

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119408.1 + phase: 0 /pseudo
(600 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
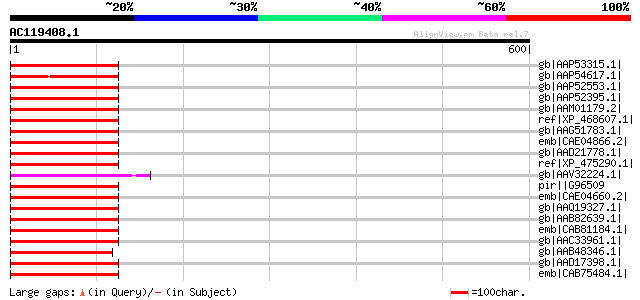
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP53315.1| putative reverse transcriptase [Oryza sativa (jap... 136 2e-30
gb|AAP54617.1| putative non-LTR retroelement reverse transcripta... 128 6e-28
gb|AAP52553.1| putative reverse transcriptase [Oryza sativa (jap... 122 3e-26
gb|AAP52395.1| putative retroelement [Oryza sativa (japonica cul... 122 3e-26
gb|AAM01179.2| Putative retroelement [Oryza sativa (japonica cul... 122 3e-26
ref|XP_468607.1| putative reverse transcriptase [Oryza sativa (j... 120 1e-25
gb|AAG51783.1| reverse transcriptase, putative; 16838-20266 [Ara... 119 4e-25
emb|CAE04866.2| OSJNBa0086O06.14 [Oryza sativa (japonica cultiva... 117 1e-24
gb|AAD21778.1| putative non-LTR retroelement reverse transcripta... 115 3e-24
ref|XP_475290.1| hypothetical protein [Oryza sativa (japonica cu... 115 3e-24
gb|AAV32224.1| hypothetical protein [Oryza sativa (japonica cult... 115 5e-24
pir||G96509 protein F27F5.21 [imported] - Arabidopsis thaliana g... 114 9e-24
emb|CAE04660.2| OSJNBa0061G20.16 [Oryza sativa (japonica cultiva... 114 1e-23
gb|AAQ19327.1| bZIP-like protein [Oryza sativa (japonica cultiva... 112 3e-23
gb|AAB82639.1| putative non-LTR retroelement reverse transcripta... 111 6e-23
emb|CAB81184.1| putative protein [Arabidopsis thaliana] gi|45393... 110 1e-22
gb|AAC33961.1| contains similarity to reverse trancriptase (Pfam... 110 1e-22
gb|AAB48346.1| reverse transcriptase 110 1e-22
gb|AAD17398.1| putative non-LTR retroelement reverse transcripta... 109 2e-22
emb|CAB75484.1| putative protein [Arabidopsis thaliana] gi|11357... 109 2e-22
>gb|AAP53315.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|37533452|ref|NP_921028.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|20279456|gb|AAM18736.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1509
Score = 136 bits (342), Expect = 2e-30
Identities = 62/125 (49%), Positives = 89/125 (70%), Gaps = 1/125 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ANR+K ILP++ + QS FVPGRLI+DN LI +E HY+R R+ GY KLD++KA
Sbjct: 722 LANRLKKILPDVISPAQSAFVPGRLISDNILIAYEMTHYMRNKRSGQVGYAAFKLDMSKA 781
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++W+F+ D + +GF + +N IM C+ VT+ I +NG+ +E FSP+RG+RQGDP
Sbjct: 782 YDRVEWSFLHDMMLKLGFHTDWVNLIMKCVSTVTYRIRVNGELSESFSPERGLRQGDPLS 841
Query: 121 PLIFL 125
P +FL
Sbjct: 842 PYLFL 846
>gb|AAP54617.1| putative non-LTR retroelement reverse transcriptase [Oryza sativa
(japonica cultivar-group)] gi|37536056|ref|NP_922330.1|
putative non-LTR retroelement reverse transcriptase
[Oryza sativa (japonica cultivar-group)]
gi|10140689|gb|AAG13524.1| putative non-LTR retroelement
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1382
Score = 128 bits (321), Expect = 6e-28
Identities = 60/125 (48%), Positives = 88/125 (70%), Gaps = 2/125 (1%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ANR+K LP+I ++FQS FVPGRLITD++L+ +E H IRK N N + +K+D+ KA
Sbjct: 537 LANRLKPFLPDIVSEFQSAFVPGRLITDSALVAYECLHTIRKQH-NKNPFFALKIDMMKA 595
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++W ++ L+ +GF + INT+M C+ V +++ ING+ T+P P RGIRQGDP
