

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
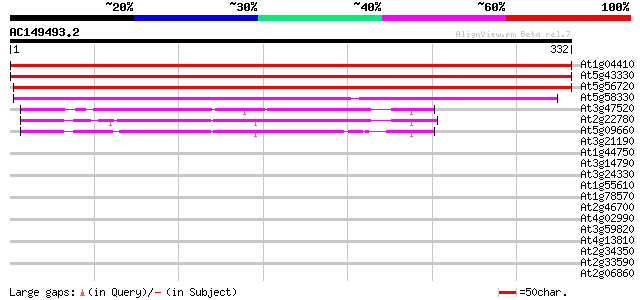
Query= AC149493.2 + phase: 0
(332 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g04410 malate dehydrogenase like protein 602 e-173
At5g43330 cytosolic malate dehydrogenase 600 e-172
At5g56720 cytosolic malate dehydrogenase 515 e-146
At5g58330 NADP-dependent malate dehydrogenase 240 7e-64
At3g47520 chloroplast NAD-dependent malate dehydrogenase 70 2e-12
At2g22780 putative glyoxysomal malate dehydrogenase precursor 67 1e-11
At5g09660 microbody NAD-dependent malate dehydrogenase 62 3e-10
At3g21190 unknown protein 33 0.28
At1g44750 Unknown protein (At1g44750) 31 0.82
At3g14790 dTDP-glucose 4,6-dehydratase, putative 31 1.1
At3g24330 beta-1,3-glucanase, putative 30 1.4
At1g55610 receptor kinase, putative 30 1.4
At1g78570 Similar to dTDP-D-glucose 4,6-dehydratase 30 1.8
At2g46700 calcium/calmodulin-dependent protein kinase CaMK4 30 2.4
At4g02990 unknown protein 29 3.1
At3g59820 putative protein 29 3.1
At4g13810 putative disease resistance protein 29 4.1
At2g34350 nodulin-like protein 29 4.1
At2g33590 putative cinnamoyl-CoA reductase 29 4.1
At2g06860 hypothetical protein 29 4.1
>At1g04410 malate dehydrogenase like protein
Length = 332
Score = 602 bits (1553), Expect = e-173
Identities = 300/332 (90%), Positives = 317/332 (95%)
Query: 1 MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVD 60
MAK+PVRVLVTGAAGQIGYALVPMIARG+MLG DQPVILHMLDI PAAE+LNGVKMEL+D
Sbjct: 1 MAKEPVRVLVTGAAGQIGYALVPMIARGIMLGADQPVILHMLDIPPAAEALNGVKMELID 60
Query: 61 AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKH 120
AAFPLLKGVVATTD VE CTGVN+AVMVGGFPRKEGMERKDVMSKNVSIYKSQA+ALEKH
Sbjct: 61 AAFPLLKGVVATTDAVEGCTGVNVAVMVGGFPRKEGMERKDVMSKNVSIYKSQAAALEKH 120
Query: 121 AAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVK 180
AA NCKVLVVANPANTNALILKEFAPSIPE+NISCLTRLDHNRALGQISERL+V VSDVK
Sbjct: 121 AAPNCKVLVVANPANTNALILKEFAPSIPEKNISCLTRLDHNRALGQISERLSVPVSDVK 180
Query: 181 NVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARK 240
NVIIWGNHSS+QYPDVNHA V T +GEKPVRELV DDAWL+GEFISTVQQRGAAIIKARK
Sbjct: 181 NVIIWGNHSSSQYPDVNHAKVQTSSGEKPVRELVKDDAWLDGEFISTVQQRGAAIIKARK 240
Query: 241 LSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIV 300
LSSALSAAS+ACDHIRDWVLGTP+GTFVSMGVYSDGSY+VPSGLIYSFPVTC NG+W IV
Sbjct: 241 LSSALSAASSACDHIRDWVLGTPEGTFVSMGVYSDGSYSVPSGLIYSFPVTCRNGDWSIV 300
