

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.3 + phase: 0
(194 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
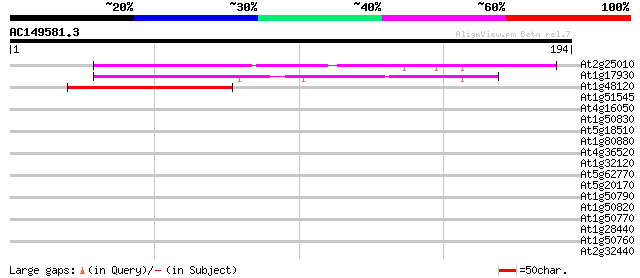
Score E
Sequences producing significant alignments: (bits) Value
At2g25010 unknown protein 54 4e-08
At1g17930 unknown protein 50 7e-07
At1g48120 serine/threonine phosphatase PP7, putative 47 5e-06
At1g51545 unknown protein 33 0.12
At4g16050 hypothetical protein 32 0.28
At1g50830 hypothetical protein 32 0.28
At5g18510 putative protein 31 0.36
At1g80880 hypothetical protein 31 0.47
At4g36520 trichohyalin like protein 30 0.61
At1g32120 hypothetical protein 30 0.80
At5g62770 putative protein 29 1.4
At5g20170 putative protein 29 1.4
At1g50790 hypothetical protein 29 1.8
At1g50820 hypothetical protein 28 2.3
At1g50770 hypothetical protein 28 3.0
At1g28440 unknown protein 27 5.2
At1g50760 hypothetical protein 27 6.8
At2g32440 putative cytochrome P450 27 8.9
>At2g25010 unknown protein
Length = 509
Score = 54.3 bits (129), Expect = 4e-08
Identities = 49/183 (26%), Positives = 84/183 (45%), Gaps = 27/183 (14%)
Query: 30 GYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHDEAMNQD 89
G ++ +++I A+ ERW ET++FHL +GEMTI LD++ +L + I G + + + +
Sbjct: 67 GPMSLNNSLISALVERWRRETNTFHLPLGEMTITLDEVALVLGLEIDGDPIVGSK-VGDE 125
Query: 90 CAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEAR-RLEEPQTREKL-- 146
A+D+ RLLG EV + + LKR + +A + + TR L
Sbjct: 126 VAMDMCGRLLGKLPSAANKEV---NCSRVKLNWLKRTFSECPEDASFDVVKCHTRAYLLY 182
Query: 147 --------QERGRKRAI------------GKWSWGGMALAYLYDYLDDSVILNNRTMAGS 186
G K ++ G+++WG ALA LY L ++ + + + G
Sbjct: 183 LIGSTIFATTDGDKVSVKYLPLFEDFDQAGRYAWGAAALACLYRALGNASLKSQSNICGC 242
Query: 187 TIL 189
L
Sbjct: 243 LTL 245
>At1g17930 unknown protein
Length = 478
Score = 50.1 bits (118), Expect = 7e-07
Identities = 44/162 (27%), Positives = 74/162 (45%), Gaps = 28/162 (17%)
Query: 30 GYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGR----ILDHDEA 85
G ++ +++I A+ ERW ET++FH GEMTI LD++ +L + + G+ + + DE
Sbjct: 58 GSISLNNSLISALVERWRRETNTFHFPCGEMTITLDEVSLILGLAVDGKPVVGVKEKDED 117
Query: 86 MNQDCAIDLMIRLLG------MSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEE 139
+Q C +RLLG +S V + ES + E+H T A +
Sbjct: 118 PSQVC-----LRLLGKLPKGELSGNRVTAKWLKESFAECPKGATMKEIEYH-TRAYLIYI 171
Query: 140 PQTREKLQERGRKRAI------------GKWSWGGMALAYLY 169
+ K ++ G+++WG ALA+LY
Sbjct: 172 VGSTIFATTDPSKISVDYLILFEDFEKAGEYAWGAAALAFLY 213
>At1g48120 serine/threonine phosphatase PP7, putative
Length = 1338
Score = 47.4 bits (111), Expect = 5e-06
Identities = 21/57 (36%), Positives = 35/57 (60%)
Query: 21 FDLHDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHG 77
F L+ + + + +A+I A+ ERW ET +FHL GE+T+ L D++ LL + + G
Sbjct: 67 FGLYGVYKVAFIQLDYALITALVERWRPETHTFHLPAGEITVTLQDVNILLGLRVDG 123
>At1g51545 unknown protein
Length = 629
Score = 32.7 bits (73), Expect = 0.12
Identities = 16/41 (39%), Positives = 24/41 (58%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHG 77
