

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.4 + phase: 0 /pseudo
(548 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
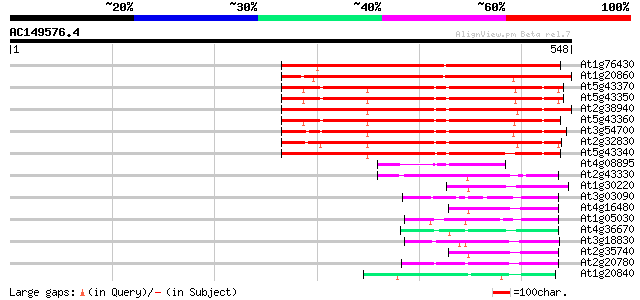
Score E
Sequences producing significant alignments: (bits) Value
At1g76430 putative phosphate transporter 353 1e-97
At1g20860 putative inorganic phosphate transporter protein 349 3e-96
At5g43370 inorganic phosphate transporter (dbj|BAA24282.1) 276 3e-74
At5g43350 phosphate transporter (gb|AAB17265.1) 275 6e-74
At2g38940 phosphate transporter (AtPT2) 273 2e-73
At5g43360 inorganic phosphate transporter (dbj|BAA24281.1) 269 3e-72
At3g54700 phosphate transport protein 265 4e-71
At2g32830 phosphate transporter like protein 248 8e-66
At5g43340 inorganic phosphate transporter (dbj|BAA34390.1) 239 4e-63
At4g08895 putative protein 104 1e-22
At2g43330 membrane transporter like protein 69 9e-12
At1g30220 unknown protein 50 2e-06
At3g03090 hypothetical protein 49 7e-06
At4g16480 membrane transporter like protein 49 9e-06
At1g05030 sugar transporter like protein 47 2e-05
At4g36670 sugar transporter like protein 46 6e-05
At3g18830 sugar transporter like protein 46 6e-05
At2g35740 putative sugar transporter 44 2e-04
At2g20780 putative sugar transporter 44 2e-04
At1g20840 putative sugar transporter protein 43 4e-04
>At1g76430 putative phosphate transporter
Length = 532
Score = 353 bits (906), Expect = 1e-97
Identities = 175/277 (63%), Positives = 214/277 (77%), Gaps = 5/277 (1%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSM-SQITEEHPLP---PATNVAYPLLSREFLWRHGRDL 321
YTALVE NV+QAAKDM++V+ VSM SQITE+ P ++ +Y L SR FL HGRDL
Sbjct: 238 YTALVENNVVQAAKDMQRVMSVSMISQITEDSSSELEQPPSSSSYKLFSRRFLSLHGRDL 297
Query: 322 FACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
FA SANWFL+D+VFY+ L S+I+ + +VY AF +A++ AI+A CSTIPGY
Sbjct: 298 FAASANWFLVDVVFYTSNLLLSQIFNFSNKPLNSTNVYDSAFEVAKLAAIVAACSTIPGY 357
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTVYFID++GRVKIQMMGFF MAV + G PY +W+K E NKGFMV+YGL FFF+
Sbjct: 358 WFTVYFIDKIGRVKIQMMGFFLMAVVYLVAGIPYSWYWSKHEK-TNKGFMVLYGLIFFFS 416
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIG 501
NFGPNTTTFI+PAELFPARFRSTCHGISGA GK GAI+G+VGFLWA+ +E+G+P
Sbjct: 417 NFGPNTTTFIIPAELFPARFRSTCHGISGAAGKFGAIVGTVGFLWATRHHEEDGFPDVKR 476
Query: 502 MKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDE 538
++ + +ILGGVCI GM VTY FT+ETMGRSLEENEDE
Sbjct: 477 VRIAFLILGGVCIAGMIVTYLFTRETMGRSLEENEDE 513
>At1g20860 putative inorganic phosphate transporter protein
Length = 534
Score = 349 bits (895), Expect = 3e-96
Identities = 177/291 (60%), Positives = 219/291 (74%), Gaps = 11/291 (3%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE---HPLPPATNVAYPLLSREFLWRHGRDLF 322
YTALVE N++QAAKDM++V+ S S I++E P PP +Y L SR F HGRDLF
Sbjct: 230 YTALVENNIVQAAKDMQRVM--SRSHISDEATTDPPPPPPPPSYKLFSRCFFRLHGRDLF 287
Query: 323 ACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDD-EDVYQEAFHLARIQAILAVCSTIPGY 381
A S NWFL+DIVFY+ L S I+ Y + E+VY AF +A + AI+A CSTIPGY
Sbjct: 288 AASFNWFLVDIVFYTSNLLLSHIFSHYSKKPSTAENVYDAAFEVAELGAIIAACSTIPGY 347
