

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC087771.6 + phase: 0 /pseudo/partial
(461 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
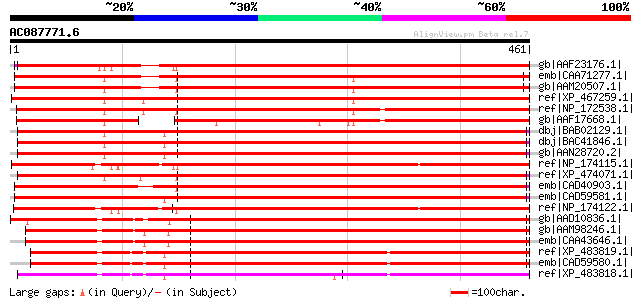
Score E
Sequences producing significant alignments: (bits) Value
gb|AAF23176.1| P-glycoprotein [Gossypium hirsutum] 610 e-173
emb|CAA71277.1| P-glycoprotein-2 [Arabidopsis thaliana] gi|21082... 604 e-171
gb|AAM20507.1| P-glycoprotein-2 [Arabidopsis thaliana] 604 e-171
ref|XP_467259.1| MDR-like ABC transporter [Oryza sativa (japonic... 574 e-162
ref|NP_172538.1| P-glycoprotein, putative [Arabidopsis thaliana] 563 e-159
gb|AAF17668.1| F20B24.12 [Arabidopsis thaliana] gi|25297460|pir|... 461 e-128
dbj|BAB02129.1| P-glycoprotein; multi-drug resistance related; A... 393 e-108
dbj|BAC41846.1| putative P-glycoprotein [Arabidopsis thaliana] 393 e-108
gb|AAN28720.2| MDR-like p-glycoprotein [Arabidopsis thaliana] 393 e-108
ref|NP_174115.1| multidrug resistance P-glycoprotein, putative [... 380 e-104
ref|XP_474071.1| OSJNBb0079B02.13 [Oryza sativa (japonica cultiv... 377 e-103
emb|CAD40903.1| OSJNBa0036B21.21 [Oryza sativa (japonica cultiva... 377 e-103
emb|CAD59581.1| MDR-like ABC transporter [Oryza sativa (japonica... 371 e-101
ref|NP_174122.1| multidrug resistance P-glycoprotein, putative [... 371 e-101
gb|AAD10836.1| P-glycoprotein [Solanum tuberosum] 357 6e-97
gb|AAM98246.1| putative ABC transporter [Arabidopsis thaliana] 355 2e-96
emb|CAA43646.1| P-glycoprotein [Arabidopsis thaliana] gi|4883607... 355 2e-96
ref|XP_483819.1| putative P-glycoprotein 1 [Oryza sativa (japoni... 348 2e-94
emb|CAD59580.1| MDR-like ABC transporter [Oryza sativa (japonica... 344 4e-93
ref|XP_483818.1| putative P-glycoprotein 1 [Oryza sativa (japoni... 342 1e-92
>gb|AAF23176.1| P-glycoprotein [Gossypium hirsutum]
Length = 1249
Score = 610 bits (1574), Expect = e-173
Identities = 323/477 (67%), Positives = 372/477 (77%), Gaps = 38/477 (7%)
Query: 8 ERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFP 67
++KK+ KV +LKLFSFAD YD+VLM +GS+GA VHGASVP+FFIFFGKLIN+IG+AYLFP
Sbjct: 21 KKKKQRKVPLLKLFSFADFYDHVLMGLGSLGACVHGASVPVFFIFFGKLINIIGMAYLFP 80
Query: 68 KEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y*GI 124
KEASHKVAK + + IL + A W T + + Y+KS
Sbjct: 81 KEASHKVAKYSLDFVYLSVAILFSSWIEVACWMHTGERQAAKMRMAYLKSMLN------- 133
Query: 125 HWRGHFCYYK*HYYCSRCSIRE--------------------GNFLHYISRFIAGFTIGF 164
+ + S E GNF+HYISRFIAGF+IGF
Sbjct: 134 --------QDISLFDTEASTGEVISAITSDIIVVQDALSEKVGNFMHYISRFIAGFSIGF 185
Query: 165 VRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFA 224
RVWQISLVTLSIVP IALAGG YAYV GLIA+VR +YV+AGEIAEEVIGNVRTVQAFA
Sbjct: 186 ARVWQISLVTLSIVPLIALAGGIYAYVATGLIARVRNSYVKAGEIAEEVIGNVRTVQAFA 245
Query: 225 GEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANG 284
GEERAV+SYK ALM TY G+KAGL KGLGLGS+HCVLF+SWALLVW+TS+VVHKNIANG
Sbjct: 246 GEERAVKSYKDALMNTYTYGKKAGLTKGLGLGSLHCVLFVSWALLVWFTSIVVHKNIANG 305
Query: 285 GESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKL 344
G+SFTTMLNVVISGLSLGQAAPDISAFIRA+AAAYPIFEMIER+TVSK SSKTGRKLSK+
Sbjct: 306 GDSFTTMLNVVISGLSLGQAAPDISAFIRARAAAYPIFEMIERNTVSKTSSKTGRKLSKV 365
Query: 345 DGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPIS 404
+G+I+ +V FSYPSRPDV IF L+IP GKIVALVGGSGSGKSTV+SLIERFYEP++
Sbjct: 366 EGNIELKNVSFSYPSRPDVVIFDRFCLNIPTGKIVALVGGSGSGKSTVISLIERFYEPLA 425
Query: 405 GQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
G+ILLD N+I+ LDLKWLRQQIGLVNQEPALFAT+I+ENILYGKDDAT++E+ RA K
Sbjct: 426 GEILLDGNNIKGLDLKWLRQQIGLVNQEPALFATTIRENILYGKDDATVDEITRAAK 482
Score = 217 bits (553), Expect = 5e-55
Identities = 149/479 (31%), Positives = 246/479 (51%), Gaps = 41/479 (8%)
Query: 5 EGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAY 64
+G + K+ VS +L+S D+ F G++ A++ GA +P+F + + + +AY
Sbjct: 658 DGIDAGKQPYVSPGRLYSMIGP-DWYYGFFGTVTALIAGAQMPLFALGVSQAL----VAY 712
Query: 65 LFPKEAS-HKVAKSK----CSNIIFILDRG------GLLDAYWRKTSSKDENGLFKVYVK 113
E + H+V K C+++I ++ G++ + + + G+F +K
Sbjct: 713 YMDWETTCHEVKKIAILFCCASVITVIVHAIEHLCFGIMG---ERLTLRVREGMFSAILK 769
Query: 114 SRYKFI*Y*GIHWRGHFCYYK*HYYCSRCSI-----------REGNFLHYISRFIAGFTI 162
+ I W SR R + + IA F I
Sbjct: 770 NE--------IGWFDDLNNAS-SMLASRLETDATFLRGVVVDRTSILIQNVGLVIAAFII 820
Query: 163 GFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQA 222
F+ W+I+L+ L+ P I G + KAY++A IA E + N+RTV A
Sbjct: 821 AFILNWRITLIILATFPLIISGHISEKLFMQGYGGNLSKAYLKANMIAGEAVSNMRTVAA 880
Query: 223 FAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIA 282
F EE+ + Y L++ K G G+ G +F S+ L +WY SV++ K +A
Sbjct: 881 FCAEEKILDLYARELIEPSERSFKRGQIAGIFYGISQFFIFSSYGLALWYGSVLMGKELA 940
Query: 283 NGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLS 342
+ + + ++++ L++G+ + ++ +FE+++R T + G +L+
Sbjct: 941 SFKSVMKSFMVLIVTALAMGETLALVPDLLKGNQMVASVFEIMDRKT--QVVGDAGEELT 998
Query: 343 KLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEP 402
++G I+ V FSYPSRPDV IF + +L + +GK +ALVG SGSGKS+V++LI RFY+P
Sbjct: 999 NVEGTIELKGVHFSYPSRPDVVIFKDFDLKVRSGKSMALVGQSGSGKSSVLALILRFYDP 1058
Query: 403 ISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
SG++++D D+++L LK LR+ IGLV QEPALFATSI ENILYGK+ A+ E+ A K
Sbjct: 1059 TSGKVMIDGRDVKKLKLKSLRKHIGLVQQEPALFATSIYENILYGKEGASESEVVEAAK 1117
>emb|CAA71277.1| P-glycoprotein-2 [Arabidopsis thaliana] gi|2108254|emb|CAA71276.1|
P-glycoprotein-2 [Arabidopsis thaliana]
gi|7269447|emb|CAB79451.1| P-glycoprotein-2 (pgp2)
[Arabidopsis thaliana] gi|4538925|emb|CAB39661.1|
P-glycoprotein-2 (pgp2) [Arabidopsis thaliana]
gi|15236067|ref|NP_194326.1| multidrug resistance
P-glycoprotein, putative [Arabidopsis thaliana]
gi|7442648|pir||T04251 P-glycoprotein 2 - Arabidopsis
thaliana
Length = 1233
Score = 604 bits (1558), Expect = e-171
Identities = 323/480 (67%), Positives = 367/480 (76%), Gaps = 38/480 (7%)
Query: 5 EGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAY 64
E ++ + KVS+LKLFSFAD YD VLM +GS+GA +HGASVPIFFIFFGKLIN+IGLAY
Sbjct: 10 EKEKEMTQPKVSLLKLFSFADFYDCVLMTLGSVGACIHGASVPIFFIFFGKLINIIGLAY 69
Query: 65 LFPKEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y 121
LFPK+ASH+VAK + + IL L A W T + + + Y++S
Sbjct: 70 LFPKQASHRVAKYSLDFVYLSVAILFSSWLEVACWMHTGERQAAKMRRAYLRSMLS---- 125
Query: 122 *GIHWRGHFCYYK*HYYCSRCSIRE--------------------GNFLHYISRFIAGFT 161
+ + S E GNFLHYISRFIAGF
Sbjct: 126 -----------QDISLFDTEASTGEVISAITSDILVVQDALSEKVGNFLHYISRFIAGFA 174
Query: 162 IGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQ 221
IGF VWQISLVTLSIVP IALAGG YA+V IGLIA+VRK+Y++AGEIAEEVIGNVRTVQ
Sbjct: 175 IGFTSVWQISLVTLSIVPLIALAGGIYAFVAIGLIARVRKSYIKAGEIAEEVIGNVRTVQ 234
Query: 222 AFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNI 281
AF GEERAVR Y+ AL TY GRKAGL KGLGLGSMHCVLFLSWALLVW+TSVVVHK+I
Sbjct: 235 AFTGEERAVRLYREALENTYKYGRKAGLTKGLGLGSMHCVLFLSWALLVWFTSVVVHKDI 294
Query: 282 ANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKL 341
A+GG+SFTTMLNVVI+GLSLGQAAPDISAF+RAKAAAYPIF+MIER+TV+K S+K+GRKL
Sbjct: 295 ADGGKSFTTMLNVVIAGLSLGQAAPDISAFVRAKAAAYPIFKMIERNTVTKTSAKSGRKL 354
Query: 342 SKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYE 401
K+DGHIQF D FSYPSRPDV IF LNL IPAGKIVALVGGSGSGKSTV+SLIERFYE
Sbjct: 355 GKVDGHIQFKDATFSYPSRPDVVIFDRLNLAIPAGKIVALVGGSGSGKSTVISLIERFYE 414
Query: 402 PISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
PISG +LLD N+I ELD+KWLR QIGLVNQEPALFAT+I+ENILYGKDDAT EE+ RA K
Sbjct: 415 PISGAVLLDGNNISELDIKWLRGQIGLVNQEPALFATTIRENILYGKDDATAEEITRAAK 474
Score = 209 bits (533), Expect = 1e-52
Identities = 116/310 (37%), Positives = 184/310 (58%), Gaps = 8/310 (2%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L + + F I F+ W+++LV L+ P + G + KAY++A +
Sbjct: 794 LQNLGLVVTSFIIAFILNWRLTLVVLATYPLVISGHISEKLFMQGYGGDLNKAYLKANML 853
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF EE+ + Y L++ + + G GL G +F S+ L
Sbjct: 854 AGESVSNIRTVAAFCAEEKILELYSRELLEPSKSSFRRGQIAGLFYGVSQFFIFSSYGLA 913
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFEMIE 326
+WY S ++ K +A T + ++++ L++G+ APD+ ++ +FE+++
