

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149491.1 + phase: 0 /pseudo
(1441 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
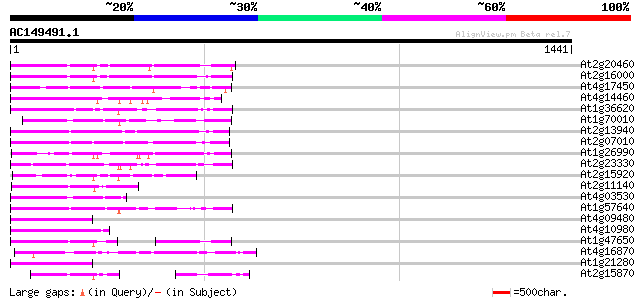
Score E
Sequences producing significant alignments: (bits) Value
At2g20460 putative retroelement pol polyprotein 259 8e-69
At2g16000 putative retroelement pol polyprotein 248 1e-65
At4g17450 retrotransposon like protein 219 9e-57
At4g14460 retrovirus-related like polyprotein 216 8e-56
At1g36620 hypothetical protein 214 4e-55
At1g70010 hypothetical protein 213 5e-55
At2g13940 putative retroelement pol polyprotein 208 2e-53
At2g07010 putative retroelement pol polyprotein 205 2e-52
At1g26990 polyprotein, putative 195 2e-49
At2g23330 putative retroelement pol polyprotein 193 7e-49
At2g15920 putative retroelement pol polyprotein 174 3e-43
At2g11140 putative retroelement pol polyprotein 171 4e-42
At4g03530 putative reverse transcriptase 168 2e-41
At1g57640 160 5e-39
At4g09480 unknown protein 156 9e-38
At4g10980 putative protein 149 1e-35
At1g47650 hypothetical protein 146 1e-34
At4g16870 retrotransposon like protein 139 2e-32
At1g21280 unknown protein (At1g21280) 127 4e-29
At2g15870 putative retroelement pol polyprotein 119 2e-26
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 259 bits (662), Expect = 8e-69
Identities = 183/610 (30%), Positives = 294/610 (48%), Gaps = 72/610 (11%)
Query: 1 SIYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRT 60
S +++H ++ P ++ L Y W+ +M+ +L KNK F+DGS+P P D N
Sbjct: 64 SPFFLHSADHPGLSIISHRLDETTYGDWSVAMRISLDAKNKLGFVDGSLPRPLESDPNFR 123
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
W RCN+++ SW++NSVSPQI ++I+ A D+W +L +RF+ + R +L I +L
Sbjct: 124 LWSRCNSMVKSWLLNSVSPQIYRSILRLNDATDIWRDLFDRFNLTNLPRTYNLTQEIQDL 183
Query: 121 KQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGL 180
+QG S+ +Y+T +K+LW++L+S + PC C + ++++FL GL
Sbjct: 184 RQGTMSLSEYYTLLKTLWDQLDSTEAL-----DDPCTCGKAVRLYQKAEKAKIMKFLAGL 238
Query: 181 NDSFSVVKTQVLLIDPLPSINKVYSMVIQEESN------IIPPTSLASNEDSSILVNASD 234
N+S+++V+ Q++ LPS+ +VY ++ Q+ S + PP + +E V+ S
Sbjct: 239 NESYAIVRRQIIAKKALPSLAEVYHILDQDNSQKGFFNVVAPPAAFQVSE-----VSHSP 293
Query: 235 ARKPFLRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPTA-SSNAVTSEH 293
P + SG ++ + C+FC R H E CY KH FP P SS+
Sbjct: 294 ITSPEIMYVQSGPNKGRPT---CSFCNRVGHIAERCYKKHGFPPGFTPKGKSSDKPPKPQ 350
Query: 294 AVDSHTSSEGTSSSSQT-----GLTQEQYVHLVSLLQQSSLVPSATPPNPASTNHVATS- 347
AV + + + Q + +Q +L++L S L P P AS+ H A+S
Sbjct: 351 AVAAQVTLSPDKMTGQLETLAGNFSPDQIQNLIALF-SSQLQPQIVSPQTASSQHEASSS 409
Query: 348 ---FPSSIDFTS------GINTIFSCSLHVPSDHWLIDSGANEHICSSLHLFHSYYRIKP 398
PS I F+ GI + SL SD W+IDSGA H+ LF +
Sbjct: 410 QSVAPSGILFSPSTYCFIGILAVSHNSL--SSDTWVIDSGATHHVSHDRKLFQTLDTSIV 467
Query: 399 ICVNLPNGSSVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQSLPYHVHFLLNT 458
VNLP G +V + GTV+ + + +VL+ P F++ LIS+S + L V F +
Sbjct: 468 SFVNLPTGPNVRISGVGTVLINKDIILQNVLFIPEFRLNLISISSLTTDLGTRVIFDPSC 527
Query: 459 YVIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNNVSS 518
IQD+ +G G LY L +PA +V++
Sbjct: 528 CQIQDLTKGLTLGEGKRIGNLYVLDTQSPAI-------------------------SVNA 562
Query: 519 IPSNAIWHFRLGHLSNQRLSMMHSLYSSITIDNK--AVCDICHFAKQRKLPY-------N 569
+ ++WH RLGH S RL + + + NK A C +CH AKQ+KL + N
Sbjct: 563 VVDVSVWHKRLGHPSFSRLDSLSEVLGTTRHKNKKSAYCHVCHLAKQKKLSFPSANNICN 622
