMLSYNDLPVHLKRCFSYCAIFPKDYPFRKEQVIHLWIANGIV 459
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIER---SNDDIVKEKFVMHDVVYDLATIASGKSCC 123
+ +++ G+ YF EL SRSL E+ + ++E F+MHD+V DLA IAS K C
Sbjct: 460 PKD--DQIIQDSGNQYFLELRSRSLFEKVPNPSKRNIEELFLMHDLVNDLAQIASSKLCI 517
Query: 124 RF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIG-SRLQESY--LSCK 178
R G + E H++Y+ F K + + LR+ LP S + Y LS +
Sbjct: 518 RLEESKGSDMLEKSRHLSYSMGRGGDFEKLTPLYKLEQLRTLLPTCISTVNYCYHPLSKR 577
Query: 179 VIDDLIPSIKRLRMLSLSNYNITVLPNSI-NKLVQLRYLNLSHTDIKCLPDTTCDLYYLQ 237
V+ ++P ++ LR+LSLS+YNI LPN + KL LR+L++S T+IK LPD+ C LY L+
Sbjct: 578 VLHTILPRLRSLRVLSLSHYNIKELPNDLFIKLKLLRFLDISQTEIKRLPDSICVLYNLE 637
Query: 238 TLLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL--TVFLVGKQK 295
LLLS C L ELP+ + KLINL HLDIS T + KMP+ + +L++LQ L FL+
Sbjct: 638 ILLLSSCDYLEELPLQMEKLINLHHLDISNTHLLKMPLHLSKLKSLQVLVGAKFLLS--- 694
Query: 296 VGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQQT--EESP 353
G + +LG+ NL G L + LQN +D EA A ++ K H++ L + W + + + S
Sbjct: 695 -GWGMEDLGEAQNLYGSLSVVELQNVVDRREAVKAKMREKNHVDMLSLEWSESSSADNSQ 753
Query: 354 TNEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQV 413
T IL+EL P N+K++ I Y G FP+WL D F +V LS+ +C+ C +LP LGQ+
Sbjct: 754 TERDILDELSPHKNIKEVKITGYRGTKFPNWLADPLFLKLVQLSVVNCKNCSSLPSLGQL 813
Query: 414 PFLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSK 473
P LK L I GM + + EFYG + S +PF SL +L F MP W++W +
Sbjct: 814 PCLKFLSISGMHGITELSEEFYG-----SLSSKKPFNSLVELRFEDMPKWKQWHVLGSGE 868
Query: 474 FPFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWC----NHFPATLSTLHWLSSVKSLD 529
F L+ L++++C EL P L ++ ++ C S L + LD
Sbjct: 869 --FATLEKLLIKNCPELSLETPIQLSCLKMFEVIGCPKVFGDAQVFRSQLEGTKQIVELD 926
Query: 530 LM-CQG--SPELSLLGNDSPCHLQVSTIFGFNKL-LSLPNMFMSSTCLQHLDLIYISSLT 585
+ C S S+L P L+ TIFG KL L +P + L++L L +
Sbjct: 927 ISDCNSVTSFPFSIL----PTTLKTITIFGCQKLKLEVP---VGEMFLEYLSLKECDCID 979
Query: 586 AFPANGLPTSLQSLRIDECQNLA-FLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVL 644
LPT+ ++L + C NL FL P T+ +L + NC + +
Sbjct: 980 DISPELLPTA-RTLYVSNCHNLTRFLIP------TATESLYIHNCEN------------V 1020
Query: 645 QILSIEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLFVLKSLTLDKL 704
+ILS+ C + + SL + CK L+ LP+RM L L SL L
Sbjct: 1021 EILSVV-CGG---------------TQMTSLTIYMCKKLKWLPERMQEL--LPSLKHLYL 1062
Query: 705 SLCCEVAC-----LPPKLQFMHIESL-GLATPVTEWGFQSLCFLSDLHIGGDNIVNTLL- 757
C E+ LP LQF+ I + L EW Q L L+ L I D ++
Sbjct: 1063 INCPEIESFPEGGLPFNLQFLQIYNCKKLVNGRKEWRLQRLPCLNVLVIEHDGSDEEIVG 1122
Query: 758 -KKKLLPPLLVSLTITNLTEMMRLKGNRLQHISTLKNLSFKCC--------STLETCKDF 808
+ LP + LTI NL + Q + +L +L + C S LE +
Sbjct: 1123 GENWELPSSIQRLTIYNLKTLSS------QVLKSLTSLQYLCIEGNLPQIQSMLEQGQFS 1176
Query: 809 FPSFLKSLVFINCPKLMSLPD-MFPSSLETLEFDDCPRLGLLPRSGFPSSLKLLSISHCP 867
+ L+SL N P L SLP+ PSSL L CP+L LP G PSSL LSI CP
Sbjct: 1177 HLTSLQSLEIRNFPNLQSLPESALPSSLSQLTIVYCPKLQSLPVKGMPSSLSELSIYQCP 1236
Query: 868 LLKSRYATKRNDHLSKISHIPVVKIN 893
LL + ++ I+ IP + I+
Sbjct: 1237 LLSPLLEFDKGEYWPNIAQIPTIDID 1262
>gb|AAB63275.1| resistance complex protein I2C-2 [Lycopersicon esculentum]
gi|7489066|pir||T06404 resistance complex protein I2C-2 -
tomato
Length = 1240
Score = 431 bits (1108), Expect = e-119
Identities = 315/919 (34%), Positives = 479/919 (51%), Gaps = 105/919 (11%)
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
IL S+IW + +++I+P+L L+Y LP+HLKRCF++C+IFPK YPF ++++I LW+A G +
Sbjct: 391 ILRSEIWELRDNDILPALMLSYNDLPAHLKRCFSFCAIFPKDYPFRKEQVIHLWIANGLV 450
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIER---SNDDIVKEKFVMHDVVYDLATIASGKSCC 123
+ + ++++G+ +F EL SRSL ER ++ +KE F+MHD+V DLA +AS K C
Sbjct: 451 --PVEDEIIQDLGNQFFLELSSRSLFERVPNPSEGNIKELFLMHDLVNDLAQLASSKLCI 508
Query: 124 RF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESY--LSCKV 179
R G + E H++Y+ F K + + LR+ LP S + Y L+ +V
Sbjct: 509 RLEESQGSHMLEQCRHLSYSMGYDGGFEKLTPLYKLEQLRTLLPTCSSVNYFYNPLTKRV 568
Query: 180 IDDLIPSIKRLRMLSLSNYNITVLPNSI-NKLVQLRYLNLSHTDIKCLPDTTCDLYYLQT 238
+ +++P+++ LR LSLS+Y + LPN + KL LR+L++S T+IK LPD+ C LY L+T
Sbjct: 569 LHNILPTLRSLRALSLSHYKMEELPNDLFIKLKLLRFLDISRTNIKRLPDSICVLYNLET 628
Query: 239 LLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL--TVFLVGKQKV 296
LLLS C KL ELP+ + KLINLRHLDIS T KMP+ + RL++LQ L FLVG +
Sbjct: 629 LLLSSC-KLEELPLQMEKLINLRHLDISNTWHLKMPLHLSRLKSLQVLVGAKFLVGVWR- 686
Query: 297 GLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQ--QTEESPT 354
+ +LG+ NL G L + L+N +D EA ++ K H+E+L + W + + S T
Sbjct: 687 ---MEDLGEAQNLYGSLSVVKLENVVDRREAVKPKMREKNHVEQLSLEWSESISADNSQT 743
Query: 355 NEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVP 414
IL+EL+P N++++ I Y G +FP+W+ D F +V LS+++C+ C +LP LGQ+P
Sbjct: 744 ERDILDELRPHKNIQEVKIIGYRGTNFPNWVADPLFLKLVKLSLRNCKDCYSLPALGQLP 803
Query: 415 FLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKF 474
LK L + GM + + EFYG S +PF LEKLEF M W++W + +
Sbjct: 804 CLKFLSVKGMHGIRVVTEEFYGRL-----SSKKPFNCLEKLEFEDMTEWKQWHALGIGE- 857
Query: 475 PFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWC-------NHFPATLSTLHWLSSVKS 527
FP L+ L + +C EL +P S+++ + C + L + + +