Sbjct: 914 LWYGSTLMDKGLAGFKSVMKTFMVLIVTALAMGETLALAPDL---LKGNQMVASVFEILD 970
Query: 327 RDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSG 386
R T + +T +L+ ++G I+ V FSYPSRPDV IF + +L + AGK +ALVG SG
Sbjct: 971 RKT--QIVGETSEELNNVEGTIELKGVHFSYPSRPDVVIFRDFDLIVRAGKSMALVGQSG 1028
Query: 387 SGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILY 446
SGKS+V+SLI RFY+P +G+++++ DI++LDLK LR+ IGLV QEPALFAT+I ENILY
Sbjct: 1029 SGKSSVISLILRFYDPTAGKVMIEGKDIKKLDLKALRKHIGLVQQEPALFATTIYENILY 1088
Query: 447 GKDDATLEEL 456
G + A+ E+
Sbjct: 1089 GNEGASQSEV 1098
>gb|AAM20507.1| P-glycoprotein-2 [Arabidopsis thaliana]
Length = 1233
Score = 604 bits (1558), Expect = e-171
Identities = 323/480 (67%), Positives = 367/480 (76%), Gaps = 38/480 (7%)
Query: 5 EGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAY 64
E ++ + KVS+LKLFSFAD YD VLM +GS+GA +HGASVPIFFIFFGKLIN+IGLAY
Sbjct: 10 EKEKEMTQPKVSLLKLFSFADFYDCVLMTLGSVGACIHGASVPIFFIFFGKLINIIGLAY 69
Query: 65 LFPKEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y 121
LFPK+ASH+VAK + + IL L A W T + + + Y++S
Sbjct: 70 LFPKQASHRVAKYSLDFVYLSVAILFSSWLEVACWMHTGERQAAKMRRAYLRSMLS---- 125
Query: 122 *GIHWRGHFCYYK*HYYCSRCSIRE--------------------GNFLHYISRFIAGFT 161
+ + S E GNFLHYISRFIAGF
Sbjct: 126 -----------QDISLFDTEASTGEVISAITSDILVVQDALSEKVGNFLHYISRFIAGFA 174
Query: 162 IGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQ 221
IGF VWQISLVTLSIVP IALAGG YA+V IGLIA+VRK+Y++AGEIAEEVIGNVRTVQ
Sbjct: 175 IGFTSVWQISLVTLSIVPLIALAGGIYAFVAIGLIARVRKSYIKAGEIAEEVIGNVRTVQ 234
Query: 222 AFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNI 281
AF GEERAVR Y+ AL TY GRKAGL KGLGLGSMHCVLFLSWALLVW+TSVVVHK+I
Sbjct: 235 AFTGEERAVRLYREALENTYKYGRKAGLTKGLGLGSMHCVLFLSWALLVWFTSVVVHKDI 294
Query: 282 ANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKL 341
A+GG+SFTTMLNVVI+GLSLGQAAPDISAF+RAKAAAYPIF+MIER+TV+K S+K+GRKL
Sbjct: 295 ADGGKSFTTMLNVVIAGLSLGQAAPDISAFVRAKAAAYPIFKMIERNTVTKTSAKSGRKL 354
Query: 342 SKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYE 401
K+DGHIQF D FSYPSRPDV IF LNL IPAGKIVALVGGSGSGKSTV+SLIERFYE
Sbjct: 355 GKVDGHIQFKDATFSYPSRPDVVIFDRLNLAIPAGKIVALVGGSGSGKSTVISLIERFYE 414
Query: 402 PISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
PISG +LLD N+I ELD+KWLR QIGLVNQEPALFAT+I+ENILYGKDDAT EE+ RA K
Sbjct: 415 PISGAVLLDGNNISELDIKWLRGQIGLVNQEPALFATTIRENILYGKDDATAEEITRAAK 474
Score = 209 bits (533), Expect = 1e-52
Identities = 116/310 (37%), Positives = 184/310 (58%), Gaps = 8/310 (2%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L + + F I F+ W+++LV L+ P + G + KAY++A +
Sbjct: 794 LQNLGLVVTSFIIAFILNWRLTLVVLATYPLVISGHISEKLFMQGYGGDLNKAYLKANML 853
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF EE+ + Y L++ + + G GL G +F S+ L
Sbjct: 854 AGESVSNIRTVAAFCAEEKILELYSRELLEPSKSSFRRGQIAGLFYGVSQFFIFSSYGLA 913
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFEMIE 326
+WY S ++ K +A T + ++++ L++G+ APD+ ++ +FE+++
Sbjct: 914 LWYGSTLMDKGLAGFKSVMKTFMVLIVTALAMGETLALAPDL---LKGNQMVASVFEILD 970
Query: 327 RDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSG 386
R T + +T +L+ ++G I+ V FSYPSRPDV IF + +L + AGK +ALVG SG
Sbjct: 971 RKT--QIVGETSEELNNVEGTIELKGVHFSYPSRPDVVIFRDFDLIVRAGKSMALVGQSG 1028
Query: 387 SGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILY 446
SGKS+V+SLI RFY+P +G+++++ DI++LDLK LR+ IGLV QEPALFAT+I ENILY
Sbjct: 1029 SGKSSVISLILRFYDPTAGKVMIEGKDIKKLDLKALRKHIGLVQQEPALFATTIYENILY 1088
Query: 447 GKDDATLEEL 456
G + A+ E+
Sbjct: 1089 GNEGASQSEV 1098
>ref|XP_467259.1| MDR-like ABC transporter [Oryza sativa (japonica cultivar-group)]
gi|27368851|emb|CAD59583.1| MDR-like ABC transporter
[Oryza sativa (japonica cultivar-group)]
gi|41053280|dbj|BAD07706.1| MDR-like ABC transporter
[Oryza sativa (japonica cultivar-group)]
gi|41052997|dbj|BAD07906.1| MDR-like ABC transporter
[Oryza sativa (japonica cultivar-group)]
Length = 1264
Score = 574 bits (1480), Expect = e-162
Identities = 303/468 (64%), Positives = 359/468 (75%), Gaps = 8/468 (1%)
Query: 2 ESKEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIG 61
E E + K KV LKLFSFAD +DYVLM +GS+GA HGASVP+FFIFFGKLIN+IG
Sbjct: 22 EQGEKEAAAKVEKVPFLKLFSFADRWDYVLMAVGSLGACAHGASVPVFFIFFGKLINIIG 81
Query: 62 LAYLFPKEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYK- 117
LAYLFP S +VAK + I IL A W T + + + Y++S
Sbjct: 82 LAYLFPTTVSGRVAKYSLDFVYLGIVILFSSWTEVACWMHTGERQAAKMRQAYLRSMLDQ 141
Query: 118 ----FI*Y*GIHWRGHFCYYK*HYYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLV 173
F + S + GNF+HYISRF+AGF IGF +VWQISLV
Sbjct: 142 DIAVFDTEASTGEVINAITSDILVVQDAISEKVGNFMHYISRFLAGFAIGFSQVWQISLV 201
Query: 174 TLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSY 233
TL+IVP IA+AGG YAYVTIGL+A+VRK+YV+AGEIAEEVIGNVRTVQAF GEE+AVR+Y
Sbjct: 202 TLAIVPLIAIAGGIYAYVTIGLMARVRKSYVKAGEIAEEVIGNVRTVQAFVGEEKAVRTY 261
Query: 234 KAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLN 293
+ AL++TY G++ GLAKGLGLGSMH VLFLSWALL+W+TSVVVHKNI+NGGESFTTMLN
Sbjct: 262 REALLRTYKYGKRGGLAKGLGLGSMHSVLFLSWALLIWFTSVVVHKNISNGGESFTTMLN 321
Query: 294 VVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDV 353
VVI+GLSLGQAAP+IS F+RA+ AAYPIF+MIER+TV+K SSK GR L +DGHIQF DV
Sbjct: 322 VVIAGLSLGQAAPNISTFLRARTAAYPIFQMIERNTVNKASSKAGRTLPSVDGHIQFRDV 381
Query: 354 CFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKND 413
F+YPSRPDV I +LD PAGKIVALVGGSGSGKSTVVSLIERFYEP++G +LLD +D
Sbjct: 382 RFAYPSRPDVVILDRFSLDFPAGKIVALVGGSGSGKSTVVSLIERFYEPLTGAVLLDGHD 441
Query: 414 IRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
I++LD+KWLRQQIGLVNQEPALFATSI+ENILYGK DA+++E+ A K
Sbjct: 442 IKDLDVKWLRQQIGLVNQEPALFATSIRENILYGKGDASMDEINHAAK 489
Score = 208 bits (530), Expect = 3e-52
Identities = 122/315 (38%), Positives = 183/315 (57%), Gaps = 8/315 (2%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L I + I F+ W+I+LV L+ P + G + K+Y++A +
Sbjct: 818 LQNIGMIVTSLIIAFIINWRITLVVLATYPLMVSGHISEKMFMKGYGGNLGKSYLKANML 877
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF EE+ ++ Y L + + G GL G LF S+AL
Sbjct: 878 AAEAVSNIRTVAAFCAEEKVIKLYADELKEPAKQSFRRGQGAGLFYGVSQFFLFSSYALA 937
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFEMIE 326
+WY S ++ K +A+ + + ++++ L++G+ APDI I+ +FE+++
Sbjct: 938 LWYGSELMSKEMASFKSVMKSFMVLIVTALAMGETLAMAPDI---IKGNQMVSSVFEILD 994
Query: 327 RDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSG 386
R T G + +++G I+ V F YP+RP+V +F L+L + AGK +ALVG SG
Sbjct: 995 RKT--DVLIDAGNDVKRVEGVIELRGVEFRYPARPEVVVFKGLDLLMKAGKSMALVGMSG 1052
Query: 387 SGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILY 446
SGKSTV+SLI RFY+PI+G++L+D DIR++ LK LR+ IGLV QEPALFAT+I +NILY
Sbjct: 1053 SGKSTVLSLILRFYDPIAGKVLIDGKDIRKVKLKSLRKHIGLVQQEPALFATTIYDNILY 1112
Query: 447 GKDDATLEELKRAVK 461
GKD AT E+ A K
Sbjct: 1113 GKDGATEAEVVDAAK 1127
>ref|NP_172538.1| P-glycoprotein, putative [Arabidopsis thaliana]
Length = 1227
Score = 563 bits (1450), Expect = e-159
Identities = 304/463 (65%), Positives = 349/463 (74%), Gaps = 12/463 (2%)
Query: 7 DERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLF 66
++ KK VS LKLFSFAD YD VLM +GSIGA +HGASVP+FFIFFGKLIN+IGLAYLF
Sbjct: 16 EKEKKRPSVSFLKLFSFADFYDCVLMALGSIGACIHGASVPVFFIFFGKLINIIGLAYLF 75
Query: 67 PKEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYK---FI* 120
P+EASHKVAK + + IL L A W T + + K Y++S +
Sbjct: 76 PQEASHKVAKYSLDFVYLSVVILFSSWLEVACWMHTGERQAAKIRKAYLRSMLSQDISLF 135
Query: 121 Y*GIHWRGHFCYYK*HYYCSRCSIRE--GNFLHYISRFIAGFTIGFVRVWQISLVTLSIV 178
I + +I E GNF+H+ISRFIAGF IGF VWQISLVTLSIV
Sbjct: 136 DTEISTGEVISAITSEILVVQDAISEKVGNFMHFISRFIAGFAIGFASVWQISLVTLSIV 195
Query: 179 PAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALM 238
P IALAGG YA+V+ GLI +VRK+YV+A EIAEEVIGNVRTVQAF GEE+AV SY+ AL
Sbjct: 196 PFIALAGGIYAFVSSGLIVRVRKSYVKANEIAEEVIGNVRTVQAFTGEEKAVSSYQGALR 255
Query: 239 KTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISG 298
TY GRKAGLAKGLGLGS+H VLFLSWALL+W+TS+VVHK IANGGESFTTMLNVVI+G
Sbjct: 256 NTYNYGRKAGLAKGLGLGSLHFVLFLSWALLIWFTSIVVHKGIANGGESFTTMLNVVIAG 315