Query: 570 LSTLLLHLNL 579
+ LLH+++
Sbjct: 623 STFELLHIDV 632
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 248 bits (634), Expect = 1e-65
Identities = 175/588 (29%), Positives = 282/588 (47%), Gaps = 58/588 (9%)
Query: 1 SIYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRT 60
S +++H ++ P ++ L NY W+ +M +L KNK FIDG++ P DLN
Sbjct: 60 SPFFLHSADHPGLNIISHRLDETNYGDWSVAMLISLDAKNKTGFIDGTLSRPLESDLNFR 119
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
W RCN+++ SW++NSVSPQI ++I+ A D+W +L RF+ + R +L I +
Sbjct: 120 LWSRCNSMVKSWLLNSVSPQIYRSILRMNDASDIWRDLNSRFNVTNLPRTYNLTQEIQDF 179
Query: 121 KQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGL 180
+QG S+ +Y+T +K+LW++L+S + PC C + + ++++FL GL
Sbjct: 180 RQGTLSLSEYYTRLKTLWDQLDSTEAL-----DEPCTCGKAMRLQQKAEQAKIVKFLAGL 234
Query: 181 NDSFSVVKTQVLLIDPLPSINKVYSMVIQEESN------IIPPTSLASNEDSSILVNASD 234
N+S+++V+ Q++ LPS+ +VY ++ Q+ S + PP + +E + S
Sbjct: 235 NESYAIVRRQIIAKKALPSLGEVYHILDQDNSQQSFSNVVAPPAAFQVSE-----ITQSP 289
Query: 235 ARKPFLRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPTASSNAVTSE-- 292
+ P + +G ++ + C+F R H E CY KH FP P + +
Sbjct: 290 SMDPTVCYVQNGPNKGR---PICSFYNRVGHIAERCYKKHGFPPGFTPKGKAGEKLQKPK 346
Query: 293 --HAVDSHTSSEGTSSSSQTG-LTQEQYVHLVSL----LQQSSLVPSATPPNPASTNHVA 345
A + +S TS S G L++EQ +++ LQ + AT S N
Sbjct: 347 PLAANVAESSEVNTSLESMVGNLSKEQLQQFIAMFSSQLQNTPPSTYATASTSQSDNLGI 406
Query: 346 TSFPSSIDFTSGINTIFSCSLHVPSDHWLIDSGANEHICSSLHLFHSYYRIKPICVNLPN 405
PS+ F GI T+ +L S W+IDSGA H+ LF S VNLP
Sbjct: 407 CFSPSTYSFI-GILTVARHTL--SSATWVIDSGATHHVSHDRSLFSSLDTSVLSAVNLPT 463
Query: 406 GSSVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVK 465
G +V + GT+ + + +VL+ P F++ LIS+S + + V F N+ IQD+
Sbjct: 464 GPTVKISGVGTLKLNDDILLKNVLFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLI 523
Query: 466 TQKMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIW 525
+M+G G LY L + S+S V+++ ++W
Sbjct: 524 KGRMLGQGRRVANLYLL---------------------DVGDQSIS----VNAVVDISMW 558
Query: 526 HFRLGHLSNQRLSMMHSLYSSITIDNKA--VCDICHFAKQRKLPYNLS 571
H RLGH S QRL + + NK C +CH AKQRKL + S
Sbjct: 559 HRRLGHASLQRLDAISDSLGTTRHKNKGSDFCHVCHLAKQRKLSFPTS 606
>At4g17450 retrotransposon like protein
Length = 1433
Score = 219 bits (558), Expect = 9e-57
Identities = 160/594 (26%), Positives = 279/594 (46%), Gaps = 82/594 (13%)
Query: 1 SIYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRT 60
S Y++H S+ P V+ +L G NY W+ +M+ +L KNK F+DGS+P P + D
Sbjct: 66 SPYFLHSSDHPGLNIVSHILDGTNYNNWSIAMRMSLDAKNKLSFVDGSLPRPDVSDRMFK 125
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
W RCN+++ +W++N V+ ++W +L RF + R L SI+ L
Sbjct: 126 IWSRCNSMVKTWLLNVVT--------------EMWNDLFSRFRVSNLPRKYQLEQSIHTL 171
Query: 121 KQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGL 180
KQG+ + Y+T K+LWE+L + R + + C CE ++ + ++IQFL GL
Sbjct: 172 KQGNLDLSTYYTKKKTLWEQLANTRVLTV----RKCNCEHVKELLEEAETSRIIQFLMGL 227
Query: 181 NDSFSVVKTQVLLIDPLPSINKVYSMVIQEESNIIPPTSLASNEDSSILVNASDARKPFL 240
ND+F+ ++ Q+L + P P + ++Y+M+ Q+ES + SN ++ V AS P +
Sbjct: 228 NDNFAHIRGQILNMKPRPGLTEIYNMLDQDESQRLVGNPTLSNPTAAFQVQAS----PII 283
Query: 241 RGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPT-----ASSNAVTSEHAV 295
+ + +Q C++C + H V+ CY KH +P +K T S+N +++
Sbjct: 284 DSQVN-MAQGSYKKPKCSYCNKLGHLVDKCYKKHGYPPGSKWTKGQTIGSTNLASTQLQP 342
Query: 296 DSHTSSEGTSSSSQTGLTQEQYVHLVSLLQQSSLVPSATPPNPASTNHVAT--SFPSSID 353
