

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119414.8 + phase: 0 /pseudo
(427 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
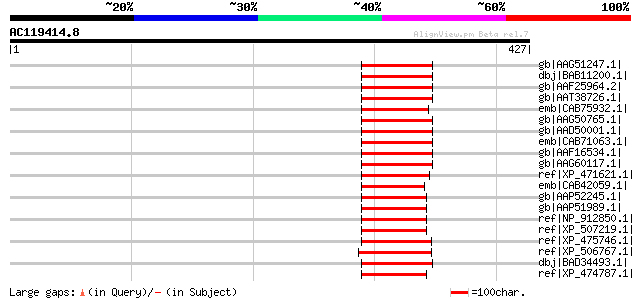
Score E
Sequences producing significant alignments: (bits) Value
gb|AAG51247.1| copia-type polyprotein, putative; 28768-32772 [Ar... 76 2e-12
dbj|BAB11200.1| copia-type polyprotein [Arabidopsis thaliana] gi... 76 2e-12
gb|AAF25964.2| F6N18.1 [Arabidopsis thaliana] 76 2e-12
gb|AAT38726.1| putative gag-pol polyprotein [Solanum demissum] 75 5e-12
emb|CAB75932.1| putative protein [Arabidopsis thaliana] gi|11278... 70 2e-10
gb|AAG50765.1| copia-type polyprotein, putative [Arabidopsis tha... 69 4e-10
gb|AAD50001.1| Hypothetical protein [Arabidopsis thaliana] gi|25... 69 4e-10
emb|CAB71063.1| copia-type polyprotein [Arabidopsis thaliana] gi... 69 4e-10
gb|AAF16534.1| T26F17.17 [Arabidopsis thaliana] 69 4e-10
gb|AAG60117.1| copia-type polyprotein, putative [Arabidopsis tha... 67 1e-09
ref|XP_471621.1| OSJNBa0029L02.7 [Oryza sativa (japonica cultiva... 60 2e-07
emb|CAB42059.1| Tpv2-1c [Phaseolus vulgaris] 59 2e-07
gb|AAP52245.1| putative pol polyprotein [Oryza sativa (japonica ... 59 3e-07
gb|AAP51989.1| putative pol polyprotein [Oryza sativa (japonica ... 59 3e-07
ref|NP_912850.1| unnamed protein product [Oryza sativa (japonica... 59 3e-07
ref|XP_507219.1| PREDICTED P0473F05.22 gene product [Oryza sativ... 59 3e-07
ref|XP_475746.1| putative polyprotein [Oryza sativa (japonica cu... 59 3e-07
ref|XP_506767.1| PREDICTED OSJNBa0009N02.26 gene product [Oryza ... 59 3e-07
dbj|BAD34493.1| Gag-Pol [Ipomoea batatas] 59 4e-07
ref|XP_474787.1| OSJNBb0026E15.10 [Oryza sativa (japonica cultiv... 58 5e-07
>gb|AAG51247.1| copia-type polyprotein, putative; 28768-32772 [Arabidopsis thaliana]
gi|25301683|pir||E86451 probable copia-type polyprotein,
28768-32772 [imported] - Arabidopsis thaliana
Length = 1334
Score = 76.3 bits (186), Expect = 2e-12
Identities = 36/59 (61%), Positives = 46/59 (77%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS YV+M+G GAI+W+S K+PIVTLSTTEAEFV+A+ +C +WLR + IG QE
Sbjct: 1194 KSTSGYVFMLGGGAIAWASKKQPIVTLSTTEAEFVSASYGACQAVWLRNVLEEIGCRQE 1252
>dbj|BAB11200.1| copia-type polyprotein [Arabidopsis thaliana]
gi|13872710|emb|CAC37622.1| polyprotein [Arabidopsis
thaliana]
Length = 1334
Score = 76.3 bits (186), Expect = 2e-12
Identities = 36/59 (61%), Positives = 46/59 (77%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS YV+M+G GAI+W+S K+PIVTLSTTEAEFV+A+ +C +WLR + IG QE
Sbjct: 1194 KSTSGYVFMLGGGAIAWASKKQPIVTLSTTEAEFVSASYGACQAVWLRNVLEEIGCRQE 1252
>gb|AAF25964.2| F6N18.1 [Arabidopsis thaliana]
Length = 1207
Score = 76.3 bits (186), Expect = 2e-12
Identities = 36/59 (61%), Positives = 46/59 (77%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS YV+M+G GAI+W+S K+PIVTLSTTEAEFV+A+ +C +WLR + IG QE
Sbjct: 1067 KSTSGYVFMLGGGAIAWASKKQPIVTLSTTEAEFVSASYGACQAVWLRNVLEEIGCRQE 1125
>gb|AAT38726.1| putative gag-pol polyprotein [Solanum demissum]
Length = 532
Score = 74.7 bits (182), Expect = 5e-12
Identities = 35/59 (59%), Positives = 45/59 (75%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS YV+++G+G +SWSS K+PIVTLSTTEAE+VAA SC+ IW+R I + QE
Sbjct: 392 KSTSGYVFLLGSGVVSWSSKKQPIVTLSTTEAEYVAATSCAYQAIWVRNILGELDFKQE 450
>emb|CAB75932.1| putative protein [Arabidopsis thaliana] gi|11278365|pir||T47841
hypothetical protein T2O9.150 - Arabidopsis thaliana
Length = 1339
Score = 69.7 bits (169), Expect = 2e-10
Identities = 28/55 (50%), Positives = 44/55 (79%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIG 344