Query: 301 QGLSIDEFSRKKLDLTAEELSEEKNLAYSCLS 332
QGL IDE SRKK+DLTAEEL EEK+LAYSCLS
Sbjct: 301 QGLPIDEVSRKKMDLTAEELKEEKDLAYSCLS 332
>At5g43330 cytosolic malate dehydrogenase
Length = 332
Score = 600 bits (1547), Expect = e-172
Identities = 300/332 (90%), Positives = 316/332 (94%)
Query: 1 MAKDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVD 60
MAK+PVRVLVTGAAGQIGYALVPMIARG+MLG DQPVILHMLDI AAE+LNGVKMELVD
Sbjct: 1 MAKEPVRVLVTGAAGQIGYALVPMIARGIMLGADQPVILHMLDIPFAAEALNGVKMELVD 60
Query: 61 AAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKH 120
AAFPLLKGVVATTD VEACTGVN+AVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKH
Sbjct: 61 AAFPLLKGVVATTDAVEACTGVNVAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKH 120
Query: 121 AAANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVK 180
AA NCKVLVVANPANTNALILKEFAPSIPE+NI+CLTRLDHNRALGQ+SERL+V VSDVK
Sbjct: 121 AAPNCKVLVVANPANTNALILKEFAPSIPEKNITCLTRLDHNRALGQVSERLSVPVSDVK 180
Query: 181 NVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARK 240
NVIIWGNHSSTQYPDVNHATV T GEKPVRELV +D WLNGEFISTVQQRGAAIIKARK
Sbjct: 181 NVIIWGNHSSTQYPDVNHATVKTSVGEKPVRELVKNDEWLNGEFISTVQQRGAAIIKARK 240
Query: 241 LSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIV 300
LSSALSAAS+ACDHIRDWV+GTP+GTFVSMGVYSDGSYNVP+GLIYSFPVTC NGEW IV
Sbjct: 241 LSSALSAASSACDHIRDWVVGTPEGTFVSMGVYSDGSYNVPAGLIYSFPVTCRNGEWTIV 300
Query: 301 QGLSIDEFSRKKLDLTAEELSEEKNLAYSCLS 332
QGL ID+ SRKK+DLTAEEL EEK+LAYSCLS
Sbjct: 301 QGLPIDDASRKKMDLTAEELKEEKDLAYSCLS 332
>At5g56720 cytosolic malate dehydrogenase
Length = 339
Score = 515 bits (1327), Expect = e-146
Identities = 247/330 (74%), Positives = 290/330 (87%)
Query: 3 KDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAA 62
KDP+RVL+TGAAG IGYA+ PMIARG+MLGPDQP+ILH+LDI PA+ SL VKMEL D+A
Sbjct: 9 KDPIRVLITGAAGNIGYAIAPMIARGIMLGPDQPMILHLLDIEPASSSLEAVKMELQDSA 68
Query: 63 FPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAA 122
FPLLKGV+ATT+VVEAC VNI +M+GGFPR GMERKDVMSKNV IYK+QASALE++A+
Sbjct: 69 FPLLKGVIATTNVVEACKDVNIVIMIGGFPRIAGMERKDVMSKNVVIYKAQASALERYAS 128
Query: 123 ANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVKNV 182
+CKVLVVANPANTNALILKEFAPSIPE NI+CLTRLDHNRAL Q++++L+V VS VKNV
Sbjct: 129 DDCKVLVVANPANTNALILKEFAPSIPEENITCLTRLDHNRALAQLADKLSVPVSSVKNV 188
Query: 183 IIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARKLS 242