+ T +E T S + + +Q ++S L + SA+P S+ ++ S P
Sbjct: 343 VNETPNEKTDSYEE--FSTDQIQTMISYLSTKLHIASASPMPTTSSASISASPSVPMISQ 400
Query: 354 FTSGINTIFSCSLH------------VPSDHWLIDSGANEHICSSLHLFHSYYRIKPICV 401
+ ++FS + + V W+IDSGA H+ + L+ ++ ++ V
Sbjct: 401 ISGTFLSLFSNAYYDMLISSVSQEPAVSPRGWVIDSGATHHVTHNRDLYLNFRSLENTFV 460
Query: 402 NLPNGSSVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVI 461
LPN +V + G + S + +VLY P FK LIS
Sbjct: 461 RLPNDCTVKIAGIGFIQLSDAISLHNVLYIPEFKFNLIS--------------------- 499
Query: 462 QDVKTQKMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNNVSSIPS 521
++ + MIG G+ LY L + H +S + T M C S ++V +
Sbjct: 500 -ELTKELMIGRGSQVGNLYVL----DFNENNHTVS--LKGTTSM-CPEFSVCSSV--VVD 549
Query: 522 NAIWHFRLGHLSNQRLSMMHSLYS-SITIDNKA------VCDICHFAKQRKLPY 568
+ WH RLGH + ++ ++ + + + NK VC +CH +KQ+ L +
Sbjct: 550 SVTWHKRLGHPAYSKIDLLSDVLNLKVKKINKEHSPVCHVCHVCHLSKQKHLSF 603
>At4g14460 retrovirus-related like polyprotein
Length = 1489
Score = 216 bits (550), Expect = 8e-56
Identities = 164/633 (25%), Positives = 274/633 (42%), Gaps = 138/633 (21%)
Query: 3 YYVHPSEGPNSVTVTP-LLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRTA 61
YY+H ++ + V+ L T ++ +W RS+ AL +NK FI+G++ PP D + A
Sbjct: 34 YYLHSADHAGLILVSDRLTTASDFHSWRRSILMALNVRNKLGFINGTITKPPEDHRDFGA 93
Query: 62 WERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLK 121
W RCN+++ +W++NSV +I Q++++ +W L RF + D R+ + ++ ++
Sbjct: 94 WSRCNDIVSTWLMNSVDKKIGQSLLYIATVQGIWNNLLSRFKQDDAPRIFDIEQKLSKIE 153
Query: 122 QGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLN 181
QG + Y+T++ +LWEE ++ +P+CTC C C++ + +V +FL LN
Sbjct: 154 QGSMDISTYYTALLTLWEEHRNYVELPVCTCG-RCECDAAVKWEHLQQRSRVTKFLKELN 212
Query: 182 DSFSVVKTQVLLIDPLPSINKVYSMVIQEE--SNIIPPTSLAS---------NEDSSILV 230
+ F + +L++ P+P+I + ++MV Q+E N+ P T + S NED + V
Sbjct: 213 EGFDQTRRHILMLKPIPTIKEAFNMVTQDERQRNVKPLTRVDSVAFQNTSMINEDENAYV 272
Query: 231 NASDARKPFLRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANK--------- 281
A + +P N CT C + HT++ CY H +P K
Sbjct: 273 AAYNTVRP-------------NQKPICTHCGKVGHTIQKCYKVHGYPPGMKTGNTGYTYK 319
Query: 282 ------------------------PTASS----NAVTSEHAVDSHTSSEGTSSS------ 307
P +S N V +A SEG S +
Sbjct: 320 PNPQLHVQPRMPMMPQPRMQFPAQPYTNSMQKANVVAQVYAETGAYPSEGYSQAPMMNPY 379
Query: 308 ---------------SQTGLTQEQYVHLVSLLQQSSLVP----SATPPNP---------- 338
S T +Q ++S Q VP S++ P+P
Sbjct: 380 GSYPMPHITHGGNNLSLQDFTPQQIEQMISQFQAQVQVPEPAASSSNPSPLATVSEHGFM 439
Query: 339 --ASTNHVATSFPSSI------DFTSGINTIFSCSLHVPSDHWLIDSGANEHICSSLHLF 390
ST+ FPS+ D +T+ + +PSD W+IDSGA+ H+CS L +F
Sbjct: 440 ALTSTSGTIIPFPSTSLKYENNDLKFQNHTLSALQKFLPSDAWIIDSGASSHVCSDLAMF 499
Query: 391 HSYYRIKPICVNLPNGSSVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQSLPY 450
+ +GTV + + +VL+ P FK L+SVS + +++
Sbjct: 500 RE-----------------LKSVSGTVHITQKLILHNVLHVPDFKFNLMSVSSLVKTISC 542
Query: 451 HVHFLLNTYVIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSV 510
HF ++ +IQ++ MIG G L LY L SP S +P + SV
Sbjct: 543 SAHFYVDCCLIQELSQGLMIGRGRLYHNLYILET-ENTSP------STSTPAACLFTGSV 595
Query: 511 SSNNNVSSIPSNAIWHFRLGHLSNQRLSMMHSL 543
++ + +WH RLGH S+ L + L
Sbjct: 596 LNDGH--------LWHQRLGHPSSVVLQKLKRL 620
>At1g36620 hypothetical protein
Length = 1152
Score = 214 bits (544), Expect = 4e-55
Identities = 164/600 (27%), Positives = 277/600 (45%), Gaps = 88/600 (14%)
Query: 1 SIYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRT 60
