

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
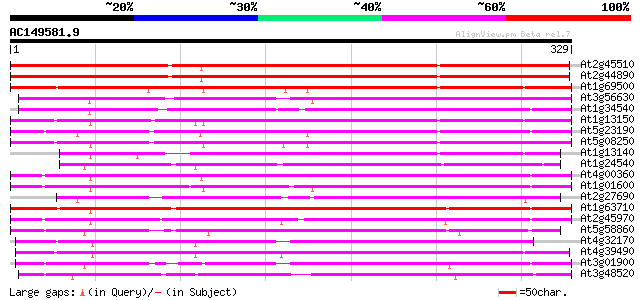
Query= AC149581.9 - phase: 0 /pseudo
(329 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g45510 putative cytochrome P450 328 2e-90
At2g44890 putative cytochrome P450 326 1e-89
At1g69500 unknown protein 259 2e-69
At3g56630 cytochrome P450-like protein 235 3e-62
At1g34540 cytochrome p450, putative 222 2e-58
At1g13150 putative cytochrome P450 monooxygenase 219 2e-57
At5g23190 cytochrome P450-like protein 216 2e-56
At5g08250 cytochrome P450-like protein 213 9e-56
At1g13140 cytochrome P450 monooxygenase like protein 210 1e-54
At1g24540 putative cytochrome P450 206 2e-53
At4g00360 probable cytochrome P450 205 3e-53
At1g01600 unknown protein 204 4e-53
At2g27690 putative cytochrome P450 202 2e-52
At1g63710 cytochrome P450 like protein 202 2e-52
At2g45970 putative cytochrome P450 200 8e-52
At5g58860 cytochrome P450 CYP86A1 196 2e-50
At4g32170 cytochrome p450 - like protein 192 3e-49
At4g39490 cytochrome P450 - like protein 189 2e-48
At3g01900 putative cytochrome P450 189 2e-48
At3g48520 cytochrome P450-like protein 188 4e-48
>At2g45510 putative cytochrome P450
Length = 511
Score = 328 bits (841), Expect = 2e-90
Identities = 171/334 (51%), Positives = 234/334 (69%), Gaps = 9/334 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M+ TLDS+FKV GVEL + G SKEG +F AF+E + +++ LWK++ F NIGS
Sbjct: 178 MRCTLDSIFKVGFGVELKCLDGFSKEGQEFMEAFDEGNVATSSRFIDPLWKLKWFFNIGS 237
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETD-----SKYL 115
++ LKK++ I+++VY++I +K ++ K Q ++ DILSRFL +E D KYL
Sbjct: 238 QSKLKKSIATIDKFVYSLITTKRKELAKEQNTV--VREDILSRFLVESEKDPENMNDKYL 295
Query: 116 KDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGST-TDELATRI 174
+D+IL+F+IAGKDTT+ LS F+Y LCK+P VQEKI QEIR+ T + +T + I
Sbjct: 296 RDIILNFMIAGKDTTAALLSWFLYMLCKNPLVQEKIVQEIRDVTFSHEKTTDVNGFVESI 355
Query: 175 TEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRM 234
EE++ +M YL AAL+ETLRL+PP+P++ + +DD LPDG+ V KGD I + Y MGRM
Sbjct: 356 NEEALDEMHYLHAALSETLRLYPPVPVDMRCAENDDVLPDGHRVSKGDNIYYIAYAMGRM 415
Query: 235 EFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLG 294
++WG+DAE+F+PERWL ++G FQ ESPFKF +F AGPRICLGK+F YRQMKI S LL
Sbjct: 416 TYIWGQDAEEFKPERWL-KDGLFQPESPFKFISFHAGPRICLGKDFAYRQMKIVSMALLH 474
Query: 295 SHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
FK+AD+N V Y+ LTL +DGGLH+ A R
Sbjct: 475 FFRFKMADENSKVYYKRMLTLHVDGGLHLCAIPR 508
>At2g44890 putative cytochrome P450
Length = 490
Score = 326 bits (835), Expect = 1e-89
Identities = 169/334 (50%), Positives = 233/334 (69%), Gaps = 9/334 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
MK TLDS+FKV GVEL + G SKEG +F AF+E + + WK++ FLNIGS
Sbjct: 157 MKCTLDSIFKVGFGVELGCLDGFSKEGEEFMKAFDEGNGATSSRVTDPFWKLKCFLNIGS 216
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETD-----SKYL 115
E+ LKK++ +I+++VY++I +K ++ K Q S ++ DILS+FL +E D KYL
Sbjct: 217 ESRLKKSIAIIDKFVYSLITTKRKELSKEQNTS--VREDILSKFLLESEKDPENMNDKYL 274
Query: 116 KDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGST-TDELATRI 174
+D+IL+ ++AGKDTT+ +LS F+Y LCK+P VQEKI QEIR+ T + +T + +
Sbjct: 275 RDIILNVMVAGKDTTAASLSWFLYMLCKNPLVQEKIVQEIRDVTSSHEKTTDVNGFIESV 334
Query: 175 TEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRM 234
TEE++ +MQYL AAL+ET+RL+PP+P + +DD LPDG+ V KGD I + Y MGRM
Sbjct: 335 TEEALAQMQYLHAALSETMRLYPPVPEHMRCAENDDVLPDGHRVSKGDNIYYISYAMGRM 394
Query: 235 EFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLG 294
++WG+DAE+F+PERWL ++G FQ ES FKF +F AGPRIC+GK+F YRQMKI S LL