I+WGNHSSTQYPD NHATV+T G++P++ELV+D WL EFI VQQRGAA+++ARK S
Sbjct: 189 IVWGNHSSTQYPDTNHATVSTKTGDRPLKELVTDHNWLKNEFIVEVQQRGAAVLRARKQS 248
Query: 243 SALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPVTCANGEWKIVQG 302
SA SAA AACDHIRDW LGTP+GT+VSMGV SDGSY +P GL+YSFPV C G WKIVQG
Sbjct: 249 SAFSAAGAACDHIRDWFLGTPKGTWVSMGVCSDGSYGIPPGLVYSFPVICEKGSWKIVQG 308
Query: 303 LSIDEFSRKKLDLTAEELSEEKNLAYSCLS 332
LSIDEFSR+K+D +A EL+EEK+LAYSCL+
Sbjct: 309 LSIDEFSREKMDDSARELAEEKDLAYSCLN 338
>At5g58330 NADP-dependent malate dehydrogenase
Length = 443
Score = 240 bits (613), Expect = 7e-64
Identities = 135/324 (41%), Positives = 196/324 (59%), Gaps = 6/324 (1%)
Query: 3 KDPVRVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAA 62
K + + V+GAAG I L+ +A G + GPDQP+ L +L + ++L GV MEL D+
Sbjct: 97 KKLINIAVSGAAGMISNHLLFKLASGEVFGPDQPIALKLLGSERSIQALEGVAMELEDSL 156
Query: 63 FPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAA 122
FPLL+ V TD E V A+++G PR GMER D++ N I+ Q AL K A+
Sbjct: 157 FPLLREVDIGTDPNEVFQDVEWAILIGAKPRGPGMERADLLDINGQIFAEQGKALNKAAS 216
Query: 123 ANCKVLVVANPANTNALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVKNV 182
N KVLVV NP NTNALI + AP+IP +N LTRLD NRA Q++ + V V N+
Sbjct: 217 PNVKVLVVGNPCNTNALICLKNAPNIPAKNFHALTRLDENRAKCQLALKAGVFYDKVSNM 276
Query: 183 IIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAIIKARKLS 242
IWGNHS+TQ PD +A +N PV+E+++D WL F +VQ+RG +I+ S
Sbjct: 277 TIWGNHSTTQVPDFLNARIN----GLPVKEVITDHKWLEEGFTESVQKRGGLLIQKWGRS 332
Query: 243 SALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGS-YNVPSGLIYSFPV-TCANGEWKIV 300
SA S A + D I+ V TP+G + S GVY+DG+ Y + GL++S P + +G++++V
Sbjct: 333 SAASTAVSIVDAIKSLVTPTPEGDWFSTGVYTDGNPYGIEEGLVFSMPCRSKGDGDYELV 392
Query: 301 QGLSIDEFSRKKLDLTAEELSEEK 324
+ + ID++ R+++ + EL EK
Sbjct: 393 KDVEIDDYLRQRIAKSEAELLAEK 416
>At3g47520 chloroplast NAD-dependent malate dehydrogenase
Length = 403
Score = 69.7 bits (169), Expect = 2e-12
Identities = 67/252 (26%), Positives = 120/252 (47%), Gaps = 29/252 (11%)
Query: 7 RVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAAFPL- 65
+V V GAAG IG L +I ++ LH+ DIA ++ GV +L P
Sbjct: 84 KVAVLGAAGGIGQPLSLLIKMSPLVST-----LHLYDIA----NVKGVAADLSHCNTPSQ 134
Query: 66 LKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAAANC 125
++ +++ + VN+ V+ G PRK GM R D+ + N +I K+ A+ ++ N
Sbjct: 135 VRDFTGPSELADCLKDVNVVVIPAGVPRKPGMTRDDLFNINANIVKTLVEAVAEN-CPNA 193
