

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.6 - phase: 0
(597 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
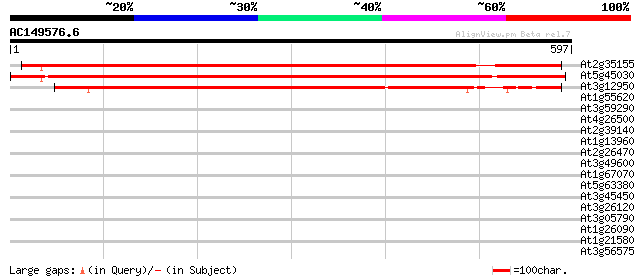
Score E
Sequences producing significant alignments: (bits) Value
At2g35155 unknown protein 812 0.0
At5g45030 putative protein 798 0.0
At3g12950 unknown protein 652 0.0
At1g55620 CLC-f chloride channel protein 35 0.091
At3g59290 epsin-like protein 35 0.15
At4g26500 unknown protein 33 0.45
At2g39140 unknown protein 32 1.0
At1g13960 WRKY DNA-binding protein 4 (WRKY4) 31 1.7
At2g26470 unknown protein 31 2.2
At3g49600 putative protein 30 3.8
At1g67070 phosphomannose isomerase (din9) 30 5.0
At5g63380 4-coumarate-CoA ligase-like protein 29 6.5
At3g45450 clpC-like protein 29 6.5
At3g26120 RNA-binding protein, putative 29 6.5
At3g05790 putative mitochondrial LON ATP-dependent protease 29 6.5
At1g26090 unknown protein 29 6.5
At1g21580 hypothetical protein 29 6.5
At3g56575 unknown protein 29 8.5
>At2g35155 unknown protein
Length = 579
Score = 812 bits (2098), Expect = 0.0
Identities = 413/584 (70%), Positives = 467/584 (79%), Gaps = 29/584 (4%)
Query: 13 SGSTQSEESALDLERNYYGH-----PSSSPLHMQTFAVGVQHSEGNAAYFSWPTLNRWND 67
+ S++SE+SALDLERN++ + SSSP +Q F + +QH+E NA YFSWPTL+R ND
Sbjct: 14 AASSESEDSALDLERNHHCNHLSLPSSSSPSPLQPFTLNIQHAESNAPYFSWPTLSRLND 73
Query: 68 AAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAIGFRI 127
EDRANYFGNLQKGVLPET+GRLPSGQQATTLLELMTIRAFHSKILRRFSLGTA+GFRI
Sbjct: 74 TVEDRANYFGNLQKGVLPETVGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAVGFRI 133
Query: 128 RGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCDVDVVEFSYYGAPAPTPK 187
GVLT++PAILVFVA KVHRQWLN +QCLP+ALEGPGGVWCDVDVVEF YYGAPA TPK
Sbjct: 134 SRGVLTNVPAILVFVARKVHRQWLNPMQCLPSALEGPGGVWCDVDVVEFQYYGAPAATPK 193
Query: 188 EQLYTELADGLRGSDSCVGSGSQVASQETYGTLGAIVRSRTGNREVGFLTNRHVAVDLDY 247
EQ+Y EL DGLRGSD C+GSGSQVASQETYGTLGAIV+SRTGN +VGFLTNRHVAVDLDY
Sbjct: 194 EQVYNELVDGLRGSDPCIGSGSQVASQETYGTLGAIVKSRTGNHQVGFLTNRHVAVDLDY 253
Query: 248 PNQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFVRADGAFIPFAEDF 307
P+QKMFHPLPPSLGPGVYLGAVERATSFITDD WYGIFAGTNPETFVRADGAFIPFAEDF
Sbjct: 254 PSQKMFHPLPPSLGPGVYLGAVERATSFITDDQWYGIFAGTNPETFVRADGAFIPFAEDF 313
Query: 308 NMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSSGLTTGTIMAYALEYNDEK 367
N +NV T I+G+G+IG+VH IDLQSPI+SLIG+QV+KVGRSSG TTGTIMAYALEYNDEK
Sbjct: 314 NTSNVTTLIKGIGEIGDVHVIDLQSPIDSLIGKQVVKVGRSSGYTTGTIMAYALEYNDEK 373