Sbjct: 395 TYIWGQDAEEFKPERWL-KDGVFQPESQFKFISFHAGPRICIGKDFAYRQMKIVSMALLH 453
Query: 295 SHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
FK+AD+N V Y+ LTL +DGGLH+ A R
Sbjct: 454 FFRFKMADENSKVSYKKMLTLHVDGGLHLCAIPR 487
>At1g69500 unknown protein
Length = 524
Score = 259 bits (661), Expect = 2e-69
Identities = 145/360 (40%), Positives = 219/360 (60%), Gaps = 34/360 (9%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M+ TLDS+ KV GVE+ T+ E F+ AF+ A+ + +++ LWK+++FLNIGS
Sbjct: 168 MRMTLDSICKVGFGVEIGTLAPELPEN-HFAKAFDTANIIVTLRFIDPLWKMKKFLNIGS 226
Query: 61 EAVLKKNLRVINEYVYTIIK--------SKIEQSQKPQKNSPELKGDILSRFLELNETDS 112
EA+L K+++V+N++ Y++I+ ++I + N+ ++K DILSRF+E+++
Sbjct: 227 EALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNNNNNNNNKVKHDILSRFIEISDDPD 286
Query: 113 -----KYLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATK------- 160
K L+D++L+F+IAG+DTT+ TL+ IY + + +V EK+ E++E K
Sbjct: 287 SKETEKSLRDIVLNFVIAGRDTTATTLTWAIYMIMMNENVAEKLYSELQELEKESAEATN 346
Query: 161 -------VEDGSTTDELATR----ITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSD 209
ED ++ +E T + +S+ K+ YL A +TETLRL+P +P + K D
Sbjct: 347 TSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHYLHAVITETLRLYPAVPQDPKGVLED 406
Query: 210 DTLPDGYSVRKGDYISFHPYVMGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQ 269
D LP+G V+ G +++ PY MGRME+ WG DA F+PERWL ++G FQ SPFKFTAFQ
Sbjct: 407 DMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAALFKPERWL-KDGVFQNASPFKFTAFQ 465
Query: 270 AGPRICLGKEFVYRQMKIFSAVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
AGPRICLGK+ Y QMK+ A+L + F L N VKYR L + GL V R+
Sbjct: 466 AGPRICLGKDSAYLQMKMAMAILCRFYKFHLV-PNHPVKYRMMTILSMAHGLKVTVSRRS 524
>At3g56630 cytochrome P450-like protein
Length = 499
Score = 235 bits (599), Expect = 3e-62
Identities = 129/333 (38%), Positives = 202/333 (59%), Gaps = 22/333 (6%)
Query: 6 DSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWY---VNFLWKVQRFLNIGSEA 62
D++ K+ V+ + G F AF A+ I + +++ WK+++ LNIGSE
Sbjct: 180 DNICKLAFNVDSACLGDDGAAGVNFMQAFETAATIISQRFQSVISYSWKIKKKLNIGSER 239
Query: 63 VLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDS-KYLKDVILS 121
VL++++ +++++ I++++IEQ + K D+LSRF+ E +S + L+D+++S
Sbjct: 240 VLRESIMIVHKFADEIVRNRIEQGKVSDH-----KEDLLSRFISKEEMNSPEILRDIVIS 294
Query: 122 FIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITE----E 177
FI+AG+DTTS LS F + L HP V++KI QE+ S + RI E E
Sbjct: 295 FILAGRDTTSSALSWFFWLLSMHPEVKDKILQELN--------SIRERTGKRIGEVYGFE 346
Query: 178 SMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEFL 237
+K M YL AA+TE+LRL+PP+P+++ C D+ LPDG + K IS++ Y MGRME +
Sbjct: 347 DLKLMNYLHAAITESLRLYPPVPVDTMSCAEDNVLPDGTFIGKDWGISYNAYAMGRMESI 406
Query: 238 WGEDAEQFRPERWLDE-NGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSH 296
WG+D ++F PERW+DE NG F+ E+P+KF AF AGPR+CLGKE Y QMK A +L
Sbjct: 407 WGKDCDRFDPERWIDETNGGFRGENPYKFPAFHAGPRMCLGKEMAYIQMKSIVAAVLERF 466
Query: 297 NFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
++ + + ++TL+I GGL+V R+
Sbjct: 467 VVEVPGKKERPEILMSVTLRIRGGLNVRVQERS 499
>At1g34540 cytochrome p450, putative
Length = 498
Score = 222 bits (566), Expect = 2e-58
Identities = 125/329 (37%), Positives = 199/329 (59%), Gaps = 15/329 (4%)
Query: 6 DSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWY---VNFLWKVQRFLNIGSEA 62
D++ K+ V+ + G F AF A+ I + + W++++ LNIGSE
Sbjct: 180 DNICKLAFNVDCACLGHDGAVGVNFMRAFETAATIISQRFRSVASCAWRIKKKLNIGSER 239
Query: 63 VLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDS-KYLKDVILS 121
VL++++ ++++ I++++I+Q + S + K D+LSRF+ E +S + L+D+++S