+STS +V+++ +GAI W+S K+P+V LSTTEAE++AAA C+C +WLR++ +G
Sbjct: 1197 RSTSGFVFLMASGAICWASKKQPVVALSTTEAEYIAAAFCACQCVWLRKVLEKLG 1251
>gb|AAG50765.1| copia-type polyprotein, putative [Arabidopsis thaliana]
gi|12321254|gb|AAG50698.1| copia-type polyprotein,
putative [Arabidopsis thaliana] gi|25301687|pir||F96614
probable copia-type polyprotein T18I24.5 [imported] -
Arabidopsis thaliana
Length = 1320
Score = 68.6 bits (166), Expect = 4e-10
Identities = 32/59 (54%), Positives = 41/59 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 1174 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1232
>gb|AAD50001.1| Hypothetical protein [Arabidopsis thaliana] gi|25301681|pir||F86246
hypothetical protein [imported] - Arabidopsis thaliana
Length = 1352
Score = 68.6 bits (166), Expect = 4e-10
Identities = 32/59 (54%), Positives = 41/59 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 1206 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1264
>emb|CAB71063.1| copia-type polyprotein [Arabidopsis thaliana] gi|11278364|pir||T47925
copia-type polyprotein - Arabidopsis thaliana
Length = 1352
Score = 68.6 bits (166), Expect = 4e-10
Identities = 32/59 (54%), Positives = 41/59 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 1206 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1264
>gb|AAF16534.1| T26F17.17 [Arabidopsis thaliana]
Length = 1291
Score = 68.6 bits (166), Expect = 4e-10
Identities = 32/59 (54%), Positives = 41/59 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 1145 KSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1203
>gb|AAG60117.1| copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1352
Score = 66.6 bits (161), Expect = 1e-09
Identities = 31/59 (52%), Positives = 40/59 (67%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KSTS +V+ +G+ A +W S K+PIV LST EAE+VAA SC C IWLR + + QE
Sbjct: 1206 KSTSGFVFYIGDTAFTWMSKKQPIVVLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQE 1264
>ref|XP_471621.1| OSJNBa0029L02.7 [Oryza sativa (japonica cultivar-group)]
gi|39546217|emb|CAE04466.3| OSJNBa0029L02.7 [Oryza
sativa (japonica cultivar-group)]
Length = 856
Score = 59.7 bits (143), Expect = 2e-07
Identities = 28/57 (49%), Positives = 40/57 (70%), Gaps = 1/57 (1%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSH-IGE 345
KST+ +YM+G ISW S K+ +V LS+ EAE++AA + +C GIWL R+ + IGE
Sbjct: 715 KSTTGVLYMLGKSLISWQSQKQKVVALSSCEAEYIAATTAACQGIWLTRLLAELIGE 771
>emb|CAB42059.1| Tpv2-1c [Phaseolus vulgaris]
Length = 374
Score = 59.3 bits (142), Expect = 2e-07
Identities = 29/52 (55%), Positives = 34/52 (64%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITS 341
KSTS YV+ +GN A SW S K+PIVTLST E E+V C IWLR + S
Sbjct: 234 KSTSGYVFFMGNTAFSWLSKKQPIVTLSTCEREYVEHLGVVCHAIWLRNLLS 285
>gb|AAP52245.1| putative pol polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37531312|ref|NP_919958.1| putative pol polyprotein
[Oryza sativa (japonica cultivar-group)]
gi|18652506|gb|AAL77140.1| Putative pol polyprotein
[Oryza sativa]
Length = 1283
Score = 58.9 bits (141), Expect = 3e-07
Identities = 24/54 (44%), Positives = 38/54 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KSTS ++ V G I+W S K+ +V LS+ EAE++AAA+ +C G+WL R+ + +
Sbjct: 1139 KSTSGQIFFVNGGPITWQSSKQKVVALSSCEAEYIAAAAATCQGVWLARLLAEV 1192
>gb|AAP51989.1| putative pol polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37530800|ref|NP_919702.1| putative pol polyprotein
[Oryza sativa (japonica cultivar-group)]
