

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.5 + phase: 0
(221 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
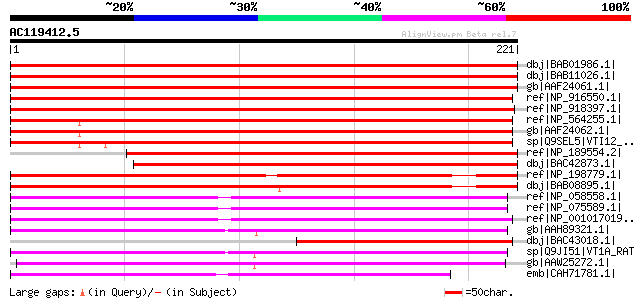
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB01986.1| vesicle transport v-SNARE (vesicle soluble NSF a... 339 4e-92
dbj|BAB11026.1| v-SNARE AtVTI1a [Arabidopsis thaliana] gi|201471... 338 6e-92
gb|AAF24061.1| v-SNARE AtVTI1a [Arabidopsis thaliana] 337 2e-91
ref|NP_916550.1| putative vesicle transport v-SNARE protein [Ory... 323 3e-87
ref|NP_918397.1| similar to vesicle transport protein [Oryza sat... 301 1e-80
ref|NP_564255.1| vesical transport v-SNARE 12 (VTI12) / vesicle ... 291 9e-78
gb|AAF24062.1| v-SNARE AtVTI1b [Arabidopsis thaliana] 290 2e-77
sp|Q9SEL5|VTI12_ARATH Vesicle transport v-SNARE 12 (AtVTI12) (Ve... 286 2e-76
ref|NP_189554.2| vesicle transport v-SNARE 13 (VTI13) / vesicle ... 260 2e-68
dbj|BAC42873.1| putative vesicle transport protein [Arabidopsis ... 256 3e-67
ref|NP_198779.1| vesicle transport v-SNARE family protein [Arabi... 179 7e-44
dbj|BAB08895.1| v-SNARE AtVTI1a-like protein [Arabidopsis thaliana] 175 7e-43
ref|NP_058558.1| vesicle transport through interaction with t-SN... 133 4e-30
ref|NP_075589.1| vesicle transport through interaction with t-SN... 129 5e-29
ref|NP_001017019.1| hypothetical protein LOC549773 [Xenopus trop... 129 8e-29
gb|AAH89321.1| Vti1a protein [Mus musculus] 127 2e-28
dbj|BAC43018.1| putative v-SNARE AtVTI1b [Arabidopsis thaliana] ... 126 4e-28
sp|Q9JI51|VT1A_RAT Vesicle transport through interaction with t-... 124 3e-27
gb|AAW25272.1| unknown [Schistosoma japonicum] 120 2e-26
emb|CAH71781.1| vesicle transport through interaction with t-SNA... 120 3e-26
>dbj|BAB01986.1| vesicle transport v-SNARE (vesicle soluble NSF attachment protein
receptor) protein [Arabidopsis thaliana]
gi|27805771|sp|Q9LVP9|VT13_ARATH Vesicle transport
v-SNARE 13 (AtVTI13) (Vesicle transport v-SNARE protein
VTI13) (Vesicle soluble NSF attachment protein receptor
13)
Length = 221
Score = 339 bits (869), Expect = 4e-92
Identities = 166/221 (75%), Positives = 196/221 (88%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS FE YERQYCE+SANLSKKCT+A AL+GEQKKQ +SE+KSG++EAEAL++KMDLEAR
Sbjct: 1 MSQGFERYERQYCEISANLSKKCTSAIALDGEQKKQNLSEIKSGVEEAEALVKKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
++ PN+K LL KLREYKSDLNN K+EVK+I SGNLN +ARDELLE+GMADT+TASADQR
Sbjct: 61 NLPPNVKSSLLVKLREYKSDLNNFKTEVKRITSGNLNATARDELLEAGMADTLTASADQR 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
+RLM ST+ L +T +R+KDSRRT+LETEELGVSILQDLH QRQSLL AH TLHGVDDN+G
Sbjct: 121 SRLMMSTDHLGRTTDRIKDSRRTILETEELGVSILQDLHGQRQSLLRAHETLHGVDDNVG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
KSKKI+T M+RRMN+NKW IG I+ VLV+AII ILYFKL +
Sbjct: 181 KSKKILTTMTRRMNRNKWTIGAIITVLVLAIIFILYFKLTR 221