Query: 126 KVLVVANPANTN----ALILKEFAPSIPERNISCLTRLDHNRALGQISERLNVQVSDVKN 181
+ +++NP N+ A +LK+ P++ + +T LD RA +S++ N+++ DV
Sbjct: 194 FIHIISNPVNSTVPIAAEVLKKKGVYDPKK-LFGVTTLDVVRANTFVSQKKNLKLIDVDV 252
Query: 182 VIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAII--KAR 239
+I G+ T P ++ + ++ ++EL +Q G ++ KA
Sbjct: 253 PVIGGHAGITILPLLSKTKPSVNFTDEEIQELT-----------VRIQNAGTEVVDAKAG 301
Query: 240 KLSSALSAASAA 251
S+ LS A AA
Sbjct: 302 AGSATLSMAYAA 313
>At2g22780 putative glyoxysomal malate dehydrogenase precursor
Length = 354
Score = 67.4 bits (163), Expect = 1e-11
Identities = 62/254 (24%), Positives = 118/254 (46%), Gaps = 29/254 (11%)
Query: 7 RVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMEL--VDAAFP 64
+V + GAAG IG L ++ ++ +LH+ D+A A GV ++ +D +
Sbjct: 44 KVAILGAAGGIGQPLAMLMKMNPLVS-----VLHLYDVANAP----GVTADISHMDTS-A 93
Query: 65 LLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAAAN 124
+++G + + EA TG+++ ++ G PRK GM R D+ + N I ++ + A+ K
Sbjct: 94 VVRGFLGQPQLEEALTGMDLVIIPAGVPRKPGMTRDDLFNINAGIVRTLSEAIAK-CCPK 152
Query: 125 CKVLVVANPANTNALILKEF---APSIPERNISCLTRLDHNRALGQISERLNVQVSDVKN 181
V +++NP N+ I E A + + + +T LD RA ++E +++ +V+
Sbjct: 153 AIVNIISNPVNSTVPIAAEVFKKAGTFDPKKLMGVTMLDVVRANTFVAEVMSLDPREVEV 212
Query: 182 VIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAII--KAR 239
++ G+ T P ++ +K + L +Q G ++ KA
Sbjct: 213 PVVGGHAGVTILPLLSQVKPPCSFTQKEIEYLT-----------DRIQNGGTEVVEAKAG 261
Query: 240 KLSSALSAASAACD 253
S+ LS A AA +
Sbjct: 262 AGSATLSMAYAAVE 275
>At5g09660 microbody NAD-dependent malate dehydrogenase
Length = 354
Score = 62.4 bits (150), Expect = 3e-10
Identities = 64/251 (25%), Positives = 115/251 (45%), Gaps = 27/251 (10%)
Query: 7 RVLVTGAAGQIGYALVPMIARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVDAAFPLL 66
+V + GAAG IG +L ++ ++ +LH+ D+ A V A ++
Sbjct: 44 KVAILGAAGGIGQSLSLLMKMNPLVS-----LLHLYDVVNAPGVTADVSHMDTGA---VV 95
Query: 67 KGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASALEKHAAANCK 126
+G + + +A TG+++ ++ G PRK GM R D+ N I K+ + K N
Sbjct: 96 RGFLGAKQLEDALTGMDLVIIPAGIPRKPGMTRDDLFKINAGIVKTLCEGVAK-CCPNAI 154
Query: 127 VLVVANPANTNALILKEF---APSIPERNISCLTRLDHNRALGQISERLNVQVSDVKNVI 183
V +++NP N+ I E A + + + +T LD RA ++E L + +V +
Sbjct: 155 VNLISNPVNSTVPIAAEVFKKAGTYDPKKLLGVTTLDVARANTFVAEVLGLDPREVDVPV 214
Query: 184 IWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFIST-VQQRGAAII--KARK 240