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTVYFID++GRVKIQ+MGFFFMAV + G PY +W+K E H+NKGFMV+YGL FFF
Sbjct: 348 WFTVYFIDKIGRVKIQIMGFFFMAVIYLVAGIPYSWYWSKHE-HNNKGFMVLYGLVFFFC 406
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHK----EKEEGYP 497
NFGPNTTTFI+PAE FPARFRSTCHGISGA GK+GAI+G+VGFLWA+ K +K + YP
Sbjct: 407 NFGPNTTTFIIPAEHFPARFRSTCHGISGAAGKLGAIVGTVGFLWATKKMESDDKNQIYP 466
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAEEYND 548
+ M+ + +ILGGVCI G+ VTYFFTKETMGRSLEENE +Q ++AE ++
Sbjct: 467 EVNRMRIAFLILGGVCIAGILVTYFFTKETMGRSLEENEHDQDNNAESEDE 517
>At5g43370 inorganic phosphate transporter (dbj|BAA24282.1)
Length = 524
Score = 276 bits (705), Expect = 3e-74
Identities = 151/286 (52%), Positives = 189/286 (65%), Gaps = 16/286 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVL--DVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFA 323
YTALV +N+ QA DM KVL D+ + + E+ P N Y L S+EFL RHG L
Sbjct: 242 YTALVAKNIKQATADMSKVLQTDIELEERVEDDVKDPRQN--YGLFSKEFLRRHGLHLLG 299
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
++ WFLLDI FYSQ LFQ +I+ ++ + + E F +AR Q ++A+CST+PGY
Sbjct: 300 TTSTWFLLDIAFYSQNLFQKDIFSAIGWIPKAATMNATHEVFRIARAQTLIALCSTVPGY 359
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV FID +GR KIQ+ GFF M V FA+ FPY +HW K EN GF+V+Y L FFFA
Sbjct: 360 WFTVAFIDTIGRFKIQLNGFFMMTVFMFAIAFPY-NHWIKPENRI--GFVVMYSLTFFFA 416
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEK----EEGYP 497
NFGPN TTFIVPAE+FPAR RSTCHGIS A GK GAIIG+ GFL+A+ + + GYP
Sbjct: 417 NFGPNATTFIVPAEIFPARLRSTCHGISAAAGKAGAIIGAFGFLYAAQNQDKAKVDAGYP 476
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE--NEDEQSH 541
GIG+K SLI+LG + +GM T F E G+SLEE E E SH
Sbjct: 477 PGIGVKNSLIVLGVLNFIGMLFT-FLVPEPKGKSLEELSGEAEVSH 521
Score = 30.0 bits (66), Expect = 3.5
Identities = 20/81 (24%), Positives = 34/81 (41%), Gaps = 8/81 (9%)
Query: 364 HLARIQAILAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGE 423
H+A +A+C T+ G F + D++GR K+ + M + A G +
Sbjct: 68 HVAAAVNGVALCGTLSGQLFFGWLGDKLGRKKVYGLTLIMMILCSVASGLSF-------- 119
Query: 424 NHDNKGFMVIYGLAFFFANFG 444
++ KG M F+ FG
Sbjct: 120 GNEAKGVMTTLCFFRFWLGFG 140
>At5g43350 phosphate transporter (gb|AAB17265.1)
Length = 524
Score = 275 bits (702), Expect = 6e-74
Identities = 150/286 (52%), Positives = 189/286 (65%), Gaps = 16/286 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVL--DVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFA 323
YTALV +N+ QA DM KVL D+ + + E+ P N Y L S+EFL RHG L
Sbjct: 242 YTALVAKNIKQATADMSKVLQTDIELEERVEDDVKDPKQN--YGLFSKEFLRRHGLHLLG 299
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
++ WFLLDI FYSQ LFQ +I+ ++ + + E F +AR Q ++A+CST+PGY
Sbjct: 300 TTSTWFLLDIAFYSQNLFQKDIFSAIGWIPKAATMNATHEVFRIARAQTLIALCSTVPGY 359
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV FID +GR KIQ+ GFF M V FA+ FPY +HW K EN GF+V+Y L FFFA
Sbjct: 360 WFTVAFIDTIGRFKIQLNGFFMMTVFMFAIAFPY-NHWIKPENRI--GFVVMYSLTFFFA 416
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEK----EEGYP 497