>dbj|BAB11026.1| v-SNARE AtVTI1a [Arabidopsis thaliana] gi|20147179|gb|AAM10306.1|
AT5g39510/MUL8_190 [Arabidopsis thaliana]
gi|17979270|gb|AAL49951.1| AT5g39510/MUL8_190
[Arabidopsis thaliana] gi|15241780|ref|NP_198767.1|
vesicle transport v-SNARE 11 (VTI11) / vesicle soluble
NSF attachment protein receptor VTI1a (VTI1A)
[Arabidopsis thaliana] gi|27805773|sp|Q9SEL6|VTI11_ARATH
Vesicle transport v-SNARE 11 (AtVTI11) (Vesicle
transport v-SNARE protein VTI1a) (Vesicle soluble NSF
attachment protein receptor VTI1a) (AtVTI1a)
Length = 221
Score = 338 bits (867), Expect = 6e-92
Identities = 166/221 (75%), Positives = 195/221 (88%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+VF+GYERQYCELSA+LSKKC++A +L+GEQKKQK+SE+KSG++ AE LIRKMDLEAR
Sbjct: 1 MSDVFDGYERQYCELSASLSKKCSSAISLDGEQKKQKLSEIKSGLENAEVLIRKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
++ PN+K LL KLRE+KSDLNN K+EVK+I SG LN +ARDELLE+GMADT TASADQR
Sbjct: 61 TLPPNLKSSLLVKLREFKSDLNNFKTEVKRITSGQLNAAARDELLEAGMADTKTASADQR 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
RLM STERL +T +RVKDSRRTM+ETEE+GVSILQDLH QRQSLL AH TLHGVDDNIG
Sbjct: 121 ARLMMSTERLGRTTDRVKDSRRTMMETEEIGVSILQDLHGQRQSLLRAHETLHGVDDNIG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
KSKKI+T+M+RRMNKNKW IG I++ L+ AI ILYFKL K
Sbjct: 181 KSKKILTDMTRRMNKNKWTIGAIIIALIAAIFIILYFKLTK 221
>gb|AAF24061.1| v-SNARE AtVTI1a [Arabidopsis thaliana]
Length = 221
Score = 337 bits (863), Expect = 2e-91
Identities = 165/221 (74%), Positives = 194/221 (87%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ F+GYERQYCELSA+LSKKC++A +L+GEQKKQK+SE+KSG++ AE LIRKMDLEAR
Sbjct: 1 MSDAFDGYERQYCELSASLSKKCSSAISLDGEQKKQKLSEIKSGLENAEVLIRKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
++ PN+K LL KLRE+KSDLNN K+EVK+I SG LN +ARDELLE+GMADT TASADQR
Sbjct: 61 TLPPNLKSSLLVKLREFKSDLNNFKTEVKRITSGQLNAAARDELLEAGMADTKTASADQR 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
RLM STERL +T +RVKDSRRTM+ETEE+GVSILQDLH QRQSLL AH TLHGVDDNIG
Sbjct: 121 ARLMMSTERLGRTTDRVKDSRRTMMETEEIGVSILQDLHGQRQSLLRAHETLHGVDDNIG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
KSKKI+T+M+RRMNKNKW IG I++ L+ AI ILYFKL K
Sbjct: 181 KSKKILTDMTRRMNKNKWTIGAIIIALIAAIFIILYFKLTK 221
>ref|NP_916550.1| putative vesicle transport v-SNARE protein [Oryza sativa (japonica
cultivar-group)] gi|19571103|dbj|BAB86528.1| vesicle
transport v-SNARE (vesicle soluble NSF attachment
protein receptor) protein-like [Oryza sativa (japonica
cultivar-group)] gi|20804648|dbj|BAB92337.1| vesicle
transport v-SNARE (vesicle soluble NSF attachment
protein receptor) protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 221
Score = 323 bits (827), Expect = 3e-87
Identities = 157/219 (71%), Positives = 195/219 (88%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS VFEGYERQYCE+SA+L +KCT A+AL+GE+KKQK+SE++SG++EAE+LIRKMDLEAR