Sbjct: 596 YDRVEWAYLSGCLSKLGFSQDWINTVMRCVSSVRYAVKINGELTKPVVPSRGIRQGDPIS 655
Query: 121 PLIFL 125
P +FL
Sbjct: 656 PYLFL 660
>gb|AAP52553.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|37531928|ref|NP_920266.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|22138478|gb|AAM93462.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 566
Score = 122 bits (306), Expect = 3e-26
Identities = 60/125 (48%), Positives = 79/125 (63%), Gaps = 1/125 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ANR+K ILP I D QS FVPGRLITDN LI +E H+++ R +GY IKLD++KA
Sbjct: 42 LANRLKSILPEIILDTQSAFVPGRLITDNVLIAYEATHFLQNKRNGKDGYAAIKLDMSKA 101
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++W F+ L +GF IM C+ V + I +NG TE P RG+RQGDP
Sbjct: 102 YDRVEWPFLLHMLRRLGFDEKWNRLIMNCVTTVNYKIKVNGDYTEVIKPDRGLRQGDPLS 161
Query: 121 PLIFL 125
P +F+
Sbjct: 162 PYLFV 166
>gb|AAP52395.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37531612|ref|NP_920108.1| putative retroelement [Oryza
sativa (japonica cultivar-group)]
Length = 1694
Score = 122 bits (306), Expect = 3e-26
Identities = 56/126 (44%), Positives = 82/126 (64%), Gaps = 1/126 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ NR++ IL ++ + QS FVPGRLITDN+LI FE FH+I++ + N Y KLD++KA
Sbjct: 1135 LVNRLRPILDDLVSQNQSAFVPGRLITDNALIAFEYFHHIQRNKNPENAYSAYKLDLSKA 1194
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD + W F+E + +GF + IM CI V +++ +NG F+P RG+RQGDP
Sbjct: 1195 YDRVDWEFLEQAMVKLGFAHCWVKWIMACITLVRYAVKLNGTLLNTFAPSRGLRQGDPLS 1254
Query: 121 PLIFLY 126
P +FL+
Sbjct: 1255 PFLFLF 1260
>gb|AAM01179.2| Putative retroelement [Oryza sativa (japonica cultivar-group)]
Length = 1888
Score = 122 bits (306), Expect = 3e-26
Identities = 56/126 (44%), Positives = 82/126 (64%), Gaps = 1/126 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ NR++ IL ++ + QS FVPGRLITDN+LI FE FH+I++ + N Y KLD++KA
Sbjct: 1092 LVNRLRPILDDLVSQNQSAFVPGRLITDNALIAFEYFHHIQRNKNPENAYSAYKLDLSKA 1151
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD + W F+E + +GF + IM CI V +++ +NG F+P RG+RQGDP
Sbjct: 1152 YDRVDWEFLEQAMVKLGFAHCWVKWIMACITLVRYAVKLNGTLLNTFAPSRGLRQGDPLS 1211
Query: 121 PLIFLY 126
P +FL+
Sbjct: 1212 PFLFLF 1217
>ref|XP_468607.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|30017567|gb|AAP12989.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 818
Score = 120 bits (302), Expect = 1e-25
Identities = 55/126 (43%), Positives = 83/126 (65%), Gaps = 1/126 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ NR++ +L ++ ++ QS FVPGR+ITDN+LI FE FHYI+K + + +KLD++KA
Sbjct: 42 LVNRLRPLLHDLISENQSAFVPGRMITDNALIAFECFHYIQKCNRESQMFCALKLDLSKA 101
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
+D + W+F++ L +GF IM C+ V +S+ NG EPF P RG+RQGDP
Sbjct: 102 FDRVDWDFLDGVLLRLGFAEKWRKWIMSCVTFVQYSVRFNGNLLEPFKPSRGLRQGDPLS 161
Query: 121 PLIFLY 126
P +FL+
Sbjct: 162 PYLFLF 167
>gb|AAG51783.1| reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
gi|25405244|pir||E96519 probable reverse transcriptase,
16838-20266 [imported] - Arabidopsis thaliana
Length = 1142
Score = 119 bits (297), Expect = 4e-25
Identities = 59/125 (47%), Positives = 82/125 (65%), Gaps = 1/125 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ R+K +LPN+ ++ QS FV GRLI+DN LI E FH +R + + ++ IK D++KA
Sbjct: 305 LCQRLKTVLPNLISETQSAFVDGRLISDNILIAQEMFHGLRTNSSCKDKFMAIKTDMSKA 364