Sbjct: 858 -FPTLEKLSIINCPELSLEIPIQFSSLKRFRVFGCPVVFYDAQVLRSQLEGMKQIEEIYI 916
Query: 528 LDLMCQGSPELSLLGNDSPCHLQVSTIFGFNKL-LSLPNMFMSSTCLQHLDLIYISSLTA 586
D S S+L P L+ I G KL L P MS L+ +
Sbjct: 917 RDCNSVTSFPFSIL----PTTLKTIDISGCPKLKLEAPVCEMS----MFLEEFSVEECGC 968
Query: 587 FPANGLPTSLQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQI 646
LPT+ + LRI C N+ FL P T+ TL ++N C+++
Sbjct: 969 VSPEFLPTA-RELRIGNCHNVRFLIP------TATETLHIRN-CENVEKLS--------- 1011
Query: 647 LSIEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLFVLKSLTLDKLSL 706
++ G + L S L +S CK L+ LP+ +L SL +L+
Sbjct: 1012 MACGGAAQLTS-----------------LDISGCKKLKCLPE------LLPSLKELQLTN 1048
Query: 707 CCEV-ACLPPKLQFMHI-ESLGLATPVTEWGFQSLCFLSDLHIGGDNIVNTLLKKKLLPP 764
C E+ LP LQ ++I + L EW Q L L H G D + + LP
Sbjct: 1049 CPEIEGELPFNLQKLYIRDCKKLVNGRKEWHLQRLTKLVIYHDGSDEDI----EHWELPC 1104
Query: 765 LLVSLTITNLTEMMRLKGNRLQHISTLKNLSFKC----CSTLETCKDFFPSF-----LKS 815
+ L + NL + QH+ +L +L + C S +++ + SF L++
Sbjct: 1105 SITRLEVFNLITLSS------QHLKSLTSLQYLCIDGNLSPIQS-QGQISSFSHLTSLQT 1157
Query: 816 LVFINCPKLMSLPD-MFPSSLETLEFDDCPRLGLLPRSGFPSSLKLLSISHCPLLKSRYA 874
L N L SL + PSSL LE CP L LP +G PSSL L IS CPLL
Sbjct: 1158 LQIWNFHNLQSLSESALPSSLSQLEIFHCPNLQSLPLNGMPSSLSKLLISGCPLLTPLLE 1217
Query: 875 TKRNDHLSKISHIPVVKIN 893
+ ++ +I+HIP + I+
Sbjct: 1218 FDKGEYWPQIAHIPTILID 1236
>gb|AAT40545.1| putative plant disease resistant protein [Solanum demissum]
Length = 1315
Score = 429 bits (1104), Expect = e-118
Identities = 330/954 (34%), Positives = 488/954 (50%), Gaps = 110/954 (11%)
Query: 7 ILNSDIWNIPN--DNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEG 64
IL S+IW +P+ + I+P+L L+Y LP+HLKRCFA+C+I+PK Y F ++++I LW+A G
Sbjct: 383 ILRSEIWELPSHSNGILPALMLSYNDLPAHLKRCFAFCAIYPKDYMFCKEQVIHLWIANG 442
Query: 65 FLEHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIV--KEKFVMHDVVYDLATIASGKSC 122
+ + G+ YF EL SRSL ER + E+F+MHD+V DLA IAS C
Sbjct: 443 LVPQL-------DSGNQYFLELRSRSLFERIPESSKWNSEEFLMHDLVNDLAQIASSNLC 495
Query: 123 CRF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYL---SC 177
R G + E H++Y+ E D F K + F + LR+ LPI +Q YL S
Sbjct: 496 IRLEENQGSHMLEQSRHISYSTGEGD-FEKLKPLFKSEQLRTLLPIS--IQRDYLFKLSK 552
Query: 178 KVIDDLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYL 236
+V+ +++P + LR LSLS Y I LPN + KL LR+L++S T IK LPD+ C LY L
Sbjct: 553 RVLHNVLPRLTSLRALSLSPYKIVELPNDLFIKLKLLRFLDISRTKIKKLPDSICVLYNL 612
Query: 237 QTLLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLT--VFLVGKQ 294
+ LLLS C L ELP+ + KLINL +LDI+ T KMP+ + +L++L L FL+G +
Sbjct: 613 EILLLSSCDDLEELPLQMEKLINLHYLDINNTSRLKMPLHLSKLKSLHVLVGAKFLLGGR 672
Query: 295 KVGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQQTEESPT 354
G + +LG+ NL G L I LQN +D EA AN+K K H+E L + W + ++
Sbjct: 673 G-GSRMDDLGEVHNLFGSLSILELQNVVDRWEALKANMKEKNHVEMLSLEWSRSIADNSK 731
Query: 355 NEV-ILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQV 413
NE IL+ LQP+ N+ +L I Y G FP+WL D SF +V LS+ +C+ C +LP LGQ+
Sbjct: 732 NEKDILDGLQPNTNINELQIGGYRGTKFPNWLADQSFLKLVQLSLSNCKDCDSLPALGQL 791
Query: 414 PFLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSK 473
P LK L I M R+ + EFYG + S +PF SLEKLEF MP W+ W +
Sbjct: 792 PSLKFLAIRRMRRIIEVTEEFYG-----SLSSKKPFNSLEKLEFAEMPEWKRWHVLGNGE 846
Query: 474 FPFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQ 533
FP LK L + DC +L P +L S+ + I C +L T LS++K +++
Sbjct: 847 --FPALKILSVEDCPKLIEKFPENLSSLTGLRISKCPEL--SLETSIQLSTLKIFEVI-- 900
Query: 534 GSPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLP 593
SP++ +L +D+ L S + ++ + +F + +SLT+ P + LP
Sbjct: 901 SSPKVGVLFDDT--ELFTSQL---QEMKHIVELFFTD----------CNSLTSLPISILP 945
Query: 594 TSLQSLRIDECQNLAFLRP--ETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEG 651
++L+ + I +C+ L P E +N L L+L CDS+ P + L +
Sbjct: 946 STLKRIHIYQCEKLKLKTPVGEMITNNMFLEELKLDG-CDSIDDISPELVPRVGTLIVGR 1004
Query: 652 CSSLKSIFISEKNSSLS-----------------LSTLQSLKVSNCKSLRSLPQRMDTLF 694
C SL + I + SL+ + +L+ L + NC+ L+ LP+ M L
Sbjct: 1005 CHSLTRLLIPTETKSLTIWSCENLEILSVACGARMMSLRFLNIENCEKLKWLPECMQEL- 1063
Query: 695 VLKSLTLDKLSLCCEVAC-----LPPKLQFMHI-ESLGLATPVTEWGFQSLCFLSDLHIG 748
L SL +L C E+ LP LQ + I L W Q L L +L I
Sbjct: 1064 -LPSLNTLELFNCPEMMSFPEGGLPFNLQVLLIWNCKKLVNGRKNWRLQRLPCLRELRIE 1122
Query: 749 GDNIVNTLL--KKKLLPPLLVSLTITNLTEMMRLKGNRLQHISTLKNLSFKCCSTLET-C 805
D +L + LP + L I+NL L L+ +++L L +++
Sbjct: 1123 HDGSDEEILAGENWELPCSIQRLYISNL---KTLSSQVLKSLTSLAYLDTYYLPQIQSLL 1179
Query: 806 KDFFPS-------------------------FLKSLVFINCPKLMSLPD-MFPSSLETLE 839
++ PS L+ L +C +L SL + PSS+ L
Sbjct: 1180 EEGLPSSLYELRLDDHHELHSLPTKGLRHLTSLRRLEIRHCNQLQSLAESTLPSSVSELT 1239
Query: 840 FDDCPRLGLLPRSGFPSSLKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKIN 893
CP L LP G PSSL L I +CPLL+ + ++ KI+HI ++I+
Sbjct: 1240 IGYCPNLQSLPVKGMPSSLSKLHIYNCPLLEPLLECDKGEYWQKITHISTIEID 1293
>gb|AAU90295.1| putative disease resistance protein I2 [Solanum demissum]
Length = 1190
Score = 429 bits (1102), Expect = e-118
Identities = 299/898 (33%), Positives = 458/898 (50%), Gaps = 124/898 (13%)
Query: 7 ILNSDIWNIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEGFL 66