S YY+HPS+ P+ V LL G NY W + + L K K FIDG++ P D +
Sbjct: 23 SPYYLHPSDHPHHVLTPMLLNGENYERWAKLTRNNLQAKQKLGFIDGTLTKPSSDSPDYP 82
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
W + N++++ W+ S+ PQ+ ++I + A +W L+ R+S + RV L+ I
Sbjct: 83 RWLQTNSMLVGWLYASLDPQVQKSISVVDNARVMWESLRTRYSVGNASRVHQLKYDIVAC 142
Query: 121 KQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGL 180
+Q ++ +YF +K +W++L+ + P+ C C P +R ++ R +++ QFL GL
Sbjct: 143 RQDGQTAANYFGKLKVMWDDLDDYEPLLTCCCNRPSCTHRVRQSQR-RDHERIHQFLMGL 201
Query: 181 NDS-FSVVKTQV---LLIDPLPSINKVYSMVIQEESNIIPPTSLASNEDSSILVNASDAR 236
+ + F +T + L D S++ +YS +I EE ++ T S E+ V +
Sbjct: 202 DAAKFGTSRTNILGRLSRDDNISLDSIYSEIIAEERHL---TITRSKEERVDAVGFA--- 255
Query: 237 KPFLRGKSSGTSQSK-NNSRYCTFCRRNNHTVEYCYLKHDFPN----------------- 278
G ++ S ++ NN CT C R+NH+ + C+ H P
Sbjct: 256 --VQTGVNAIASVTRVNNMGPCTHCGRSNHSADTCFKLHGVPEWYTEKYGDTSSGRGRGR 313
Query: 279 ----ANKPTASSNAVTSEHAVDSHTSSEGTSSSSQTGLTQEQYVHLVSLLQQSSLVPSAT 334
+ N+ + +A SH SS + S G+++E + + +LL+Q +
Sbjct: 314 SSTPRGRGRGHGNSYKANNAQTSHPSSSASEFSDIPGVSKEAWSAIRNLLKQDT------ 367
Query: 335 PPNPASTNHVATSFPSSIDFTSGINTIFSCSLHVPSDHWLIDSGANEHICSSLHLFHSYY 394
A+++ + + +DF LIDSGA+ H+ L L Y
Sbjct: 368 ----ATSSEKLSGKTNCVDF-------------------LIDSGASHHMTGFLDLLTEIY 404
Query: 395 RIKPICVNLPNGSSVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQSLPYHVHF 454
I V LPN I GT++ + +THVL+ P LISV+++ + L F
Sbjct: 405 EIPHSVVVLPNAKHTIATKKGTLILGANMKLTHVLFVPDLSCTLISVARLLRELHCFAIF 464
Query: 455 LLNTYVIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNN 514
VIQD ++ +IG+G +G+Y L +A +++ S N V
Sbjct: 465 TDKVCVIQDRTSKMLIGVGTESNGVYHLQ-------RAEVVAT--------SANVVKWKT 509
Query: 515 NVSSIPSNAIWHFRLGHLSNQRL-SMMHSL--YSSITIDNKAVCDICHFAKQRKLPYNLS 571
N A+WH RLGH S++ L S++ SL + S + D K +CD+C AKQ + ++ S
Sbjct: 510 N------KALWHMRLGHPSSKVLSSVLPSLEDFDSCSSDLKTICDVCVRAKQTRASFSES 563
>At1g70010 hypothetical protein
Length = 1315
Score = 213 bits (543), Expect = 5e-55
Identities = 159/549 (28%), Positives = 245/549 (43%), Gaps = 109/549 (19%)
Query: 32 MKRALGTKNKFVFIDGSVPIPPMDDLNRTAWERCNNLILSWIINSVSPQIAQTIVFHEYA 91
M ++ KNK F+DGS+P P DD W RCN+++ SW++NSVS +I +I++ A
Sbjct: 1 MTTSIEAKNKLGFVDGSIPKPDDDDPYCKIWRRCNSMVKSWLLNSVSKEIYTSILYFPTA 60
Query: 92 IDVWIELQERFSKVDRIRVASLRSSINNLKQGDKSVLDYFTSIKSLWEELNSHRPMPMCT 151
+W +L RF K R+ LR I++L+QG+ + Y T ++LWEEL S + +P
Sbjct: 61 AAIWKDLYTRFHKSSLPRLYKLRQQIHSLRQGNLDLSSYHTRTQTLWEELTSLQAVP--- 117
Query: 152 CPYPCRCESMRAARDFRME---DQVIQFLTGLNDSFSVVKTQVLLIDPLPSINKVYSMVI 208
R D +E ++VI FL GLND + V++Q+L+ LPS+++V++M+
Sbjct: 118 ----------RTVEDLLIERETNRVIDFLMGLNDCYDTVRSQILMKKTLPSLSEVFNMID 167
Query: 209 QEESNIIPPTSLASNEDSSILVNASDARKPFLRGKSSGTSQSKNNSRYCTFCRRNNHTVE 268
Q+E+ S SS+ ++ + + L +G + K C++C R H +
Sbjct: 168 QDETQRSARISTTPGMTSSVFPVSNQSSQSAL----NGDTYQKKERPVCSYCSRPGHVED 223
Query: 269 YCYLKHDFPNA-------NKPTASSNAVTSEHAVDSHTSSEGTSSSSQTGLTQEQYVHLV 321
CY KH +P + KP+ S+NA V ++T S S LT Q LV
Sbjct: 224 TCYKKHGYPTSFKSKQKFVKPSISANAAIGSEEVVNNT------SVSTGDLTTSQIQQLV 277
Query: 322 SLLQQSSLVPSATPPNPASTNHVATSFPSSIDFTSGINTIFSCSLHVPSDHWLIDSGANE 381
S L S L P +TP P + +S PSS S
Sbjct: 278 SFL-SSKLQPPSTPVQPEVHSISVSSDPSS-------------------------SSTVC 311
Query: 382 HICSSLHLFHSYYRIKPICVNLPNGSSVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISV 441
I S+HL H + VL+ P FK L+SV