NFGPN TTFIVPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + + GYP
Sbjct: 417 NFGPNATTFIVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSQDKAKVDAGYP 476
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE--NEDEQSH 541
GIG+K SLI+LG + +GM T F E G+SLEE E E SH
Sbjct: 477 PGIGVKNSLIMLGVLNFIGMLFT-FLVPEPKGKSLEELSGEAEVSH 521
Score = 32.3 bits (72), Expect = 0.70
Identities = 21/81 (25%), Positives = 34/81 (41%), Gaps = 8/81 (9%)
Query: 364 HLARIQAILAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGE 423
H+A +A+C T+ G F + D++GR K+ + M + A G +
Sbjct: 68 HVAAAVNGVALCGTLSGQLFFGWLGDKLGRKKVYGLTLVMMILCSVASGLSF-------- 119
Query: 424 NHDNKGFMVIYGLAFFFANFG 444
H+ KG M F+ FG
Sbjct: 120 GHEAKGVMTTLCFFRFWLGFG 140
>At2g38940 phosphate transporter (AtPT2)
Length = 534
Score = 273 bits (698), Expect = 2e-73
Identities = 146/290 (50%), Positives = 193/290 (66%), Gaps = 11/290 (3%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE-HPLPPATNVAYPLLSREFLWRHGRDLFAC 324
YTALV ++ QAA DM KVL V + ++ + + A+ L S+EF+ RHG L
Sbjct: 242 YTALVAKDAKQAASDMSKVLQVEIEPEQQKLEEISKEKSKAFGLFSKEFMSRHGLHLLGT 301
Query: 325 SANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYF 382
++ WFLLDI FYSQ LFQ +I+ ++ + QE F +AR Q ++A+CST+PGY+
Sbjct: 302 TSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAQSMNAIQEVFKIARAQTLIALCSTVPGYW 361
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFAN 442
FTV FID +GR IQMMGFFFM V FAL PY +HWT EN GF+++Y L FFFAN
Sbjct: 362 FTVAFIDVIGRFAIQMMGFFFMTVFMFALAIPY-NHWTHKENRI--GFVIMYSLTFFFAN 418
Query: 443 FGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLW-ASHKEKEE---GYPK 498
FGPN TTF+VPAE+FPARFRSTCHGIS A GK+GA++G+ GFL+ A + +K++ GYP
Sbjct: 419 FGPNATTFVVPAEIFPARFRSTCHGISAASGKLGAMVGAFGFLYLAQNPDKDKTDAGYPP 478
Query: 499 GIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAEEYND 548
GIG++ SLI+LG V +G+ T F E+ G+SLEE E + ND
Sbjct: 479 GIGVRNSLIVLGVVNFLGILFT-FLVPESKGKSLEEMSGENEDNENSNND 527
Score = 30.0 bits (66), Expect = 3.5
Identities = 19/73 (26%), Positives = 29/73 (39%), Gaps = 8/73 (10%)
Query: 372 LAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFM 431
+A C T+ G F + D++GR K+ M M + A G + H+ K M
Sbjct: 76 VAFCGTLAGQLFFGWLGDKLGRKKVYGMTLMVMVLCSIASGLSF--------GHEPKAVM 127
Query: 432 VIYGLAFFFANFG 444
F+ FG
Sbjct: 128 ATLCFFRFWLGFG 140
>At5g43360 inorganic phosphate transporter (dbj|BAA24281.1)
Length = 521
Score = 269 bits (687), Expect = 3e-72
Identities = 146/281 (51%), Positives = 185/281 (64%), Gaps = 14/281 (4%)
Query: 266 YTALVEQNVLQAAKDMEKVL--DVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFA 323
YTALV +N+ QA DM KVL D+ + + E+ P N Y L S+EFL RHG L
Sbjct: 242 YTALVAKNIKQATADMSKVLQTDLELEERVEDDVKDPKKN--YGLFSKEFLRRHGLHLLG 299
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
++ WFLLDI FYSQ LFQ +I+ ++ + + E F +AR Q ++A+CST+PGY
Sbjct: 300 TTSTWFLLDIAFYSQNLFQKDIFSAIGWIPKAATMNAIHEVFKIARAQTLIALCSTVPGY 359
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV FID +GR IQ+MGFF M V FA+ FPY +HW +N GF+V+Y L FFFA
Sbjct: 360 WFTVAFIDIIGRFAIQLMGFFMMTVFMFAIAFPY-NHWILPDNRI--GFVVMYSLTFFFA 416