IL S+IW +P+++I+P+L L+Y LP+HLKRCF+YC+IFPK Y F ++++I LW+A G +
Sbjct: 400 ILRSEIWELPHNDIVPALMLSYNDLPAHLKRCFSYCAIFPKDYSFRKEQVIHLWIANGLV 459
Query: 67 EHSMVGKAVEEVGDDYFNELLSRSLIER-SNDDI--VKEKFVMHDVVYDLATIASGKSCC 123
+ + +E+ G+ YF EL SRSL E+ N + ++E F+MHD++ DLA IAS K C
Sbjct: 460 QKE--DEIIEDSGNQYFLELRSRSLFEKVPNPSVGNIEELFLMHDLINDLAQIASSKLCI 517
Query: 124 RF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSCKVID 181
R G + E H++Y+ E F K T + + LR+ LPI + LS +V+
Sbjct: 518 RLEESQGSHMLEKSRHLSYSMGEGGEFEKLTTLYKLEQLRTLLPIYIDVNYYSLSKRVLY 577
Query: 182 DLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYLQTLL 240
+++P ++ LR+LSLS YNI LPN + +L LR+L++S T IK LPD+ C LY L+TLL
Sbjct: 578 NILPRLRSLRVLSLSYYNIKELPNDLFIELKLLRFLDISRTKIKRLPDSICVLYNLETLL 637
Query: 241 LSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLT--VFLVGKQKVGL 298
LS C L ELP+ + KLINLRHLDIS T + KMP+ + +L++LQ L FL+ G
Sbjct: 638 LSSCADLEELPLQMEKLINLRHLDISNTSLLKMPLHLSKLKSLQVLVGAKFLLS----GW 693
Query: 299 SIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQQT--EESPTNE 356
+ +LG+ NL G + + L+N +D EA A ++ K H+++L + W + + + S T
Sbjct: 694 RMEDLGEAQNLYGSVSVVELENVVDRREAVKAKMREKNHVDKLSLEWSESSSADNSQTER 753
Query: 357 VILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFL 416
IL+EL+P N+K++ I Y G FP+WL D F +V LSI +C+ C TLP LGQ+P L
Sbjct: 754 DILDELRPHKNIKEVEITGYRGTKFPNWLADPLFLKLVQLSIDNCKDCYTLPALGQLPCL 813
Query: 417 KELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFPF 476
K L I GM + + EFYG + S +PF LEKL F MP W++W +FP
Sbjct: 814 KFLSISGMHGITEVTEEFYG-----SFSSKKPFNCLEKLAFEDMPEWKQWHVLGSGEFPI 868
Query: 477 PRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQGSP 536
L+ L +++C EL +L T LSS+KS ++ G P
Sbjct: 869 --LEKLFIKNCPEL------------------------SLETPIQLSSLKSFEV--SGCP 900
Query: 537 ELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTSL 596
++ ++ +D+ L S + G +++ L + Y +S+T P + LPT+L
Sbjct: 901 KVGVVFDDA--QLFRSQLEGMKQIVELY-------------ISYCNSVTFLPFSILPTTL 945
Query: 597 QSLRIDECQNLAFLRPE-TWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCSSL 655
+ + I C+ L P S + + +E +C D ++ L P + L + C +L
Sbjct: 946 KRIEISRCRKLKLEAPVGEMSMFLEELRVEGSDCIDVISPELL---PRARNLRVVSCHNL 1002
Query: 656 KSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLFVLKSLTLDKLSLCCEVACLPP 715
+ I + L + +C+++ L ++ SLT+ C ++ CLP
Sbjct: 1003 TRVLIPTATAFLC--------IWDCENVEKLSVACGGT-LMTSLTI---GCCSKLKCLPE 1050
Query: 716 KLQFMHIESLGLATPVTEWGFQSLCFLSDLHIGGDNIVNTLLKKKLLPPLLVSLTITNLT 775
++Q L + E + + GG L +L I ++
Sbjct: 1051 RMQ-------ELLPSLKELDLRKCPEIESFPQGG---------------LPFNLQILEIS 1088
Query: 776 EMMRLKGNRLQHISTLKNLSFKCCSTLETCKDFFPSFLKSLVFINCPKLMSLPDM-FPSS 834
E +L R K++ L L CP L SL + PSS
Sbjct: 1089 ECKKLVNGR---------------------KEWRLQRLSQLAIYGCPNLQSLSESALPSS 1127
Query: 835 LETLEFDDCPRLGLLPRSGFPSSLKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKI 892
L L CP L LP G PSSL L IS CPLL + + ++ I+ P + I
Sbjct: 1128 LSKLTIIGCPNLQSLPVKGMPSSLSELHISECPLLTALLEFDKGEYWPNIAQFPTIDI 1185
>gb|AAS49215.1| disease resistance protein [Glycine max]
Length = 1189
Score = 428 bits (1101), Expect = e-118
Identities = 312/902 (34%), Positives = 453/902 (49%), Gaps = 122/902 (13%)
Query: 7 ILNSDIWNIPNDN--IMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEG 64
+L S IW + + I+P+L L+Y HLPSHLKRCFAYC++FPK + F + LI LW+AE
Sbjct: 391 VLKSKIWELTKEESKIIPALLLSYYHLPSHLKRCFAYCALFPKDHEFYKDSLIQLWVAEN 450
Query: 65 FLEHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVKEKFVMHDVVYDLATIASGKSCCR 124
F++ S + EE+G+ YFN+LLSRS +RS+ +++ F MHD++ DLA G C R
Sbjct: 451 FVQCSQQSNSQEEIGEQYFNDLLSRSFFQRSS---IEKCFFMHDLLNDLAKYVCGDICFR 507
Query: 125 FGSGGRIS-EDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQ-ESYLSCKVIDD 182
S V H ++ E F+ + + + + LR+F+P+ L ++ K++D+
Sbjct: 508 LEVDKPKSISKVRHFSFVTEIDQYFDGYGSLYHAQRLRTFMPMTRPLLLTNWGGRKLVDE 567
Query: 183 LIPSIKRLRMLSLSNYNITVLPNSINKLVQLRYLNLSHTDIKCLPDTTCDLYYLQTLLLS 242
L K LR+LSL ++ +P+S+ L LR L+LS+T IK LPD+ C L LQ L L+
Sbjct: 568 LCSKFKFLRILSLFRCDLKEMPDSVGNLNHLRSLDLSYTFIKKLPDSMCFLCNLQVLKLN 627
Query: 243 GCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLTVFLVGKQKVGLSIRE 302
C L ELP ++ KL NLR L+ TK++KMPM + +L+NLQ L+ F VGK SI++
Sbjct: 628 YCVHLEELPSNLHKLTNLRCLEFMCTKVRKMPMHMGKLKNLQVLSPFYVGKGIDNCSIQQ 687
Query: 303 LGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYW--DQQTEESPTNEVILN 360
LG+ NL G L I+ LQN ++ +A A+LK+K HL +L + W D+ ++S +L
Sbjct: 688 LGEL-NLHGSLSIEELQNIVNPLDALAABLKNKTHLLDLRLEWNEDRNLDDSIKERQVLE 746
Query: 361 ELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFLKELK 420
LQPS +L+KLSI+ YGG FPSWL D S N+V L++ +C+Y + LPPLG +P LKEL
Sbjct: 747 NLQPSRHLEKLSIRNYGGTQFPSWLSDNSLCNVVSLTLMNCKYFLCLPPLGLLPILKELS 806
Query: 421 IDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFPFPRLK 480
I+G+ + +I +F+G + S F SLE L+F+ M W EW +G FPRL+
Sbjct: 807 IEGLDGIVSINADFFGSSSCS-------FTSLESLKFSDMKEWEEW-ECKGVTGAFPRLQ 858
Query: 481 TLMLRDCTELRGHLPSHLPSIEKITILWCNHF-PATLSTLHWLSSVKSLDLMCQGSPELS 539
L ++ C +L+GHLP L + + I C P+ LS + L L G ++
Sbjct: 859 RLSIKRCPKLKGHLPEQLCHLNGLKISGCEQLVPSALSA----PDIHQLYLGDCGKLQI- 913
Query: 540 LLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTSLQSL 599