+++ A+ E+W ET SF GE TI L+D+ LL + G
Sbjct: 30 SLLLALVEKWCPETKSFLFPWGEATITLEDVLVLLGFSVQG 70
>At4g16050 hypothetical protein
Length = 900
Score = 31.6 bits (70), Expect = 0.28
Identities = 15/41 (36%), Positives = 25/41 (60%)
Query: 37 AMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHG 77
++I ++ + W ET++F GE TI L+D++ LL I G
Sbjct: 362 SLILSLAQNWCPETNTFVFPWGEATITLEDVNVLLGFSISG 402
>At1g50830 hypothetical protein
Length = 768
Score = 31.6 bits (70), Expect = 0.28
Identities = 17/45 (37%), Positives = 25/45 (54%)
Query: 33 TVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHG 77
T ++I ++ E+W ET SF GE TI L+D+ LL + G
Sbjct: 114 TKNPSLILSVSEKWCPETKSFVFPWGEATITLEDVMVLLGFSVLG 158
>At5g18510 putative protein
Length = 702
Score = 31.2 bits (69), Expect = 0.36
Identities = 16/39 (41%), Positives = 23/39 (58%)
Query: 39 ICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHG 77
I ++ E+W +ET SF GE TI L+D+ LL + G
Sbjct: 98 ILSIAEKWCSETKSFIFPWGEATITLEDVMVLLGFSVLG 136
>At1g80880 hypothetical protein
Length = 540
Score = 30.8 bits (68), Expect = 0.47
Identities = 13/51 (25%), Positives = 24/51 (46%), Gaps = 1/51 (1%)
Query: 16 EPVEGFDLHDLI-YTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLD 65
+ + FD+ D +T Y ++CA+C H E + +L + P+D
Sbjct: 207 QAIRTFDIMDKFKHTPYDEAFQGLLCALCRHGHIEKAEEFMLASKKLFPVD 257
>At4g36520 trichohyalin like protein
Length = 1432
Score = 30.4 bits (67), Expect = 0.61
Identities = 17/63 (26%), Positives = 34/63 (52%), Gaps = 3/63 (4%)
Query: 84 EAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVYEHHLTEARRLEEPQTR 143
EA+ + +++ + L G + ++E+RS+S ++ P + E + EAR EE R
Sbjct: 594 EALGNESTLEVSVELNGNGK---KMEMRSQSETKLNEPLKRMEEETRIKEARLREENDRR 650
Query: 144 EKL 146
E++
Sbjct: 651 ERV 653
Score = 28.5 bits (62), Expect = 2.3
Identities = 21/84 (25%), Positives = 38/84 (45%), Gaps = 12/84 (14%)
Query: 68 HNLLHIPIHGRILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTLKRVY 127
+ H G I + +NQD ++ + S V V+ E +E LKR
Sbjct: 1095 YKFTHQQERGNIYETQAGLNQDAKVERPLP----SRVSVQREKEAER--------LKRER 1142
Query: 128 EHHLTEARRLEEPQTREKLQERGR 151
+ + + R++EE + RE+ +E+ R
Sbjct: 1143 DLEMEQLRKVEEEREREREREKDR 1166
>At1g32120 hypothetical protein
Length = 1206
Score = 30.0 bits (66), Expect = 0.80
Identities = 13/30 (43%), Positives = 20/30 (66%)
Query: 38 MICAMCERWHTETSSFHLLVGEMTIPLDDM 67
+I A+ E+W ET++F GE T+ L+DM
Sbjct: 121 LIVALVEKWCIETNTFVFPWGEATLTLEDM 150
>At5g62770 putative protein
Length = 268
Score = 29.3 bits (64), Expect = 1.4
Identities = 21/91 (23%), Positives = 40/91 (43%), Gaps = 14/91 (15%)
Query: 65 DDMHNLLHIPIHGRILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGH--ISYPT 122
DD+ L P H +I H ++ + ++ VRS S G+ + +P
Sbjct: 158 DDLQRLSSSPSHSKIKSHSAGFSKRWKLRNLLY------------VRSSSEGNDKLVFPA 205
Query: 123 LKRVYEHHLTEARRLEEPQTREKLQERGRKR 153
+ + +++ R EEP ++ +E GR+R
Sbjct: 206 PVKKNDETVSDQREEEEPPSKVDGEEEGRER 236