Sbjct: 240 VLRESIATVHKFADEIVRNRIDQGR-----SSDHKEDLLSRFISKEEMNSPEILRDIVIS 294
Query: 122 FIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITEESMKK 181
FI+AG+DTTS LS F + L HP V++KI QE+ + + G E+ E +K
Sbjct: 295 FILAGRDTTSSALSWFFWLLSMHPEVEDKILQELN-SIRARTGKRIGEV---YGFEHLKM 350
Query: 182 MQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEFLWGED 241
M YL AA+TE+LRL+PP+P++ K C D+ LPDG V KG I+++ + MGRME +WG+D
Sbjct: 351 MNYLHAAITESLRLYPPVPVDIKSCAEDNVLPDGTFVGKGWAITYNIFAMGRMESIWGKD 410
Query: 242 AEQFRPERWLDE-NGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSHNFKL 300
++F PERW+DE NG F+ E P KF AF AGPR+C+GK+ Y QMK A +L ++
Sbjct: 411 CDRFDPERWIDETNGCFRGEDPSKFPAFHAGPRMCVGKDMAYIQMKSIVAAVLERFVVEV 470
Query: 301 ADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
+ R + ++TL+I GGL R+
Sbjct: 471 PGKER-PEILLSMTLRIKGGLFARVQERS 498
>At1g13150 putative cytochrome P450 monooxygenase
Length = 529
Score = 219 bits (558), Expect = 2e-57
Identities = 128/340 (37%), Positives = 196/340 (57%), Gaps = 16/340 (4%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYV--NFLWKVQRFLNI 58
++ T D + LG + +T+ + F+ AF EA+ + +F ++ F+WK RFL+
Sbjct: 184 LRLTFDIICLAGLGADPETLAVDLPQ-VPFAKAFEEATESTLFRFMIPPFIWKPMRFLDT 242
Query: 59 GSEAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLEL------NETDS 112
G E L+ + V++ +V +I +I + ++ + + + D+LSR +++ NE D
Sbjct: 243 GYEKGLRIAVGVVHGFVDKMIVDRI--CELKEEETLDNRSDVLSRIIQIESHKRENEIDP 300
Query: 113 ---KYLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDE 169
++ + SFI+AG+DT+S+ LS F + + KHP V+ KI EIRE + S T +
Sbjct: 301 STIRFFRQFCTSFILAGRDTSSVALSWFCWVIQKHPEVENKIICEIREILRQRGDSPTSK 360
Query: 170 LATRITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPY 229
+ T + + M YL AAL+ETLRL PPIPME K DD LPDG VRKG + F Y
Sbjct: 361 NESLFTVKELNNMVYLQAALSETLRLFPPIPMEMKQAIEDDVLPDGTFVRKGSRVYFSIY 420
Query: 230 VMGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFS 289
MGRME +WG+D E FRPERW+ + G F + FK+ F AGPR+C+GK F Y QMK+ +
Sbjct: 421 AMGRMESIWGKDCEIFRPERWI-QAGKFVSDDQFKYVVFNAGPRLCIGKTFAYLQMKMIA 479
Query: 290 AVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
A +L ++ K+ Q+ ++ R T L + GL V R+
Sbjct: 480 ASVLLRYSIKVV-QDHVIAPRVTTNLYMKYGLKVTITPRS 518
>At5g23190 cytochrome P450-like protein
Length = 559
Score = 216 bits (549), Expect = 2e-56
Identities = 120/339 (35%), Positives = 202/339 (59%), Gaps = 15/339 (4%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEAS--ATIMFWYVNFLWKVQRFLNI 58
++ T D+V + GV+ + G + F+ AF +A+ A + F +WK R+L+I
Sbjct: 208 LRLTFDNVCMIAFGVDPGCL-GPDQPVIPFAKAFEDATEAAVVRFVMPTCVWKFMRYLDI 266
Query: 59 GSEAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNET-----DSK 113
G+E LK++++ ++++ +I+++ + + + + D+L+ F+ L + K
Sbjct: 267 GTEKKLKESIKGVDDFADEVIRTR--KKELSLEGETTKRSDLLTVFMGLRDEKGESFSDK 324
Query: 114 YLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATR 173
+L+D+ ++FI+AG+DT+S+ LS F + L K+P V+EKI E+ + + D E +
Sbjct: 325 FLRDICVNFILAGRDTSSVALSWFFWLLEKNPEVEEKIMVEMCKILRQRDDHGNAEKSDY 384
Query: 174 ---ITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYV 230
E +KKM YL AAL+E LRL+P +P++ K DD PDG ++KGD + + Y
Sbjct: 385 EPVFGPEEIKKMDYLQAALSEALRLYPSVPVDHKEVQEDDVFPDGTMLKKGDKVIYAIYA 444
Query: 231 MGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSA 290
MGRME +WG+D +FRPERWL +G F ES +KFTAF GPR+CLGK+F Y QMK +A
Sbjct: 445 MGRMEAIWGKDCLEFRPERWL-RDGRFMSESAYKFTAFNGGPRLCLGKDFAYYQMKSTAA 503
Query: 291 VLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
++ + K+ + ++ V+ + LT+ + GL VN +R+
Sbjct: 504 AIVYRYKVKVVNGHK-VEPKLALTMYMKHGLMVNLINRS 541
>At5g08250 cytochrome P450-like protein