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKE----KEEGYP 497
NFGPN TTFIVPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + + GYP
Sbjct: 417 NFGPNATTFIVPAEIFPARLRSTCHGISAATGKAGAIVGAFGFLYAAQPQDKTKTDAGYP 476
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDE 538
GIG+K SLI+LG + VGM T F E G+SLEE E
Sbjct: 477 PGIGVKNSLIMLGVINFVGMLFT-FLVPEPKGKSLEELSGE 516
>At3g54700 phosphate transport protein
Length = 433
Score = 265 bits (678), Expect = 4e-71
Identities = 144/285 (50%), Positives = 186/285 (64%), Gaps = 13/285 (4%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFACS 325
YTALV ++ AA +M KVL V + E+ +N ++ L S+EF+ RHG L +
Sbjct: 140 YTALVAKDAKLAASNMSKVLQVEIE--AEQQGTEDKSN-SFGLFSKEFMKRHGLHLLGTT 196
Query: 326 ANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFF 383
+ WFLLDI FYSQ LFQ +I+ ++ + QE F +AR Q ++A+CST+PGY+F
Sbjct: 197 STWFLLDIAFYSQNLFQKDIFSAIGWIPPAQTMNAIQEVFKIARAQTLIALCSTVPGYWF 256
Query: 384 TVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANF 443
TV FID +GR IQMMGFFFM V FAL PY HWT EN GF+ +Y L FFFANF
Sbjct: 257 TVAFIDVIGRFAIQMMGFFFMTVFMFALAIPY-DHWTHKENRI--GFVAMYSLTFFFANF 313
Query: 444 GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHK----EKEEGYPKG 499
GPN TTF+VPAE+FPARFRSTCHGIS A GK+GA++G+ GFL+ + + E GYP G
Sbjct: 314 GPNATTFVVPAEIFPARFRSTCHGISAASGKLGAMVGAFGFLYLAQSPDKTKTEHGYPPG 373
Query: 500 IGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAE 544
IG+K SLI+LG V ++GM T E+ G+SLEE E + E
Sbjct: 374 IGVKNSLIVLGVVNLLGMVFT-LLVPESKGKSLEEMSGENEQNDE 417
>At2g32830 phosphate transporter like protein
Length = 542
Score = 248 bits (632), Expect = 8e-66
Identities = 137/286 (47%), Positives = 178/286 (61%), Gaps = 18/286 (6%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPAT----NVAYPLLSREFLWRHGRDL 321
YTALV +N QAA DM KVL V + I EE + N + L +REF RHG L
Sbjct: 242 YTALVARNTKQAASDMSKVLQVDL--IAEEEAQSNSNSSNPNFTFGLFTREFARRHGLHL 299
Query: 322 FACSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIP 379
+ WFLLDI +YS LFQ +IY ++ + + E F +++ Q ++A+C T+P
Sbjct: 300 LGTTTTWFLLDIAYYSSNLFQKDIYTAIGWIPAAETMNAIHEVFTVSKAQTLIALCGTVP 359
Query: 380 GYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFF 439
GY+FTV FID +GR IQ+MGF FM + FAL PY HW EN GF+++Y L F
Sbjct: 360 GYWFTVAFIDILGRFFIQLMGFIFMTIFMFALAIPY-DHWRHRENRI--GFLIMYSLTMF 416
Query: 440 FANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEE----G 495
FANFGPN TTF+VPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ E G
Sbjct: 417 FANFGPNATTFVVPAEIFPARLRSTCHGISAASGKAGAIVGAFGFLYAAQSSDSEKTDAG 476
Query: 496 YPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE--NEDEQ 539
YP GIG++ SL++L V +G+ T E+ G+SLEE EDE+
Sbjct: 477 YPPGIGVRNSLLMLACVNFLGIVFT-LLVPESKGKSLEEISREDEE 521
>At5g43340 inorganic phosphate transporter (dbj|BAA34390.1)
Length = 516
Score = 239 bits (609), Expect = 4e-63
Identities = 131/275 (47%), Positives = 171/275 (61%), Gaps = 15/275 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFACS 325
YTALV +N QAA DM KVL+V + ++ ++ + L S +FL RHG L +