Query: 299 LSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYP 358
LSLGQAAPDIS F+RA AAAYPIF+MIER+T KTGRKL ++G I F DV F+YP
Sbjct: 316 LSLGQAAPDISTFMRASAAAYPIFQMIERNT----EDKTGRKLGNVNGDILFKDVTFTYP 371
Query: 359 SRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELD 418
SRPDV IF LN IPAGK+VALVGGSGSGKST++SLIERFYEP G ++LD NDIR LD
Sbjct: 372 SRPDVVIFDKLNFVIPAGKVVALVGGSGSGKSTMISLIERFYEPTDGAVMLDGNDIRYLD 431
Query: 419 LKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
LKWLR IGLVNQEP LFAT+I+ENI+YGKDDAT EE+ A K
Sbjct: 432 LKWLRGHIGLVNQEPVLFATTIRENIMYGKDDATSEEITNAAK 474
Score = 217 bits (553), Expect = 5e-55
Identities = 122/315 (38%), Positives = 187/315 (58%), Gaps = 8/315 (2%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L + + F I F+ W+++LV L+ P I G + KAY++A +
Sbjct: 786 LENLGLVVTAFIISFILNWRLTLVVLATYPLIISGHISEKIFMQGYGGNLSKAYLKANML 845
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E I N+RTV AF EE+ + Y L++ + G G+ G +F S+ L
Sbjct: 846 AGESISNIRTVVAFCAEEKVLDLYSKELLEPSERSFRRGQMAGILYGVSQFFIFSSYGLA 905
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQA---APDISAFIRAKAAAYPIFEMIE 326
+WY S+++ K +++ T + ++++ L +G+ APD+ ++ +FE+++
Sbjct: 906 LWYGSILMEKGLSSFESVMKTFMVLIVTALVMGEVLALAPDL---LKGNQMVVSVFELLD 962
Query: 327 RDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSG 386
R T + TG +LS ++G I+ V FSYPSRPDV IF++ NL +P+GK +ALVG SG
Sbjct: 963 RRT--QVVGDTGEELSNVEGTIELKGVHFSYPSRPDVTIFSDFNLLVPSGKSMALVGQSG 1020
Query: 387 SGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILY 446
SGKS+V+SL+ RFY+P +G I++D DI++L LK LR+ IGLV QEPALFAT+I ENILY
Sbjct: 1021 SGKSSVLSLVLRFYDPTAGIIMIDGQDIKKLKLKSLRRHIGLVQQEPALFATTIYENILY 1080
Query: 447 GKDDATLEELKRAVK 461
GK+ A+ E+ A K
Sbjct: 1081 GKEGASESEVMEAAK 1095
>gb|AAF17668.1| F20B24.12 [Arabidopsis thaliana] gi|25297460|pir||B86240 protein
F20B24.12 [imported] - Arabidopsis thaliana
Length = 1316
Score = 461 bits (1185), Expect = e-128
Identities = 237/324 (73%), Positives = 266/324 (81%), Gaps = 13/324 (4%)
Query: 147 GNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRA 206
GNF+H+ISRFIAGF IGF VWQISLVTLSIVP IALAGG YA+V+ GLI +VRK+YV+A
Sbjct: 192 GNFMHFISRFIAGFAIGFASVWQISLVTLSIVPFIALAGGIYAFVSSGLIVRVRKSYVKA 251
Query: 207 GEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSW 266
EIAEEVIGNVRTVQAF GEE+AV SY+ AL TY GRKAGLAKGLGLGS+H VLFLSW
Sbjct: 252 NEIAEEVIGNVRTVQAFTGEEKAVSSYQGALRNTYNYGRKAGLAKGLGLGSLHFVLFLSW 311
Query: 267 ALLVWYTSVVVHKNIANGGESFTTMLNVVISGL---------SLGQAAPDISAFIRAKAA 317
ALL+W+TS+VVHK IANGGESFTTMLNVVI+G SLGQAAPDIS F+RA AA
Sbjct: 312 ALLIWFTSIVVHKGIANGGESFTTMLNVVIAGFHNKALFLYRSLGQAAPDISTFMRASAA 371
Query: 318 AYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGK 377
AYPIF+MIER+T KTGRKL ++G I F DV F+YPSRPDV IF LN IPAGK
Sbjct: 372 AYPIFQMIERNT----EDKTGRKLGNVNGDILFKDVTFTYPSRPDVVIFDKLNFVIPAGK 427
Query: 378 IVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFA 437
+VALVGGSGSGKST++SLIERFYEP G ++LD NDIR LDLKWLR IGLVNQEP LFA
Sbjct: 428 VVALVGGSGSGKSTMISLIERFYEPTDGAVMLDGNDIRYLDLKWLRGHIGLVNQEPVLFA 487
Query: 438 TSIKENILYGKDDATLEELKRAVK 461
T+I+ENI+YGKDDAT EE+ A K
Sbjct: 488 TTIRENIMYGKDDATSEEITNAAK 511
Score = 204 bits (519), Expect = 5e-51
Identities = 124/340 (36%), Positives = 189/340 (55%), Gaps = 33/340 (9%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIA----------------LAGGCYAYVTI 193
L + + F I F+ W+++LV L+ P I L G
Sbjct: 850 LENLGLVVTAFIISFILNWRLTLVVLATYPLIISGHISEVKRSFLRFYILFFGRQKIFMQ 909
Query: 194 GLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGL 253
G + KAY++A +A E I N+RTV AF EE+ + Y L++ + G G+
Sbjct: 910 GYGGNLSKAYLKANMLAGESISNIRTVVAFCAEEKVLDLYSKELLEPSERSFRRGQMAGI 969
Query: 254 GLGSMHCVLFLSWALLVWYT---------SVVVHKNIANGGESFTTMLNVVISGLSLGQA 304
G +F S+ L +WY S+++ K +++ T + ++++ L +G+
Sbjct: 970 LYGVSQFFIFSSYGLALWYIYKLFHTKYGSILMEKGLSSFESVMKTFMVLIVTALVMGEV 1029
Query: 305 ---APDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRP 361
APD+ ++ +FE+++R T + TG +LS ++G I+ V FSYPSRP
Sbjct: 1030 LALAPDL---LKGNQMVVSVFELLDRRT--QVVGDTGEELSNVEGTIELKGVHFSYPSRP 1084
Query: 362 DVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKW 421
DV IF++ NL +P+GK +ALVG SGSGKS+V+SL+ RFY+P +G I++D DI++L LK
Sbjct: 1085 DVTIFSDFNLLVPSGKSMALVGQSGSGKSSVLSLVLRFYDPTAGIIMIDGQDIKKLKLKS 1144
Query: 422 LRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
LR+ IGLV QEPALFAT+I ENILYGK+ A+ E+ A K
Sbjct: 1145 LRRHIGLVQQEPALFATTIYENILYGKEGASESEVMEAAK 1184
Score = 118 bits (296), Expect = 3e-25
Identities = 63/111 (56%), Positives = 76/111 (67%), Gaps = 3/111 (2%)
Query: 7 DERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLF 66
++ KK VS LKLFSFAD YD VLM +GSIGA +HGASVP+FFIFFGKLIN+IGLAYLF
Sbjct: 16 EKEKKRPSVSFLKLFSFADFYDCVLMALGSIGACIHGASVPVFFIFFGKLINIIGLAYLF 75
Query: 67 PKEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKS 114
P+EASHKVAK + + IL L A W T + + K Y++S
Sbjct: 76 PQEASHKVAKYSLDFVYLSVVILFSSWLEVACWMHTGERQAAKIRKAYLRS 126
>dbj|BAB02129.1| P-glycoprotein; multi-drug resistance related; ABC transporter-like
protein [Arabidopsis thaliana]
gi|15228506|ref|NP_189528.1| multidrug resistance
P-glycoprotein, putative [Arabidopsis thaliana]
Length = 1252
Score = 393 bits (1009), Expect = e-108
Identities = 205/460 (44%), Positives = 298/460 (64%), Gaps = 8/460 (1%)
Query: 8 ERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFP 67
E+KKE + KLFSFAD +DY+LMF+GS+GAIVHG+S+P+FF+ FG+++N G +
Sbjct: 17 EKKKEQSLPFFKLFSFADKFDYLLMFVGSLGAIVHGSSMPVFFLLFGQMVNGFGKNQMDL 76
Query: 68 KEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKF-I*Y*G 123
+ H+V++ + + + A W + + L K Y+++ K + +
Sbjct: 77 HQMVHEVSRYSLYFVYLGLVVCFSSYAEIACWMYSGERQVAALRKKYLEAVLKQDVGFFD 136
Query: 124 IHWRGHFCYYK*H----YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVP 179
R + S + GNF+HY+S F+AG +GFV W+++L++++++P
Sbjct: 137 TDARTGDIVFSVSTDTLLVQDAISEKVGNFIHYLSTFLAGLVVGFVSAWKLALLSVAVIP 196
Query: 180 AIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMK 239
IA AGG YAY G+ +K R++Y AG IAE+ I VRTV ++ GE +A+ +Y A+
Sbjct: 197 GIAFAGGLYAYTLTGITSKSRESYANAGVIAEQAIAQVRTVYSYVGESKALNAYSDAIQY 256
Query: 240 TYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGL 299
T G KAG+AKGLGLG + + +SWAL+ WY V + +GG++FT + + ++ G+
Sbjct: 257 TLKLGYKAGMAKGLGLGCTYGIACMSWALVFWYAGVFIRNGQTDGGKAFTAIFSAIVGGM 316
Query: 300 SLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPS 359
SLGQ+ ++ AF + KAA Y + E+I + + G+ L ++ G+I+F DV FSYPS
Sbjct: 317 SLGQSFSNLGAFSKGKAAGYKLMEIINQRPTIIQDPLDGKCLDQVHGNIEFKDVTFSYPS 376
Query: 360 RPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDL 419
RPDV IF N N+ P+GK VA+VGGSGSGKSTVVSLIERFY+P SGQILLD +I+ L L
Sbjct: 377 RPDVMIFRNFNIFFPSGKTVAVVGGSGSGKSTVVSLIERFYDPNSGQILLDGVEIKTLQL 436
Query: 420 KWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
K+LR+QIGLVNQEPALFAT+I ENILYGK DAT+ E++ A
Sbjct: 437 KFLREQIGLVNQEPALFATTILENILYGKPDATMVEVEAA 476
Score = 192 bits (488), Expect = 2e-47
Identities = 109/312 (34%), Positives = 166/312 (52%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L ++ + F + F+ W++SL+ L P + LA G KA+ + I
Sbjct: 812 LQNMTSLLTSFIVAFIVEWRVSLLILGTFPLLVLANFAQQLSLKGFAGDTAKAHAKTSMI 871
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF + + + + L G G L+ S AL+
Sbjct: 872 AGEGVSNIRTVAAFNAQSKILSLFCHELRVPQKRSLYRSQTSGFLFGLSQLALYGSEALI 931
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT 329
+WY + +V K ++ + + +VI+ S+ + IR A +F +++R T
Sbjct: 932 LWYGAHLVSKGVSTFSKVIKVFVVLVITANSVAETVSLAPEIIRGGEAVGSVFSVLDRQT 991
Query: 330 VSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGK 389
+ + G I+F V F+YPSRPDV +F + NL I AG ALVG SGSGK
Sbjct: 992 RIDPDDADADPVETIRGDIEFRHVDFAYPSRPDVMVFRDFNLRIRAGHSQALVGASGSGK 1051
Query: 390 STVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKD 449
S+V+++IERFY+P++G++++D DIR L+LK LR +IGLV QEPALFA +I +NI YGKD
Sbjct: 1052 SSVIAMIERFYDPLAGKVMIDGKDIRRLNLKSLRLKIGLVQQEPALFAATIFDNIAYGKD 1111
Query: 450 DATLEELKRAVK 461
AT E+ A +
Sbjct: 1112 GATESEVIDAAR 1123
>dbj|BAC41846.1| putative P-glycoprotein [Arabidopsis thaliana]