Length = 550
Score = 213 bits (543), Expect = 9e-56
Identities = 117/341 (34%), Positives = 202/341 (58%), Gaps = 16/341 (4%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYV--NFLWKVQRFLNI 58
++ T D+V + GV+ + E F+ AF +A+ + +V F+WK+ R LN+
Sbjct: 205 LRLTFDNVCMIAFGVDPGCLSPKLPE-IPFAKAFEDATEATVVRFVMPKFVWKLMRSLNL 263
Query: 59 GSEAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDS-----K 113
G+E LK+++ ++++ +I+++ + + + + D+L+ F+ L + + K
Sbjct: 264 GTEKKLKESINGVDDFAEEVIRTR--KKEMSLETEIAKRPDLLTIFMGLRDENGQKFSDK 321
Query: 114 YLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREAT--KVEDGSTTDELA 171
+L+D+ ++FI+AG+DT+S+ LS F + + K+P V+EKI I + +V+ G T +
Sbjct: 322 FLRDICVNFILAGRDTSSVALSWFFWLIEKNPEVEEKIMMGICKILEQRVDHGDTKKNME 381
Query: 172 TR--ITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPY 229
E +KKM YL AAL+ETLRL+P +P++ K DD PDG ++KG+ + + Y
Sbjct: 382 YEPVFRPEEIKKMDYLQAALSETLRLYPSVPVDHKEVLEDDVFPDGTKLKKGEKVIYAIY 441
Query: 230 VMGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFS 289
MGRME +WG+D +F+PERWL +G + ES +KFTAF GPR+CLGK+F Y QM+ +
Sbjct: 442 AMGRMETIWGKDCREFKPERWL-RDGRYMSESAYKFTAFNGGPRLCLGKDFAYYQMRYVA 500
Query: 290 AVLLGSHNFKLADQ-NRLVKYRTTLTLQIDGGLHVNAFHRN 329
A ++ + ++ D+ V+ + LT+ + GL VN R+
Sbjct: 501 AAIIYRYKVRVDDKGGHKVEPKMALTMYMKHGLKVNMVKRS 541
>At1g13140 cytochrome P450 monooxygenase like protein
Length = 519
Score = 210 bits (534), Expect = 1e-54
Identities = 118/302 (39%), Positives = 172/302 (56%), Gaps = 24/302 (7%)
Query: 30 FSNAFNEASATIMFWYV--NFLWKVQRFLNIGSEAVLKKNLRVINE------YVYTIIKS 81
F+ AF EA+ + MF ++ F+WK +F +IG E L+K + V ++ +
Sbjct: 219 FAQAFEEATESTMFRFMIPPFIWKPLKFFDIGYEKGLRKAVDVSMSLSTRWLWIVSASSK 278
Query: 82 KIEQSQKPQKNSPELKGDILSRFLELNETDSKYLKDVILSFIIAGKDTTSITLS*FIYQL 141
K EQS K E + + K+ + SFI+AG+DT+S+ L+ F + +
Sbjct: 279 KKEQSHKTTD--------------EKDPSTIKFFRQFCTSFILAGRDTSSVALTWFFWVI 324
Query: 142 CKHPHVQEKIAQEIREATKVEDGSTTDELATRITEESMKKMQYLDAALTETLRLHPPIPM 201
KHP V+ KI +EI E + S T + + T + + M YL AAL+ET+RL+PPIPM
Sbjct: 325 QKHPEVENKIIREISEILRQRGDSPTSKNESLFTVKELNDMVYLQAALSETMRLYPPIPM 384
Query: 202 ESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEFLWGEDAEQFRPERWLDENGNFQRES 261
E K DD PDG +RKG + F Y MGRME +WG+D E F+PERW+ ++GNF +
Sbjct: 385 EMKQAIEDDVFPDGTFIRKGSRVYFATYAMGRMESIWGKDCESFKPERWI-QSGNFVNDD 443
Query: 262 PFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSHNFKLADQNRLVKYRTTLTLQIDGGL 321
FK+ F AGPR+CLGK F Y QMK +A +L ++ K+A ++ +V R T TL + GL
Sbjct: 444 QFKYVVFNAGPRLCLGKTFAYLQMKTIAASVLSRYSIKVA-KDHVVVPRVTTTLYMRHGL 502
Query: 322 HV 323
V
Sbjct: 503 KV 504
>At1g24540 putative cytochrome P450
Length = 522
Score = 206 bits (523), Expect = 2e-53
Identities = 116/307 (37%), Positives = 175/307 (56%), Gaps = 20/307 (6%)
Query: 30 FSNAFNEASATIM--FWYVNFLWKVQRFLNIGSEAVLKKNLRVINEYVYTIIKSKIEQSQ 87
F+ AF +A+ + F F+WK RFL IG E L +R+++ + ++ + + +
Sbjct: 216 FAKAFEDATEYTLARFLIPPFVWKPMRFLGIGYERKLNNAVRIVHAFANKTVRERRNKMR 275
Query: 88 KPQKNSPELKGDILSRFLEL-----------NETDSKYLKDVILSFIIAGKDTTSITLS* 136
K + D+LSR ++ N KY ++ SFIIAG+DTTS+ L
Sbjct: 276 KLGNLNDY--ADLLSRLMQREYEKEEDTTRGNYFSDKYFREFCTSFIIAGRDTTSVALVW 333
Query: 137 FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITEESMKKMQYLDAALTETLRLH 196
F + + KHP V+++I +EIRE ++ TT E + E ++M YL AALTE+LRL+
Sbjct: 334 FFWLVQKHPEVEKRILREIRE---IKRKLTTQETEDQFEAEDFREMVYLQAALTESLRLY 390
Query: 197 PPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEFLWGEDAEQFRPERWLDENGN 256
P +PME K DD LPDG V+KG I + Y MGR+E +WG+D E+F+PERW+ E G
Sbjct: 391 PSVPMEMKQALEDDVLPDGTRVKKGARIHYSVYSMGRIESIWGKDWEEFKPERWIKE-GR 449