Sbjct: 243 YTALVSKNAEQAALDMTKVLNVDIEASAAKNDQARVSSDEFGLFSMKFLRRHGLHLLGTA 302
Query: 326 ANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFF 383
+ WFLLDI FYSQ LFQ +I+ +L + QE + +A+ Q I+A CST+PGYFF
Sbjct: 303 STWFLLDIAFYSQNLFQKDIFTTIGWLPSAKTMNAIQELYMIAKAQTIIACCSTVPGYFF 362
Query: 384 TVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANF 443
TV FID +GR KIQ+MGF M + +L PY+ HWT N GF+V+Y FFF+NF
Sbjct: 363 TVGFIDYMGRKKIQIMGFAMMTIFMLSLAIPYH-HWTLPANRI--GFVVLYSFTFFFSNF 419
Query: 444 GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMK 503
GPN TTFIVPAE+FPAR RSTCHGIS A GK GA++GS GF K +GM
Sbjct: 420 GPNATTFIVPAEIFPARIRSTCHGISAASGKAGAMVGSFGF---------SALVKALGMS 470
Query: 504 ASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDE 538
+L I+ G+ ++G+ +T F ET G+SLEE E
Sbjct: 471 NTLYIMAGINLLGLLLT-FTIPETNGKSLEELSGE 504
>At4g08895 putative protein
Length = 155
Score = 104 bits (260), Expect = 1e-22
Identities = 57/125 (45%), Positives = 67/125 (53%), Gaps = 35/125 (28%)
Query: 360 QEAFHLARIQAILAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHW 419
QE F + R Q I+A CST+P PY+ HW
Sbjct: 41 QELFMIVRAQTIIACCSTVPAV--------------------------------PYH-HW 67
Query: 420 TKGENHDNKGFMVIYGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAII 479
T N GF++ Y FFF+NFGPN TTFIVPAE+FPAR RSTCHGIS A GK GA++
Sbjct: 68 TLPANRI--GFVIFYSFTFFFSNFGPNATTFIVPAEIFPARIRSTCHGISAASGKAGAMV 125
Query: 480 GSVGF 484
GS GF
Sbjct: 126 GSFGF 130
>At2g43330 membrane transporter like protein
Length = 509
Score = 68.6 bits (166), Expect = 9e-12
Identities = 46/179 (25%), Positives = 79/179 (43%), Gaps = 15/179 (8%)
Query: 360 QEAFHLARIQAILAVCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHW 419
Q A L+ I A + T+ G +YFID GR K+ + F + +S L ++
Sbjct: 311 QLALFLSLIVAAMNAAGTVVG----IYFIDHCGRKKLALSSLFGVIISLLILSVSFFKQS 366
Query: 420 TKGENHDNKGFMVIYGLAFFFANFGP--NTTTFIVPAELFPARFRSTCHGISGAVGKVGA 477
+ G++ + GLA + F P + V +E++P ++R C G+S V +
Sbjct: 367 ETSSDGGLYGWLAVLGLALYIVFFAPGMGPVPWTVNSEIYPQQYRGICGGMSATVNWISN 426
Query: 478 IIGSVGFLWASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
+I + FL + G GM + +IL G+ ++ + F ET G + E E
Sbjct: 427 LIVAQTFLTIAE-------AAGTGM--TFLILAGIAVLAVIFVIVFVPETQGLTFSEVE 476
>At1g30220 unknown protein
Length = 580
Score = 50.4 bits (119), Expect = 2e-06
Identities = 31/122 (25%), Positives = 52/122 (42%), Gaps = 11/122 (9%)
Query: 427 NKGFMVIYGLAFFFANFGPN--TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGF 484
N G+ + GL + F P T +IV +E++P RFR C GI+ + +I + F
Sbjct: 451 NFGWFALLGLGLYIIFFSPGMGTVPWIVNSEIYPLRFRGICGGIAATANWISNLIVAQSF 510
Query: 485 LWASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAE 544
L + IG + +I G + ++ + ET G +EE E +
Sbjct: 511 L---------SLTEAIGTSWTFLIFGVISVIALLFVMVCVPETKGMPMEEIEKMLERRSM 561
Query: 545 EY 546
E+
Sbjct: 562 EF 563
Score = 30.8 bits (68), Expect = 2.0
Identities = 13/37 (35%), Positives = 22/37 (59%)
Query: 380 GYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYY 416
G ++YFIDR+GR K+ ++ F + +S L +Y
Sbjct: 327 GSIISIYFIDRIGRKKLLIISLFGVIISLGILTGVFY 363