Query: 368 GICFLTDFLVVGENQQTFDLEGDSGSLILLTGQNREKPRPVGIIWGGTANRGRLKLRVGQ 427
GICFLTDFLV+GENQQTFDLEGDSGSLILLTG N +KPRPVGIIWGGTANRGRLKL GQ
Sbjct: 374 GICFLTDFLVIGENQQTFDLEGDSGSLILLTGPNGQKPRPVGIIWGGTANRGRLKLIAGQ 433
Query: 428 PPENWTSGVDLGRLLDLLELDLVTTNETLQ--DSGQEQMNGSTAGIGSTVGESSP--TVP 483
PENWTSGVDLGRLLDLLELDL+T+N L+ + +E+ N S + STV +SSP VP
Sbjct: 434 EPENWTSGVDLGRLLDLLELDLITSNHELEAAAAAREERNTSVTALDSTVSQSSPPDPVP 493
Query: 484 IKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETVPNVEHQFIRTSFAG 543
+K +ESFEPF P EFHI I+ VE +
Sbjct: 494 SGDKQDESFEPFI--------------------PPEFHIEEAIKPTLEVEEHIFIAPISV 533
Query: 544 KSPVHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRK 587
+E + +L L+N +E+ +SLHLGEP+ K+ K
Sbjct: 534 NESTSAIKGQEIPKLDNLMALKNSSEEEVNISLHLGEPKLKKPK 577
>At5g45030 putative protein
Length = 607
Score = 798 bits (2060), Expect = 0.0
Identities = 424/605 (70%), Positives = 486/605 (80%), Gaps = 21/605 (3%)
Query: 1 MNRNRLGLSAHHSGSTQSEESA-LDLERNYYGH---PSSSPLHMQTFAVGVQHSEGNAA- 55
M RL L HHS S+QS ESA LDL++N Y H SSSPL Q F G QH E +AA
Sbjct: 1 MEGKRLDLRFHHSTSSQSVESAALDLDKNVYNHIKLASSSPL--QPFPSGAQHPETSAAA 58
Query: 56 -YFSWPTLNRWNDAAEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKIL 114
YFSWPT +R ND+AEDRANYF NLQKGVLPE+ LP+G++ATTLLELM IRAFHSK L
Sbjct: 59 AYFSWPTSSRLNDSAEDRANYFANLQKGVLPESFDGLPTGKKATTLLELMMIRAFHSKNL 118
Query: 115 RRFSLGTAIGFRIRGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAALEGPGGVWCDVDVV 174
RRFSLGTAIGFRIR GVLT+I AILVFVA KVH+QWLN +QCLP ALEGPGGVWCDVDVV
Sbjct: 119 RRFSLGTAIGFRIRRGVLTNIAAILVFVARKVHKQWLNPLQCLPTALEGPGGVWCDVDVV 178
Query: 175 EFSYYGAPAPTPKEQLYTELADGLRGSDSCVGSGSQVASQETYGTLGAIVRSRTGNREVG 234
EF YYGAPA TPKEQ+YTEL D LRGS S +GSGSQVASQETYGTLGAIV+S+TG R+VG
Sbjct: 179 EFQYYGAPAQTPKEQVYTELVDDLRGSGSSIGSGSQVASQETYGTLGAIVKSKTGIRQVG 238
Query: 235 FLTNRHVAVDLDYPNQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFV 294
FLTNRHVAVDLDYP+QKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFV
Sbjct: 239 FLTNRHVAVDLDYPSQKMFHPLPPSLGPGVYLGAVERATSFITDDLWYGIFAGTNPETFV 298
Query: 295 RADGAFIPFAEDFNMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVGRSSGLTTG 354
RADGAFIPFAEDFN NNV T+++G+G+IG++H DLQSP+NSLIGR+V+KVGRSSGLTTG
Sbjct: 299 RADGAFIPFAEDFNTNNVTTTVKGIGEIGDIHATDLQSPVNSLIGRKVVKVGRSSGLTTG 358
Query: 355 TIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEGDSGSLILLTG--QNREKPRPVGIIW 412
TIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEGDSGSLILL + EKPRPVGIIW