Length = 1252
Score = 393 bits (1009), Expect = e-108
Identities = 205/460 (44%), Positives = 298/460 (64%), Gaps = 8/460 (1%)
Query: 8 ERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFP 67
E+KKE + KLFSFAD +DY+LMF+GS+GAIVHG+S+P+FF+ FG+++N G +
Sbjct: 17 EKKKEQSLPFFKLFSFADKFDYLLMFVGSLGAIVHGSSMPVFFLLFGQMVNGFGKNQMDL 76
Query: 68 KEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKF-I*Y*G 123
+ H+V++ + + + A W + + L K Y+++ K + +
Sbjct: 77 HQMVHEVSRYSLYFVYLGLVVCFSSYAEIACWMYSGERQVAALRKKYLEAVLKQDVGFFD 136
Query: 124 IHWRGHFCYYK*H----YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVP 179
R + S + GNF+HY+S F+AG +GFV W+++L++++++P
Sbjct: 137 TDARTGDIVFSVSTDTLLVQDAISEKVGNFIHYLSTFLAGLVVGFVSAWKLALLSVAVIP 196
Query: 180 AIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMK 239
IA AGG YAY G+ +K R++Y AG IAE+ I VRTV ++ GE +A+ +Y A+
Sbjct: 197 GIAFAGGLYAYTLTGITSKSRESYANAGVIAEQAIAQVRTVYSYVGESKALNAYSDAIQY 256
Query: 240 TYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGL 299
T G KAG+AKGLGLG + + +SWAL+ WY V + +GG++FT + + ++ G+
Sbjct: 257 TLKLGYKAGMAKGLGLGCTYGIACMSWALVFWYAGVFIRNGQTDGGKAFTAIFSAIVGGM 316
Query: 300 SLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPS 359
SLGQ+ ++ AF + KAA Y + E+I + + G+ L ++ G+I+F DV FSYPS
Sbjct: 317 SLGQSFSNLGAFSKGKAAGYKLMEIINQRPTIIQDPLDGKCLDQVHGNIEFKDVTFSYPS 376
Query: 360 RPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDL 419
RPDV IF N N+ P+GK VA+VGGSGSGKSTVVSLIERFY+P SGQILLD +I+ L L
Sbjct: 377 RPDVMIFRNFNIFFPSGKTVAVVGGSGSGKSTVVSLIERFYDPNSGQILLDGVEIKTLQL 436
Query: 420 KWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
K+LR+QIGLVNQEPALFAT+I ENILYGK DAT+ E++ A
Sbjct: 437 KFLREQIGLVNQEPALFATTILENILYGKPDATMVEVEAA 476
Score = 192 bits (488), Expect = 2e-47
Identities = 109/312 (34%), Positives = 166/312 (52%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L ++ + F + F+ W++SL+ L P + LA G KA+ + I
Sbjct: 812 LQNMTSLLTSFIVAFIVEWRVSLLILGTFPLLVLANFAQQLSLKGFAGDTAKAHAKTSMI 871
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF + + + + L G G L+ S AL+
Sbjct: 872 AGEGVSNIRTVAAFNAQSKILSLFCHELRVPQKRSLYRSQTSGFLFGLSQLALYGSEALI 931
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT 329
+WY + +V K ++ + + +VI+ S+ + IR A +F +++R T
Sbjct: 932 LWYGAHLVSKGVSTFSKVIKVFVVLVITANSVAETVSLAPEIIRGGEAVGSVFSVLDRQT 991
Query: 330 VSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGK 389
+ + G I+F V F+YPSRPDV +F + NL I AG ALVG SGSGK
Sbjct: 992 RIDPDDADADPVETIRGDIEFRHVDFAYPSRPDVMVFRDFNLRIRAGHSQALVGASGSGK 1051
Query: 390 STVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKD 449
S+V+++IERFY+P++G++++D DIR L+LK LR +IGLV QEPALFA +I +NI YGKD
Sbjct: 1052 SSVIAMIERFYDPLAGKVMIDGKDIRRLNLKSLRLKIGLVQQEPALFAATIFDNIAYGKD 1111
Query: 450 DATLEELKRAVK 461
AT E+ A +
Sbjct: 1112 GATESEVIDAAR 1123
>gb|AAN28720.2| MDR-like p-glycoprotein [Arabidopsis thaliana]
Length = 1252
Score = 393 bits (1009), Expect = e-108
Identities = 205/460 (44%), Positives = 298/460 (64%), Gaps = 8/460 (1%)
Query: 8 ERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFP 67
E+KKE + KLFSFAD +DY+LMF+GS+GAIVHG+S+P+FF+ FG+++N G +
Sbjct: 17 EKKKEQSLPFFKLFSFADKFDYLLMFVGSLGAIVHGSSMPVFFLLFGQMVNGFGKNQMDL 76
Query: 68 KEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKF-I*Y*G 123
+ H+V++ + + + A W + + L K Y+++ K + +
Sbjct: 77 HQMVHEVSRYSLYFVYLGLVVCFSSYAEIACWMYSGERQVAALRKKYLEAVLKQDVGFFD 136
Query: 124 IHWRGHFCYYK*H----YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVP 179
R + S + GNF+HY+S F+AG +GFV W+++L++++++P
Sbjct: 137 TDARTGDIVFSVSTDTLLVQDAISEKVGNFIHYLSTFLAGLVVGFVSAWKLALLSVAVIP 196
Query: 180 AIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMK 239
IA AGG YAY G+ +K R++Y AG IAE+ I VRTV ++ GE +A+ +Y A+
Sbjct: 197 GIAFAGGLYAYTLTGITSKSRESYANAGVIAEQAIAQVRTVYSYVGESKALNAYSDAIQY 256
Query: 240 TYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGL 299
T G KAG+AKGLGLG + + +SWAL+ WY V + +GG++FT + + ++ G+
Sbjct: 257 TLKLGYKAGMAKGLGLGCTYGIACMSWALVFWYAGVFIRNGQTDGGKAFTAIFSAIVGGM 316
Query: 300 SLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPS 359
SLGQ+ ++ AF + KAA Y + E+I + + G+ L ++ G+I+F DV FSYPS
Sbjct: 317 SLGQSFSNLGAFSKGKAAGYKLMEIINQRPTIIQDPLDGKCLDQVHGNIEFKDVTFSYPS 376
Query: 360 RPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDL 419
RPDV IF N N+ P+GK VA+VGGSGSGKSTVVSLIERFY+P SGQILLD +I+ L L
Sbjct: 377 RPDVMIFRNFNIFFPSGKTVAVVGGSGSGKSTVVSLIERFYDPNSGQILLDGVEIKTLQL 436
Query: 420 KWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
K+LR+QIGLVNQEPALFAT+I ENILYGK DAT+ E++ A
Sbjct: 437 KFLREQIGLVNQEPALFATTILENILYGKPDATMVEVEAA 476
Score = 188 bits (478), Expect = 3e-46
Identities = 108/312 (34%), Positives = 165/312 (52%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L ++ + F + F+ W++SL+ L P + LA G KA+ + I
Sbjct: 812 LQNMTSLLTSFIVAFIVEWRVSLLILGTFPLLVLANFAQQLSLKGFAGDTAKAHAKTSMI 871
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF + + + + L G G L+ S AL+
Sbjct: 872 AGEGVSNIRTVAAFNAQSKILSLFCHELRVPQKRSLYRSQTSGFLFGLSQLALYGSEALI 931
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT 329
+WY + +V K ++ + + +VI+ S+ + IR A +F +++R T
Sbjct: 932 LWYGAHLVSKGVSTFSKVIKVFVVLVITANSVAETVSLAPEIIRGGEAVGSVFSVLDRQT 991
Query: 330 VSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGK 389
+ + G I+F V F+YPSRPDV +F + NL I AG ALVG SGSGK
Sbjct: 992 RIDPDDADADPVETIRGDIEFRHVDFAYPSRPDVMVFRDFNLRIRAGHSQALVGASGSGK 1051
Query: 390 STVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKD 449
S+V+++IERFY+ ++G++++D DIR L+LK LR +IGLV QEPALFA +I +NI YGKD
Sbjct: 1052 SSVIAMIERFYDLLAGKVMIDGKDIRRLNLKSLRLKIGLVQQEPALFAATIFDNIAYGKD 1111
Query: 450 DATLEELKRAVK 461
AT E+ A +
Sbjct: 1112 GATESEVIDAAR 1123
>ref|NP_174115.1| multidrug resistance P-glycoprotein, putative [Arabidopsis
thaliana] gi|12322992|gb|AAG51482.1| P-glycoprotein,
putative [Arabidopsis thaliana] gi|25297458|pir||G86404
probable P-glycoprotein [imported] - Arabidopsis
thaliana
Length = 1245
Score = 380 bits (975), Expect = e-104
Identities = 210/475 (44%), Positives = 305/475 (64%), Gaps = 22/475 (4%)
Query: 2 ESKEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIG 61
E+KE + K+ VS++ LFS AD DY LM +G +GA +HGA++P+FF+FFGK+++ +G
Sbjct: 17 EAKEEKKNIKKESVSLMGLFSAADKLDYFLMLLGGLGACIHGATLPLFFVFFGKMLDSLG 76
Query: 62 LAYLFPKEASHKVAKSKCSNIIFILDRG--GLLDAY-----WRKTSSKDENGLFKVYVKS 114
PK S +V++ N ++++ G + A+ W +T + L Y+KS
Sbjct: 77 NLSTDPKAISSRVSQ----NALYLVYLGLVNFVSAWIGVSCWMQTGERQTARLRINYLKS 132
Query: 115 RY-KFI*Y*GIHWRGHFCYYK*HYYCSRCSIREG------NFLHYISRFIAGFTIGFVRV 167
K I + R + H +++ + L Y+S+FIAGF IGF+ V
Sbjct: 133 ILAKDITFFDTEARDSNLIF--HISSDAILVQDAIGDKTDHVLRYLSQFIAGFVIGFLSV 190
Query: 168 WQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEE 227
WQ++L+TL +VP IA+AGG YA V + K AY AG++AEEV+ VRTV AF GEE
Sbjct: 191 WQLTLLTLGVVPLIAIAGGGYAIVMSTISEKSETAYADAGKVAEEVMSQVRTVYAFVGEE 250
Query: 228 RAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGES 287
+AV+SY +L K G+++GLAKGLG+G + +LF +WALL+WY S++V NG ++
Sbjct: 251 KAVKSYSNSLKKALKLGKRSGLAKGLGVGLTYSLLFCAWALLLWYASLLVRHGKTNGAKA 310
Query: 288 FTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMI-ERDTVSKKSSKTGRKLSKLDG 346
FTT+LNV+ SG +LGQAAP +SA + + AA IF MI ++ S + G L + G
Sbjct: 311 FTTILNVIFSGFALGQAAPSLSAIAKGRVAAANIFRMIGNNNSESSQRLDEGTTLQNVAG 370
Query: 347 HIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQ 406
I+F V F+YPSRP++ +F NL+ I +GK A VG SGSGKST++S+++RFYEP SG+
Sbjct: 371 RIEFQKVSFAYPSRPNM-VFENLSFTIRSGKTFAFVGPSGSGKSTIISMVQRFYEPNSGE 429
Query: 407 ILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
ILLD NDI+ L LKW R+Q+GLV+QEPALFAT+I NIL GK++A ++++ A K
Sbjct: 430 ILLDGNDIKSLKLKWFREQLGLVSQEPALFATTIASNILLGKENANMDQIIEAAK 484
Score = 207 bits (528), Expect = 4e-52
Identities = 136/476 (28%), Positives = 227/476 (47%), Gaps = 25/476 (5%)
Query: 2 ESKEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIG 61
++K D +K SM+ +S ++ +GSIGA++ GA P+F + G
Sbjct: 651 KTKNDDSKKDFSSSSMIWELIKLNSPEWPYALLGSIGAVLAGAQTPLFSM---------G 701