Query: 257 FQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSHNFKLADQNRLVKYRTTLTLQ 316
E FK+ F GPR+C+GK+F Y QMK+ +A +L ++ K+ +V TT TL
Sbjct: 450 IVSEDQFKYVVFNGGPRLCVGKKFAYTQMKMVAAAILMRYSVKVVQGQEIVPKLTT-TLY 508
Query: 317 IDGGLHV 323
+ G++V
Sbjct: 509 MKNGMNV 515
>At4g00360 probable cytochrome P450
Length = 553
Score = 205 bits (521), Expect = 3e-53
Identities = 113/335 (33%), Positives = 194/335 (57%), Gaps = 8/335 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYV--NFLWKVQRFLNI 58
++ T D++ + G + T C F++AF+ A+ + ++ FLW+++++L +
Sbjct: 180 LRLTFDNICGLAFGKDTRT-CAPGLPENGFASAFDRATEASLQRFILPEFLWRLKKWLGL 238
Query: 59 GSEAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETD--SKYLK 116
G E L ++L I+ Y+ +I ++ ++ +++ + D+LSRF++ + +L+
Sbjct: 239 GLEVSLSRSLGEIDGYLDAVINTRKQELLSQRESGVQRHDDLLSRFMKKKDQSYSETFLR 298
Query: 117 DVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITE 176
V L+FI+AG+DT+S+ LS F + + HP V++KI +EI G+ E
Sbjct: 299 HVALNFILAGRDTSSVALSWFFWLITTHPTVEDKIVREICSVLIETRGTDVSSWTAEPLE 358
Query: 177 -ESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRME 235
+ + ++ YL AAL+ETLRL+P +P +SK+ +DD LPDG V G +++ Y GRM+
Sbjct: 359 FDEVDRLVYLKAALSETLRLYPSVPEDSKHVVNDDILPDGTFVPAGSSVTYSIYAAGRMK 418
Query: 236 FLWGEDAEQFRPERWLD-ENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLG 294
WGED +F+PERW+ ++G F ++F AF AGPRICLGK+ Y QMK +A +L
Sbjct: 419 STWGEDCLEFKPERWISPDDGKFVNHDQYRFVAFNAGPRICLGKDLAYLQMKTIAAAVLL 478
Query: 295 SHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
H +A ++ V+ + +LTL + GL VN R+
Sbjct: 479 RHRLTVAPGHK-VEQKMSLTLFMKNGLLVNVHKRD 512
>At1g01600 unknown protein
Length = 554
Score = 204 bits (520), Expect = 4e-53
Identities = 118/340 (34%), Positives = 201/340 (58%), Gaps = 16/340 (4%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYV--NFLWKVQRFLNI 58
++ T D++ + G + T C F++AF+ A+ + ++ F+WK++++L +
Sbjct: 180 LRLTFDNICGLAFGKDTRT-CAPGLPENGFASAFDRATEASLQRFIIPKFMWKLKKWLGL 238
Query: 59 GSEAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELK-GDILSRFLELNETDS---KY 114
G E L ++L I+EY+ +I ++ ++ Q++ + D+LSRF+ + +T+S +
Sbjct: 239 GLEVSLSRSLGEIDEYLAAVINTRKQELMSQQESGTHQRHDDLLSRFM-MKKTESYSDTF 297
Query: 115 LKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRI 174
L+ V L+FI+AG+DT+S+ LS F + + HP V++KI +EI G TD++A+
Sbjct: 298 LQHVALNFILAGRDTSSVALSWFFWLITMHPTVEDKIVREICSVLIETRG--TDDVASWT 355
Query: 175 TE----ESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYV 230
E + + ++ YL AA++ETLRL+P +P +SK+ +DD LPDG V G +++ Y
Sbjct: 356 EEPLGFDEIDRLVYLKAAISETLRLYPSVPEDSKHVENDDVLPDGTFVPAGSSVTYSIYA 415
Query: 231 MGRMEFLWGEDAEQFRPERWLDE-NGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFS 289
GRM+ WGED +F PERW+ +G F ++F AF AGPRICLGK+ Y QMK +
Sbjct: 416 AGRMKSTWGEDCLEFNPERWISPIDGKFINHDQYRFVAFNAGPRICLGKDLAYLQMKTIA 475
Query: 290 AVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
A +L H + ++ V+ + +LTL + GL VN + R+
Sbjct: 476 AAVLLRHRLTVVPGHK-VEQKMSLTLFMKNGLLVNLYKRD 514
>At2g27690 putative cytochrome P450
Length = 495
Score = 202 bits (514), Expect = 2e-52
Identities = 109/302 (36%), Positives = 176/302 (58%), Gaps = 19/302 (6%)
Query: 28 TQFSNAFNEAS---ATIMFWYVNFLWKVQRFLNIGSEAVLKKNLRVINEYVYTIIKSKIE 84
++F+ AF+ AS A LWK +R L IGSE L++++ VIN +IK +
Sbjct: 201 SEFAVAFDTASLLSAKRALAPFPLLWKTKRLLRIGSEKKLQESINVINRLAGDLIKQR-- 258
Query: 85 QSQKPQKNSPELKGDILSRFLEL-NETDSKYLKDVILSFIIAGKDTTSITLS*FIYQLCK 143
+ K D++SRF+ + E D +YL+D+++SF++AG+DT + L+ F + L +
Sbjct: 259 -----RLTGLMGKNDLISRFMAVVAEDDDEYLRDIVVSFLLAGRDTVAAGLTGFFWLLTR 313