Sbjct: 359 TIMAYALEYNDEKGICFLTDFLVVGENQQTFDLEGDSGSLILLAAGDEKNEKPRPVGIIW 418
Query: 413 GGTANRGRLKLRVGQPPENWTSGVDLGRLLDLLELDLVTTNETLQDSGQEQMNG-STAGI 471
GGTANRGRLKL+VG+ PENWTSGVDLGR+L+LLELDL+T+NE LQ + EQ NG A +
Sbjct: 419 GGTANRGRLKLKVGEQPENWTSGVDLGRVLNLLELDLITSNEGLQAAVLEQRNGIMCAAV 478
Query: 472 GSTVGESSPTV--PIKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEFHIRNEIETV 529
STV ESSP V + K E+FEP LN++ V +E+ ++ + P EF I + +E+V
Sbjct: 479 DSTVVESSPGVCNISRCKTGENFEPINLNVQQVLIEDDNSNIHP-----EFQIEDVLESV 533
Query: 530 PNV-EHQFIRTSFAGKSPVHQS-FLKEDMQFKSLSELRNEPDEDNF-VSLHLGEPEAKRR 586
+ EHQFI +S S +HQ E+++ K+LS L+ D SL LGE + K+R
Sbjct: 534 AVIEEHQFIPSSSNNGSALHQKPNGPENLESKNLSSLKTSSSGDEIGFSLQLGESDTKKR 593
Query: 587 KHSNS 591
K ++S
Sbjct: 594 KRTDS 598
>At3g12950 unknown protein
Length = 558
Score = 652 bits (1683), Expect = 0.0
Identities = 353/564 (62%), Positives = 429/564 (75%), Gaps = 50/564 (8%)
Query: 48 QHSEGNAA-YFSWPTLNRWNDAAEDRANYFGNLQKG------VLPETLGRLPSGQQATTL 100
QH E AA YFSWPT +R ++AAE+RANYF NLQK V PE + P GQ+ATTL
Sbjct: 9 QHCEFTAASYFSWPTSSRLSNAAEERANYFSNLQKEEDDDDEVSPEPVSTEPKGQRATTL 68
Query: 101 LELMTIRAFHSKILRRFSLGTAIGFRIRGGVLTDIPAILVFVAHKVHRQWLNHVQCLPAA 160
LELMTIRAFHSK+LR +SLGTAIGFRIR GVLTDIPAI+VFV+ KVH+QWL+ +QCLP A
Sbjct: 69 LELMTIRAFHSKMLRCYSLGTAIGFRIRRGVLTDIPAIIVFVSRKVHKQWLSPLQCLPTA 128
Query: 161 LEGPGGVWCDVDVVEFSYYGAP--APTPKEQLYTELADGLRGSDSCVGSGSQVASQETYG 218
LEG GG+WCDVDVVEFSY+G P PTPK+ T++ D L+GSD +GSGSQVASQET G
Sbjct: 129 LEGAGGIWCDVDVVEFSYFGEPDHQPTPKQTFTTDIVDHLQGSDPFIGSGSQVASQETCG 188
Query: 219 TLGAIVRSRTGNREVGFLTNRHVAVDLDYPNQKMFHPLPPSLGPGVYLGAVERATSFITD 278
TLGAIVRS+TG R+VGF+TNRHVAV+LDYP+QKMFHPLPP+LGPGVYLGAVERATSFITD
Sbjct: 189 TLGAIVRSQTGGRQVGFVTNRHVAVNLDYPSQKMFHPLPPALGPGVYLGAVERATSFITD 248
Query: 279 DLWYGIFAGTNPETFVRADGAFIPFAEDFNMNNVITSIR-GVGDIGEVHRIDLQSPINSL 337
DLW+GIFAGTNPETFVRADGAFIPFA+D++++ V TS++ GVG+IGEV I+LQSP+ SL
Sbjct: 249 DLWFGIFAGTNPETFVRADGAFIPFADDYDLSRVTTSVKGGVGEIGEVKAIELQSPVGSL 308
Query: 338 IGRQVIKVGRSSGLTTGTIMAYALEYNDEKGICFLTDFLVVGENQQT-FDLEGDSGSLIL 396
+G+QV+KVGRSSGLTTGT++AYALEYNDE+G+CFLTDFLVVGEN ++ FDLEGDSGSLI+
Sbjct: 309 VGKQVVKVGRSSGLTTGTVLAYALEYNDERGVCFLTDFLVVGENHRSPFDLEGDSGSLIV 368
Query: 397 LTGQNREKPRPVGIIWGGTANRGRLKLRVGQPPENWTSGVDLGRLLDLLELDLVTTNETL 456
+ G+ EK RP+GIIWGGT +RGRLKL+VG+ PE+WT+GVDLGRLL L+LDL+TT+E L
Sbjct: 369 MKGE--EKARPIGIIWGGTGSRGRLKLKVGECPESWTTGVDLGRLLTHLQLDLITTDEGL 426
Query: 457 QDSGQEQMNGSTAGIGSTVGESSPT-VPIK-------EKLEESFEPFCLNMEHVPVEEPS 508