+ G+ T P ++ V P+ P +E+ E+++ +Q G ++ KA
Sbjct: 215 VGGHAGVTILPLLSQ--VKPPSSFTP-QEI---------EYLTNRIQNGGTEVVEAKAGA 262
Query: 241 LSSALSAASAA 251
S+ LS A AA
Sbjct: 263 GSATLSMAYAA 273
>At3g21190 unknown protein
Length = 422
Score = 32.7 bits (73), Expect = 0.28
Identities = 27/99 (27%), Positives = 46/99 (46%), Gaps = 12/99 (12%)
Query: 234 AIIKARKLSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGLIYSFPV--T 291
AI+ A K S L + S+ +++ D+ + + FV SGL Y+ V
Sbjct: 332 AIMPASKRSKYLESVSSEYENVIDFYISSRSDVFVP----------AISGLFYANTVGKR 381
Query: 292 CANGEWKIVQGLSIDEFSRKKLDLTAEELSEEKNLAYSC 330
A G+ +++ I E S D + +S++ +LAYSC
Sbjct: 382 IALGKPQVLVPAEISETSGLATDFISPYISKKNHLAYSC 420
>At1g44750 Unknown protein (At1g44750)
Length = 379
Score = 31.2 bits (69), Expect = 0.82
Identities = 14/15 (93%), Positives = 15/15 (99%)
Query: 1 MAKDPVRVLVTGAAG 15
MAK+PVRVLVTGAAG
Sbjct: 1 MAKEPVRVLVTGAAG 15
>At3g14790 dTDP-glucose 4,6-dehydratase, putative
Length = 664
Score = 30.8 bits (68), Expect = 1.1
Identities = 24/73 (32%), Positives = 36/73 (48%), Gaps = 7/73 (9%)
Query: 5 PVRVLVTGAAGQIGYALVPMIARGVMLGPD-QPVILHMLDIAPAAESLNGVKMELVDAAF 63
P +L+TGAAG I + + R PD + V+L LD ++LN K F
Sbjct: 6 PKNILITGAAGFIASHVANRLVRSY---PDYKIVVLDKLDYCSNLKNLNPSKS---SPNF 59
Query: 64 PLLKGVVATTDVV 76
+KG +A+ D+V
Sbjct: 60 KFVKGDIASADLV 72
>At3g24330 beta-1,3-glucanase, putative
Length = 500
Score = 30.4 bits (67), Expect = 1.4
Identities = 24/79 (30%), Positives = 37/79 (46%), Gaps = 2/79 (2%)
Query: 54 VKMELVDAAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQ 113
VKM L+D +F LK A ++++A G +I VM+ G P + E S S +
Sbjct: 57 VKM-LMDNSFTKLKLFEADQNILDALIGSDIEVMI-GIPNRFLKEMAQDTSVAASWVEEN 114
Query: 114 ASALEKHAAANCKVLVVAN 132
+A + N K + V N
Sbjct: 115 VTAYSYNGGVNIKYIAVGN 133
>At1g55610 receptor kinase, putative
Length = 1166
Score = 30.4 bits (67), Expect = 1.4
Identities = 45/180 (25%), Positives = 73/180 (40%), Gaps = 38/180 (21%)
Query: 124 NCKVLVVANPANTNALILKEFAPSIPE-------RNISCLTRLDHNRALGQISERLNVQV 176
NCK L N + N A IP +N+ L+ L HNR G+I L++
Sbjct: 249 NCKFLETLNISRNN------LAGKIPNGEYWGSFQNLKQLS-LAHNRLSGEIPPELSLLC 301
Query: 177 SDVKNVIIWGNHSSTQYPDVNHATVNTPAGEKPVRELVSDDAWLNGEFISTVQQRGAAI- 235
+ + + GN S + P A V ++ L + +L+G+F++TV + I
Sbjct: 302 KTLVILDLSGNTFSGELPSQFTACV-------WLQNLNLGNNYLSGDFLNTVVSKITGIT 354
Query: 236 ---IKARKLSSALSAASAACDHIRDWVLGTPQGTFVSMGVYSDGSYNVPSGL--IYSFPV 290