Query: 62 LAYLFPKEASH--KVAKSKCSNIIFILDRGGLLDA------------YWRKTSSKDENGL 107
+AY+ S V K + I G++ A + +S+ L
Sbjct: 702 IAYVLTAFYSPFPNVIKRDVEKVAIIFAGAGIVTAPIYLLQHYFYTLMGERLTSRVRLSL 761
Query: 108 FKVYVKSRYKFI*Y*GIHWRGHFCYYK*HYYCSRCSI--REGNFLHYISRFIAGFTIGFV 165
F + + + + R ++ R + +S + + F
Sbjct: 762 FSAILSNEIGWFDLDENNTGSLTSILAADATLVRSALADRLSTIVQNLSLTVTALALAFF 821
Query: 166 RVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAG 225
W+++ V + P + A G +AY RA +A E I N+RTV A+
Sbjct: 822 YSWRVAAVVTACFPLLIAASLTEQLFLKGFGGDYTRAYSRATSVAREAIANIRTVAAYGA 881
Query: 226 EERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGG 285
E++ + L K N G G G G + F S+AL +WY SV+++ N G
Sbjct: 882 EKQISEQFTCELSKPTKNAFVRGHISGFGYGLSQFLAFCSYALGLWYVSVLINHKETNFG 941
Query: 286 ESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLD 345
+S + + ++++ S+ + ++ A +F ++ R+T R +S++
Sbjct: 942 DSIKSFMVLIVTAFSVSETLALTPDIVKGTQALGSVFRVLHRETKISPDQPNSRMVSQVK 1001
Query: 346 GHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISG 405
G I+F +V F YP+RP++ IF NLNL + AGK +A+VG SGSGKSTV+ LI RFY+P +G
Sbjct: 1002 GDIEFRNVSFVYPTRPEIDIFKNLNLRVSAGKSLAVVGPSGSGKSTVIGLIMRFYDPSNG 1061
Query: 406 QILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
+ +D DI+ L+L+ LR+++ LV QEPALF+T+I ENI YG ++A+ E+ A K
Sbjct: 1062 NLCIDGQDIKTLNLRSLRKKLALVQQEPALFSTTIYENIKYGNENASEAEIMEAAK 1117
>ref|XP_474071.1| OSJNBb0079B02.13 [Oryza sativa (japonica cultivar-group)]
gi|38347317|emb|CAE05967.2| OSJNBa0063C18.8 [Oryza
sativa (japonica cultivar-group)]
gi|38344910|emb|CAD41854.2| OSJNBb0079B02.13 [Oryza
sativa (japonica cultivar-group)]
gi|27368849|emb|CAD59582.1| MDR-like ABC transporter
[Oryza sativa (japonica cultivar-group)]
Length = 1268
Score = 377 bits (968), Expect = e-103
Identities = 202/460 (43%), Positives = 291/460 (62%), Gaps = 8/460 (1%)
Query: 8 ERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFP 67
+++ + V+ +LF+FAD +D VLM GS+GA+ HGA++P+FF+ FG LIN G
Sbjct: 32 KKRADQAVAFHELFTFADKWDLVLMAAGSLGALAHGAAMPLFFLLFGDLINGFGKNQTDL 91
Query: 68 KEASHKVAKSKCSNI---IFILDRGGLLDAYWRKTSSKDENGLFKVYVKS--RYKFI*Y* 122
+ + +V+K + + + A W T + L K Y+ + R +
Sbjct: 92 RTMTDEVSKYALYFVYLGLVVCASSYAEIACWMYTGERQVIALRKAYLDAVLRQDVGFFD 151
Query: 123 GIHWRGHFCY-YK*HYYCSRCSIRE--GNFLHYISRFIAGFTIGFVRVWQISLVTLSIVP 179
G + + +I E GNF+HYI+ F+AG +GFV W+++L++++++P
Sbjct: 152 TDARTGDIVFGVSTDTLLVQDAIGEKVGNFIHYIATFLAGLVVGFVAAWRLALLSVAVIP 211
Query: 180 AIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMK 239
AIA AGG YAY GL +K R++Y AG +AE+ I VRTV +FAGE +A+ SY A+
Sbjct: 212 AIAFAGGLYAYTLTGLTSKSRESYANAGVVAEQAIAQVRTVYSFAGESKALNSYSEAIQN 271
Query: 240 TYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGL 299
T G KAG+AKGLG+G + + +SWAL+ WY V + +GG++FT + + ++ G+
Sbjct: 272 TLKLGYKAGMAKGLGIGCTYGIACMSWALVFWYAGVFIRNGQTDGGKAFTAIFSAIVGGM 331
Query: 300 SLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPS 359
SLGQA ++ AF + K A Y + E+I + K G+ L+++ G+I+F DV FSYPS
Sbjct: 332 SLGQAFSNLGAFSKGKIAGYKLLEVIRQKPSIVHDHKDGKLLAEVHGNIEFKDVTFSYPS 391
Query: 360 RPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDL 419
RPDV IF + +L PA K VA+VGGSGSGKSTVV+LIERFY+P GQ+LLD DI+ L L
Sbjct: 392 RPDVMIFRDFSLFFPAAKTVAVVGGSGSGKSTVVALIERFYDPNEGQVLLDNVDIKTLQL 451
Query: 420 KWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
+WLR QIGLVNQEPALFAT+I ENILYGK DAT+ E++ A
Sbjct: 452 RWLRDQIGLVNQEPALFATTIHENILYGKPDATMAEVEAA 491
Score = 196 bits (498), Expect = 1e-48
Identities = 110/312 (35%), Positives = 170/312 (54%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L ++ + F +GF+ W+++L+ L+ P + LA G KA+ ++ +
Sbjct: 829 LQNMTSLMTSFIVGFIIEWRVALLILATFPLLVLANFAQQLSMKGFAGDTAKAHAKSSMV 888
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF + + + + L + GL G L+ S AL+
Sbjct: 889 AGEGVSNIRTVAAFNAQNKILSLFSYELRIPEQQILRRSQTSGLLFGLSQLCLYSSEALI 948
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT 329
+WY S +V + + + + +V++ S+ + +R + IF ++ R T
Sbjct: 949 LWYGSHLVRSHGSTFSKVIKVFVVLVVTANSVAETVSLAPEIVRGGESIRSIFGILNRAT 1008
Query: 330 VSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGK 389
+ +++ + G I+ V F+YP+RPD+ IF + NL I AG+ ALVG SGSGK
Sbjct: 1009 RIEPDDPESERVTNVRGDIELRHVDFAYPARPDIQIFKDFNLKIQAGRSQALVGASGSGK 1068
Query: 390 STVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKD 449
STV++LIERFY+P G++ +D DIR L+LK LR +IGLV QEP LFA SI ENI YGKD
Sbjct: 1069 STVIALIERFYDPTGGKVTIDGKDIRRLNLKALRLKIGLVQQEPVLFAASILENIAYGKD 1128
Query: 450 DATLEELKRAVK 461
AT EE+ +A K
Sbjct: 1129 GATEEEVIQAAK 1140
>emb|CAD40903.1| OSJNBa0036B21.21 [Oryza sativa (japonica cultivar-group)]
gi|50924782|ref|XP_472741.1| OSJNBa0036B21.21 [Oryza
sativa (japonica cultivar-group)]
Length = 1252
Score = 377 bits (968), Expect = e-103
Identities = 199/459 (43%), Positives = 291/459 (63%), Gaps = 17/459 (3%)
Query: 5 EGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAY 64
E +++ + V+ +LF FAD D++LM GS GA+VHGA++P+FF+ FG+LIN G
Sbjct: 19 EAVKKRVDQSVAFHELFGFADPLDWLLMAAGSAGAVVHGAAMPVFFLLFGELINGFGKNQ 78
Query: 65 LFPKEASHKVAKSKCSNIIFILDRG-GLLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y*G 123
+ + +V+K++ + ++ +R G L + + + + G F ++
Sbjct: 79 HSLRRMTDEVSKAQIACWMYTGERQVGALRRRYLEAVLRQDVGFFDTDART--------- 129
Query: 124 IHWRGHFCY-YK*HYYCSRCSIRE--GNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPA 180
G + + +I E GNF+HY+S F+AG +GFV W+++L++++++P
Sbjct: 130 ----GDVVFSVSTDTLLVQDAIGEKVGNFIHYLSTFLAGLVVGFVSAWRLALLSIAVIPG 185
Query: 181 IALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKT 240
IA AGG YAY GL +K R +Y AG IAE+ I VRTV ++ GE +A+ SY A+ T
Sbjct: 186 IAFAGGLYAYTLTGLTSKSRDSYANAGIIAEQAIAQVRTVYSYVGESKALNSYSEAIQNT 245
Query: 241 YVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLS 300
G KAG+AKGLG+G + + +SWAL+ WY V + +GG++FT + + ++ GLS
Sbjct: 246 LKLGYKAGMAKGLGIGCTYGIACMSWALVFWYAGVFIRNGQTDGGKAFTAIFSAIVGGLS 305
Query: 301 LGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSR 360
LGQ+ ++ AF + K A Y + E+I + + GR L ++ G+I+F +V FSYPSR
Sbjct: 306 LGQSFSNLGAFSKGKIAGYKLLEVIRQRPTIVQDPADGRCLDEVHGNIEFKEVAFSYPSR 365
Query: 361 PDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLK 420
PDV IF + +L PAGK A+VGGSGSGKSTVV+LIERFY+P GQ+LLD DI+ L LK
Sbjct: 366 PDVMIFRDFSLFFPAGKTAAVVGGSGSGKSTVVALIERFYDPNQGQVLLDNVDIKTLQLK 425
Query: 421 WLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
WLR QIGLVNQEPALFAT+I ENILYGK DAT+ E++ A
Sbjct: 426 WLRDQIGLVNQEPALFATTILENILYGKPDATMAEVEAA 464
Score = 192 bits (488), Expect = 2e-47
Identities = 106/312 (33%), Positives = 170/312 (53%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L ++ + F +GF+ W+++++ L P + LA G KA+ + I
Sbjct: 812 LQNMTSLLVSFVVGFIIEWRVAVLILVTFPLLVLANFAQQLSMKGFAGDTAKAHAKTSMI 871
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF +++ + + L ++ + G G L+ S AL+
Sbjct: 872 AGEGVSNIRTVAAFNAQDKVLSLFCTELRVPQMHSLRRSQISGALFGLSQLSLYASEALI 931
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT 329
+WY + +V +++ + + +VI+ ++ + +R + +F ++ T
Sbjct: 932 LWYGAHLVRHHVSTFSKVIKVFVVLVITANTVAETVSLAPEIVRGGESIRSVFAILNYRT 991
Query: 330 VSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGK 389
+ + G I F V F+YPSRPDV +F + +L I AG+ ALVG SGSGK
Sbjct: 992 RIDPDEPETEPVESVRGDIDFRHVDFAYPSRPDVMVFKDFSLRIRAGQSQALVGASGSGK 1051
Query: 390 STVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKD 449
STV++LIERFY+P++G++++D DIR L+++ LR +IGLV QEP LFATSI ENI YGKD
Sbjct: 1052 STVIALIERFYDPLAGKVMIDGKDIRRLNVRSLRLKIGLVQQEPVLFATSIFENIAYGKD 1111
Query: 450 DATLEELKRAVK 461
AT EE+ A K
Sbjct: 1112 GATEEEVIEAAK 1123
>emb|CAD59581.1| MDR-like ABC transporter [Oryza sativa (japonica cultivar-group)]
Length = 1256
Score = 371 bits (953), Expect = e-101
Identities = 196/460 (42%), Positives = 285/460 (61%), Gaps = 5/460 (1%)
Query: 5 EGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAY 64
E +++ + V+ +LF FAD D++LM GS GA+VHGA++P+FF+ FG+LIN G
Sbjct: 19 EAVKKRVDQSVAFHELFGFADPLDWLLMAAGSAGAVVHGAAMPVFFLLFGELINGFGKNQ 78
Query: 65 LFPKEASHKVAKSKCSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKF-I*Y*G 123