Sbjct: 1 MSEVFEGYERQYCEVSASLFRKCTTASALDGEKKKQKLSEIQSGVEEAESLIRKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
S+QP++K LLAKLREYKSDLNNLKSE+K+I + N + R+ELLESG+ADT+ AS DQR
Sbjct: 61 SLQPSVKAGLLAKLREYKSDLNNLKSELKRISAPNARQATREELLESGLADTLAASTDQR 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
RLM +TERLN++ +++K+SRRT+LETEELGVSILQDLH QRQSLLHAH TLHGVDDN+G
Sbjct: 121 GRLMMTTERLNQSNDKIKESRRTILETEELGVSILQDLHQQRQSLLHAHTTLHGVDDNVG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
KSKKI+ MS+RM++NKWIIG I+ LV+AI+ ILYFKL
Sbjct: 181 KSKKILAAMSKRMDRNKWIIGGIIAALVLAILLILYFKL 219
>ref|NP_918397.1| similar to vesicle transport protein [Oryza sativa (japonica
cultivar-group)] gi|18844938|dbj|BAB85407.1| vesicle
transport v-SNARE (vesicle soluble NSF attachment
protein receptor) protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 221
Score = 301 bits (770), Expect = 1e-80
Identities = 145/220 (65%), Positives = 186/220 (83%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
M++VFEGYERQYCE+SA+L++KCTAA+AL GE+ KQK SE+KSGID AEALI+KMDLEAR
Sbjct: 1 MADVFEGYERQYCEISASLARKCTAASALQGEKLKQKASEIKSGIDGAEALIKKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ QP+++ LLAKLREYKSDLNNLK +K++ +GN +R+ELLESGMA+T+ SADQ+
Sbjct: 61 NQQPSVRAGLLAKLREYKSDLNNLKGTLKRVTTGNAQQGSREELLESGMAETLGVSADQK 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
+RL+ TE+ NKT +R++DS RTMLETE+LGVS+LQDLH QR+ L+HAH TL VDDNIG
Sbjct: 121 SRLLRITEKQNKTTDRIRDSHRTMLETEDLGVSLLQDLHQQRERLIHAHGTLDNVDDNIG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLV 220
KS++IM M RRM++NKWIIG I+ +LV+ I+ ILYFK V
Sbjct: 181 KSRRIMGAMVRRMDRNKWIIGFIIALLVLVILVILYFKFV 220
>ref|NP_564255.1| vesical transport v-SNARE 12 (VTI12) / vesicle soluble NSF
attachment protein receptor VTI1b (VTI1B) receptor VTI1b
[Arabidopsis thaliana] gi|9295720|gb|AAF87026.1|
T24P13.5 [Arabidopsis thaliana]
Length = 222
Score = 291 bits (745), Expect = 9e-78
Identities = 144/220 (65%), Positives = 182/220 (82%), Gaps = 1/220 (0%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATAL-NGEQKKQKVSEVKSGIDEAEALIRKMDLEA 59
MS+VFEGYERQYCELS NLS+KC +A+ L NGE+KK K++E+KSGIDEA+ LIRKMDLEA
Sbjct: 1 MSDVFEGYERQYCELSTNLSRKCHSASVLSNGEEKKGKIAEIKSGIDEADVLIRKMDLEA 60
Query: 60 RSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQ 119
RS+QP+ K V L+KLREYKSDLN LK E K++ S + PS+R+EL+ESGMAD SADQ
Sbjct: 61 RSLQPSAKAVCLSKLREYKSDLNQLKKEFKRVSSADAKPSSREELMESGMADLHAVSADQ 120
Query: 120 RTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNI 179
R RL S ERL+++ +R+++SRR MLETEE+G+SI+QDL QRQ+LLHAHN LHGVDD I
Sbjct: 121 RGRLAMSVERLDQSSDRIRESRRLMLETEEVGISIVQDLSQQRQTLLHAHNKLHGVDDAI 180
Query: 180 GKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
KSKK++T MSRRM +NKWII +++ LV+AII I+ +KL
Sbjct: 181 DKSKKVLTAMSRRMTRNKWIITSVIVALVLAIILIISYKL 220