+ + QEQ ST G+ S V +SSP V +K EKLE S P L ++H+ +EE
Sbjct: 427 KAAVQEQRAASTTGMSSMVADSSPPYVNLKKEKRSPEEKLEASLGP--LQVQHIDLEE-- 482
Query: 509 TIVKPSLRPCEFHIRNEIET---VPNVEHQFIRTSFAGK--SPVHQSFLKEDMQFKSLSE 563
IET P+VEHQF+ T F+G+ + +ED+
Sbjct: 483 ----------------RIETKGGAPSVEHQFMPT-FSGQCSASAWPETAREDL---VAGF 522
Query: 564 LRNEPDEDNFVSLHLGEPEAKRRK 587
D D V L LG+ AKRR+
Sbjct: 523 TNGSCDGDLCVGLRLGDDGAKRRR 546
>At1g55620 CLC-f chloride channel protein
Length = 781
Score = 35.4 bits (80), Expect = 0.091
Identities = 28/101 (27%), Positives = 42/101 (40%), Gaps = 20/101 (19%)
Query: 412 WGGTANRGRLKLRVGQPPENW--------TSGVDLGRLLDLLE-LDLVTTNETLQDSGQE 462
W GT N G LR+ + + W T GV +G + LLE LD + + + Q G +
Sbjct: 161 WAGTPNEGAAWLRLQRLADTWHRILLIPVTGGVIVGMMHGLLEILDQIRQSNSSQRQGLD 220
Query: 463 QMNG-----------STAGIGSTVGESSPTVPIKEKLEESF 492
+ G T G G ++G P+V I + F
Sbjct: 221 FLAGIYPVIKAIQAAVTLGTGCSLGPEGPSVDIGKSCANGF 261
>At3g59290 epsin-like protein
Length = 1024
Score = 34.7 bits (78), Expect = 0.15
Identities = 33/112 (29%), Positives = 44/112 (38%), Gaps = 23/112 (20%)
Query: 393 SLILLTG------QNREKPRPVGIIWGGTANRGRLKLRVGQPPENWTS--GVD---LGRL 441
SL LTG QN++K P IW T +RG + + P N + GVD + R
Sbjct: 805 SLTPLTGAIEIVPQNQKKFEPKSTIWADTLSRGLVNFNISGPKTNPLADIGVDFEAINRK 864
Query: 442 LDLLELDLVTTNETLQDSGQEQMNGSTAGIG------------STVGESSPT 481
LE +T + + GS G+G S VG S PT
Sbjct: 865 EKRLEKPTITQQQVTSTINMGKAMGSGTGLGRAGAGAMRPPTNSMVGSSMPT 916
>At4g26500 unknown protein
Length = 371
Score = 33.1 bits (74), Expect = 0.45
Identities = 19/57 (33%), Positives = 34/57 (59%), Gaps = 1/57 (1%)
Query: 446 ELDLVTTNETLQDSGQEQMNGSTAGIGSTVGESSPTVPIKEKLEESFEPFCLNMEHV 502
E+DL + + ++D G E+++ S +G + V S + I+EKLE+ +P L +E V
Sbjct: 248 EVDLESKPDLVEDLGTEKIDDSESG-SNVVALGSRGMRIREKLEKELDPVELEVEDV 303
>At2g39140 unknown protein
Length = 303
Score = 32.0 bits (71), Expect = 1.0
Identities = 26/93 (27%), Positives = 43/93 (45%), Gaps = 12/93 (12%)
Query: 405 PRPVGIIWGGTANRGRLKLRVGQPPENWTSGVDLGRLLDLLELDLVTTNETLQDSGQEQM 464
P+ + I W ++ G LKL+ QPP RL+ L +D T N +D +++
Sbjct: 68 PQQLFIPWIVRSDDGTLKLQ-SQPP---------ARLIHNLAIDATTQNPKKKDKSKKKQ 117
Query: 465 NGSTAGIGSTVGESSPT--VPIKEKLEESFEPF 495
+T+ +T SSP +K KL ++ F
Sbjct: 118 PQATSSSSATTTASSPASHSEVKPKLSKAARRF 150
>At1g13960 WRKY DNA-binding protein 4 (WRKY4)
Length = 514
Score = 31.2 bits (69), Expect = 1.7
Identities = 17/76 (22%), Positives = 31/76 (40%)
Query: 191 YTELADGLRGSDSCVGSGSQVASQETYGTLGAIVRSRTGNREVGFLTNRHVAVDLDYPNQ 250