+ +S ++ + C ++R VL F NVPSG + S PV
Sbjct: 355 YLYVAYNNISGSVPISLTNCSNLR--VLDLSSNGFTG---------NVPSGFCSLQSSPV 403
>At1g78570 Similar to dTDP-D-glucose 4,6-dehydratase
Length = 669
Score = 30.0 bits (66), Expect = 1.8
Identities = 24/73 (32%), Positives = 36/73 (48%), Gaps = 7/73 (9%)
Query: 5 PVRVLVTGAAGQIGYALVPMIARGVMLGPD-QPVILHMLDIAPAAESLNGVKMELVDAAF 63
P +L+TGAAG I + + R PD + V+L LD ++LN K F
Sbjct: 6 PKNILITGAAGFIASHVANRLIRSY---PDYKIVVLDKLDYCSNLKNLNPSKH---SPNF 59
Query: 64 PLLKGVVATTDVV 76
+KG +A+ D+V
Sbjct: 60 KFVKGDIASADLV 72
>At2g46700 calcium/calmodulin-dependent protein kinase CaMK4
Length = 595
Score = 29.6 bits (65), Expect = 2.4
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 8/55 (14%)
Query: 210 VRELVSDDAWLNGEFISTV-----QQRGAAIIKARKLSSALSAASAACDHIRDWV 259
+ +L + DAW E I+T + G +I +L+ L+ ++A H+RDWV
Sbjct: 515 IHQLEAVDAW---EEIATAGFQHFETEGNRVITIEELARELNVGASAYGHLRDWV 566
>At4g02990 unknown protein
Length = 541
Score = 29.3 bits (64), Expect = 3.1
Identities = 51/246 (20%), Positives = 101/246 (40%), Gaps = 29/246 (11%)
Query: 36 PVILH---MLDIAPAAESLNGVKMELVDAAFPLLKGVVATTDVVEACTGVNIAVMVG-GF 91
P +LH ++D+AP + L G+ ++ D L + +E ++A +VG G
Sbjct: 204 PQVLHSSVVIDLAPVVKYLQGLDIKPSDVPRVLERYPEVLGFKLEGTMSTSVAYLVGIGV 263
Query: 92 PRKEGMERKDVMSKNVSIYKSQASALEKHAAANCKVLVVANPANTNALILKEFAPSIPER 151
R+E ++++ I + + + K +VL + A A L E P I
Sbjct: 264 ARRE---IGGILTRYPEILGMRVARIIKPLVEYLEVLGIPRLA---AARLIEKRPHILGF 317
Query: 152 NISCLTRLDHNRALGQISERLNVQVSDVKNVIIWGNHSSTQYPDVNHATVNTPAGEKPVR 211
+ D + QI + NV+ + + ++I QYP++ + + R
Sbjct: 318 ELD-----DTVKPNVQILQDFNVRETSLPSII-------AQYPEIIGIDLKPKLDTQ--R 363
Query: 212 ELVSDDAWLNGEFISTVQQRGAAIIKAR-----KLSSALSAASAACDHIRDWVLGTPQGT 266
+L+ LN E + ++ +R + K L+ + D R+ V+G PQ
Sbjct: 364 KLLCSAIHLNPEDLGSLIERMPQFVSLSESPMLKHIDFLTKCGFSIDQTREMVIGCPQVL 423
Query: 267 FVSMGV 272
+++G+
Sbjct: 424 ALNLGI 429
>At3g59820 putative protein
Length = 755
Score = 29.3 bits (64), Expect = 3.1
Identities = 17/46 (36%), Positives = 24/46 (51%), Gaps = 1/46 (2%)
Query: 168 ISERLNVQVSDVKNVIIWGNHSSTQYPDVNHATVNTPAGE-KPVRE 212
IS+ LNV +++ GN S T + H+ +N P E KPV E
Sbjct: 13 ISDYLNVYARSIQSFQYIGNSSQTVHSHAYHSGINRPPVETKPVTE 58
>At4g13810 putative disease resistance protein
Length = 391
Score = 28.9 bits (63), Expect = 4.1
Identities = 50/217 (23%), Positives = 83/217 (38%), Gaps = 43/217 (19%)