D P L+ TI G N M + L+ + Y S P +
Sbjct: 914 ----DHPTTLKELTITGHN---------MEAALLEQIGRNYSCSNKNIPMH--------- 951
Query: 600 RIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCSSLKSIF 659
S Y LV L + CDSLT+ L+ FP L+ L I C +L+ I
Sbjct: 952 ----------------SCYDFLVWLLINGGCDSLTTIHLDIFPKLKELYICQCPNLQRIS 995
Query: 660 ISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLFVLKSLTLDKLSLCCEVACLPP---- 715
+ ++ LQ L + C L SLP+ M L L SL + C +V P
Sbjct: 996 QGQAHNH-----LQDLSMRECPQLESLPEGMHVL--LPSLDSLWIIHCPKVEMFPEGGLP 1048
Query: 716 ---KLQFMHIESLGLATPVTEWGFQSLCFLSDLHIGGDNIVNTLLKKKLLPPLLVSLTIT 772
K+ +H S L + + L L IGG + V L + +LP LV+L I
Sbjct: 1049 SNLKVMSLHGGSYKLIY-LLKSALGGNHSLESLSIGGVD-VECLPDEGVLPHSLVTLMIN 1106
Query: 773 NLTEMMRLKGNRLQHISTLKNLSFKCCSTLETCKDFFPSFLKSLVFINCPKLMSLPDMFP 832
++ RL L H+S+LK LS CP+L LP+
Sbjct: 1107 KCGDLKRLDYKGLCHLSSLKRLS----------------------LWECPRLQCLPE--- 1141
Query: 833 SSLETLEFDDCPRLGLLPRSGFPSSLKLLSISHCPLLKSRYATKRNDHLSKISHIPVVKI 892
G P S+ L I +CPLLK R + KI+HI V +
Sbjct: 1142 -------------------EGLPKSISTLRILNCPLLKQRCREPEGEDWPKIAHIKRVWL 1182
Query: 893 NG 894
G
Sbjct: 1183 LG 1184
>gb|AAT40525.1| putative disease resistance complex protein I2C-1, 5'-partial
[Solanum demissum]
Length = 1097
Score = 426 bits (1094), Expect = e-117
Identities = 336/989 (33%), Positives = 484/989 (47%), Gaps = 157/989 (15%)
Query: 7 ILNSDIWNIPN--DNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEG 64
IL S+IW + + + I+P+L L+Y L LKRCFA+C+I+PK Y F ++++I LW+A G
Sbjct: 162 ILRSEIWELQSCSNGILPALMLSYNDLHPQLKRCFAFCAIYPKDYLFCKEQVIHLWIANG 221
Query: 65 FLE--HSMVGKAVEEVGDDYFNELLSRSLIER--SNDDIVKEKFVMHDVVYDLATIASGK 120
++ HS + YF EL SRSL E+ + + +F+MHD+V DLA IAS
Sbjct: 222 LVQQLHS---------ANHYFLELRSRSLFEKVQESSEWNPGEFLMHDLVNDLAQIASSN 272
Query: 121 SCCRFGS--GGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSCK 178
C R G + E H++Y+ D F K + + + LR+ LPI + LS +
Sbjct: 273 LCIRLEENLGSHMLEQSRHISYSMG-LDDFKKLKPLYKLEQLRTLLPINIQQHSYCLSKR 331
Query: 179 VIDDLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYLQ 237
++ D++P + LR LSLS+Y+I LPN + KL LR+L+ S T IK LPD+ C LY L+
Sbjct: 332 ILHDILPRLTSLRALSLSHYSIEELPNDLFIKLKYLRFLDFSWTKIKKLPDSICLLYNLE 391
Query: 238 TLLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLT-VFLVGKQKV 296
TLLLS C L ELP+H+ KLINLRHLDIS + P+ + +L++L L L+ +
Sbjct: 392 TLLLSHCSYLKELPLHMEKLINLRHLDISEAYL-TTPLHLSKLKSLHALVGANLILSGRG 450
Query: 297 GLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWD-QQTEESPTN 355
GL + +LG+ NL G L I LQN +D E+ AN++ K H+E L + W + S T
Sbjct: 451 GLRMEDLGEVHNLYGSLSILELQNVVDRRESLKANMREKKHVERLSLEWSGSNADNSQTE 510
Query: 356 EVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPF 415
IL+ELQP+ N+K++ I Y G FP+ D SF ++ +S+ C+ C +LP LGQ+P
Sbjct: 511 REILDELQPNTNIKEVQIIRYRGTKFPT---DHSFHKLIEMSLSYCKDCDSLPALGQLPC 567
Query: 416 LKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFP 475
LK L I GM ++ + EFYG + S +PF SLEKLEF MP W++W K
Sbjct: 568 LKFLTIRGMHQITEVSEEFYG-----SLSSTKPFKSLEKLEFAGMPEWKQWHVL--GKGE 620
Query: 476 FPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQGS 535
FP L+ L + C +L G LP +L S+ ++ I C +L T LS++K +L
Sbjct: 621 FPILEELWINGCPKLIGKLPENLSSLRRLRISECPEL--SLETPIQLSNLK--ELKVADC 676
Query: 536 PELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTS 595
P++ +L ++ L S + G +++ L + + C SLT+ P ++
Sbjct: 677 PKVGVLFANA--QLFTSQLEGMKQIVKL----VITDC---------KSLTSLPI----ST 717
Query: 596 LQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCSSL 655
L+S I C L+ E N L L LK C S +L FP + LS+ C++L
Sbjct: 718 LKSREISGCGE---LKLEASMNAMFLEDLSLKGC----DSPEL--FPRARNLSVRSCNNL 768
Query: 656 KSIFISEKNSSLSLS--------------TLQSLKVSNCKSLRSLPQRMDTLF-VLKSLT 700
+ I + +LS + SL + NC+ L+SLP+ M L LK LT
Sbjct: 769 TRLLIPTETETLSFGDCDNLEILSVACGIQMTSLNIHNCQKLKSLPEHMQELLPSLKELT 828
Query: 701 LDKLSLCCEVAC-----LPPKLQFMHIESL-GLATPVTEWGFQSLCFLSDLHIGGD---- 750
LD C E+ LP LQF+ I L EW Q L L L I D
Sbjct: 829 LDN---CPEIESFPQGGLPFNLQFLWISRCKKLVNGRKEWHLQRLPSLMQLEISHDGSDI 885
Query: 751 --------------------NIVNTLLK--------------------KKLLPPLLVSLT 770
+ + LLK ++ LP L L
Sbjct: 886 AGENWELPCSIRRLTIANLKTLSSQLLKSLTSLEYLYAINLPQIQSLLEEELPSSLSELH 945
Query: 771 ITNLTEMMRLKGNRLQHISTLKNLSFKCCSTLETCKDF-FPSFLKSLVFINCPKLMSLPD 829
+ ++ L LQ + + L C L++ + PS L L +C L SLP+
Sbjct: 946 LHQHHDLHSLPTEGLQRLMWFRCLEIWDCPNLQSLPESGMPSSLSKLTIQHCSNLQSLPE 1005
Query: 830 M-FPSSLETLEFDDCPRLGLLPRSGFPSSLKLLS-----------------------ISH 865
PSSL L +CP L LP SGFPSSL L IS
Sbjct: 1006 SGMPSSLSDLTISNCPSLQSLPESGFPSSLSELGIWNCSNLQSLPESGMPPSICNLYISE 1065
Query: 866 CPLLKSRYATKRNDHLSKISHIPVVKING 894
CPLLK + D+ KI+HIP + I+G
Sbjct: 1066 CPLLKPLLEFNKGDYWPKIAHIPTIYIDG 1094
>gb|AAU90287.1| putative disease resistance protein I2C-5 [Solanum demissum]
Length = 1255
Score = 424 bits (1089), Expect = e-117
Identities = 318/927 (34%), Positives = 469/927 (50%), Gaps = 111/927 (11%)
Query: 7 ILNSDIW--NIPNDNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEG 64
IL S+IW +I ++ I+P+L L+Y LP+ LK+CFAYC+I+PK Y F + ++I LW+A G
Sbjct: 396 ILRSEIWELSICSNGILPALMLSYNDLPARLKQCFAYCAIYPKDYQFCKDQVIHLWIANG 455