Query: 144 HPHVQEKIAQEIREATKVEDGSTTDELATRITEESMKKMQYLDAALTETLRLHPPIPMES 203
HP V+ +I +E+ G+ D + R E M++M YL A+L E++RL PP+ +S
Sbjct: 314 HPEVENRIREELDRVM----GTGFDSVTARCDE--MREMDYLHASLYESMRLFPPVQFDS 367
Query: 204 KYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEFLWGEDAEQFRPERWLDENGNFQRESPF 263
K+ +DD L DG V G +++H Y MGRM+ +WG D E+F+PERWLD G F+ E+P
Sbjct: 368 KFALNDDVLSDGTFVNSGTRVTYHAYAMGRMDRIWGPDYEEFKPERWLDNEGKFRPENPV 427
Query: 264 KFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSHNFKLA--DQNRLVKYRTTLTLQIDGGL 321
K+ FQAG R+C+GKE +MK + ++ ++A + +++ LT ++GGL
Sbjct: 428 KYPVFQAGARVCIGKEMAIMEMKSIAVAIIRRFETRVASPETTETLRFAPGLTATVNGGL 487
Query: 322 HV 323
V
Sbjct: 488 PV 489
>At1g63710 cytochrome P450 like protein
Length = 523
Score = 202 bits (514), Expect = 2e-52
Identities = 120/334 (35%), Positives = 204/334 (60%), Gaps = 10/334 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYV--NFLWKVQRFLNI 58
++ T D++ + G + T+ E F+ AF+ A+ + ++ F+WK++++L +
Sbjct: 180 LRLTFDNICGLTFGKDPRTLSPEFPENG-FAVAFDGATEATLQRFIMPEFIWKIRKWLRL 238
Query: 59 GSEAVLKKNLRVINEYVYTIIKS-KIEQSQKPQKNSPELKGDILSRFLELNETDS-KYLK 116
G E + +++ ++ Y+ II + K+E + Q S D+LSRF++ E+ S KYLK
Sbjct: 239 GLEDDMSRSISHVDNYLSEIINTRKLELLGQQQDESRH--DDLLSRFMKKKESYSDKYLK 296
Query: 117 DVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREAT-KVEDGSTTDELATRIT 175
V L+FI+AG+DT+S+ +S F + + +P V+EKI EI K D + + +T
Sbjct: 297 YVALNFILAGRDTSSVAMSWFFWLVSLNPRVEEKIINEICTILIKTRDTNVSKWTDEPLT 356
Query: 176 EESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRME 235
+ + ++ YL AAL+ETLRL+P +P +SK+ ++D LPDG V G +++ Y +GRM+
Sbjct: 357 FDEIDQLVYLKAALSETLRLYPSVPEDSKFVVANDVLPDGTFVPSGSNVTYSIYSVGRMK 416
Query: 236 FLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGS 295
F+WGED +F+PERWL+E+ + ++ + +KF AF AGPRICLGK+ Y QMK +A +L
Sbjct: 417 FIWGEDCLEFKPERWLEESRD-EKCNQYKFVAFNAGPRICLGKDLAYLQMKSITASILLR 475
Query: 296 HNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
H +A +R V+ + +LTL + GL ++ R+
Sbjct: 476 HRLTVAPGHR-VEQKMSLTLFMKFGLKMDVHKRD 508
>At2g45970 putative cytochrome P450
Length = 537
Score = 200 bits (509), Expect = 8e-52
Identities = 123/338 (36%), Positives = 199/338 (58%), Gaps = 15/338 (4%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYV--NFLWKVQRFLNI 58
++ T D++ + G + T C F+ AF+ A+ + ++ LWK +R+L +
Sbjct: 180 LRLTFDNICGLTFGKDPRT-CAPGLPVNTFAVAFDRATEASLQRFILPEILWKFKRWLRL 238
Query: 59 GSEAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDS-KYLKD 117
G E L ++L ++ Y+ II ++ E+ Q N+ + D+LSRF++ E+ S + L+
Sbjct: 239 GLEVSLTRSLVQVDNYLSEIITTRKEEMMT-QHNNGKHHDDLLSRFIKKKESYSDETLQR 297
Query: 118 VILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREA---TKVEDGST-TDELATR 173
V L+FI+AG+DT+S+ LS F + + +HP +++KI +EI T+ +D + TDE
Sbjct: 298 VALNFILAGRDTSSVALSWFFWLITQHPAIEDKILREICTVLVETRGDDVALWTDE---P 354
Query: 174 ITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGR 233
++ E + ++ +L AAL+ETLRL+P +P +SK DD LPDG V G I++ Y GR
Sbjct: 355 LSCEELDRLVFLKAALSETLRLYPSVPEDSKRAVKDDVLPDGTFVPAGSSITYSIYSAGR 414
Query: 234 MEFLWGEDAEQFRPERWLDEN--GNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAV 291
M+ WGED +F+PERW+ ++ G F PFKF AF AGPRICLGK+ Y QMK ++
Sbjct: 415 MKSTWGEDCLEFKPERWISQSDGGRFINHDPFKFVAFNAGPRICLGKDLAYLQMKSIASA 474
Query: 292 LLGSHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
+L H + ++ V+ + +LTL + GL VN R+
Sbjct: 475 VLLRHRLTVVTGHK-VEQKMSLTLFMKYGLLVNVHERD 511
>At5g58860 cytochrome P450 CYP86A1
Length = 513
Score = 196 bits (497), Expect = 2e-50
Identities = 122/335 (36%), Positives = 187/335 (55%), Gaps = 26/335 (7%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIM--FWYVNFLWKVQRFLNI 58