Query: 53 GVKMELVDAAFPLLKGVVATTDVVEACTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKS 112
G+KMELV + F + K + + + +E G P G + K V +
Sbjct: 202 GLKMELVGSGFTIYKTIDVSGNRLE-----------GDIPESIG------LLKEVIVLSM 244
Query: 113 QASALEKHAAANCKVLVVANPANTNALILKE--FAPSIPERNISCLTRLD-----HNRAL 165
+A H + ++N +N +L L + + SIP + LT L+ HNR
Sbjct: 245 SNNAFTGHIPPS-----LSNLSNLQSLDLSQNRLSGSIP-GELGKLTFLEWMNFSHNRLE 298
Query: 166 GQISERLNVQVSDVKNVIIWGNHSSTQYPDVNHATVNTPAG--EKPVRELVSDDAWLNGE 223
G I E +Q D + S T+ P + A + G E+ ++ +D +
Sbjct: 299 GPIPETTQIQTQD--------SSSFTENPGLCGAPLLKKCGGEEEATKQEQDEDKEEEDQ 350
Query: 224 FISTVQQRGAAIIKARKLSSALSAASAACDHIRDWVL 260
S + AAI + L+ H RDW +
Sbjct: 351 VFSWI---AAAIGYVPGVVCGLTIGHILVSHKRDWFM 384
>At2g34350 nodulin-like protein
Length = 2301
Score = 28.9 bits (63), Expect = 4.1
Identities = 20/57 (35%), Positives = 30/57 (52%), Gaps = 6/57 (10%)
Query: 79 CTGVNIAVMVGGFPRKEGMERKDVMSKNVSIYKSQASAL--EKHAAANCKVLVVANP 133
C +NI + KE +E K+V + + S +A+A +HAAAN KVL + P
Sbjct: 1658 CASLNILIQ----QNKEVVEGKEVPTNDASPAMQRATARYDSQHAAANLKVLRLCAP 1710
>At2g33590 putative cinnamoyl-CoA reductase
Length = 321
Score = 28.9 bits (63), Expect = 4.1
Identities = 22/82 (26%), Positives = 37/82 (44%), Gaps = 14/82 (17%)
Query: 7 RVLVTGAAGQIGYALVPMI------ARGVMLGPDQPVILHMLDIAPAAESLNGVKMELVD 60
+V VTGA G +G +V ++ G + PD H+ + A + L K +L+D
Sbjct: 8 KVCVTGAGGFLGSWVVDLLLSKDYFVHGTVRDPDNEKYAHLKKLEKAGDKLKLFKADLLD 67
Query: 61 AAFPLLKGVVATTDVVEACTGV 82
+ L+ +A C+GV
Sbjct: 68 --YGSLQSAIA------GCSGV 81
>At2g06860 hypothetical protein
Length = 938
Score = 28.9 bits (63), Expect = 4.1
Identities = 18/46 (39%), Positives = 23/46 (49%), Gaps = 2/46 (4%)
Query: 163 RALGQISERLNVQVSD--VKNVIIWGNHSSTQYPDVNHATVNTPAG 206
R+L I + + V+D KN II G+HS VNH PAG
Sbjct: 288 RSLKLIRKTIISAVTDWFTKNTIIEGDHSKDSDSGVNHPKETAPAG 333
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.132 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,110,041
Number of Sequences: 26719
Number of extensions: 285730
Number of successful extensions: 708
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 694
Number of HSP's gapped (non-prelim): 32
length of query: 332
length of database: 11,318,596
effective HSP length: 100
effective length of query: 232
effective length of database: 8,646,696
effective search space: 2006033472
effective search space used: 2006033472
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC149493.2