Query: 65 FLE--HSMVGKAVEEVGDDYFNELLSRSLIERSND--DIVKEKFVMHDVVYDLATIASGK 120
++ HS G+ YF EL SRSL E ++ + EKF+MHD+V DLA IAS
Sbjct: 456 LVQQFHS---------GNQYFLELRSRSLFEMVSESSESNSEKFLMHDLVNDLAQIASSN 506
Query: 121 SCCRF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQ--ESYLS 176
C R G + E H++Y E F K ++ F + +R+ LPI +L LS
Sbjct: 507 LCIRLEENKGLHMLEQCRHMSYLIGEDGDFEKLKSLFKSEQVRTLLPINIQLYYYNIQLS 566
Query: 177 CKVIDDLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYY 235
+V+ +++P + LR LSL Y I LPN + KL LRYL++S T IK LPD+ C LY
Sbjct: 567 RRVLHNILPRLTSLRALSLLGYKIVELPNDLFIKLKLLRYLDISQTKIKRLPDSICVLYN 626
Query: 236 LQTLLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL--TVFLVGK 293
L+TLLLS C L ELP+ + KLINLRHLDIS T++ KMP+ + +L++LQ L FL+G
Sbjct: 627 LETLLLSSCDCLEELPLQMEKLINLRHLDISNTRLLKMPLHLSKLKSLQVLLGAKFLLG- 685
Query: 294 QKVGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQQT--EE 351
GLS+ +LG+ NL G L + LQN +D EA A ++ K H+++L + W + + +
Sbjct: 686 ---GLSMEDLGEAQNLYGSLSVVELQNVVDRREAVKAKMREKNHVDKLSLEWSESSSADN 742
Query: 352 SPTNEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLG 411
S T IL+EL+P N+K++ I Y G +FP+WL D F + LSI +C+ C +LP LG
Sbjct: 743 SQTERDILDELRPHKNIKEVKIIGYRGTTFPNWLADPLFLKLEQLSIDNCKNCFSLPALG 802
Query: 412 QVPFLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRG 471
Q+P LK L I GM + + EFYG +PF LEKLEF MP W++W
Sbjct: 803 QLPCLKILSIRGMHGITEVTEEFYGSLSSK-----KPFNCLEKLEFVDMPVWKQWHVLGS 857
Query: 472 SKFPFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLM 531
FP L+ L +++C EL P L S+++ ++
Sbjct: 858 GDFPI--LEKLFIKNCPELSLETPIQLSSLKRFQVV------------------------ 891
Query: 532 CQGSPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANG 591
GS ++ ++ +D+ L S + G ++ + L++ +S+ +FP +
Sbjct: 892 --GSSKVGVVFDDA--QLFRSQLEGMKQI-------------EALNISDCNSVISFPYSI 934
Query: 592 LPTSLQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEG 651
LPT+L+ + I CQ L L P L L LK C D + P + L +E
Sbjct: 935 LPTTLKRITISRCQKLK-LDPPVGEMSMFLEYLSLKEC-DCIDDISPELLPRARELWVEN 992
Query: 652 CSSLKSIFISEKNSSLSLSTLQSLKVS---------------NCKSLRSLPQRMDTLFVL 696
C +L I L++ ++L++ C+ L+ LP+RM L L
Sbjct: 993 CHNLTRFLIPTATERLNIQNCENLEILLVASEGTQMTYLNIWGCRKLKWLPERMQEL--L 1050
Query: 697 KSLTLDKLSLCCEVAC-----LPPKLQFMHIESLG-LATPVTEWGFQSLCFLSDLHIGGD 750
SL +L C E+ LP LQ + I + L EW Q L L++L I D
Sbjct: 1051 PSLKELRLFNCPEIESFPQGGLPFNLQALWIRNCKKLVNGQKEWHLQRLPCLTELWISHD 1110
Query: 751 NIVNTLL--KKKLLPPLLVSLTITNLTEMMRLKGNRLQHISTLKNLSF-KCCSTLETCKD 807
++ + LP + L I N+ + QH+ +L +L + S LE +
Sbjct: 1111 GSDEEIVGGENWELPSSIQRLRINNVKTLSS------QHLKSLTSLQYLDIPSMLEQGRF 1164
Query: 808 FFPSFLKSLVFINCPKLMSLPDM-FPSSLETLEFDDCPRLGLLPRSGFPSSLKLLSISHC 866
S L SL SL + PSSL L CP+L LP G PSSL L I C
Sbjct: 1165 SSFSQLTSLQSQLIGNFQSLSESALPSSLSQLTIIYCPKLQSLPVKGMPSSLSKLVIYKC 1224
Query: 867 PLLKSRYATKRNDHLSKISHIPVVKIN 893
PLL + ++ I+HI ++I+
Sbjct: 1225 PLLSPLLEFDKGEYWPNIAHISTIEID 1251
>gb|AAB63274.1| resistance complex protein I2C-1 [Lycopersicon esculentum]
gi|7489065|pir||T06403 resistance complex protein I2C-1 -
tomato
Length = 1220
Score = 422 bits (1084), Expect = e-116
Identities = 320/912 (35%), Positives = 477/912 (52%), Gaps = 124/912 (13%)
Query: 7 ILNSDIWNIPN--DNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEG 64
IL S+IW +P+ + I+P+L L+Y LP+HLK+CFAYC+I+PK Y F ++++I LW+A G
Sbjct: 404 ILRSEIWELPSCSNGILPALMLSYNDLPAHLKQCFAYCAIYPKDYQFRKEQVIHLWIANG 463
Query: 65 FLE--HSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVK--EKFVMHDVVYDLATIASGK 120
+ HS G+ YF EL SRSL E +++ + E+F+MHD+V DLA IAS
Sbjct: 464 LVHQFHS---------GNQYFIELRSRSLFEMASEPSERDVEEFLMHDLVNDLAQIASSN 514
Query: 121 SCCRF--GSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESY-LSC 177
C R G + E H++Y+ + F K ++ F + LR+ LPI + S LS
Sbjct: 515 HCIRLEDNKGSHMLEQCRHMSYSIGQDGEFEKLKSLFKSEQLRTLLPIDIQFHYSKKLSK 574
Query: 178 KVIDDLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYL 236
+V+ +++P+++ LR LSLS+Y I VLPN + KL LR+L+LS T I LPD+ LY L
Sbjct: 575 RVLHNILPTLRSLRALSLSHYQIEVLPNDLFIKLKLLRFLDLSETSITKLPDSIFVLYNL 634
Query: 237 QTLLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL--TVFLVGKQ 294
+TLLLS C L ELP+ + KLINLRHLDIS T+ KMP+ + RL++LQ L FLVG
Sbjct: 635 ETLLLSSCEYLEELPLQMEKLINLRHLDISNTRRLKMPLHLSRLKSLQVLVGAKFLVG-- 692
Query: 295 KVGLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQ--QTEES 352
G + LG+ NL G L I L+N +D EA A ++ K H+E+L + W + + S
Sbjct: 693 --GWRMEYLGEAHNLYGSLSILELENVVDRREAVKAKMREKNHVEQLSLEWSESISADNS 750
Query: 353 PTNEVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQ 412
T IL+EL+P N+K + I Y G +FP+W+ D F +V+L +++C+ C +LP LGQ
Sbjct: 751 QTERDILDELRPHKNIKAVEITGYRGTNFPNWVADPLFVKLVHLYLRNCKDCYSLPALGQ 810
Query: 413 VPFLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGS 472
+P L+ L I GM + + EFYG S +PF SL KL F MP W++W +
Sbjct: 811 LPCLEFLSIRGMHGIRVVTEEFYGRL-----SSKKPFNSLVKLRFEDMPEWKQWHTLGIG 865
Query: 473 KFPFPRLKTLMLRDCTELRGHLPSHLPSIEKITILWC---NHFPATLSTLHWLSSVKSLD 529
+ FP L+ L +++C EL +P S++++ I C FP ++ +++K +
Sbjct: 866 E--FPTLEKLSIKNCPELSLEIPIQFSSLKRLDICDCKSVTSFPFSILP----TTLKRIK 919
Query: 530 LMCQGSPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPA 589