+++L G S + + + A+ Y LG S +G+ + F NR + +
Sbjct: 61 FSQLLAGAMSSPATAAAAAAAATASDYQRLGEGTNSSSGDVDPRFKQNRPTGLMISQSQS 120
Query: 251 KMFHPLPPSLGPGVYL 266
+PP L P + L
Sbjct: 121 PSMFTVPPGLSPAMLL 136
>At2g26470 unknown protein
Length = 487
Score = 30.8 bits (68), Expect = 2.2
Identities = 30/154 (19%), Positives = 60/154 (38%), Gaps = 24/154 (15%)
Query: 456 LQDSGQEQMNGSTAGIGSTVGESSPTVPIKEKLEESFEPFCLNMEHVPVEEPSTIVKPSL 515
+Q+ +E+ G+ GESS ++P E + L+++ + EEP T
Sbjct: 335 VQEGTKEEGKSLKTGLTKETGESSASIPA--------ESYILDLQRISKEEPETQSHHFF 386
Query: 516 RPCE--FHIRNEIETVPNVEHQ-------------FIRTSFAGKSPVHQSFLKEDMQFKS 560
E + +E +P E+Q + + +G ++ F ++ K
Sbjct: 387 TADERLNQLHEAVEDIPGNENQKTVLTSPTTKEGGLVFSEKSGTKRDYEVFSAQEKPQKH 446
Query: 561 LSELRNEPDEDNFVSLH-LGEPEAKRRKHSNSSL 593
L+N +H +G+ K+ K + S+L
Sbjct: 447 YERLQNVKSSGKQGKMHGMGKSNIKKSKETQSTL 480
>At3g49600 putative protein
Length = 1672
Score = 30.0 bits (66), Expect = 3.8
Identities = 18/62 (29%), Positives = 32/62 (51%), Gaps = 1/62 (1%)
Query: 533 EHQFIRTSFAGKSPVHQSFLKEDMQFKSLSELRNEPDEDNFVSLHLGEPEAKRRKHSNSS 592
EH F+ +G+ V + ++D + K + R D+D V + E+K+R+H +SS
Sbjct: 169 EHSFLDRD-SGRKKVDEDVDEKDAKVKESKKQRGGDDDDVDVVKRHKKKESKKRRHDDSS 227
Query: 593 LS 594
S
Sbjct: 228 ES 229
>At1g67070 phosphomannose isomerase (din9)
Length = 441
Score = 29.6 bits (65), Expect = 5.0
Identities = 31/125 (24%), Positives = 53/125 (41%), Gaps = 24/125 (19%)
Query: 3 RNRLGLSAHHSGSTQSEESALDLERNYYGHPSSSPLHMQTFAVGVQHSEGNAAYFSWPTL 62
+NRL L H ++ E+ L+LE+ Y G +GV +A +F++ L
Sbjct: 239 KNRLLLETKHRELSEKEKLVLELEKQYTGD------------IGVI----SAFFFNYVKL 282
Query: 63 N----RWNDAAEDRANYFGNLQKGVLPE----TLGRLPSGQQATTLLELMTIRAFHSKIL 114
N + DA E A G+ + + G P + TL ++T + + +IL
Sbjct: 283 NPGEALYLDANEPHAYISGDCVECMAASDNVVRAGLTPKHRDVQTLCSMLTYKLGYPEIL 342
Query: 115 RRFSL 119
+ F L
Sbjct: 343 KGFPL 347
>At5g63380 4-coumarate-CoA ligase-like protein
Length = 562
Score = 29.3 bits (64), Expect = 6.5
Identities = 14/53 (26%), Positives = 29/53 (54%), Gaps = 1/53 (1%)
Query: 312 VITSIRGVGDIGEV-HRIDLQSPINSLIGRQVIKVGRSSGLTTGTIMAYALEY 363
V++ +G EV H++++ P+ + Q +K +SS L GT++ + E+
Sbjct: 130 VVSPANPIGSESEVSHQVEVSEPVIAFATSQTVKKLQSSSLPLGTVLMDSTEF 182
>At3g45450 clpC-like protein
Length = 341
Score = 29.3 bits (64), Expect = 6.5
Identities = 12/31 (38%), Positives = 18/31 (57%)
Query: 506 EPSTIVKPSLRPCEFHIRNEIETVPNVEHQF 536
+ + I+KP+L CE R IE P +E +F
Sbjct: 263 DAANILKPALERCELQYRKHIENDPALERRF 293
>At3g26120 RNA-binding protein, putative