+ + + + + + L A W T + L + Y+++ + + +
Sbjct: 79 HSLRRMTDEYSLYFVYLGLVVCASSYLEIACWMYTGERQVGALRRRYLEAVLRQDVGFFD 138
Query: 124 IHWRGHFCYYK*H----YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVP 179
R + + GNF+HY+S F+AG +GFV W+++L++++++P
Sbjct: 139 TDARTGDVVFSVSTDTLLVQDAIGEKVGNFIHYLSTFLAGLVVGFVSAWRLALLSIAVIP 198
Query: 180 AIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMK 239
IA AGG YAY GL +K R +Y AG IAE+ I VRTV ++ GE +A+ SY A+
Sbjct: 199 GIAFAGGLYAYTLTGLTSKSRDSYANAGIIAEQAIAQVRTVYSYVGESKALNSYSEAIQN 258
Query: 240 TYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGL 299
T G KAG+AKGLG+G + + +SWAL+ WY V + +GG++FT + + ++ GL
Sbjct: 259 TLKLGYKAGMAKGLGIGCTYGIACMSWALVFWYAGVFIRNGQTDGGKAFTAIFSAIVGGL 318
Query: 300 SLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPS 359
SLGQ+ ++ AF + K A Y + E+I + + GR L ++ G+I+F +V FSYPS
Sbjct: 319 SLGQSFSNLGAFSKGKIAGYKLLEVIRQRPTIVQDPADGRCLDEVHGNIEFKEVAFSYPS 378
Query: 360 RPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDL 419
RPDV IF + +L PAGK A+VGGSGSGKSTVV+LIERFY+P GQ+LLD DI+ L L
Sbjct: 379 RPDVMIFRDFSLFFPAGKTAAVVGGSGSGKSTVVALIERFYDPNQGQVLLDNVDIKTLQL 438
Query: 420 KWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
KWLR QIGLVNQEPALFAT+I ENILYGK DAT+ E++ A
Sbjct: 439 KWLRDQIGLVNQEPALFATTILENILYGKPDATMAEVEAA 478
Score = 192 bits (488), Expect = 2e-47
Identities = 106/312 (33%), Positives = 170/312 (53%)
Query: 150 LHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEI 209
L ++ + F +GF+ W+++++ L P + LA G KA+ + I
Sbjct: 816 LQNMTSLLVSFVVGFIIEWRVAVLILVTFPLLVLANFAQQLSMKGFAGDTAKAHAKTSMI 875
Query: 210 AEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALL 269
A E + N+RTV AF +++ + + L ++ + G G L+ S AL+
Sbjct: 876 AGEGVSNIRTVAAFNAQDKVLSLFCTELRVPQMHSLRRSQISGALFGLSQLSLYASEALI 935
Query: 270 VWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT 329
+WY + +V +++ + + +VI+ ++ + +R + +F ++ T
Sbjct: 936 LWYGAHLVRHHVSTFSKVIKVFVVLVITANTVAETVSLAPEIVRGGESIRSVFAILNYRT 995
Query: 330 VSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGK 389
+ + G I F V F+YPSRPDV +F + +L I AG+ ALVG SGSGK
Sbjct: 996 RIDPDEPETEPVESVRGDIDFRHVDFAYPSRPDVMVFKDFSLRIRAGQSQALVGASGSGK 1055
Query: 390 STVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKD 449
STV++LIERFY+P++G++++D DIR L+++ LR +IGLV QEP LFATSI ENI YGKD
Sbjct: 1056 STVIALIERFYDPLAGKVMIDGKDIRRLNVRSLRLKIGLVQQEPVLFATSIFENIAYGKD 1115
Query: 450 DATLEELKRAVK 461
AT EE+ A K
Sbjct: 1116 GATEEEVIEAAK 1127
>ref|NP_174122.1| multidrug resistance P-glycoprotein, putative [Arabidopsis
thaliana] gi|12322986|gb|AAG51476.1| P-glycoprotein,
putative [Arabidopsis thaliana] gi|25297457|pir||F86405
probable P-glycoprotein [imported] - Arabidopsis
thaliana
Length = 1247
Score = 371 bits (952), Expect = e-101
Identities = 207/474 (43%), Positives = 304/474 (63%), Gaps = 24/474 (5%)
Query: 4 KEGDERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLA 63
KE ++ K+ VS++ LFS AD+ DY LMF+G +G +HG ++P+FF+FFG +++ +G
Sbjct: 20 KEEKKKMKKESVSLMGLFSAADNVDYFLMFLGGLGTCIHGGTLPLFFVFFGGMLDSLGKL 79
Query: 64 YLFPKEASHKVAKSKCSNIIFILDRG--GLLDAY-----WRKTSSKDENGLFKVYVKSRY 116
P S +V++ N ++++ G L+ A+ W +T + L Y+KS
Sbjct: 80 STDPNAISSRVSQ----NALYLVYLGLVNLVSAWIGVACWMQTGERQTARLRINYLKSIL 135
Query: 117 -KFI*Y*GIHWR-GHFCYYK*HYYCSRCSIRE------GNFLHYISRFIAGFTIGFVRVW 168
K I + R +F + H +++ G+ L Y+ +FIAGF IGF+ VW
Sbjct: 136 AKDITFFDTEARDSNFIF---HISSDAILVQDAIGDKTGHVLRYLCQFIAGFVIGFLSVW 192
Query: 169 QISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEER 228
Q++L+TL +VP IA+AGG YA V + K AY AG++AEEV+ VRTV AF GEE+
Sbjct: 193 QLTLLTLGVVPLIAIAGGGYAIVMSTISEKSEAAYADAGKVAEEVMSQVRTVYAFVGEEK 252
Query: 229 AVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESF 288
AV+SY +L K +++GLAKGLG+G + +LF +WALL WY S++V NG ++F
Sbjct: 253 AVKSYSNSLKKALKLSKRSGLAKGLGVGLTYSLLFCAWALLFWYASLLVRHGKTNGAKAF 312
Query: 289 TTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTV-SKKSSKTGRKLSKLDGH 347
TT+LNV+ SG +LGQA P +SA + + AA IF+MI + + S + + G L + G
Sbjct: 313 TTILNVIYSGFALGQAVPSLSAISKGRVAAANIFKMIGNNNLESSERLENGTTLQNVVGK 372
Query: 348 IQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQI 407
I+F V F+YPSRP++ +F NL+ I +GK A VG SGSGKST++S+++RFYEP SG+I
Sbjct: 373 IEFCGVSFAYPSRPNM-VFENLSFTIHSGKTFAFVGPSGSGKSTIISMVQRFYEPRSGEI 431
Query: 408 LLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
LLD NDI+ L LKWLR+Q+GLV+QEPALFAT+I NIL GK+ A ++++ A K
Sbjct: 432 LLDGNDIKNLKLKWLREQMGLVSQEPALFATTIASNILLGKEKANMDQIIEAAK 485
Score = 207 bits (526), Expect = 7e-52
Identities = 109/317 (34%), Positives = 176/317 (55%)
Query: 145 REGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYV 204
R + +S I + F W+++ V + P + A G +AY
Sbjct: 803 RLSTIVQNLSLTITALALAFFYSWRVAAVVTACFPLLIAASLTEQLFLKGFGGDYTRAYS 862
Query: 205 RAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFL 264
RA +A E I N+RTV AF+ E++ + L K + G G G G C+ F
Sbjct: 863 RATSLAREAISNIRTVAAFSAEKQISEQFTCELSKPTKSALLRGHISGFGYGLSQCLAFC 922
Query: 265 SWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEM 324
S+AL +WY SV++ +N N +S + + ++++ S+ + ++ A +F +
Sbjct: 923 SYALGLWYISVLIKRNETNFEDSIKSFMVLLVTAYSVAETLALTPDIVKGTQALGSVFRV 982
Query: 325 IERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGG 384
+ R+T R ++ + G I+F +V F+YP+RP++ IF NLNL + AGK +A+VG
Sbjct: 983 LHRETEIPPDQPNSRLVTHIKGDIEFRNVSFAYPTRPEIAIFKNLNLRVSAGKSLAVVGP 1042
Query: 385 SGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENI 444
SGSGKSTV+ LI RFY+P +G + +D +DI+ ++L+ LR+++ LV QEPALF+TSI ENI
Sbjct: 1043 SGSGKSTVIGLIMRFYDPSNGNLCIDGHDIKSVNLRSLRKKLALVQQEPALFSTSIHENI 1102
Query: 445 LYGKDDATLEELKRAVK 461
YG ++A+ E+ A K
Sbjct: 1103 KYGNENASEAEIIEAAK 1119
>gb|AAD10836.1| P-glycoprotein [Solanum tuberosum]
Length = 1313
Score = 357 bits (915), Expect = 6e-97
Identities = 202/480 (42%), Positives = 290/480 (60%), Gaps = 26/480 (5%)
Query: 1 MESKEGDERKKEHK----VSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKL 56
++ +EG + +K V +LF FAD D VLM IGS+GA VHG S+P+F FF L
Sbjct: 35 VKKEEGGDVEKPSSPPPAVGFGELFRFADGLDCVLMIIGSLGAFVHGCSLPLFLRFFADL 94
Query: 57 INVIGLAYLFPKEASHKVAKSKCSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRY 116
+N G + + +V K F++ + + W + S G + K R
Sbjct: 95 VNSFGSYANDVDKMTQEVLKYA---FYFLVVGAAIWASSWAEISCWMWTGERQT-TKMRI 150
Query: 117 KFI*Y*GIHWRGHFCYYK*HYYCS---------------RCSIREGNFLHYISRFIAGFT 161
K++ Y+ S S + GNF+HY++ F++GF
Sbjct: 151 KYL---EAALNQDIQYFDTEVRTSDVVSAINTDAVVVQDAISEKLGNFIHYMATFLSGFV 207
Query: 162 IGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQ 221
+GF VWQ++LVTL++VP IA+ G Y + L ++ ++A +AG I E+ + +RTV
Sbjct: 208 VGFTAVWQLALVTLAVVPLIAVIGAIYTVTSAKLSSQSQEALSKAGNIVEQTVVQIRTVL 267
Query: 222 AFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNI 281
F GE +A+++Y AAL + G K+G +KGLGLG+ + +F +ALL+WY +V +
Sbjct: 268 VFVGEAKALQAYTAALRVSQKIGYKSGFSKGLGLGATYFTVFCCYALLLWYGGYLVRHHF 327
Query: 282 ANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKL 341
NGG + TM V+I GL+LGQ+AP ++AF +A+ AA IF +I+ +++KTG +L
Sbjct: 328 TNGGLAIATMFAVMIGGLALGQSAPSMTAFAKARVAAAKIFRIIDHKPSVDRNAKTGLEL 387
Query: 342 SKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYE 401
+ G ++ +V FSYPSRP++ I N NL +PAGK +ALVG SGSGKSTVVSLIERFY+
Sbjct: 388 DTVSGQLELKNVEFSYPSRPEIKILNNFNLVVPAGKTIALVGSSGSGKSTVVSLIERFYD 447
Query: 402 PISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
P SGQ++LD NDI+ L LKWLRQQIGLV+QEPALFATSIKENIL G+ DAT E++ A +
Sbjct: 448 PTSGQLMLDGNDIKTLKLKWLRQQIGLVSQEPALFATSIKENILLGRPDATQIEIEEAAR 507
Score = 199 bits (506), Expect = 2e-49
Identities = 113/300 (37%), Positives = 169/300 (55%), Gaps = 1/300 (0%)
Query: 161 TIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTV 220
T GFV W+++LV + + P + A G + A+ +A ++A E + NVRTV
Sbjct: 861 TAGFVLQWRLALVLIGVFPVVVAATVLQKMFMKGFSGDLEAAHAKATQLAGEAVANVRTV 920
Query: 221 QAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKN 280
AF E + V + ++L G G G G +L+ S+AL +WY S +V
Sbjct: 921 AAFNSETKIVNLFDSSLQTPLRRCFWKGQIAGSGYGIAQFLLYSSYALGLWYASWLVKHG 980
Query: 281 IANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRK 340