Sbjct: 312 PISGSVHL------------------------------GRHLILNDVLFIPQFKFNLLSV 341
Query: 442 SKICQSLPYHVHFLLNTYVIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSP 501
S + +S+ + F + V+QD + M+G+G LY +
Sbjct: 342 SSLTKSMGCRIWFDETSCVLQDATRELMVGMGKQVANLY------------------IVD 383
Query: 502 TNKMSCNSVSSNNNVSSIPSNAIWHFRLGHLSNQRLSMMHSLYSSITIDNKA--VCDICH 559
+ +S S+ V+S+ S+ +WH RLGH S Q+L M SL S N C +CH
Sbjct: 384 LDSLSHPGTDSSITVASVTSHDLWHKRLGHPSVQKLQPMSSLLSFPKQKNNTDFHCRVCH 443
Query: 560 FAKQRKLPY 568
+KQ+ LP+
Sbjct: 444 ISKQKHLPF 452
>At2g13940 putative retroelement pol polyprotein
Length = 1501
Score = 208 bits (530), Expect = 2e-53
Identities = 154/576 (26%), Positives = 258/576 (44%), Gaps = 46/576 (7%)
Query: 1 SIYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRT 60
S Y + S+ P +V + L G NY W M AL K K FI+G++P PP +D N
Sbjct: 30 SPYTLASSDNPGAVISSVELNGDNYNQWATEMLNALQAKRKTGFINGTIPRPPPNDPNYE 89
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
W N++I+ WI S+ P++ T+ F A +W +L++RFS +++R+ +R+ +++
Sbjct: 90 NWTAVNSMIVGWIRTSIEPKVKATVTFISDAHLLWKDLKQRFSVGNKVRIHQIRAQLSSC 149
Query: 121 KQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGL 180
+Q ++V++Y+ + +LWEE N ++P+ +CTC CRC + R E+++ QF+ GL
Sbjct: 150 RQDGQAVIEYYGRLSNLWEEYNIYKPVTVCTCGL-CRCGATSEPTKEREEEKIHQFVLGL 208
Query: 181 NDS-FSVVKTQVLLIDPLPSINKVYSMVIQEESNIIPPTSLASNEDSSILVNASD----- 234
++S F + ++ +DPLPS+ ++YS VI+EE + E++ + +
Sbjct: 209 DESRFGGLCATLINMDPLPSLGEIYSRVIREEQRLASVHVREQKEEAVGFLARREQLDHH 268
Query: 235 ARKPFLRGKSSGTSQSKNNSRY-----CTFCRRNNHTVEYCYLKHDFPNANKPTASSNAV 289
+R +S T S++NS C+ C R H + C+ FP+
Sbjct: 269 SRVDASSSRSEHTGGSRSNSIIKGRVTCSNCGRTGHEKKECWQIVGFPDWWSERNGGRG- 327
Query: 290 TSEHAVDSHTSSEGTSSSSQTGLTQEQYVHLVSLLQQSSLVPSATPPNPASTNHVATSFP 349
+ G S+ G Q H S SS+ P T + + +
Sbjct: 328 ------SNGRGRGGRGSNGGRGQGQVMAAHATS--SNSSVFPEFTEEHMRVLSQLVKE-- 377
Query: 350 SSIDFTSGINTIFSCSLHVPSDHWLIDSGANEHICSSLHLFHSYYRIKPICVNLPNGSSV 409
S ++ N S ++DSGA+ H+ +L + + P V +GS
Sbjct: 378 KSNSGSTSNNNSDRLSGKTKLGDIILDSGASHHMTGTLSSLTNVVPVPPCPVGFADGSKA 437
Query: 410 IVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKM 469
G + S +T+VL+ PS LISVSK+ + F +QD ++ +
Sbjct: 438 FALSVGVLTLSNTVSLTNVLFVPSLNCTLISVSKLLKQTQCLATFTDTLCFLQDRSSKTL 497
Query: 470 IGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRL 529
IG G G+Y L PA K+ +V S+ A+WH RL
Sbjct: 498 IGSGEERGGVYYLTDVTPA---------------KIHTANVDSD--------QALWHQRL 534
Query: 530 GHLSNQRLSMMHSLYSSITIDNKAVCDICHFAKQRK 565
GH S LS + + + CD+C AKQ +
Sbjct: 535 GHPSFSVLSSLPLFSKTSSTVTSHSCDVCFRAKQTR 570
>At2g07010 putative retroelement pol polyprotein
Length = 1413
Score = 205 bits (521), Expect = 2e-52
Identities = 148/567 (26%), Positives = 254/567 (44%), Gaps = 40/567 (7%)
Query: 1 SIYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRT 60
S Y + S+ P ++ + +LTG NY W+ M AL K K FI+GS+ PP+D+ +
Sbjct: 25 SPYTLASSDNPGAMISSVMLTGDNYNEWSTKMLNALQAKRKTGFINGSISKPPLDNPDYE 84
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
W+ N++I+ WI S+ P++ T+ F A +W EL++RFS +++ V +++ +
Sbjct: 85 NWQAVNSMIVGWIRASIEPKVKSTVTFICDAHQLWSELKQRFSVGNKVHVHQIKTQLAAC 144
Query: 121 KQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGL 180
+Q + V+DY+ + LWEE ++P+ +C C C C + R E+++ QF+ GL
Sbjct: 145 RQDGQPVIDYYGRLCKLWEEFQIYKPITVCKCGL-CTCGATLEPSKEREEEKIHQFVLGL 203
Query: 181 NDS-FSVVKTQVLLIDPLPSINKVYSMVIQEESNIIPPTSLASNEDSSILVNASDARKPF 239