Length = 615
Score = 29.3 bits (64), Expect = 6.5
Identities = 18/59 (30%), Positives = 26/59 (43%), Gaps = 3/59 (5%)
Query: 462 EQMNGSTAGIGSTVGESSPTVPIKEKLEESFEPFCLNMEHVPVEEPSTIVKPSLRPCEF 520
++MNG G V E S IK + S +P + P+ EP ++ P RP F
Sbjct: 267 DRMNGKEIGGKQVVIEFSRPGGIKNRFRSSRQP---QLPFQPLREPPILIPPLRRPVSF 322
>At3g05790 putative mitochondrial LON ATP-dependent protease
Length = 942
Score = 29.3 bits (64), Expect = 6.5
Identities = 14/54 (25%), Positives = 26/54 (47%)
Query: 293 FVRADGAFIPFAEDFNMNNVITSIRGVGDIGEVHRIDLQSPINSLIGRQVIKVG 346
F+ D A + N++ ++G I +H + + I+S+ G QVI +G
Sbjct: 122 FLLKDDASSDSSSSSETENILEKLKGKELINRIHEVGTLAQISSIQGEQVILIG 175
>At1g26090 unknown protein
Length = 327
Score = 29.3 bits (64), Expect = 6.5
Identities = 19/66 (28%), Positives = 30/66 (44%)
Query: 69 AEDRANYFGNLQKGVLPETLGRLPSGQQATTLLELMTIRAFHSKILRRFSLGTAIGFRIR 128
A+ R N + +GV+ E LG LP ++LEL + F + R+ G I
Sbjct: 12 ADARLNMTQGVLEGVVGEELGVLPGMDSIFSMLELERLVGFFRQATRKNHKGKPFDVIIY 71
Query: 129 GGVLTD 134
G+ T+
Sbjct: 72 DGISTE 77
>At1g21580 hypothetical protein
Length = 1696
Score = 29.3 bits (64), Expect = 6.5
Identities = 25/86 (29%), Positives = 34/86 (39%), Gaps = 6/86 (6%)
Query: 429 PENWTS-----GVDLGRLLDLLELDLVTTNETLQDSGQ-EQMNGSTAGIGSTVGESSPTV 482
PEN +S G+D LD+ + L N SG N T G +SP
Sbjct: 709 PENDSSRGRPMGLDSPASLDIANVSLDLANSNNSASGDLANANSFTVGTYMNPMVTSPDK 768
Query: 483 PIKEKLEESFEPFCLNMEHVPVEEPS 508
+ ++E P C N + PVE S
Sbjct: 769 SVVFQMESKNLPHCKNTVNAPVENVS 794
>At3g56575 unknown protein
Length = 320
Score = 28.9 bits (63), Expect = 8.5
Identities = 21/91 (23%), Positives = 37/91 (40%), Gaps = 3/91 (3%)
Query: 464 MNGSTAGIGSTVGESSPTVPIKEKLEESFEPFCLNMEH---VPVEEPSTIVKPSLRPCEF 520
+NG+ GIG G ++ LEE E H P + S P+++ +
Sbjct: 118 VNGAAPGIGIARGTNAGDYFFGPGLEELIEQLSSGTHHRGPPPAPKSSIDALPTIKITQK 177
Query: 521 HIRNEIETVPNVEHQFIRTSFAGKSPVHQSF 551
H+++ P + +F S A + P H +
Sbjct: 178 HLKSSDSHCPVCKDEFELKSEAKQMPCHHIY 208
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,148,793
Number of Sequences: 26719
Number of extensions: 653124
Number of successful extensions: 1542
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 1516
Number of HSP's gapped (non-prelim): 21
length of query: 597
length of database: 11,318,596
effective HSP length: 105
effective length of query: 492
effective length of database: 8,513,101
effective search space: 4188445692
effective search space used: 4188445692
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149576.6