+ G P+L L ++P V +F +++L +I +
Sbjct: 920 I--SGCPKLKL---EAP----VGEMF-----------------VEYLSVIDCGCVDDISP 953
Query: 590 NGLPTSLQSLRIDECQNLA-FLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILS 648
LPT+ Q L I+ C N+ FL P T+ +L ++N C+ L+ ++
Sbjct: 954 EFLPTARQ-LSIENCHNVTRFLIP------TATESLHIRN-CEKLS------------MA 993
Query: 649 IEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLFVLKSLTLDKLSLCC 708
G + L S L + CK L+ LP+ +L SL +L+ C
Sbjct: 994 CGGAAQLTS-----------------LNIWGCKKLKCLPE------LLPSLKELRLTYCP 1030
Query: 709 EV-ACLPPKLQFMHIE-SLGLATPVTEWGFQSLCFLSDLHIGGDNIVNTLLKKKLLPPLL 766
E+ LP LQ + I L EW Q L L H G D + + LP +
Sbjct: 1031 EIEGELPFNLQILDIRYCKKLVNGRKEWHLQRLTELWIKHDGSDEHI----EHWELPSSI 1086
Query: 767 VSLTITNLTEM--MRLKG-NRLQHISTLKNLS-FKCCSTLETCKDFFPSFLKSLVFINCP 822
L I NL + LK LQ + + NLS F+ L + + L++L N
Sbjct: 1087 QRLFIFNLKTLSSQHLKSLTSLQFLRIVGNLSQFQSQGQLSSFSHL--TSLQTLQIWNFL 1144
Query: 823 KLMSLPD-MFPSSLETLEFDDCPRLGLLPRSGFPSSLKLLSISHCPLLKSRYATKRNDHL 881
L SLP+ PSSL L +CP L LP G PSSL LSIS CPLL + ++
Sbjct: 1145 NLQSLPESALPSSLSHLIISNCPNLQSLPLKGMPSSLSTLSISKCPLLTPLLEFDKGEYW 1204
Query: 882 SKISHIPVVKIN 893
++I+HIP ++I+
Sbjct: 1205 TEIAHIPTIQID 1216
>gb|AAR19096.1| NBS-LRR type disease resistance protein RPG1-B [Glycine max]
Length = 1217
Score = 409 bits (1050), Expect = e-112
Identities = 308/909 (33%), Positives = 462/909 (49%), Gaps = 118/909 (12%)
Query: 6 AILNSDIWNIPND--NIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAE 63
+IL S+IW + +I+P+L L+Y HLPSHLKRCFAYC++FPK Y F+++ LI LWMAE
Sbjct: 404 SILQSEIWEFSTERSDIVPALALSYHHLPSHLKRCFAYCALFPKDYEFDKECLIQLWMAE 463
Query: 64 GFLEHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVKEKFVMHDVVYDLATIASGKSCC 123
FL+ S GK+ EVG+ YFN+LLSR ++S+ + + FVMHD++ DLA G C
Sbjct: 464 KFLQCSQQGKSPGEVGEQYFNDLLSRCFFQQSS-NTERTDFVMHDLLNDLARFICGDICF 522
Query: 124 RFGSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSCKV-IDD 182
R G + + + F+ F T D K LR+++P + Y C++ I +
Sbjct: 523 RL-DGNQTKGTPKATRHFLIDVKCFDGFGTLCDTKKLRTYMPTSYK----YWDCEMSIHE 577
Query: 183 LIPSIKRLRMLSLSN-YNITVLPNSINKLVQLRYLNLSHTDIKCLPDTTCDLYYLQTLLL 241
L LR+LSL + +++ +P+S+ L LR L+LS+T I+ LP++ C LY LQ L L
Sbjct: 578 LFSKFNYLRVLSLFDCHDLREVPDSVGNLKYLRSLDLSNTKIEKLPESICSLYNLQILKL 637
Query: 242 SGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL-TVFLVGKQKVGLSI 300
+GC L ELP ++ KL +L L++ T ++K+P + +LE LQ L + F VGK + SI
Sbjct: 638 NGCRHLKELPSNLHKLTDLHRLELIETGVRKVPAHLGKLEYLQVLMSSFNVGKSR-EFSI 696
Query: 301 RELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQ--QTEESPTNEVI 358
++LG+ NL G L I+ LQN + S+A +LK+K HL E+E+ WD ++S +
Sbjct: 697 QQLGEL-NLHGSLSIRQLQNVENPSDALAVDLKNKTHLVEVELEWDSDWNPDDSTKERDV 755
Query: 359 LNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFLKE 418
+ LQPS +L+KL ++ YGG FP WL + S ++V L++K+C+YC+ LPPLG +P LKE
Sbjct: 756 IENLQPSKHLEKLRMRNYGGTQFPRWLFNNSSCSVVSLTLKNCKYCLCLPPLGLLPSLKE 815
Query: 419 LKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFPFPR 478
L I G+ + +I +F+G + S F SL+ LEF M W EW +G FPR
Sbjct: 816 LSIKGLDGIVSINADFFGSSSCS-------FTSLKSLEFYHMKEWEEW-ECKGVTGAFPR 867
Query: 479 LKTLMLRDCTELRGHLPSHLPSIEKITILWCNHF-PATLSTLHWLSSVKSLDLMCQGSPE 537
L+ L + C +L+GHLP L + + I C P+ LS + L L G +
Sbjct: 868 LQRLSIERCPKLKGHLPEQLCHLNSLKISGCEQLVPSALSA----PDIHKLYLGDCGELQ 923
Query: 538 LSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTSLQ 597
+ D L+ TI G N + + + + Y S P +
Sbjct: 924 I-----DHGTTLKELTIEGHN---------VEAALFEEIGRNYSCSNNNIPMH------- 962
Query: 598 SLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCSSLKS 657
S Y LV+L +K CDSLT+F L+ F +L+ L I C +L+
Sbjct: 963 ------------------SCYDFLVSLRIKGGCDSLTTFPLDMFTILRELCIWKCPNLRR 1004
Query: 658 IFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLF-VLKSLTLDKLSLCCEVACLPPK 716
I + ++ LQ+L + C L SLP+ M L L SL +D C +V P
Sbjct: 1005 ISQGQAHNH-----LQTLDIKECPQLESLPEGMHVLLPSLDSLCIDD---CPKVEMFPEG 1056
Query: 717 LQFMHIESLGLATPVTEWGFQSLCFLSDLHIGGDNIVNTLL----------KKKLLPPLL 766
+++ +GL G L L +GG++ + L+ ++ +LP L
Sbjct: 1057 GLPSNLKEMGLF-----GGSYKLISLLKSALGGNHSLERLVIGKVDFECLPEEGVLPHSL 1111
Query: 767 VSLTITNLTEMMRLKGNRLQHISTLKNLSFKCCSTLETCKDFFPSFLKSLVFINCPKLMS 826
VSL I + ++ RL + H+S+LK LS + +CP+L
Sbjct: 1112 VSLQINSCGDLKRLDYKGICHLSSLKELSLE----------------------DCPRLQC 1149
Query: 827 LPDM-FPSSLETL-EFDDCPRLGLLPRSGFPSSLKLLSISH-CPLLKSRYATKRNDHLSK 883
LP+ P S+ TL + DC L R P I+H CPLL R + K
Sbjct: 1150 LPEEGLPKSISTLWIWGDCQL--LKQRCREPEGEDWPKIAHFCPLLNQRCREPGGEDWPK 1207
Query: 884 ISHIPVVKI 892
I+ I V I
Sbjct: 1208 IADIENVYI 1216
>dbj|BAB01339.1| disease resistance comples protein [Arabidopsis thaliana]
gi|29839649|sp|Q9LRR4|R13L1_ARATH Putative disease
resistance RPP13-like protein 1
gi|15231862|ref|NP_188065.1| disease resistance protein
(NBS-LRR class), putative [Arabidopsis thaliana]
Length = 1054
Score = 408 bits (1049), Expect = e-112
Identities = 246/630 (39%), Positives = 365/630 (57%), Gaps = 61/630 (9%)
Query: 7 ILNSDIWNIPND--NIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEG 64
+L+S IW++P D N++P L ++Y +LP+HLKRCFAYCSIFPKG+ F + K++LLWMAEG
Sbjct: 396 VLSSRIWDLPADKSNLLPVLRVSYYYLPAHLKRCFAYCSIFPKGHAFEKDKVVLLWMAEG 455