>gb|AAF24062.1| v-SNARE AtVTI1b [Arabidopsis thaliana]
Length = 222
Score = 290 bits (742), Expect = 2e-77
Identities = 143/220 (65%), Positives = 182/220 (82%), Gaps = 1/220 (0%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATAL-NGEQKKQKVSEVKSGIDEAEALIRKMDLEA 59
MS+VFEGYERQYCELS NLS+KC +A+ L NGE+KK K++++KSGIDEA+ LIRKMDLEA
Sbjct: 1 MSDVFEGYERQYCELSTNLSRKCHSASVLSNGEEKKGKIAQIKSGIDEADVLIRKMDLEA 60
Query: 60 RSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQ 119
RS+QP+ K V L+KLREYKSDLN LK E K++ S + PS+R+EL+ESGMAD SADQ
Sbjct: 61 RSLQPSAKAVCLSKLREYKSDLNQLKKEFKRVSSADAKPSSREELMESGMADLHAVSADQ 120
Query: 120 RTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNI 179
R RL S ERL+++ +R+++SRR MLETEE+G+SI+QDL QRQ+LLHAHN LHGVDD I
Sbjct: 121 RGRLAMSVERLDQSSDRIRESRRLMLETEEVGISIVQDLSQQRQTLLHAHNKLHGVDDAI 180
Query: 180 GKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
KSKK++T MSRRM +NKWII +++ LV+AII I+ +KL
Sbjct: 181 DKSKKVLTAMSRRMTRNKWIITSVIVALVLAIILIISYKL 220
>sp|Q9SEL5|VTI12_ARATH Vesicle transport v-SNARE 12 (AtVTI12) (Vesicle transport v-SNARE
protein VTI1b) (Vesicle soluble NSF attachment protein
receptor VTI1b) (AtVTI1b)
Length = 223
Score = 286 bits (733), Expect = 2e-76
Identities = 144/221 (65%), Positives = 182/221 (82%), Gaps = 2/221 (0%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATAL-NGEQKKQKVSE-VKSGIDEAEALIRKMDLE 58
MS+VFEGYERQYCELS NLS+KC +A+ L NGE+KK K++E +KSGIDEA+ LIRKMDLE
Sbjct: 1 MSDVFEGYERQYCELSTNLSRKCHSASVLSNGEEKKGKIAEQIKSGIDEADVLIRKMDLE 60
Query: 59 ARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASAD 118
ARS+QP+ K V L+KLREYKSDLN LK E K++ S + PS+R+EL+ESGMAD SAD
Sbjct: 61 ARSLQPSAKAVCLSKLREYKSDLNQLKKEFKRVSSADAKPSSREELMESGMADLHAVSAD 120
Query: 119 QRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDN 178
QR RL S ERL+++ +R+++SRR MLETEE+G+SI+QDL QRQ+LLHAHN LHGVDD
Sbjct: 121 QRGRLAMSVERLDQSSDRIRESRRLMLETEEVGISIVQDLSQQRQTLLHAHNKLHGVDDA 180
Query: 179 IGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
I KSKK++T MSRRM +NKWII +++ LV+AII I+ +KL
Sbjct: 181 IDKSKKVLTAMSRRMTRNKWIITSVIVALVLAIILIISYKL 221
>ref|NP_189554.2| vesicle transport v-SNARE 13 (VTI13) / vesicle soluble NSF
attachment protein receptor 13 [Arabidopsis thaliana]
Length = 195
Score = 260 bits (664), Expect = 2e-68
Identities = 127/170 (74%), Positives = 150/170 (87%)
Query: 52 IRKMDLEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMAD 111
++KMDLEAR++ PN+K LL KLREYKSDLNN K+EVK+I SGNLN +ARDELLE+GMAD
Sbjct: 26 VKKMDLEARNLPPNVKSSLLVKLREYKSDLNNFKTEVKRITSGNLNATARDELLEAGMAD 85
Query: 112 TMTASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNT 171
T+TASADQR+RLM ST+ L +T +R+KDSRRT+LETEELGVSILQDLH QRQSLL AH T
Sbjct: 86 TLTASADQRSRLMMSTDHLGRTTDRIKDSRRTILETEELGVSILQDLHGQRQSLLRAHET 145
Query: 172 LHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