+DS F + ++ +DP PS+ ++YS V++EE + + + S+I +
Sbjct: 204 DDSRFGGLSATLIAMDPFPSLGEIYSRVVREEQR-LASVQIREQQQSAIGFLTRQSEVTA 262
Query: 240 LRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPTASSNAVTSEHAVDSHT 299
S +S++ S C+ C R+ H + C+ FP+ + S S
Sbjct: 263 DGRTDSSIIKSRDRSVLCSHCGRSGHEKKDCWQIVGFPDWWTERTNGGGRGS----SSRG 318
Query: 300 SSEGTSSSSQTGLTQEQYVHLVSLLQQSSLVPSATPPN-PASTNHVATSFPSSIDFTSGI 358
+S S+ +G + Q + S P TP T + + D SG
Sbjct: 319 RGGRSSGSNNSGRGRGQVTAAHATTSNLSPFPEFTPDQLRVITQMIQNKNNGTSDKLSG- 377
Query: 359 NTIFSCSLHVPSDHWLIDSGANEHICSSLHLFHSYYRIKPICVNLPNGSSVIVQYAGTVV 418
+ ++D+GA+ H+ L L + I V +G GT
Sbjct: 378 --------KMKLGDVILDTGASHHMTGQLSLLTNIVTIPSCSVGFADGRKTFAISMGTFK 429
Query: 419 FSPHFHITHVLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKMIGLGNLCDG 478
S +++VLY P+ LISVSK+ + + F V+QD ++ +IG G DG
Sbjct: 430 LSETVSLSNVLYVPALNCSLISVSKLVKQIKCLALFTDTICVLQDRFSRTLIGTGEERDG 489
Query: 479 LYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRLGHLSNQRLS 538
+Y +++ A + T + V +A+WH RLGH S LS
Sbjct: 490 VY-------------YLTDAATTT----------VHKVDITTDHALWHQRLGHPSFSVLS 526
Query: 539 MMHSLYSSITIDNKAVCDICHFAKQRK 565
+ S + CD+C AKQ +
Sbjct: 527 SLPLFSGSSCSVSSRSCDVCFRAKQTR 553
>At1g26990 polyprotein, putative
Length = 1436
Score = 195 bits (495), Expect = 2e-49
Identities = 161/621 (25%), Positives = 271/621 (42%), Gaps = 133/621 (21%)
Query: 5 VHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRTAWER 64
+H S+ P V +L G NY +W+ +M+ +L KNK F+DGS+ P +DD W R
Sbjct: 60 LHSSDHPGLSIVAHILDGSNYNSWSIAMRISLDAKNKLGFVDGSLLRPSVDDSTFRIWSR 119
Query: 65 CNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLKQGD 124
CN+++ + L R+ L ++ L+QG
Sbjct: 120 CNSMVNN--------------------------LPRRYQ---------LEQAVMTLQQGK 144
Query: 125 KSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLNDSF 184
+ YFT K+LWE+L + + + C C+ ++ + +VIQFL GL+D F
Sbjct: 145 LDLSTYFTKKKTLWEQLANTKSRSVKKCD----CDQVKELLEEAETSRVIQFLMGLSDDF 200
Query: 185 SVVKTQVLLIDPLPSINKVYSMVIQEESN--------IIPPTSLAS-------NEDSSIL 229
+ +++Q+ + P P +N++Y+M+ Q+ES +P S A+ N+ ++IL
Sbjct: 201 NTIRSQIFNMKPRPGLNEIYNMLDQDESQRLVGFAAKSVPSPSPAAFQTQGVLNDQNTIL 260
Query: 230 VNASDARKPFLRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPTASSNAV 289
+ + +KP CT C R HTV+ CY H +P + + V
Sbjct: 261 LAQGNFKKP-----------------KCTHCNRIGHTVDKCYKVHGYPPGHPRAKENTYV 303
Query: 290 TSEH-AVDSHTSSEGTSSSSQTG---LTQEQYVHLVSLLQ------------------QS 327
S + A ++ + S TG ++ + L+S L S
Sbjct: 304 GSTNLASTDQIETQAPPTMSATGHETMSNDHIQQLISYLSTKLQSPSITSCFDKAIASSS 363
Query: 328 SLVPS---------ATPPNPA-STNHVATSFPSSID------FTSGINTIFSCSLHVPSD 371
+ VPS A+ NP S + + +F S D TS I SL
Sbjct: 364 NPVPSISQITDKAIASSSNPVPSISQITGTFFSLYDSTYYEMLTSSIPIETELSLRA--- 420
Query: 372 HWLIDSGANEHICSSLHLFHSYYRIKPICVNLPNGSSVIVQYAGTVVFSPHFHITHVLYS 431
W+IDSGA+ H+ +L+H+Y + V LPNG +V ++ G + + + +VL+
Sbjct: 421 -WVIDSGASHHVTHERNLYHTYKALDRTFVRLPNGHTVKIEGTGFIQLTDALSLHNVLFI 479
Query: 432 PSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQ 491
P FK L+SVS + ++L V F + +IQ + + M+G G+ LY L+
Sbjct: 480 PEFKFNLLSVSVLTKTLQSKVSFTSDECMIQALTKELMLGKGSQVGNLYILNLDKSLVDV 539
Query: 492 AHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRLGHLSNQRLSMMHSLY----SSI 547
+ F +V C+SV + + +WH RLGH S ++ + + I
Sbjct: 540 SSFPGKSV-------CSSVKNESE--------MWHKRLGHPSFAKIDTLSDVLMLPKQKI 584
Query: 548 TIDNKAVCDICHFAKQRKLPY 568