Query: 65 FLEHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVKEKFVMHDVVYDLATIASGKSCCR 124
FL+ + K +EE+G++YF+EL SRSL++++ K +++MHD + +LA ASG+ +
Sbjct: 456 FLQQTRSSKNLEELGNEYFSELESRSLLQKT-----KTRYIMHDFINELAQFASGEFSSK 510
Query: 125 FGSGGR--ISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSC----K 178
F G + +SE +++Y ++ Y +FE + K LR+FLP+ L S SC
Sbjct: 511 FEDGCKLQVSERTRYLSYLRDNYAEPMEFEALREVKFLRTFLPLS--LTNSSRSCCLDQM 568
Query: 179 VIDDLIPSIKRLRMLSLSNYNITVLPNSINKLVQ-LRYLNLSHTDIKCLPDTTCDLYYLQ 237
V + L+P++ RLR+LSLS+Y I LP K + R+L+LS T+++ LP + C +Y LQ
Sbjct: 569 VSEKLLPTLTRLRVLSLSHYKIARLPPDFFKNISHARFLDLSRTELEKLPKSLCYMYNLQ 628
Query: 238 TLLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLTVFLVGKQKVG 297
TLLLS C L ELP + LINLR+LD+ TK+++MP + RL++LQTLT F V G
Sbjct: 629 TLLLSYCSSLKELPTDISNLINLRYLDLIGTKLRQMPRRFGRLKSLQTLTTFFVSASD-G 687
Query: 298 LSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQQTEESPTNE- 356
I ELG +L GKL I LQ +DV++A +ANL K HL E++ W + S N
Sbjct: 688 SRISELGGLHDLHGKLKIVELQRVVDVADAAEANLNSKKHLREIDFVWRTGSSSSENNTN 747
Query: 357 --------VILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLP 408
+ +L+P +++KL+I+ Y G FP WL D SFS +V + ++ C+YC +LP
Sbjct: 748 PHRTQNEAEVFEKLRPHRHIEKLAIERYKGRRFPDWLSDPSFSRIVCIRLRECQYCTSLP 807
Query: 409 PLGQVPFLKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWIS 468
LGQ+P LKEL I GM +++IG +FY + QPF SLE L F+++P W+EW+
Sbjct: 808 SLGQLPCLKELHISGMVGLQSIGRKFYFSDQQLRDQDQQPFRSLETLRFDNLPDWQEWLD 867
Query: 469 FRGSKFP-FPRLKTLMLRDCTELRGHLPSHLPSIEKITILWC-------NHFPATLSTLH 520
R ++ FP LK L + C EL G LP+ LPS+ + I C +H + L
Sbjct: 868 VRVTRGDLFPSLKKLFILRCPELTGTLPTFLPSLISLHIYKCGLLDFQPDHHEYSYRNLQ 927
Query: 521 WLSSVKSLDLMCQGSPELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIY 580
LS S D + + F N +L + + C +Y
Sbjct: 928 TLSIKSSCDTLVK---------------------FPLNHFANLDKLEVDQ-CTS----LY 961
Query: 581 ISSLTAFPANGLPTSLQSLRIDECQNLAFL 610
L+ G P +L++LRI++CQNL L
Sbjct: 962 SLELSNEHLRG-PNALRNLRINDCQNLQLL 990
Score = 58.9 bits (141), Expect = 7e-07
Identities = 39/116 (33%), Positives = 66/116 (56%), Gaps = 2/116 (1%)
Query: 573 LQHLDLIYISSLTAFPANGLPTSLQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDS 632
L+ L ++ LT LP SL SL I +C L F +Y +L TL +K+ CD+
Sbjct: 879 LKKLFILRCPELTGTLPTFLP-SLISLHIYKCGLLDFQPDHHEYSYRNLQTLSIKSSCDT 937
Query: 633 LTSFQLNGFPVLQILSIEGCSSLKSIFISEKNSSLSLSTLQSLKVSNCKSLRSLPQ 688
L F LN F L L ++ C+SL S+ +S ++ + L++L++++C++L+ LP+
Sbjct: 938 LVKFPLNHFANLDKLEVDQCTSLYSLELSNEHLR-GPNALRNLRINDCQNLQLLPK 992
>gb|AAR19097.1| NBS-LRR type disease resistance protein RPG1-B [Glycine max]
gi|38373621|gb|AAR19095.1| NBS-LRR type disease
resistance protein RPG1-B [Glycine max]
Length = 1217
Score = 408 bits (1049), Expect = e-112
Identities = 308/909 (33%), Positives = 462/909 (49%), Gaps = 118/909 (12%)
Query: 6 AILNSDIWNIPND--NIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAE 63
+IL S+IW + +I+P+L L+Y HLPSHLKRCFAYC++FPK Y F+++ LI LWMAE
Sbjct: 404 SILQSEIWEFSTERSDIVPALALSYHHLPSHLKRCFAYCALFPKDYEFDKECLIQLWMAE 463
Query: 64 GFLEHSMVGKAVEEVGDDYFNELLSRSLIERSNDDIVKEKFVMHDVVYDLATIASGKSCC 123
FL+ S GK+ EVG+ YFN+LLSR ++S+ + + FVMHD++ DLA G C
Sbjct: 464 KFLQCSQQGKSPGEVGEQYFNDLLSRCFFQQSS-NTERTDFVMHDLLNDLARFICGDICF 522
Query: 124 RFGSGGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSCKV-IDD 182
R G + + + F+ F T D K LR+++P + Y C++ I +
Sbjct: 523 RL-DGNQTKGTPKATRHFLIDVKCFDGFGTLCDTKKLRTYMPTSYK----YWDCEMSIHE 577
Query: 183 LIPSIKRLRMLSLSN-YNITVLPNSINKLVQLRYLNLSHTDIKCLPDTTCDLYYLQTLLL 241
L LR+LSL + +++ +P+S+ L LR L+LS+T I+ LP++ C LY LQ L L
Sbjct: 578 LFSKFNYLRVLSLFDCHDLREVPDSVGNLKYLRSLDLSNTKIEKLPESICSLYNLQILKL 637
Query: 242 SGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTL-TVFLVGKQKVGLSI 300
+GC L ELP ++ KL +L L++ T ++K+P + +LE LQ L + F VGK + SI
Sbjct: 638 NGCRHLKELPSNLHKLTDLHRLELIETGVRKVPAHLGKLEYLQVLMSSFNVGKSR-EFSI 696
Query: 301 RELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWDQ--QTEESPTNEVI 358
++LG+ NL G L I+ LQN + S+A +LK+K HL ELE+ WD ++S +
Sbjct: 697 QQLGEL-NLHGSLSIRQLQNVENPSDALAVDLKNKTHLVELELEWDSDWNPDDSTKERDV 755
Query: 359 LNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPFLKE 418
+ LQPS +L+KL ++ YGG FP WL + S ++V L++K+C+YC+ LPPLG +P LKE
Sbjct: 756 IENLQPSKHLEKLRMRNYGGTQFPRWLFNNSSCSVVSLTLKNCKYCLCLPPLGLLPSLKE 815
Query: 419 LKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFPFPR 478
L I G+ + +I +F+G + S F SL+ LEF M W EW +G FPR
Sbjct: 816 LSIKGLDGIVSINADFFGSSSCS-------FTSLKSLEFYHMKEWEEW-ECKGVTGAFPR 867
Query: 479 LKTLMLRDCTELRGHLPSHLPSIEKITILWCNHF-PATLSTLHWLSSVKSLDLMCQGSPE 537
L+ L + C +L+GHLP L + + I C P+ LS + L L G +
Sbjct: 868 LQRLSIERCPKLKGHLPEQLCHLNSLKISGCEQLVPSALSA----PDIHKLYLGDCGELQ 923
Query: 538 LSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTSLQ 597
+ D L+ TI G N + + + + Y S P +
Sbjct: 924 I-----DHGTTLKELTIEGHN---------VEAALFEEIGRNYSCSNNNIPMH------- 962
Query: 598 SLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCSSLKS 657
S Y LV+L +K CDSLT+F L+ F +L+ L I C +L+
Sbjct: 963 ------------------SCYDFLVSLRIKGGCDSLTTFPLDMFTILRELCIWKCPNLRR 1004