>At5g20170 putative protein
Length = 692
Score = 29.3 bits (64), Expect = 1.4
Identities = 16/59 (27%), Positives = 28/59 (47%), Gaps = 9/59 (15%)
Query: 138 EEPQTREKLQERGRKRAIGKWSWGGMA---------LAYLYDYLDDSVILNNRTMAGST 187
EE + ++K Q++ K ++ +W W GM L + D +D + T+AG T
Sbjct: 53 EEDELKKKKQKKSSKDSVEQWKWKGMVENLQLAHQELTVIIDLIDTVQANDAVTVAGMT 111
>At1g50790 hypothetical protein
Length = 812
Score = 28.9 bits (63), Expect = 1.8
Identities = 13/40 (32%), Positives = 23/40 (57%)
Query: 38 MICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHG 77
++ + E+W +T++F GE TI L+D+ LL + G
Sbjct: 101 LVMGIAEKWCPDTNTFVFSWGEATITLEDVMVLLGFSVLG 140
>At1g50820 hypothetical protein
Length = 528
Score = 28.5 bits (62), Expect = 2.3
Identities = 13/40 (32%), Positives = 22/40 (54%)
Query: 38 MICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHG 77
++ + E+W +T +F GE TI L+D+ LL + G
Sbjct: 97 LVLGLAEKWCPDTKTFIFPWGEATITLEDVMVLLGFSVLG 136
>At1g50770 hypothetical protein
Length = 632
Score = 28.1 bits (61), Expect = 3.0
Identities = 13/40 (32%), Positives = 22/40 (54%)
Query: 38 MICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHG 77
++ + E+W +T +F GE TI L+D+ LL + G
Sbjct: 100 LVLGIAEKWCPDTKTFVFPWGETTITLEDVMLLLGFSVLG 139
>At1g28440 unknown protein
Length = 996
Score = 27.3 bits (59), Expect = 5.2
Identities = 17/60 (28%), Positives = 26/60 (43%)
Query: 24 HDLIYTGYSTVTHAMICAMCERWHTETSSFHLLVGEMTIPLDDMHNLLHIPIHGRILDHD 83
H +Y T + A C+ T S +LL GE+ L D+ L+H+ + G D
Sbjct: 88 HLSLYNNSINSTLPLNIAACKSLQTLDLSQNLLTGELPQTLADIPTLVHLDLTGNNFSGD 147
>At1g50760 hypothetical protein
Length = 649
Score = 26.9 bits (58), Expect = 6.8
Identities = 10/30 (33%), Positives = 18/30 (59%)
Query: 38 MICAMCERWHTETSSFHLLVGEMTIPLDDM 67
++ + E+W +T +F GE TI L+D+
Sbjct: 90 LVLGVAEKWSPDTKTFVFSWGEATITLEDV 119
>At2g32440 putative cytochrome P450
Length = 489
Score = 26.6 bits (57), Expect = 8.9
Identities = 26/113 (23%), Positives = 50/113 (44%), Gaps = 20/113 (17%)
Query: 65 DDMHNLLHIPI-HGRILDHDEAMNQDCAIDLMIRLLGMSEVDVRVEVRSESAGHISYPTL 123
D + NL+ + +GR+LD +E IDL++ L ES+GH++
Sbjct: 268 DMLDNLIDVKDENGRVLDDEEI------IDLLLMYLNAGH---------ESSGHLTMWAT 312
Query: 124 KRVYEHHLTEARRLEEPQTREKLQERGRKRAIGKWSWGGMALAYLYDYLDDSV 176
+ EH + + EE + K + G+K + + + YL +D+++
Sbjct: 313 ILMQEHPMILQKAKEEQERIVKKRAPGQKLTLKE----TREMVYLSQVIDETL 361
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.139 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,386,744
Number of Sequences: 26719
Number of extensions: 175391
Number of successful extensions: 437
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 417
Number of HSP's gapped (non-prelim): 23
length of query: 194
length of database: 11,318,596
effective HSP length: 94
effective length of query: 100
effective length of database: 8,807,010
effective search space: 880701000
effective search space used: 880701000
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC149581.3