++ T D++ + G + +T+ FS AF+ A+ + Y FLW++Q+ + I
Sbjct: 183 LRLTFDNICGLTFGKDPETL-SLDLPDNPFSVAFDTATEATLKRLLYTGFLWRIQKAMGI 241
Query: 59 GSEAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDSKYL--- 115
GSE LKK+L V+ Y+ I ++ KNSP D+LSRFL+ + + L
Sbjct: 242 GSEDKLKKSLEVVETYMNDAIDAR--------KNSPS--DDLLSRFLKKRDVNGNVLPTD 291
Query: 116 --KDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATR 173
+ + L+F++AG+DT+S+ LS F + + + V+ KI E+ K G+ ++
Sbjct: 292 VLQRIALNFVLAGRDTSSVALSWFFWLVMNNREVETKIVNELSMVLKETRGNDQEKWTEE 351
Query: 174 ITE-ESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMG 232
E + ++ YL AAL ETLRL+P +P + KY DD LPDG V +G +++ Y +G
Sbjct: 352 PLEFDEADRLVYLKAALAETLRLYPSVPQDFKYVVDDDVLPDGTFVPRGSTVTYSIYSIG 411
Query: 233 RMEFLWGEDAEQFRPERWLDENGNFQRESP---FKFTAFQAGPRICLGKEFVYRQMK-IF 288
RM+ +WGED +FRPERWL +G + E+P +KF AF AGPR CLGK+ Y QMK +
Sbjct: 412 RMKTIWGEDCLEFRPERWLTADGE-RFETPKDGYKFVAFNAGPRTCLGKDLAYNQMKSVA 470
Query: 289 SAVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHV 323
SAVLL F + V+ + +LTL + GL V
Sbjct: 471 SAVLLRYRVFPVPGHR--VEQKMSLTLFMKNGLRV 503
>At4g32170 cytochrome p450 - like protein
Length = 506
Score = 192 bits (487), Expect = 3e-49
Identities = 110/314 (35%), Positives = 170/314 (54%), Gaps = 19/314 (6%)
Query: 4 TLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVN--FLWKVQRFLNIGSE 61
T D++F ++ G + ++ E +F+ A ++ I++ + FLWK+Q ++ G E
Sbjct: 179 TFDTIFILVTGSDPRSLSIEMPED-EFAKALDDVGEGILYRHFKPRFLWKLQNWIGFGQE 237
Query: 62 AVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLEL--------NETDSK 113
L + + I +K E+ ++ Q S D+L+ F++L N +D K
Sbjct: 238 KKLTEANATFDRVCAKYISAKREEIKRSQGTSNGGSQDLLTSFIKLDTTKYKLLNPSDDK 297
Query: 114 YLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATR 173
+L+D IL+FI+AG+DTT+ LS F + L ++PHV KI QEI TD T
Sbjct: 298 FLRDNILAFILAGRDTTATALSWFFWLLSENPHVVAKIHQEIN--------INTDLSRTG 349
Query: 174 ITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGR 233
++E++ K+ YL AL E +RL+PP+ K D LP G+ V I Y +GR
Sbjct: 350 NSQENVDKLVYLHGALCEAMRLYPPVSFGRKSPIKSDVLPSGHKVDANSKIIICLYALGR 409
Query: 234 MEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLL 293
M +WGEDA QF+PERW+ ENG + E FKF +F AGPR CLGK QMKI + +L
Sbjct: 410 MRAVWGEDASQFKPERWISENGGIKHEPSFKFLSFNAGPRTCLGKHLAMTQMKIVAVEIL 469
Query: 294 GSHNFKLADQNRLV 307
+++ K+ ++V
Sbjct: 470 RNYDIKVLQGQKIV 483
>At4g39490 cytochrome P450 - like protein
Length = 479
Score = 189 bits (479), Expect = 2e-48
Identities = 109/339 (32%), Positives = 180/339 (52%), Gaps = 16/339 (4%)
Query: 4 TLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVN--FLWKVQRFLNIGSE 61
T D+ F + G + + E +F+ A ++A I F +V W++Q L +G E
Sbjct: 140 TFDTTFVLATGYDPGCLSVEMPE-VEFARALDDAEEAIFFRHVKPEIFWRLQGLLGLGDE 198
Query: 62 AVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLEL--------NETDSK 113
+ K ++ I K ++ + N D+L+ ++ L N +D +
Sbjct: 199 KKMTKARSTLDRVCSKYIAIKRDEVSRGTNNVDSHSKDLLTSYMNLDTTKYKLLNPSDER 258
Query: 114 YLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREA----TKVEDGSTTDE 169
+L+D IL+F++AG+DTT L+ F + L K+P V KI QEI A +KV+D ++ +
Sbjct: 259 FLRDTILTFMLAGRDTTGSGLTWFFWLLIKNPEVIAKIRQEINTALFQRSKVDDDASNNN 318
Query: 170 LATRITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPY 229
+ + + +KK+ YL A+ E+LRL+PP+P + K D LP G+ V I F Y
Sbjct: 319 DSDSFSPQELKKLVYLHGAICESLRLYPPVPFQHKSPTKPDVLPSGHKVDANSKILFCLY 378
Query: 230 VMGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFS 289
+GRM+ +WGEDA +F+PERW+ E+GN E +KF +F AGPR CLGKE QMK +