LHGVDDN+GKSKKI+T M+RRMN+NKW IG I+ VLV+AII ILYFKL +
Sbjct: 146 LHGVDDNVGKSKKILTTMTRRMNRNKWTIGAIITVLVLAIIFILYFKLTR 195
>dbj|BAC42873.1| putative vesicle transport protein [Arabidopsis thaliana]
Length = 167
Score = 256 bits (654), Expect = 3e-67
Identities = 126/167 (75%), Positives = 147/167 (87%)
Query: 55 MDLEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMT 114
MDLEAR++ PN+K LL KLREYKSDLNN K+EVK+I SGNLN +ARDELLE+GMADT+T
Sbjct: 1 MDLEARNLPPNVKSSLLVKLREYKSDLNNFKTEVKRITSGNLNATARDELLEAGMADTLT 60
Query: 115 ASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHG 174
ASADQR+RLM ST+ L +T +R+KDSRRT+LETEELGVSILQDLH QRQSLL AH TLHG
Sbjct: 61 ASADQRSRLMMSTDHLGRTTDRIKDSRRTILETEELGVSILQDLHGQRQSLLRAHETLHG 120
Query: 175 VDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
VDDN+GKSKKI+T M+RRMN+NKW IG I+ VLV+AII ILYFKL +
Sbjct: 121 VDDNVGKSKKILTTMTRRMNRNKWTIGAIITVLVLAIIFILYFKLTR 167
>ref|NP_198779.1| vesicle transport v-SNARE family protein [Arabidopsis thaliana]
Length = 207
Score = 179 bits (453), Expect = 7e-44
Identities = 99/221 (44%), Positives = 143/221 (63%), Gaps = 14/221 (6%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MSNVF Y++QY ELS NLSKKC+ A +L G +KK+K+SE+ ++ AE LI KMD A
Sbjct: 1 MSNVFHEYDQQYSELSINLSKKCSLAFSLKGGEKKEKLSEITYDLENAEKLISKMDHAAS 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
++ PNIK +LL KL+E KS L L++E+K+ S NL + R+E+LE+ AD ADQR
Sbjct: 61 NLPPNIKSILLEKLKESKSSLKRLRNEIKRNTSENLKVTTREEVLEAEKADL----ADQR 116
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
+RLM STE L +T E +KDS+R +LETE +G+SIL++L Q++SL ++ LH +DD +
Sbjct: 117 SRLMKSTEGLVRTREMIKDSQRKLLETENIGISILENLQRQKESLQNSQAMLHEIDDTVK 176
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
+S+ I+ ++ + + II FKLVK
Sbjct: 177 ESRSIVRSIKIKE----------FFTVTAPIIIYFLFKLVK 207
>dbj|BAB08895.1| v-SNARE AtVTI1a-like protein [Arabidopsis thaliana]
Length = 212
Score = 175 bits (444), Expect = 7e-43
Identities = 98/222 (44%), Positives = 142/222 (63%), Gaps = 11/222 (4%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MSNVF Y++QY ELS NLSKKC+ A +L G +KK+K+SE+ ++ AE LI KMD A
Sbjct: 1 MSNVFHEYDQQYSELSINLSKKCSLAFSLKGGEKKEKLSEITYDLENAEKLISKMDHAAS 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTAS-ADQ 119
++ PNIK +LL KL+E KS L L++E+K+ S NL + R+E+LE+ A ADQ
Sbjct: 61 NLPPNIKSILLEKLKESKSSLKRLRNEIKRNTSENLKVTTREEVLEAEKAVCKDMDLADQ 120
Query: 120 RTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNI 179
R+RLM STE L +T E +KDS+R +LETE +G+SIL++L Q++SL ++ LH +DD +
Sbjct: 121 RSRLMKSTEGLVRTREMIKDSQRKLLETENIGISILENLQRQKESLQNSQAMLHEIDDTV 180
Query: 180 GKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
+S+ I+ ++ + + II FKLVK
Sbjct: 181 KESRSIVRSIKIKE----------FFTVTAPIIIYFLFKLVK 212
>ref|NP_058558.1| vesicle transport through interaction with t-SNAREs homolog 1A [Mus