gi|21326494|gb|AAM47622.1| Putative pol polyprotein
[Oryza sativa (japonica cultivar-group)]
gi|20514806|gb|AAM23251.1| Putative pol polyprotein
[Oryza sativa (japonica cultivar-group)]
Length = 1426
Score = 58.9 bits (141), Expect = 3e-07
Identities = 25/54 (46%), Positives = 39/54 (71%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KST+ +YM+G+ ISW S K+ +V LS+ EAE++AA + +C GIWL R+ + +
Sbjct: 1285 KSTTGVLYMLGDSLISWQSQKQKVVALSSCEAEYIAATTGACQGIWLNRLLAEL 1338
>ref|NP_912850.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
Length = 1330
Score = 58.9 bits (141), Expect = 3e-07
Identities = 25/54 (46%), Positives = 39/54 (71%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KST+ +YM+G+ ISW S K+ +V LS+ EAE++AA + +C GIWL R+ + +
Sbjct: 1189 KSTTGVLYMLGDSLISWQSQKQKVVALSSCEAEYIAATTGACQGIWLNRLLAEL 1242
>ref|XP_507219.1| PREDICTED P0473F05.22 gene product [Oryza sativa (japonica
cultivar-group)]
Length = 1453
Score = 58.9 bits (141), Expect = 3e-07
Identities = 23/54 (42%), Positives = 37/54 (67%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KSTS +Y +G ++W S K+ +V LS+ EAE++A A+ +C G+WLRR+ +
Sbjct: 1314 KSTSGIIYFLGGNPVAWQSQKQRVVALSSCEAEYIAGAAAACQGVWLRRLLQDV 1367
>ref|XP_475746.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|48843818|gb|AAT47077.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1136
Score = 58.9 bits (141), Expect = 3e-07
Identities = 29/58 (50%), Positives = 39/58 (67%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQ 347
KSTS Y + +G+G SWS+ K+ IV LS+ EAE+VAA+ +WLRRI +GE Q
Sbjct: 998 KSTSGYAFSLGSGMWSWSTKKQNIVALSSAEAEYVAASKAVSQVVWLRRIMEDLGEKQ 1055
>ref|XP_506767.1| PREDICTED OSJNBa0009N02.26 gene product [Oryza sativa (japonica
cultivar-group)]
Length = 349
Score = 58.9 bits (141), Expect = 3e-07
Identities = 23/60 (38%), Positives = 40/60 (66%)
Query: 288 TEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQ 347
T KSTS ++ +G+ +SW S K+ +V LS+ EAE++AA + +C G+W+ ++ I + Q
Sbjct: 203 TRKSTSGVMFFLGSNPVSWQSQKQKVVALSSCEAEYIAARTAACQGVWMAQLLGEINQAQ 262
>dbj|BAD34493.1| Gag-Pol [Ipomoea batatas]
Length = 1298
Score = 58.5 bits (140), Expect = 4e-07
Identities = 28/59 (47%), Positives = 37/59 (62%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KST+ YV+ V GA+SW S + +V STTEAE+VAA S IWL+ + +G QE
Sbjct: 1161 KSTTGYVFKVAGGAVSWVSKLQAVVATSTTEAEYVAATQASKEAIWLKMLLEELGHKQE 1219
>ref|XP_474787.1| OSJNBb0026E15.10 [Oryza sativa (japonica cultivar-group)]
gi|38344222|emb|CAE03692.2| OSJNBb0026E15.10 [Oryza
sativa (japonica cultivar-group)]
Length = 1449
Score = 58.2 bits (139), Expect = 5e-07
Identities = 22/54 (40%), Positives = 38/54 (69%)
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KSTS ++ + G ++W S K+ +V LS+ EAE++AAA+ +C G+WL R+ + +
Sbjct: 1305 KSTSGQIFFINGGPVTWQSSKQKVVALSSCEAEYIAAAAATCQGVWLARLLAEV 1358
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.372 0.166 0.671
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 557,214,820
Number of Sequences: 2540612
Number of extensions: 17678533
Number of successful extensions: 61803
Number of sequences better than 10.0: 542
Number of HSP's better than 10.0 without gapping: 532
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 61178
Number of HSP's gapped (non-prelim): 631
length of query: 427
length of database: 863,360,394
effective HSP length: 131
effective length of query: 296
effective length of database: 530,540,222
effective search space: 157039905712
effective search space used: 157039905712
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC119414.8