I++ ++ + +++S + FI+ A +FE+++R T +
Sbjct: 981 ISDFSKTIRVFMVLMVSANGAAETLTLAPDFIKGGRAMRSVFELLDRKTEVEPDDPDATA 1040
Query: 341 L-SKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERF 399
+ +L G ++F V FSYP+RPDV IF +LNL AGK +ALVG SG GKS+V+SLIERF
Sbjct: 1041 VPDRLRGEVEFKHVDFSYPTRPDVSIFRDLNLRARAGKTLALVGPSGCGKSSVISLIERF 1100
Query: 400 YEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
YEP SG++++D DIR+ +LK LR+ I +V QEP LFAT+I ENI YG + AT E+ A
Sbjct: 1101 YEPSSGRVIIDGKDIRKYNLKSLRRHIAVVPQEPCLFATTIYENIAYGHESATEAEITEA 1160
>gb|AAM98246.1| putative ABC transporter [Arabidopsis thaliana]
Length = 1286
Score = 355 bits (911), Expect = 2e-96
Identities = 200/459 (43%), Positives = 284/459 (61%), Gaps = 16/459 (3%)
Query: 15 VSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKEASHKV 74
V+ +LF FAD DYVLM IGS+GA VHG S+P+F FF L+N G ++ +V
Sbjct: 27 VAFKELFRFADGLDYVLMGIGSVGAFVHGCSLPLFLRFFADLVNSFGSNSNNVEKMMEEV 86
Query: 75 AKSKCSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKF--------I*Y*GIHW 126
K + F++ + + W + S +G + K R K+ I +
Sbjct: 87 LKYA---LYFLVVGAAIWASSWAEISCWMWSGERQT-TKMRIKYLEAALNQDIQFFDTEV 142
Query: 127 RGHFCYYK*HYYC----SRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIA 182
R + + S + GNF+HY++ F++GF +GF VWQ++LVTL++VP IA
Sbjct: 143 RTSDVVFAINTDAVMVQDAISEKLGNFIHYMATFVSGFIVGFTAVWQLALVTLAVVPLIA 202
Query: 183 LAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYV 242
+ GG + L K +++ +AG I E+ + +R V AF GE RA ++Y +AL
Sbjct: 203 VIGGIHTTTLSKLSNKSQESLSQAGNIVEQTVVQIRVVMAFVGESRASQAYSSALKIAQK 262
Query: 243 NGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLG 302
G K GLAKG+GLG+ + V+F +ALL+WY +V ++ NGG + TM V+I GL+LG
Sbjct: 263 LGYKTGLAKGMGLGATYFVVFCCYALLLWYDGYLVRHHLTNGGLAIATMFAVMIGGLALG 322
Query: 303 QAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPD 362
Q+AP ++AF +AK AA IF +I+ +++S++G +L + G ++ +V FSYPSRPD
Sbjct: 323 QSAPSMAAFAKAKVAAAKIFRIIDHKPTIERNSESGVELDSVTGLVELKNVDFSYPSRPD 382
Query: 363 VGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWL 422
V I N L +PAGK +ALVG SGSGKSTVVSLIERFY+P SGQ+LLD D++ L L+WL
Sbjct: 383 VKILNNFCLSVPAGKTIALVGSSGSGKSTVVSLIERFYDPNSGQVLLDGQDLKTLKLRWL 442
Query: 423 RQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
RQQIGLV+QEPALFATSIKENIL G+ DA E++ A +
Sbjct: 443 RQQIGLVSQEPALFATSIKENILLGRPDADQVEIEEAAR 481
Score = 193 bits (490), Expect = 1e-47
Identities = 111/300 (37%), Positives = 167/300 (55%), Gaps = 1/300 (0%)
Query: 161 TIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTV 220
T GFV W+++LV +++ P + A G + A+ + ++A E I NVRTV
Sbjct: 836 TAGFVLQWRLALVLVAVFPVVVAATVLQKMFMTGFSGDLEAAHAKGTQLAGEAIANVRTV 895
Query: 221 QAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKN 280
AF E + VR Y A L G G G G L+ S+AL +WY S +V
Sbjct: 896 AAFNSEAKIVRLYTANLEPPLKRCFWKGQIAGSGYGVAQFCLYASYALGLWYASWLVKHG 955
Query: 281 IANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT-VSKKSSKTGR 339
I++ ++ + +++S + FI+ A +FE+++R T + T
Sbjct: 956 ISDFSKTIRVFMVLMVSANGAAETLTLAPDFIKGGQAMRSVFELLDRKTEIEPDDPDTTP 1015
Query: 340 KLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERF 399
+L G ++ + FSYPSRPD+ IF +L+L AGK +ALVG SG GKS+V+SLI+RF
Sbjct: 1016 VPDRLRGEVELKHIDFSYPSRPDIQIFRDLSLRARAGKTLALVGPSGCGKSSVISLIQRF 1075
Query: 400 YEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
YEP SG++++D DIR+ +LK +R+ I +V QEP LF T+I ENI YG + AT E+ +A
Sbjct: 1076 YEPSSGRVMIDGKDIRKYNLKAIRKHIAIVPQEPCLFGTTIYENIAYGHECATEAEIIQA 1135
>emb|CAA43646.1| P-glycoprotein [Arabidopsis thaliana] gi|4883607|gb|AAD31576.1|
putative ABC transporter [Arabidopsis thaliana]
gi|15228052|ref|NP_181228.1| multidrug resistance
P-glycoprotein (PGP1) [Arabidopsis thaliana]
gi|419760|pir||A42150 P-glycoprotein pgp1 - Arabidopsis
thaliana
Length = 1286
Score = 355 bits (910), Expect = 2e-96
Identities = 200/459 (43%), Positives = 284/459 (61%), Gaps = 16/459 (3%)
Query: 15 VSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKEASHKV 74
V+ +LF FAD DYVLM IGS+GA VHG S+P+F FF L+N G ++ +V
Sbjct: 27 VAFKELFRFADGLDYVLMGIGSVGAFVHGCSLPLFLRFFADLVNSFGSNSNNVEKMMEEV 86
Query: 75 AKSKCSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKF--------I*Y*GIHW 126
K + F++ + + W + S +G + K R K+ I +
Sbjct: 87 LKYA---LYFLVVGAAIWASSWAEISCWMWSGERQT-TKMRIKYLEAALNQDIQFFDTEV 142
Query: 127 RGHFCYYK*HYYC----SRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIA 182
R + + S + GNF+HY++ F++GF +GF VWQ++LVTL++VP IA
Sbjct: 143 RTSDVVFAINTDAVMVQDAISEKLGNFIHYMATFVSGFIVGFTAVWQLALVTLAVVPLIA 202
Query: 183 LAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYV 242
+ GG + L K +++ +AG I E+ + +R V AF GE RA ++Y +AL
Sbjct: 203 VIGGIHTTTLSKLSNKSQESLSQAGNIVEQTVVQIRVVMAFVGESRASQAYSSALKIAQK 262
Query: 243 NGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLG 302
G K GLAKG+GLG+ + V+F +ALL+WY +V ++ NGG + TM V+I GL+LG
Sbjct: 263 LGYKTGLAKGMGLGATYFVVFCCYALLLWYGGYLVRHHLTNGGLAIATMFAVMIGGLALG 322
Query: 303 QAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPD 362
Q+AP ++AF +AK AA IF +I+ +++S++G +L + G ++ +V FSYPSRPD
Sbjct: 323 QSAPSMAAFAKAKVAAAKIFRIIDHKPTIERNSESGVELDSVTGLVELKNVDFSYPSRPD 382
Query: 363 VGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWL 422
V I N L +PAGK +ALVG SGSGKSTVVSLIERFY+P SGQ+LLD D++ L L+WL
Sbjct: 383 VKILNNFCLSVPAGKTIALVGSSGSGKSTVVSLIERFYDPNSGQVLLDGQDLKTLKLRWL 442
Query: 423 RQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
RQQIGLV+QEPALFATSIKENIL G+ DA E++ A +
Sbjct: 443 RQQIGLVSQEPALFATSIKENILLGRPDADQVEIEEAAR 481
Score = 193 bits (490), Expect = 1e-47
Identities = 111/300 (37%), Positives = 167/300 (55%), Gaps = 1/300 (0%)
Query: 161 TIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTV 220
T GFV W+++LV +++ P + A G + A+ + ++A E I NVRTV
Sbjct: 836 TAGFVLQWRLALVLVAVFPVVVAATVLQKMFMTGFSGDLEAAHAKGTQLAGEAIANVRTV 895
Query: 221 QAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKN 280
AF E + VR Y A L G G G G L+ S+AL +WY S +V
Sbjct: 896 AAFNSEAKIVRLYTANLEPPLKRCFWKGQIAGSGYGVAQFCLYASYALGLWYASWLVKHG 955
Query: 281 IANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDT-VSKKSSKTGR 339
I++ ++ + +++S + FI+ A +FE+++R T + T
Sbjct: 956 ISDFSKTIRVFMVLMVSANGAAETLTLAPDFIKGGQAMRSVFELLDRKTEIEPDDPDTTP 1015
Query: 340 KLSKLDGHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERF 399
+L G ++ + FSYPSRPD+ IF +L+L AGK +ALVG SG GKS+V+SLI+RF
Sbjct: 1016 VPDRLRGEVELKHIDFSYPSRPDIQIFRDLSLRARAGKTLALVGPSGCGKSSVISLIQRF 1075
Query: 400 YEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
YEP SG++++D DIR+ +LK +R+ I +V QEP LF T+I ENI YG + AT E+ +A
Sbjct: 1076 YEPSSGRVMIDGKDIRKYNLKAIRKHIAIVPQEPCLFGTTIYENIAYGHECATEAEIIQA 1135
>ref|XP_483819.1| putative P-glycoprotein 1 [Oryza sativa (japonica cultivar-group)]
gi|45735908|dbj|BAD12940.1| putative P-glycoprotein 1
[Oryza sativa (japonica cultivar-group)]
Length = 1344
Score = 348 bits (894), Expect = 2e-94
Identities = 195/456 (42%), Positives = 279/456 (60%), Gaps = 20/456 (4%)
Query: 19 KLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKEASHKVAKSK 78
+LFSFAD DYVLM +G++GA+VHG S+P+F FF L++ G P V K
Sbjct: 97 QLFSFADGLDYVLMTLGTLGALVHGCSLPVFLRFFADLVDSFGSHAAHPDTMLRLVVKYA 156
Query: 79 CSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y*GIHWRGHFCYYK*H-- 136
F++ + + W + S G + + R +++ + +H F
Sbjct: 157 ---FYFLVVGAAIWASSWAEISCWMWTGE-RQSTRMRIRYL-HAALHQDVSFFDTDVRTS 211
Query: 137 -----------YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAG 185
S + GN +HY++ F++GF +GF WQ++LVTL++VP IA+ G
Sbjct: 212 DVIHAINADAVVVQDAISEKLGNLIHYLATFVSGFVVGFTAAWQLALVTLAVVPLIAVIG 271
Query: 186 GCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGR 245
G A L ++ + A A IAE+ + +R VQ+F GEER +R+Y AAL G
Sbjct: 272 GLSAAALAKLSSRSQDALSDASGIAEQALAQIRIVQSFVGEERVMRAYSAALAVAQRIGY 331
Query: 246 KAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAA 305
++G AKG+GLG + +F +ALL+WY +V + NGG + TM +V+I GL+LGQ+A
Sbjct: 332 RSGFAKGIGLGGTYFTVFCCYALLLWYGGHLVRRAHTNGGLAIATMFSVMIGGLALGQSA 391
Query: 306 PDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGI 365