musculus] gi|18043878|gb|AAH19386.1| Vesicle transport
through interaction with t-SNAREs homolog 1A [Mus
musculus] gi|13124607|sp|O89116|VTI1A_MOUSE Vesicle
transport through interaction with t-SNAREs homolog 1A
(Vesicle transport v-SNARE protein Vti1-like 2)
(Vti1-rp2) gi|3421062|gb|AAC32049.1| 29-kDa Golgi SNARE
[Mus musculus] gi|3213227|gb|AAC23482.1| putative
v-SNARE Vti1a [Mus musculus] gi|26324584|dbj|BAC26046.1|
unnamed protein product [Mus musculus]
gi|12836163|dbj|BAB23532.1| unnamed protein product [Mus
musculus]
Length = 217
Score = 133 bits (334), Expect = 4e-30
Identities = 67/217 (30%), Positives = 128/217 (58%), Gaps = 5/217 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FEGYE+ + L+A ++ K L ++KKQ V+ V+ ++EA L+ +MDLE R
Sbjct: 1 MSSDFEGYEQDFAVLTAEITSKIARVPRLPPDEKKQMVANVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ P +G+ ++R YK ++ L+++ K+ + DE+ + D +S +QR
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKRS-----RIAYSDEVRNELLGDAGNSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +L++L R+ + A + L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDREKIQRARDRLRDADANLG 175
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYF 217
KS +I+T M RR+ +N+ ++ + +++V+AI+ + F
Sbjct: 176 KSSRILTGMLRRIIQNRILLVILGIIVVIAILTAIAF 212
>ref|NP_075589.1| vesicle transport through interaction with t-SNAREs homolog 1A
[Rattus norvegicus] gi|9719420|gb|AAF97790.1| SNARE
Vti1a protein [Rattus norvegicus]
Length = 217
Score = 129 bits (325), Expect = 5e-29
Identities = 66/217 (30%), Positives = 126/217 (57%), Gaps = 5/217 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS FEGYE+ + L+A ++ K + L ++KKQ V+ V+ ++EA L+ +MDLE R
Sbjct: 1 MSADFEGYEQDFAVLTAEITSKISRVPRLPPDEKKQMVANVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ P +G+ ++R YK ++ L+++ K+ + DE+ + D +S +QR
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKRS-----RIAYSDEVRNELLGDAGNSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +L++L R+ + A L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDRERIQRARERLRETDANLG 175
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYF 217
KS +I+T M RR+ +N+ ++ + +++V+ I+ + F
Sbjct: 176 KSSRILTGMLRRIIQNRILLVILGIIVVITILTAITF 212
>ref|NP_001017019.1| hypothetical protein LOC549773 [Xenopus tropicalis]
Length = 217
Score = 129 bits (323), Expect = 8e-29
Identities = 66/219 (30%), Positives = 124/219 (56%), Gaps = 5/219 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FEGYE+ Y L+A ++ + L G++KKQ + V+ ++EA L+ +MDLE R
Sbjct: 1 MSSDFEGYEQDYAVLTAEITGRIGKVPRLMGDEKKQMILNVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ + + ++R YK ++ L ++ K+ + DE+ + D +S QR
Sbjct: 61 EIPVQSRAMYSNRMRSYKQEMGKLDADFKRS-----RIAYSDEVRNELLGDEGNSSESQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +IL++L+ R+ + A L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQVAVETEQIGQNILENLNQDREKIQRARERLRETDANLG 175