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++WNFIE L MGF I+ IM CI V + +LINGQP P+RG+RQGDP
Sbjct: 365 YDQVEWNFIEALLRKMGFCEKWISWIMWCITTVQYKVLINGQPKGLIIPERGLRQGDPLS 424
Query: 121 PLIFL 125
P +F+
Sbjct: 425 PYLFI 429
>emb|CAE04866.2| OSJNBa0086O06.14 [Oryza sativa (japonica cultivar-group)]
gi|50928373|ref|XP_473714.1| OSJNBa0086O06.14 [Oryza
sativa (japonica cultivar-group)]
Length = 1205
Score = 117 bits (292), Expect = 1e-24
Identities = 54/126 (42%), Positives = 80/126 (62%), Gaps = 1/126 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ NR++ +L I ++ QS FVPGR+ITDN+L+ FE FH I K + + +KLD++KA
Sbjct: 509 LVNRLRPLLQEIISETQSAFVPGRMITDNALVAFECFHSIHKCTRESQDFCALKLDLSKA 568
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD + W F++ L +GF IM C+ V +S+ +NG EPF P RG+R+GDP
Sbjct: 569 YDRVDWGFLDGALQKLGFGNIWRKWIMSCVTSVRYSVRLNGNMLEPFYPTRGLREGDPLN 628
Query: 121 PLIFLY 126
P +FL+
Sbjct: 629 PYLFLF 634
>gb|AAD21778.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25410938|pir||G84429 hypothetical protein
At2g01840 [imported] - Arabidopsis thaliana
Length = 1715
Score = 115 bits (289), Expect = 3e-24
Identities = 53/126 (42%), Positives = 79/126 (62%), Gaps = 1/126 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ R+K L ++ +D Q+ FVPG+ I+DN L+ E H ++ R +GYV +K DI+KA
Sbjct: 883 LIKRLKQCLGDVISDSQAAFVPGQNISDNVLVAHELLHSLKSRRECQSGYVAVKTDISKA 942
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++WNF+E + +GF + IM C+ V++ +LING P P RGIRQGDP
Sbjct: 943 YDRVEWNFLEKVMIQLGFAPRWVKWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGDPLS 1002
Query: 121 PLIFLY 126
P +FL+
Sbjct: 1003 PYLFLF 1008
>ref|XP_475290.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|49328177|gb|AAT58873.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 1264
Score = 115 bits (289), Expect = 3e-24
Identities = 54/125 (43%), Positives = 80/125 (63%), Gaps = 1/125 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
I NR+K +L I + QS FVP RLIT N L+ +E HY+ + + NG IKLD++KA
Sbjct: 869 IVNRLKVVLLEIISPSQSAFVPRRLITYNVLLAYELTHYLNQRKKGKNGVAAIKLDMSKA 928
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++W+F+ + +GF +N +M C+ VT+ I ING+ ++ PQRG+RQGDP
Sbjct: 929 YDRVEWDFLRHMMLRLGFHDQWVNLVMKCVTSVTYRIKINGEHSDQIYPQRGLRQGDPLS 988
Query: 121 PLIFL 125
P +F+
Sbjct: 989 PYLFI 993
>gb|AAV32224.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|45267888|gb|AAS55787.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 1936
Score = 115 bits (287), Expect = 5e-24
Identities = 60/163 (36%), Positives = 95/163 (57%), Gaps = 4/163 (2%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ NR++ IL ++ + QS F+ GR+ITDN+L+ FE FH I+K + N+ KLD++KA
Sbjct: 1161 LVNRLRPILDDLVSKEQSAFIQGRMITDNALLAFECFHSIQKNKKANSAACAYKLDLSKA 1220
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD + W F+E L +GF ++ IMLC+ V +S+ NG F+P RG+RQG+P
Sbjct: 1221 YDRVDWRFLELALNKLGFAHRWVSWIMLCVTTVRYSVKFNGTLLRSFAPTRGLRQGEPLS 1280
Query: 121 PLIFLYYVLRFFLV*SQNVKMKV*FMVSPLPLMPQLYLIYFML 163
P +FL+ L+ + V ++PL + Q I ++L
Sbjct: 1281 PFLFLFVADGLSLLLKEKVAQN---SLTPLKICRQAPGISYLL 1320
>pir||G96509 protein F27F5.21 [imported] - Arabidopsis thaliana
gi|7767672|gb|AAF69169.1| F27F5.21 [Arabidopsis
thaliana]
Length = 1023
Score = 114 bits (285), Expect = 9e-24