Query: 658 IFISEKNSSLSLSTLQSLKVSNCKSLRSLPQRMDTLF-VLKSLTLDKLSLCCEVACLPPK 716
I + ++ LQ+L + C L SLP+ M L L SL +D C +V P
Sbjct: 1005 ISQGQAHNH-----LQTLDIKECPQLESLPEGMHVLLPSLDSLCIDD---CPKVEMFPEG 1056
Query: 717 LQFMHIESLGLATPVTEWGFQSLCFLSDLHIGGDNIVNTLL----------KKKLLPPLL 766
+++ +GL G L L +GG++ + L+ ++ +LP L
Sbjct: 1057 GLPSNLKEMGLF-----GGSYKLMSLLKSALGGNHSLERLVIGKVDFECLPEEGVLPHSL 1111
Query: 767 VSLTITNLTEMMRLKGNRLQHISTLKNLSFKCCSTLETCKDFFPSFLKSLVFINCPKLMS 826
VSL I + ++ RL + H+S+LK LS + +CP+L
Sbjct: 1112 VSLQINSCGDLKRLDYKGICHLSSLKELSLE----------------------DCPRLQC 1149
Query: 827 LPDM-FPSSLETL-EFDDCPRLGLLPRSGFPSSLKLLSISH-CPLLKSRYATKRNDHLSK 883
LP+ P S+ +L + DC L R P I+H CPLL R + K
Sbjct: 1150 LPEEGLPKSISSLWIWGDCQL--LKERCREPEGEDWPKIAHFCPLLNQRCREPGGEDWPK 1207
Query: 884 ISHIPVVKI 892
I+ I V I
Sbjct: 1208 IADIENVYI 1216
>gb|AAT66780.1| putative disease resistance protein [Solanum demissum]
Length = 1312
Score = 405 bits (1042), Expect = e-111
Identities = 329/989 (33%), Positives = 474/989 (47%), Gaps = 176/989 (17%)
Query: 7 ILNSDIWNIPN--DNIMPSLFLTYQHLPSHLKRCFAYCSIFPKGYPFNRKKLILLWMAEG 64
IL S+IW + + + I+P+L L+Y L LKRCFA+C+I+PK Y F ++++I LW+A G
Sbjct: 396 ILRSEIWELQSCSNGILPALMLSYNDLHPQLKRCFAFCAIYPKDYLFCKEQVIHLWIANG 455
Query: 65 FLE--HSMVGKAVEEVGDDYFNELLSRSLIER--SNDDIVKEKFVMHDVVYDLATIASGK 120
++ HS + YF EL SRSL E+ + + +F+MHD+V DLA IAS
Sbjct: 456 LVQQLHS---------ANHYFLELRSRSLFEKVQESSEWNPGEFLMHDLVNDLAQIASSN 506
Query: 121 SCCRFGS--GGRISEDVHHVTYNQEEYDIFNKFETFFDFKCLRSFLPIGSRLQESYLSCK 178
C R G + E H++Y+ D F K + + + LR+ LPI + LS +
Sbjct: 507 LCIRLEENLGSHMLEQSRHISYSMG-LDDFKKLKPLYKLEQLRTLLPINIQQHSYCLSKR 565
Query: 179 VIDDLIPSIKRLRMLSLSNYNITVLPNSIN-KLVQLRYLNLSHTDIKCLPDTTCDLYYLQ 237
++ D++P + LR LSLS+Y+I LPN + KL LR+L+ S T IK LPD+ C LY L+
Sbjct: 566 ILHDILPRLTSLRALSLSHYSIEELPNDLFIKLKYLRFLDFSWTKIKKLPDSICLLYNLE 625
Query: 238 TLLLSGCWKLIELPIHVGKLINLRHLDISYTKIKKMPMQIVRLENLQTLT-VFLVGKQKV 296
TLLLS C L ELP+H+ KLINLRHLDIS + P+ + +L++L L L+ +
Sbjct: 626 TLLLSHCSYLKELPLHMEKLINLRHLDISEAYL-TTPLHLSKLKSLHALVGANLILSGRG 684
Query: 297 GLSIRELGKFPNLRGKLCIKNLQNAIDVSEACDANLKHKVHLEELEVYWD-QQTEESPTN 355
GL + +LG+ NL G L I LQN +D E+ AN++ K H+E L + W + S T
Sbjct: 685 GLRMEDLGEVHNLYGSLSILELQNVVDRRESLKANMREKKHVERLSLEWSGSNADNSQTE 744
Query: 356 EVILNELQPSINLKKLSIKFYGGISFPSWLGDCSFSNMVYLSIKSCEYCITLPPLGQVPF 415
IL+ELQP+ N+K+L + +S+ C+ C +LP LGQ+P
Sbjct: 745 REILDELQPNTNIKEL----------------------IEMSLSYCKDCDSLPALGQLPC 782
Query: 416 LKELKIDGMSRVETIGPEFYGMTGGSTNSPFQPFPSLEKLEFNSMPSWREWISFRGSKFP 475
LK L I GM ++ + EFYG + S +PF SLEKLEF MP W++W K
Sbjct: 783 LKFLTIRGMHQITEVSEEFYG-----SLSSTKPFKSLEKLEFAGMPEWKQWHVL--GKGE 835
Query: 476 FPRLKTLMLRDCTELRGHLPSHLPSIEKITILWCNHFPATLSTLHWLSSVKSLDLMCQGS 535
FP L+ L + C +L G LP +L S+ ++ I C +L T LS++K +L
Sbjct: 836 FPILEELWINGCPKLIGKLPENLSSLRRLRISECPEL--SLETPIQLSNLK--ELKVADC 891
Query: 536 PELSLLGNDSPCHLQVSTIFGFNKLLSLPNMFMSSTCLQHLDLIYISSLTAFPANGLPTS 595
P++ +L ++ L S + G +++ L + + C SLT+ P ++
Sbjct: 892 PKVGVLFANA--QLFTSQLEGMKQIVKL----VITDC---------KSLTSLPI----ST 932
Query: 596 LQSLRIDECQNLAFLRPETWSNYTSLVTLELKNCCDSLTSFQLNGFPVLQILSIEGCSSL 655
L+S I C L+ E N L L LK C S +L FP + LS+ C++L
Sbjct: 933 LKSREISGCGE---LKLEASMNAMFLEDLSLKGC----DSPEL--FPRARNLSVRSCNNL 983
Query: 656 KSIFISEKNSSLSLS--------------TLQSLKVSNCKSLRSLPQRMDTLF-VLKSLT 700
+ I + +LS + SL + NC+ L+SLP+ M L LK LT
Sbjct: 984 TRLLIPTETETLSFGDCDNLEILSVACGIQMTSLNIHNCQKLKSLPEHMQELLPSLKELT 1043
Query: 701 LDKLSLCCEVAC-----LPPKLQFMHIESL-GLATPVTEWGFQSLCFLSDLHIGGD---- 750
LD C E+ LP LQF+ I L EW Q L L L I D
Sbjct: 1044 LDN---CPEIESFPQGGLPFNLQFLWISRCKKLVNGRKEWHLQRLPSLMQLEISHDGSDI 1100
Query: 751 --------------------NIVNTLLK--------------------KKLLPPLLVSLT 770
+ + LLK ++ LP L L
Sbjct: 1101 AGENWELPCSIRRLTIANLKTLSSQLLKSLTSLEYLYAINLPQIQSLLEEELPSSLSELH 1160
Query: 771 ITNLTEMMRLKGNRLQHISTLKNLSFKCCSTLETCKDF-FPSFLKSLVFINCPKLMSLPD 829
+ ++ L LQ + + L C L++ + PS L L +C L SLP+
Sbjct: 1161 LHQHHDLHSLPTEGLQRLMWFRCLEIWDCPNLQSLPESGMPSSLSKLTIQHCSNLQSLPE 1220
Query: 830 M-FPSSLETLEFDDCPRLGLLPRSGFPSSLKLLS-----------------------ISH 865
PSSL L +CP L LP SGFPSSL L IS
Sbjct: 1221 SGMPSSLSDLTISNCPSLQSLPESGFPSSLSELGIWNCSNLQSLPESGMPPSICNLYISE 1280
Query: 866 CPLLKSRYATKRNDHLSKISHIPVVKING 894
CPLLK + D+ KI+HIP + I+G
Sbjct: 1281 CPLLKPLLEFNKGDYWPKIAHIPTIYIDG 1309
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.139 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,524,159,585
Number of Sequences: 2540612
Number of extensions: 64722067
Number of successful extensions: 155621
Number of sequences better than 10.0: 2694
Number of HSP's better than 10.0 without gapping: 1395
Number of HSP's successfully gapped in prelim test: 1328
Number of HSP's that attempted gapping in prelim test: 136479
Number of HSP's gapped (non-prelim): 11258
length of query: 898
length of database: 863,360,394
effective HSP length: 137
effective length of query: 761
effective length of database: 515,296,550
effective search space: 392140674550
effective search space used: 392140674550
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 80 (35.4 bits)
Medicago: description of AC119417.2