>At3g03090 hypothetical protein
Length = 342
Score = 48.9 bits (115), Expect = 7e-06
Identities = 45/153 (29%), Positives = 68/153 (44%), Gaps = 14/153 (9%)
Query: 384 TVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANF 443
+V IDRVGR + + G M +S F LG YY + ++ G + +F
Sbjct: 200 SVIVIDRVGRRPLLLCGVSGMVISLFLLG-SYYMFYKNVPAVAVAALLLYVGC--YQLSF 256
Query: 444 GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIGMK 503
GP +++ +E+FP + R GIS AV V F ++ KE +G
Sbjct: 257 GP--IGWLMISEIFPLKLRG--RGISLAVLVNFGANALVTFAFSPLKEL-------LGAG 305
Query: 504 ASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
G +C+V +F Y+ ET G +LEE E
Sbjct: 306 ILFCAFGVICVVSLFFIYYIVPETKGLTLEEIE 338
>At4g16480 membrane transporter like protein
Length = 582
Score = 48.5 bits (114), Expect = 9e-06
Identities = 30/110 (27%), Positives = 48/110 (43%), Gaps = 11/110 (10%)
Query: 429 GFMVIYGLAFFFANFGPN--TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLW 486
GF+ I L + + P T +IV +E++P R+R GI+ V +I S FL
Sbjct: 457 GFLAIVFLGLYIVVYAPGMGTVPWIVNSEIYPLRYRGLGGGIAAVSNWVSNLIVSESFLS 516
Query: 487 ASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
+H +G + ++ G +G+F + ET G EE E
Sbjct: 517 LTH---------ALGSSGTFLLFAGFSTIGLFFIWLLVPETKGLQFEEVE 557
>At1g05030 sugar transporter like protein
Length = 524
Score = 47.4 bits (111), Expect = 2e-05
Identities = 47/157 (29%), Positives = 66/157 (41%), Gaps = 23/157 (14%)
Query: 386 YFIDRVGRVKIQMMGFFFMAVSFF----ALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
Y ID+ GR K+ + + MAVS F A+GFP + D + I G +
Sbjct: 375 YLIDKQGRKKLLIGSYLGMAVSMFLIVYAVGFPL--------DEDLSQSLSILGTLMYIF 426
Query: 442 NF--GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKG 499
+F G T ++ EL R R G S +V V + VG + EK
Sbjct: 427 SFAIGAGPVTGLIIPELSSNRTRGKIMGFSFSVHWVSNFL--VGLFFLDLVEK------- 477
Query: 500 IGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
G+ G V ++ ++ FT ET GRSLEE E
Sbjct: 478 YGVGTVYASFGSVSLLAAAFSHLFTVETKGRSLEEIE 514
>At4g36670 sugar transporter like protein
Length = 493
Score = 45.8 bits (107), Expect = 6e-05
Identities = 43/163 (26%), Positives = 65/163 (39%), Gaps = 23/163 (14%)
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNK-------GFMVIY 434
F +D+VGR K+ + M ++ LGF T +N K + Y
Sbjct: 329 FTATLLLDKVGRRKLLLTSVGGMVIALTMLGFGL----TMAQNAGGKLAWALVLSIVAAY 384
Query: 435 G-LAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKE 493
+AFF GP T++ +E+FP + R+ + AV +V S+ FL
Sbjct: 385 SFVAFFSIGLGP--ITWVYSSEVFPLKLRAQGASLGVAVNRVMNATVSMSFL-------- 434
Query: 494 EGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
I + + GV V +F ET G+SLEE E
Sbjct: 435 -SLTSAITTGGAFFMFAGVAAVAWNFFFFLLPETKGKSLEEIE 476
>At3g18830 sugar transporter like protein
Length = 539
Score = 45.8 bits (107), Expect = 6e-05
Identities = 40/157 (25%), Positives = 68/157 (42%), Gaps = 17/157 (10%)
Query: 386 YFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAF---FFAN 442
+ +DR+GR + + M +S ALG S ++ + V+ +A + A
Sbjct: 352 FLLDRIGRRPLLLTSVGGMVLSLAALGT---SLTIIDQSEKKVMWAVVVAIATVMTYVAT 408