Identities = 57/125 (45%), Positives = 81/125 (64%), Gaps = 1/125 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ R+K ILP++ ++ S FV GRLI+DN LI E FH +R + ++ IK D++KA
Sbjct: 371 LCQRLKKILPSLISETHSAFVEGRLISDNILIAQEMFHGLRTNPSCKGKFMAIKTDMSKA 430
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++WNFIE+ L MGF I+ IM CI V + +L+NGQP P+RG+RQGDP
Sbjct: 431 YDRVEWNFIEELLKKMGFCEKWISWIMWCITTVQYRVLLNGQPRGLIVPKRGLRQGDPLS 490
Query: 121 PLIFL 125
P +F+
Sbjct: 491 PYLFI 495
>emb|CAE04660.2| OSJNBa0061G20.16 [Oryza sativa (japonica cultivar-group)]
Length = 1285
Score = 114 bits (284), Expect = 1e-23
Identities = 53/126 (42%), Positives = 80/126 (63%), Gaps = 1/126 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ NR++ IL ++ + QS F+ GR+ITDN+L+ FE FH I+K + N+ KLD++KA
Sbjct: 1073 LVNRLRPILDDLVSKEQSAFIQGRMITDNALLAFECFHSIQKNKKANSAACAYKLDLSKA 1132
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD + W F+E L +GF ++ IM C+ V +S+ NG F+P RG+RQGDP
Sbjct: 1133 YDRVDWRFLELALNKLGFAHRWVSWIMSCVTTVRYSVKFNGTLLRSFAPTRGLRQGDPLS 1192
Query: 121 PLIFLY 126
P +FL+
Sbjct: 1193 PFLFLF 1198
>gb|AAQ19327.1| bZIP-like protein [Oryza sativa (japonica cultivar-group)]
Length = 2367
Score = 112 bits (281), Expect = 3e-23
Identities = 52/126 (41%), Positives = 80/126 (63%), Gaps = 1/126 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ NR++ IL ++ + QS FV GR+ITDN+L+ FE FH ++K + N+ KLD++KA
Sbjct: 1255 LVNRLRPILDDLVSVEQSAFVQGRMITDNALLAFECFHAMQKNKKANHAACAYKLDLSKA 1314
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDP-F 120
YD + W F+E + +GF +N IM C+ V + + NG + F+P RG+RQGDP
Sbjct: 1315 YDRVDWRFLEMAMNKLGFARRWVNWIMKCVTSVRYMVKFNGTLLQSFAPTRGLRQGDPLL 1374
Query: 121 PLIFLY 126
P +FL+
Sbjct: 1375 PFLFLF 1380
>gb|AAB82639.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408936|pir||A84888 hypothetical protein
At2g45230 [imported] - Arabidopsis thaliana
Length = 1374
Score = 111 bits (278), Expect = 6e-23
Identities = 53/125 (42%), Positives = 79/125 (62%), Gaps = 1/125 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ANR+K ILP++ ++ Q+ FV GRLI+DN LI E H + + ++ IK DI+KA
Sbjct: 521 MANRLKKILPSLISETQAAFVKGRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKA 580
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++W F+E + +GF + I IM C++ V + +LING P P RG+RQGDP
Sbjct: 581 YDRVEWPFLEKAMRGLGFADHWIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLS 640
Query: 121 PLIFL 125
P +F+
Sbjct: 641 PYLFV 645
>emb|CAB81184.1| putative protein [Arabidopsis thaliana] gi|4539357|emb|CAB40051.1|
putative protein [Arabidopsis thaliana]
gi|7486147|pir||T04278 hypothetical protein F25I24.40 -
Arabidopsis thaliana
Length = 1294
Score = 110 bits (275), Expect = 1e-22
Identities = 51/125 (40%), Positives = 77/125 (60%), Gaps = 1/125 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ R+K L I +D Q+ F+PGRL+ DN +I E H ++ + + Y+ +K D++KA
Sbjct: 883 LVERLKGHLDAIVSDSQAAFIPGRLVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKA 942
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++WNF+E T+ GF I IM ++ V +S+L+NG P PQRGIRQGDP
Sbjct: 943 YDRVEWNFLETTMRLFGFSETWIKWIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLS 1002
Query: 121 PLIFL 125
P +F+
Sbjct: 1003 PYLFI 1007
>gb|AAC33961.1| contains similarity to reverse trancriptase (Pfam: rvt.hmm, score:
42.57) [Arabidopsis thaliana] gi|7486711|pir||T01893
hypothetical protein F8M12.22 - Arabidopsis thaliana