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
KS +++T M RR+ +N+ ++ + + +V II +YF +
Sbjct: 176 KSSRVLTGMLRRIIQNRVLVVVLGIFIVFTIILAIYFSI 214
>gb|AAH89321.1| Vti1a protein [Mus musculus]
Length = 224
Score = 127 bits (319), Expect = 2e-28
Identities = 66/220 (30%), Positives = 129/220 (58%), Gaps = 4/220 (1%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FEGYE+ + L+A ++ K L ++KKQ V+ V+ ++EA L+ +MDLE R
Sbjct: 1 MSSDFEGYEQDFAVLTAEITSKIARVPRLPPDEKKQMVANVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLE---SGMADTMTASA 117
+ P +G+ ++R YK ++ L+++ K+ + R+ELL + + +
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKRSRIA-YSDEVRNELLGDAGNSSENQLIKLR 119
Query: 118 DQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDD 177
++R L+ +TERL ++ R++ + +ETE++G +L++L R+ + A + L D
Sbjct: 120 EERAHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDREKIQRARDRLRDADA 179
Query: 178 NIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYF 217
N+GKS +I+T M RR+ +N+ ++ + +++V+AI+ + F
Sbjct: 180 NLGKSSRILTGMLRRIIQNRILLVILGIIVVIAILTAIAF 219
>dbj|BAC43018.1| putative v-SNARE AtVTI1b [Arabidopsis thaliana]
gi|28416783|gb|AAO42922.1| At1g26670 [Arabidopsis
thaliana]
Length = 97
Score = 126 bits (317), Expect = 4e-28
Identities = 59/94 (62%), Positives = 79/94 (83%)
Query: 126 STERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKI 185
S ERL+++ +R+++SRR MLETEE+G+SI+QDL QRQ+LLHAHN LHGVDD I KSKK+
Sbjct: 2 SVERLDQSSDRIRESRRLMLETEEVGISIVQDLSQQRQTLLHAHNKLHGVDDAIDKSKKV 61
Query: 186 MTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
+T MSRRM +NKWII +++ LV+AII I+ +KL
Sbjct: 62 LTAMSRRMTRNKWIITSVIVALVLAIILIISYKL 95
>sp|Q9JI51|VT1A_RAT Vesicle transport through interaction with t-SNAREs homolog 1A
(Vesicle transport v-SNARE protein Vti1-like 2)
(Vti1-rp2) gi|9719422|gb|AAF97791.1| SNARE Vti1a-beta
protein [Rattus norvegicus]
Length = 224
Score = 124 bits (310), Expect = 3e-27
Identities = 65/220 (29%), Positives = 127/220 (57%), Gaps = 4/220 (1%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS FEGYE+ + L+A ++ K + L ++KKQ V+ V+ ++EA L+ +MDLE R
Sbjct: 1 MSADFEGYEQDFAVLTAEITSKISRVPRLPPDEKKQMVANVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELL---ESGMADTMTASA 117
+ P +G+ ++R YK ++ L+++ K+ + R+ELL + + +
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKRSRIA-YSDEVRNELLGDAGNSSENQLIKLR 119
Query: 118 DQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDD 177
++R L+ +TERL ++ R++ + +ETE++G +L++L R+ + A L D
Sbjct: 120 EERAHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDRERIQRARERLRETDA 179
Query: 178 NIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYF 217
N+GKS +I+T M RR+ +N+ ++ + +++V+ I+ + F
Sbjct: 180 NLGKSSRILTGMLRRIIQNRILLVILGIIVVITILTAITF 219