Sbjct: 379 SLGRMKSVWGEDALEFKPERWISESGNSVHEPSYKFLSFNAGPRTCLGKEVAMMQMKSVA 438
Query: 290 AVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
++ ++ K+ + + ++ ++ L + GL V R
Sbjct: 439 VKIIQNYEMKIV-EGQQIEPAPSVILHMKHGLKVTVTKR 476
>At3g01900 putative cytochrome P450
Length = 496
Score = 189 bits (479), Expect = 2e-48
Identities = 116/334 (34%), Positives = 177/334 (52%), Gaps = 28/334 (8%)
Query: 4 TLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIM---FWYVNFLWKVQRFLNIGS 60
T + V V LG++ T+ S ++F AF ASA ++F+WK +R + GS
Sbjct: 178 TFNIVCIVFLGIDRCTL-NPSSPVSEFDRAFQTASAVSAGRGSAPLSFVWKFKRLVGFGS 236
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDSKYLKDVIL 120
E L+K + ++ V II+ K +KP D LSR + E+D ++D+++
Sbjct: 237 EKELRKAVGEVHNCVDEIIRDK---KRKPANQ------DFLSRLIVAGESDET-VRDMVI 286
Query: 121 SFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITEESMK 180
S I+AG+DTTS + + + H + + EIR S +E+ ES+K
Sbjct: 287 SIIMAGRDTTSAVATRLFWLITGHEETEHDLVSEIR--------SVKEEITGGFDYESLK 338
Query: 181 KMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEFLWGE 240
K+ L A L E +RL+PP+P +SK+ +DD LPDG VR GD +++ PY MGRME LWGE
Sbjct: 339 KLSLLKACLCEVMRLYPPVPWDSKHALTDDRLPDGTLVRAGDRVTYFPYGMGRMEELWGE 398
Query: 241 DAEQFRPERWLDENGN-----FQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGS 295
D ++F+P RW + ++ +PFKF FQAGPR+CLG+E Y QMK A +L
Sbjct: 399 DWDEFKPNRWAESYDKTCCRVLKKVNPFKFPVFQAGPRVCLGEEMAYVQMKYIVASILDR 458
Query: 296 HNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
+ ++ + LT + GG+ V R+
Sbjct: 459 FEIEPIPTDK-PDFVPMLTAHMAGGMQVRVHRRD 491
>At3g48520 cytochrome P450-like protein
Length = 506
Score = 188 bits (477), Expect = 4e-48
Identities = 116/332 (34%), Positives = 180/332 (53%), Gaps = 24/332 (7%)
Query: 6 DSVFKVILGVELDTMCGTSKEGTQFSNAFN---EASATIMFWYVNFLWKVQRFLNIGSEA 62
D V KV LG + D + ++ AF+ E SA + +WK +R LN+GSE
Sbjct: 185 DVVCKVSLGWDPDCL-DLTRPVNPLVEAFDTAAEISARRATEPIYAVWKTKRVLNVGSER 243
Query: 63 VLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDSKYLKDVILSF 122
L++ +R ++ V I+++K + + E K D+LSRFL + + ++D+++SF
Sbjct: 244 KLREAIRTVHVLVSEIVRAKKKSLEIG--TGAEAKQDLLSRFLAAGH-NGEAVRDMVISF 300
Query: 123 IIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITEESMKKM 182
I+AG+DTTS ++ + L ++ V+ KI +E+ + G E +K+M
Sbjct: 301 IMAGRDTTSAAMTWLFWLLTENDDVERKILEEVDPLVSLGLGF-----------EDLKEM 349
Query: 183 QYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEFLWGEDA 242
Y A L E +RL+PP+ +SK+ +DD LPDG V++GD +++ PY MGRME LWG D+
Sbjct: 350 AYTKACLCEAMRLYPPVSWDSKHAANDDVLPDGTRVKRGDKVTYFPYGMGRMETLWGTDS 409
Query: 243 EQFRPERWLDENGNFQRE-----SPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSHN 297
E+F P RW D R SP+KF FQAGPR+C+GKE + QMK +L
Sbjct: 410 EEFNPNRWFDSEPGSTRPVLKPISPYKFPVFQAGPRVCVGKEMAFMQMKYVVGSVLSRFE 469
Query: 298 FKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
+++R V + LT + GGL V R+
Sbjct: 470 IVPVNKDRPV-FVPLLTAHMAGGLKVKIKRRS 500
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.137 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,074,874
Number of Sequences: 26719
Number of extensions: 293454
Number of successful extensions: 1356
Number of sequences better than 10.0: 252
Number of HSP's better than 10.0 without gapping: 228
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 736
Number of HSP's gapped (non-prelim): 263
length of query: 329
length of database: 11,318,596
effective HSP length: 100
effective length of query: 229
effective length of database: 8,646,696
effective search space: 1980093384
effective search space used: 1980093384
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC149581.9