P ++AF +A+ AA IF M+E ++ G +L + G ++ DV FSYPSRPDVGI
Sbjct: 392 PSMAAFAKARVAAAKIFRMMEHKPSMEREG--GVELEAVTGRVELRDVEFSYPSRPDVGI 449
Query: 366 FTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQ 425
L+L +PAGK +ALVG SGSGKSTVVSLIERFYEP +G ILLD +D+R+L+L+WLR+Q
Sbjct: 450 LRGLSLSVPAGKTIALVGSSGSGKSTVVSLIERFYEPNAGTILLDGHDLRDLNLRWLRRQ 509
Query: 426 IGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
IGLV+QEPALFAT+I+EN+L G+D AT EEL+ A +
Sbjct: 510 IGLVSQEPALFATTIRENLLLGRDGATQEELEEAAR 545
Score = 182 bits (461), Expect = 3e-44
Identities = 103/300 (34%), Positives = 168/300 (55%), Gaps = 1/300 (0%)
Query: 161 TIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTV 220
T GFV W+++LV L++ P + A G + +A+ RA +IA E + NVRTV
Sbjct: 896 TAGFVLQWRLALVLLAVFPLVVAATVLQKMFLKGFSGDLERAHARATQIAGEAVANVRTV 955
Query: 221 QAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKN 280
AF E + V ++A L G G G G +L+ S+AL +WY + +V
Sbjct: 956 AAFGSEAKIVGLFEANLAGPLRRCFWKGQIAGSGYGVAQFLLYASYALGLWYAAWLVKHG 1015
Query: 281 IANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRK 340
+++ ++ + +++S + F++ A +FE ++R T +
Sbjct: 1016 VSDFSKTIRVFMVLMVSANGAAETLTLAPDFVKGGRAMQAVFEAMDRRTEIEPDDVDAAA 1075
Query: 341 LSKLD-GHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERF 399
+ + G ++ V F+YPSRP+V +F +L+L AG+ +ALVG SG GKS+V++L++RF
Sbjct: 1076 VPERPRGEVELKHVDFAYPSRPEVQVFRDLSLRARAGRTLALVGASGCGKSSVLALVQRF 1135
Query: 400 YEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
YEP SG++LLD D+R+ +L+ LR+ + LV QEP LFA +I +NI YG++ AT E+ A
Sbjct: 1136 YEPNSGRVLLDGRDLRKFNLRSLRRAMALVPQEPFLFAATIHDNIAYGREGATEAEVVEA 1195
>emb|CAD59580.1| MDR-like ABC transporter [Oryza sativa (japonica cultivar-group)]
Length = 1349
Score = 344 bits (882), Expect = 4e-93
Identities = 193/456 (42%), Positives = 277/456 (60%), Gaps = 20/456 (4%)
Query: 19 KLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFPKEASHKVAKSK 78
+LFSF D DYVLM +G++GA+VHG S+ +F FF L++ G P V K
Sbjct: 83 QLFSFGDGLDYVLMTLGTLGALVHGCSLTVFLRFFADLVDSFGSHAAHPDTMLRLVVKYA 142
Query: 79 CSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y*GIHWRGHFCYYK*H-- 136
F++ + + W + S G + + R +++ + +H F
Sbjct: 143 ---FYFLVVGAAIWASSWAEISCWMWTGE-RQSTRMRIRYL-HAALHQDVSFFDTDVRTS 197
Query: 137 -----------YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVTLSIVPAIALAG 185
S + GN +HY++ F++GF +GF WQ++LVTL++VP IA+ G
Sbjct: 198 DVIHAINADAVVVQDAISEKLGNLIHYLATFVSGFVVGFTAAWQLALVTLAVVPLIAVIG 257
Query: 186 GCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYKAALMKTYVNGR 245
G A L ++ + A A IAE+ + +R VQ+F GEER +R+Y AAL G
Sbjct: 258 GLSAAALAKLSSRSQDALSDASGIAEQALAQIRIVQSFVGEERVMRAYSAALAVAQRIGY 317
Query: 246 KAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNVVISGLSLGQAA 305
++G AKG+GLG + +F +ALL+WY +V + NGG + TM +V+I GL+LGQ+A
Sbjct: 318 RSGFAKGIGLGGTYFTVFCCYALLLWYGGHLVRRAHTNGGLAIATMFSVMIGGLALGQSA 377
Query: 306 PDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVCFSYPSRPDVGI 365
P ++AF +A+ AA IF M+E ++ G +L + G ++ DV FSYPSRPDVGI
Sbjct: 378 PSMAAFAKARVAAAKIFRMMEHKPSMEREG--GVELEAVTGRVELRDVEFSYPSRPDVGI 435
Query: 366 FTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDIRELDLKWLRQQ 425
L+L +PAGK +ALVG SGSGKSTVVSLIERFYEP +G ILLD +D+R+L+L+WLR+Q
Sbjct: 436 LRGLSLSVPAGKTIALVGSSGSGKSTVVSLIERFYEPNAGTILLDGHDLRDLNLRWLRRQ 495
Query: 426 IGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
IGLV+QEPALFAT+I+EN+L G+D AT EEL+ A +
Sbjct: 496 IGLVSQEPALFATTIRENLLLGRDGATQEELEEAAR 531
Score = 182 bits (461), Expect = 3e-44
Identities = 103/300 (34%), Positives = 168/300 (55%), Gaps = 1/300 (0%)
Query: 161 TIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTV 220
T GFV W+++LV L++ P + A G + +A+ RA +IA E + NVRTV
Sbjct: 901 TAGFVLQWRLALVLLAVFPLVVAATVLQKMFLKGFSGDLERAHARATQIAGEAVANVRTV 960
Query: 221 QAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKN 280
AF E + V ++A L G G G G +L+ S+AL +WY + +V
Sbjct: 961 AAFGSEAKIVGLFEANLAGPLRRCFWKGQIAGSGYGVAQFLLYASYALGLWYAAWLVKHG 1020
Query: 281 IANGGESFTTMLNVVISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRK 340
+++ ++ + +++S + F++ A +FE ++R T +
Sbjct: 1021 VSDFSKTIRVFMVLMVSANGAAETLTLAPDFVKGGRAMQAVFEAMDRRTEIEPDDVDAAA 1080
Query: 341 LSKLD-GHIQFNDVCFSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERF 399
+ + G ++ V F+YPSRP+V +F +L+L AG+ +ALVG SG GKS+V++L++RF
Sbjct: 1081 VPERPRGEVELKHVDFAYPSRPEVQVFRDLSLRARAGRTLALVGASGCGKSSVLALVQRF 1140
Query: 400 YEPISGQILLDKNDIRELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRA 459
YEP SG++LLD D+R+ +L+ LR+ + LV QEP LFA +I +NI YG++ AT E+ A
Sbjct: 1141 YEPNSGRVLLDGRDLRKFNLRSLRRAMALVPQEPFLFAATIHDNIAYGREGATEAEVVEA 1200
>ref|XP_483818.1| putative P-glycoprotein 1 [Oryza sativa (japonica cultivar-group)]
gi|45735907|dbj|BAD12939.1| putative P-glycoprotein 1
[Oryza sativa (japonica cultivar-group)]
Length = 952
Score = 342 bits (877), Expect = 1e-92
Identities = 194/467 (41%), Positives = 282/467 (59%), Gaps = 20/467 (4%)
Query: 8 ERKKEHKVSMLKLFSFADSYDYVLMFIGSIGAIVHGASVPIFFIFFGKLINVIGLAYLFP 67
E + + +V +L + AD DYVLM +G++GA+VHG S+P+F FF L++ G P
Sbjct: 2 EEEIKGRVVVLGADAAADGLDYVLMTLGTLGALVHGCSLPVFLRFFADLVDSFGSHAAHP 61
Query: 68 KEASHKVAKSKCSNIIFILDRGGLLDAYWRKTSSKDENGLFKVYVKSRYKFI*Y*GIHWR 127
V K F++ + + W + S G + + R +++ + +H
Sbjct: 62 DTMLRLVVKYA---FYFLVVGAAIWASSWAEISCWMWTGE-RQSTRMRIRYL-HAALHQD 116
Query: 128 GHFCYYK*H-------------YYCSRCSIREGNFLHYISRFIAGFTIGFVRVWQISLVT 174
F S + GN +HY++ F++GF +GF WQ++LVT
Sbjct: 117 VSFFDTDVRTSDVIHAINADAVVVQDAISEKLGNLIHYLATFVSGFVVGFTAAWQLALVT 176
Query: 175 LSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTVQAFAGEERAVRSYK 234
L++VP IA+ GG A L ++ + A A IAE+ + +R VQ+F GEER +R+Y
Sbjct: 177 LAVVPLIAVIGGLSAAALAKLSSRSQDALSDASGIAEQALAQIRIVQSFVGEERVMRAYS 236
Query: 235 AALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKNIANGGESFTTMLNV 294
AAL G ++G AKG+GLG + +F +ALL+WY +V + NGG + TM +V
Sbjct: 237 AALAVAQRIGYRSGFAKGIGLGGTYFTVFCCYALLLWYGGHLVRRAHTNGGLAIATMFSV 296
Query: 295 VISGLSLGQAAPDISAFIRAKAAAYPIFEMIERDTVSKKSSKTGRKLSKLDGHIQFNDVC 354
+I GL+LGQ+AP ++AF +A+ AA IF M+E ++ G +L + G ++ DV
Sbjct: 297 MIGGLALGQSAPSMAAFAKARVAAAKIFRMMEHKPSMEREG--GVELEAVTGRVELRDVE 354
Query: 355 FSYPSRPDVGIFTNLNLDIPAGKIVALVGGSGSGKSTVVSLIERFYEPISGQILLDKNDI 414
FSYPSRPDVGI L+L +PAGK +ALVG SGSGKSTVVSLIERFYEP +G ILLD +D+
Sbjct: 355 FSYPSRPDVGILRGLSLSVPAGKTIALVGSSGSGKSTVVSLIERFYEPNAGTILLDGHDL 414
Query: 415 RELDLKWLRQQIGLVNQEPALFATSIKENILYGKDDATLEELKRAVK 461
R+L+L+WLR+QIGLV+QEPALFAT+I+EN+L G+D AT EEL+ A +
Sbjct: 415 RDLNLRWLRRQIGLVSQEPALFATTIRENLLLGRDGATQEELEEAAR 461
Score = 67.4 bits (163), Expect = 9e-10
Identities = 42/138 (30%), Positives = 70/138 (50%), Gaps = 3/138 (2%)
Query: 161 TIGFVRVWQISLVTLSIVPAIALAGGCYAYVTIGLIAKVRKAYVRAGEIAEEVIGNVRTV 220
T GFV W+++LV L++ P + A G + +A+ RA +IA E + NVRTV
Sbjct: 812 TAGFVLQWRLALVLLAVFPLVVAATVLQKMFLKGFSGDLERAHARATQIAGEAVANVRTV 871
Query: 221 QAFAGEERAVRSYKAALMKTYVNGRKAGLAKGLGLGSMHCVLFLSWALLVWYTSVVVHKN 280
AF E + V ++A L G G G G +L+ S+AL +WY + +V
Sbjct: 872 AAFGSEAKIVGLFEANLAGPLRRCFWKGQIAGSGYGVAQFLLYASYALGLWYAAWLVKHG 931
Query: 281 IANGGES---FTTMLNVV 295
+++ ++ F +L+V+
Sbjct: 932 VSDFSKTIRVFMLLLDVL 949
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.327 0.142 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 739,008,892
Number of Sequences: 2540612
Number of extensions: 30200332
Number of successful extensions: 133784
Number of sequences better than 10.0: 16213
Number of HSP's better than 10.0 without gapping: 12835
Number of HSP's successfully gapped in prelim test: 3378
Number of HSP's that attempted gapping in prelim test: 113561
Number of HSP's gapped (non-prelim): 20665
length of query: 461
length of database: 863,360,394
effective HSP length: 131
effective length of query: 330
effective length of database: 530,540,222
effective search space: 175078273260
effective search space used: 175078273260
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 77 (34.3 bits)
Medicago: description of AC087771.6