D+ + C +CH +KQ+ LP+
Sbjct: 585 NKDS-SHCHVCHLSKQKHLPF 604
>At2g23330 putative retroelement pol polyprotein
Length = 1496
Score = 193 bits (490), Expect = 7e-49
Identities = 155/593 (26%), Positives = 254/593 (42%), Gaps = 89/593 (15%)
Query: 3 YYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRTAW 62
Y ++ S+ P ++ + +L NY W+ ++ L K K FIDGS+P P D + W
Sbjct: 22 YLINASDNPGALISSVVLKENNYAEWSEELQNFLRAKQKLGFIDGSIPKPAADP-ELSLW 80
Query: 63 ERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLKQ 122
N++I+ WI S+ P I T+ F A +W L+ RFS + +R L+ I Q
Sbjct: 81 IAINSMIVGWIRTSIDPTIRSTVGFVSEASQLWENLRRRFSVGNGVRKTLLKDEIAACTQ 140
Query: 123 GDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLND 182
+ VL Y+ + LWEEL +++ C+CE+ R +D+V +FL GL+
Sbjct: 141 DGQPVLAYYGRLIKLWEELQNYK------SGRECKCEAASDIEKEREDDRVHKFLLGLDS 194
Query: 183 SFSVVKTQVLLIDPLPSINKVYSMVIQEESNIIPPTSLASNEDSSILVNASDARKPFLRG 242
FS +++ + I+PLP + +VYS V++EE N+ + + +I + + P R
Sbjct: 195 RFSSIRSSITDIEPLPDLYQVYSRVVREEQNLNASRTKDVVKTEAIGFSVQSSTTPRFRD 254
Query: 243 KSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPN--------ANKPT---------AS 285
KS + +CT C R H V C+L H +P+ N+P+ S
Sbjct: 255 KS---------TLFCTHCNRKGHEVTQCFLVHGYPDWWLEQNPQENQPSTRGRGSNGRGS 305
Query: 286 SNAVTSEHAVDSHTSSEGTSSSSQ------TGLTQEQYVHLVSLLQQSSLVPSATPPNPA 339
S+ + T G ++++Q +G +Q L+SLLQ P+
Sbjct: 306 SSGRGGNRSSAPTTRGRGRANNAQAAAPTVSGDGNDQIAQLISLLQAQ---------RPS 356
Query: 340 STNHVATSFPSSIDFTSGINTIFSCSLHVPSDHWLIDSGANEHICSSLHLFHSYYRIKPI 399
S++ + T G+ ID+GA+ H+ + + I P
Sbjct: 357 SSSE---RLSGNTCLTDGV----------------IDTGASHHMTGDCSILVDVFDITPS 397
Query: 400 CVNLPNGSSVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQSLPYHVHFLLNTY 459
V P+G + GT++ + + VL+ P F LISVSK+ + F
Sbjct: 398 PVTKPDGKASQATKCGTLLLHDSYKLHDVLFVPDFDCTLISVSKLLKQTSSIAIFTDTFC 457
Query: 460 VIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNNVSSI 519
+QD + +IG G +G+Y + +P+ H SS +
Sbjct: 458 FLQDRFLRTLIGAGEEREGVY--YFTGVLAPRVHKASSDFA------------------- 496
Query: 520 PSNAIWHFRLGHLSNQ-RLSMMHSLYSSITIDNKAVCDICHFAKQRKLPYNLS 571
S +WH RLGH S LS+ SS D CD C +KQ + + +S
Sbjct: 497 ISGDLWHRRLGHPSTSVLLSLPECNRSSQGFDKIDSCDTCFRSKQTREVFPIS 549
>At2g15920 putative retroelement pol polyprotein
Length = 1100
Score = 174 bits (442), Expect = 3e-43
Identities = 136/495 (27%), Positives = 232/495 (46%), Gaps = 66/495 (13%)
Query: 8 SEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRTAWERCNN 67
++ P ++ L NY WN +M +L KNK FIDG++P P D N W RCN+
Sbjct: 23 ADHPGLNIISHRLDETNYGDWNVAMLISLDVKNKSGFIDGTLPRPLETDKNFCLWSRCNS 82
Query: 68 LILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLKQGDKSV 127
+I ++I+ A D+W +L RF+ + R +L I +L+QG S+
Sbjct: 83 MI------------CRSILRMNDASDIWRDLNSRFNMTNLPRTYNLTQEIQDLRQGTLSL 130
Query: 128 LDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLNDSFSVV 187
+Y+T +K+LW++L+S + PC + ++++FL GLN+S++++
Sbjct: 131 SEYYTRLKTLWDQLDSTEEL-----DDPCTRQK---------RAKIVKFLAGLNESYAII 176
Query: 188 KTQVLLIDPLPSINKVYSMVIQEESN------IIPPTSLASNEDSSILVNASDARKPFLR 241
+ Q++ LPS+ +VY +V Q+ S + P + +E V +++ P +
Sbjct: 177 RRQIIAKKILPSLAEVYHIVDQDNSQQGFSNVVARPAAFQVSE-----VTITNSIDPTIY 231
Query: 242 GKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFP-------------NANKPTASSNA 288
+G ++ ++ C+FC R H E CY KH FP KP AS+ A
Sbjct: 232 YVQNGPNKGRS---MCSFCNRVGHIAERCYKKHGFPPRFTPKGKVGDKTQKPKPVASNVA 288