>gb|AAW25272.1| unknown [Schistosoma japonicum]
Length = 240
Score = 120 bits (302), Expect = 2e-26
Identities = 65/216 (30%), Positives = 125/216 (57%), Gaps = 3/216 (1%)
Query: 4 VFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEARSMQ 63
+FE YE+QY L+++++ K LNG ++ ++V +V + DE LI +M LE + M+
Sbjct: 3 LFESYEKQYGNLTSDITFKLGKIPQLNGSERTKEVRDVSALFDETRDLIDQMSLEVQEMR 62
Query: 64 PNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELL---ESGMADTMTASADQR 120
+I+ +L Y+ +L L +E K+ P R+ + E + + + D R
Sbjct: 63 HDIRPKFQNRLECYRKELEKLTTEFKRPRYAARRPENRNLGISDDEDTLHEEVLFDTDMR 122
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
+L+++TER+++T +++D R LETEE+G ILQDL QR+ L + + L D ++
Sbjct: 123 AQLLSNTERISRTTTKLEDGVRIALETEEVGGHILQDLSEQREKLQRSRDRLRQADSDLK 182
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILY 216
KS ++++ M+RR+ +NK ++ ++ ++V+ + +Y
Sbjct: 183 KSSRVLSVMTRRILQNKLLLIAVIFIMVLIFLLTIY 218
>emb|CAH71781.1| vesicle transport through interaction with t-SNAREs homolog 1A
(yeast) [Homo sapiens] gi|16877603|gb|AAH17052.1| SNARE
Vti1a-beta protein [Homo sapiens]
gi|21624648|ref|NP_660207.1| SNARE Vti1a-beta protein
[Homo sapiens] gi|29337113|sp|Q96AJ9|VT1A_HUMAN Vesicle
transport through interaction with t-SNAREs homolog 1A
(Vesicle transport v-SNARE protein Vti1-like 2)
(Vti1-rp2)
Length = 203
Score = 120 bits (301), Expect = 3e-26
Identities = 62/192 (32%), Positives = 111/192 (57%), Gaps = 5/192 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FEGYE+ + L+A ++ K L ++KKQ V+ V+ ++EA+ L+ +MDLE R
Sbjct: 1 MSSDFEGYEQDFAVLTAEITSKIARVPRLPPDEKKQMVANVEKQLEEAKELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ P +G+ ++R YK ++ L+++ K+ + DE+ + D +S +QR
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKR-----SRIAYSDEVRNELLGDDGNSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +L++L R+ + A L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDREKIQRARERLRETDANLG 175
Query: 181 KSKKIMTNMSRR 192
KS +I+T M RR
Sbjct: 176 KSSRILTGMLRR 187
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.314 0.129 0.342
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 307,467,041
Number of Sequences: 2540612
Number of extensions: 10998477
Number of successful extensions: 44832
Number of sequences better than 10.0: 791
Number of HSP's better than 10.0 without gapping: 132
Number of HSP's successfully gapped in prelim test: 662
Number of HSP's that attempted gapping in prelim test: 44333
Number of HSP's gapped (non-prelim): 1055
length of query: 221
length of database: 863,360,394
effective HSP length: 123
effective length of query: 98
effective length of database: 550,865,118
effective search space: 53984781564
effective search space used: 53984781564
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 72 (32.3 bits)
Medicago: description of AC119412.5