Length = 1662
Score = 110 bits (275), Expect = 1e-22
Identities = 51/125 (40%), Positives = 77/125 (60%), Gaps = 1/125 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ R+K L I +D Q+ F+PGRL+ DN +I E H ++ + + Y+ +K D++KA
Sbjct: 903 LVERLKGHLDAIVSDSQAAFIPGRLVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKA 962
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++WNF+E T+ GF I IM ++ V +S+L+NG P PQRGIRQGDP
Sbjct: 963 YDRVEWNFLETTMRLFGFSETWIKWIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLS 1022
Query: 121 PLIFL 125
P +F+
Sbjct: 1023 PYLFI 1027
>gb|AAB48346.1| reverse transcriptase
Length = 199
Score = 110 bits (275), Expect = 1e-22
Identities = 50/118 (42%), Positives = 73/118 (61%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ R+K L ++ +D Q+ FVPGR I+DN L+ E H ++ R +GYV +K DI+KA
Sbjct: 77 LIKRLKQCLGDVISDSQAAFVPGRNISDNVLVAHELLHSLKSRRECQSGYVAVKTDISKA 136
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDP 119
YD ++WNF+E + +GF + IM C+ V++ +LING P P RGIRQG P
Sbjct: 137 YDRVEWNFLEKVMIQLGFAPRWVKWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGYP 194
>gb|AAD17398.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25411644|pir||C84530 hypothetical protein
At2g15540 [imported] - Arabidopsis thaliana
Length = 1225
Score = 109 bits (273), Expect = 2e-22
Identities = 53/125 (42%), Positives = 78/125 (62%), Gaps = 1/125 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+++R+K +LP + ++ QS FV RLITDN LI E FH +R Y+ IK D++KA
Sbjct: 468 LSSRLKRLLPELISETQSAFVAERLITDNILIAQENFHALRTNPACKKKYMAIKTDMSKA 527
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++W+F+ + MGF ++ I+ CI V++ IL+NG P P RGIRQGDP
Sbjct: 528 YDRVEWSFLRALMLKMGFAQKWVDWIIFCISSVSYKILLNGSPKGFIKPSRGIRQGDPIS 587
Query: 121 PLIFL 125
P +F+
Sbjct: 588 PFLFI 592
>emb|CAB75484.1| putative protein [Arabidopsis thaliana] gi|11357921|pir||T47495
hypothetical protein F9K21.130 - Arabidopsis thaliana
Length = 851
Score = 109 bits (273), Expect = 2e-22
Identities = 52/125 (41%), Positives = 79/125 (62%), Gaps = 1/125 (0%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ NR+K L +I +D Q+ F+PGR+I DN +I E H ++ + + Y+ +K D++KA
Sbjct: 142 MVNRLKAHLNSIVSDSQAAFIPGRIINDNVMIAHEIMHSLKVRKRVSKTYMAVKTDVSKA 201
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
YD ++W+F+E T+ GF I IM ++ V +S+LING P SP RGIRQGDP
Sbjct: 202 YDRVEWDFLETTMRLFGFCDKWIGWIMAAVKSVHYSVLINGSPHGYISPTRGIRQGDPLS 261
Query: 121 PLIFL 125
P +F+
Sbjct: 262 PYLFI 266
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.344 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 936,949,877
Number of Sequences: 2540612
Number of extensions: 36550973
Number of successful extensions: 114487
Number of sequences better than 10.0: 666
Number of HSP's better than 10.0 without gapping: 490
Number of HSP's successfully gapped in prelim test: 176
Number of HSP's that attempted gapping in prelim test: 113610
Number of HSP's gapped (non-prelim): 759
length of query: 600
length of database: 863,360,394
effective HSP length: 134
effective length of query: 466
effective length of database: 522,918,386
effective search space: 243679967876
effective search space used: 243679967876
T: 11
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 78 (34.7 bits)
Medicago: description of AC119408.1