Query: 443 F--GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGI 500
F G T++ +E+FP R RS + V +V + + S+ FL S K +
Sbjct: 409 FSIGAGPITWVYSSEIFPLRLRSQGSSMGVVVNRVTSGVISISFLPMS---------KAM 459
Query: 501 GMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENED 537
+ + GG+ V Y F ET GR LE+ ++
Sbjct: 460 TTGGAFYLFGGIATVAWVFFYTFLPETQGRMLEDMDE 496
>At2g35740 putative sugar transporter
Length = 580
Score = 43.9 bits (102), Expect = 2e-04
Identities = 27/110 (24%), Positives = 48/110 (43%), Gaps = 11/110 (10%)
Query: 429 GFMVIYGLAFFFANFGPN--TTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLW 486
G++ I L + + P T +IV +E++P R+R GI+ + ++ S FL
Sbjct: 456 GYLAIVFLGLYIIVYAPGMGTVPWIVNSEIYPLRYRGLAGGIAAVSNWMSNLVVSETFLT 515
Query: 487 ASHKEKEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
++ +G + ++ G VG+F + ET G EE E
Sbjct: 516 LTN---------AVGSSGTFLLFAGSSAVGLFFIWLLVPETKGLQFEEVE 556
>At2g20780 putative sugar transporter
Length = 547
Score = 43.9 bits (102), Expect = 2e-04
Identities = 36/155 (23%), Positives = 65/155 (41%), Gaps = 13/155 (8%)
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYG-LAFFFA 441
F + ID VGR + + M + F L F + +G + + G +AFF
Sbjct: 376 FATFLIDSVGRKPLLYVSTIGMTLCLFCLSFTL-TFLGQGTLGITLALLFVCGNVAFFSI 434
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIG 501
GP +++ +E+FP R R+ + +V + + ++ FL S + I
Sbjct: 435 GMGP--VCWVLTSEIFPLRLRAQASALGAVGNRVCSGLVAMSFLSVS---------RAIT 483
Query: 502 MKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
+ + + V + + Y ET G+SLE+ E
Sbjct: 484 VGGTFFVFSLVSALSVIFVYVLVPETSGKSLEQIE 518
>At1g20840 putative sugar transporter protein
Length = 734
Score = 43.1 bits (100), Expect = 4e-04
Identities = 47/195 (24%), Positives = 71/195 (36%), Gaps = 20/195 (10%)
Query: 346 YKRYLNEKDDEDVYQEAFHLARIQAILAVCST-----IPGYFFTVYFIDRVGRVKIQMMG 400
Y + E+ D+ + L+ I A + +P + +D GR + +
Sbjct: 532 YTPQILERAGVDILLSSLGLSSISASFLISGLTTLLMLPAIVVAMRLMDVSGRRSLLLWT 591
Query: 401 FFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANFGPNTTTFIVPAELFPAR 460
+ VS L H +K N V+ FF +GP I+ +E+FP R
Sbjct: 592 IPVLIVSLVVLVISELIHISKVVNAALSTGCVVLYFCFFVMGYGPIPN--ILCSEIFPTR 649
Query: 461 FRSTCHGISGAVGKVGAII--GSVGFLWASHKEKEEGYPKGIGMKASLIILGGVCIVGMF 518
R C I V +G II S+ L +S IG+ I VC++
Sbjct: 650 VRGLCIAICAMVFWIGDIIVTYSLPVLLSS-----------IGLVGVFSIYAAVCVISWI 698
Query: 519 VTYFFTKETMGRSLE 533
Y ET G LE
Sbjct: 699 FVYMKVPETKGMPLE 713
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.342 0.150 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,388,280
Number of Sequences: 26719
Number of extensions: 454932
Number of successful extensions: 1678
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 33
Number of HSP's that attempted gapping in prelim test: 1586
Number of HSP's gapped (non-prelim): 65
length of query: 548
length of database: 11,318,596
effective HSP length: 104
effective length of query: 444
effective length of database: 8,539,820
effective search space: 3791680080
effective search space used: 3791680080
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149576.4


