

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.7 + phase: 0
(392 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
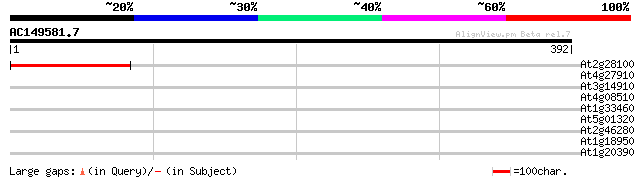
Score E
Sequences producing significant alignments: (bits) Value
At2g28100 unknown protein 125 5e-29
At4g27910 trithorax 4 (TX4) 32 0.46
At3g14910 hypothetical protein 32 0.60
At4g08510 hypothetical protein 31 1.0
At1g33460 mutator transposase MUDRA, putative 31 1.3
At5g01320 pyruvate decarboxylase-like protein 29 5.1
At2g46280 eukaryotic translation initiation factor 3 delta subunit 28 6.7
At1g18950 hypothetical protein 28 6.7
At1g20390 hypothetical protein 28 8.7
>At2g28100 unknown protein
Length = 181
Score = 125 bits (313), Expect = 5e-29
Identities = 55/84 (65%), Positives = 68/84 (80%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+ILTAKHHDGFCLWPS+YT +SV SS+W+NG GDVV E +AA + GI +G+YLSPWDRH
Sbjct: 98 VILTAKHHDGFCLWPSEYTDYSVKSSQWRNGAGDVVAELASAAKEAGIGLGLYLSPWDRH 157
Query: 61 DSRYGHDLLYNEYYLAQLQELLKK 84
+ YG L YNE+YL+Q+ ELL K
Sbjct: 158 EQCYGKTLEYNEFYLSQMTELLTK 181
>At4g27910 trithorax 4 (TX4)
Length = 954
Score = 32.3 bits (72), Expect = 0.46
Identities = 17/62 (27%), Positives = 27/62 (43%)
Query: 223 PNTTGLISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHL 282
P T L S+ + K + FH+ + +V + K+G FGPEN D +
Sbjct: 152 PKATNLKSKELDRKSKYSALCKEERFHEQHNDEARARVDEKLPNKKGTFGPENFYSGDLV 211
Query: 283 WS 284
W+
Sbjct: 212 WA 213
>At3g14910 hypothetical protein
Length = 455
Score = 32.0 bits (71), Expect = 0.60
Identities = 14/44 (31%), Positives = 28/44 (62%), Gaps = 3/44 (6%)
Query: 330 IYVDGKSIIQGTTIGYKRLHRLDGDVVHARVV---RIRFIKARG 370
++ D + ++ GT+ GY ++ + GD++H ++V RI I+ RG
Sbjct: 86 VFDDVRVVVAGTSCGYLLVYSVTGDLIHKQIVHQSRILKIRVRG 129
>At4g08510 hypothetical protein
Length = 551
Score = 31.2 bits (69), Expect = 1.0
Identities = 16/58 (27%), Positives = 26/58 (44%)
Query: 289 REDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERYEIYVDGKSIIQGTTIGYK 346
R +K++ +++ W ND S+ F I +R G + QG T+GYK
Sbjct: 89 RSREKDRMSYMDPWDNDSSMPFGTFLIGRGEEPLRRSHSMTTRKQGNHLAQGFTVGYK 146
>At1g33460 mutator transposase MUDRA, putative
Length = 826
Score = 30.8 bits (68), Expect = 1.3
Identities = 14/42 (33%), Positives = 22/42 (52%)
Query: 54 LSPWDRHDSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDG 95
+ W + S Y ++L E+ LA E+ K + E+WFDG
Sbjct: 576 IGKWKKKISEYVSEILKEEWELAVKCEVTKGTHEKFEVWFDG 617
>At5g01320 pyruvate decarboxylase-like protein
Length = 603
Score = 28.9 bits (63), Expect = 5.1
Identities = 14/37 (37%), Positives = 20/37 (53%)
Query: 219 LNVPPNTTGLISENDAHRLKEFRSAIDTIFHKNIAEN 255
L V P+T GL+ EN H + + A+ T F I E+
Sbjct: 280 LAVMPSTKGLVPENHPHFIGTYWGAVSTPFCSEIVES 316
>At2g46280 eukaryotic translation initiation factor 3 delta subunit
Length = 328
Score = 28.5 bits (62), Expect = 6.7
Identities = 12/28 (42%), Positives = 15/28 (52%)
Query: 333 DGKSIIQGTTIGYKRLHRLDGDVVHARV 360
DGKS G GY RLH D D + ++
Sbjct: 301 DGKSFSSGGEDGYVRLHHFDSDYFNIKI 328
>At1g18950 hypothetical protein
Length = 766
Score = 28.5 bits (62), Expect = 6.7
Identities = 19/73 (26%), Positives = 35/73 (47%), Gaps = 10/73 (13%)
Query: 229 ISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLW----- 283
+S+N+ EF+S D I+ + + + Y+K Q+ G G E D ++ W
Sbjct: 524 VSDNEV----EFQSD-DDIYGEAVYDEEYLKKRKQKKLSSGSEGDEEKGDEEYKWDEDNA 578
Query: 284 SYWTPREDDKEKD 296
Y E+++E+D
Sbjct: 579 EYEEEEEEEEEED 591
>At1g20390 hypothetical protein
Length = 1791
Score = 28.1 bits (61), Expect = 8.7
Identities = 18/50 (36%), Positives = 27/50 (54%), Gaps = 2/50 (4%)
Query: 322 GQRIERYEIYVDGKSIIQGTTIGYKRLHRLDGDVVHARVVRIRFIKARGV 371
G R E + +YVDG S +G+ IG RL +V+ + R+RF+ V
Sbjct: 1204 GNRGEEWSLYVDGSSSARGSGIGI-RLVSPTAEVLE-QSFRLRFVATNNV 1251
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,730,511
Number of Sequences: 26719
Number of extensions: 443969
Number of successful extensions: 1012
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 1004
Number of HSP's gapped (non-prelim): 11
length of query: 392
length of database: 11,318,596
effective HSP length: 101
effective length of query: 291
effective length of database: 8,619,977
effective search space: 2508413307
effective search space used: 2508413307
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149581.7