Query: 289 VTSEHAVDSHT---SSEGTSSSSQTGLTQEQYVHLVSLLQQSSLVPSATPPNPASTNHVA 345
+ + + D+H+ S G S Q +Q++ + S Q ++ + + +++
Sbjct: 289 LATTESNDTHSGLKSLVGNLSKEQL----QQFIAMFSSQLQPQPHSNSAVASSSQADNIG 344
Query: 346 TSFPSSIDFTSGINTIFSCSLHVPSDHWLIDSGANEHICSSLHLFHSYYRIKPICVNLPN 405
SF S GI T+ +L S W+IDSGA H+ L L S VNLP
Sbjct: 345 ISFSPSTYSFIGILTVAQHTL--SSKTWVIDSGAIHHVNLLLTLNTSVLS----SVNLPA 398
Query: 406 GSSVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVK 465
G +V + GT+ + + +VL+ P F + L+S+S + + V F + IQD+
Sbjct: 399 GPTVKISGVGTLRLNDDNLLKNVLFIPEFCLNLMSISSLTDDIGSRVIFNQHACEIQDLI 458
Query: 466 TQKMIGLGNLCDGLY 480
+M+G G LY
Sbjct: 459 KGRMLGHGRRVANLY 473
Score = 33.1 bits (74), Expect = 1.2
Identities = 21/60 (35%), Positives = 33/60 (55%)
Query: 717 IKCLPLFLNKNLLINFFMILYLT*IPSKFLVVYVLLLLFLLKGPNFNLEQESLFSLQVRI 776
I+ L F LL+ F M+L L+ S+ LV YV +L K +F+L+Q+ + S R+
Sbjct: 490 IRHLQSFFQIRLLMRFSMVLLLSMDSSEHLVAYVTVLHHRSKDISFSLDQKLVSSWDTRL 549
>At2g11140 putative retroelement pol polyprotein
Length = 411
Score = 171 bits (432), Expect = 4e-42
Identities = 100/336 (29%), Positives = 172/336 (50%), Gaps = 25/336 (7%)
Query: 5 VHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRTAWER 64
+H S+ P V +L G NY +W+ +M+ +L KNK F+DGS+ P +DD W R
Sbjct: 69 LHSSDHPGLSIVAHVLDGSNYNSWSIAMRISLDAKNKLGFVDGSLLRPSVDDSTFRIWSR 128
Query: 65 CNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLKQGD 124
CN+++ SWI+N V+ +I +I+++E A+++W +L RF + R L ++ LKQG
Sbjct: 129 CNSMVKSWILNVVNKEIYDSILYYEDAVEMWTDLFTRFRVNNLPRKYQLEQAVMTLKQGS 188
Query: 125 KSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLNDSF 184
++ YFT K+LWE+L + + + C C+ ++ + +VIQFL GLND F
Sbjct: 189 LNLSTYFTKKKTLWEQLLNTKTRSV----KKCDCDQVKELLEDAETSRVIQFLMGLNDDF 244
Query: 185 SVVKTQVLLIDPLPSINKVYSMVIQEESNII------PPTSLASNEDSSILVNASDARKP 238
+ + +Q+L + P P +N++Y+M+ Q+ES + P S A+ + +L + P
Sbjct: 245 NTIMSQILNMKPRPGLNEIYNMLDQDESQRLVGHASKPTPSPAAFQTQGLLTE----QNP 300
Query: 239 FLRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPTASSNAVTSEHAVDSH 298
L +Q CT C R HTV+ CY H +P + ++ + S
Sbjct: 301 IL------MAQGNFKKPKCTHCNRIGHTVDKCYKVHGYPPGHPRANQQSSCVGSTNLTSI 354
Query: 299 TSSEGTSSSSQTGLTQEQYV-----HLVSLLQQSSL 329
SE + Q + + ++ +L + LQ +S+
Sbjct: 355 DQSENQAPVMQDEIMSKDHIQQLISYLSTKLQSASI 390
Score = 32.0 bits (71), Expect = 2.7
Identities = 23/101 (22%), Positives = 45/101 (43%), Gaps = 11/101 (10%)
Query: 472 LGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRLGH 531
+G+ D Y++H + P P+A+ SS V TN S+ + N + + + I
Sbjct: 319 IGHTVDKCYKVHGYPPGHPRANQQSSCVGSTN---LTSIDQSENQAPVMQDEI------- 368
Query: 532 LSNQRLSMMHSLYSSITIDNKAVCDICHFAKQRKLPYNLST 572
+S + + S Y S + + ++ A Q+ + N+ T
Sbjct: 369 MSKDHIQQLIS-YLSTKLQSASITSCPDKATQKTVQLNIKT 408
>At4g03530 putative reverse transcriptase
Length = 374
Score = 168 bits (425), Expect = 2e-41
Identities = 99/302 (32%), Positives = 156/302 (50%), Gaps = 22/302 (7%)
Query: 1 SIYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRT 60
S Y++ S+ P V LL G +Y W +M ++ KNK F+DGS+P P DD
Sbjct: 68 SPYHLVSSDHPGLVLAPELLDGNSYGTWIIAMTTSIEAKNKLGFVDGSIPKPDDDDPYCK 127
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
W RCN+++ SW++NSVS +I +I++ A +W +L RF K R+ LR I++L
Sbjct: 128 IWRRCNSMVKSWLLNSVSKEIYTSILYFPTAAAIWKDLYTRFHKSSLPRLYKLRQQIHSL 187
Query: 121 KQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRME---DQVIQFL 177
+QG+ + Y T ++LWEEL S + +P R D +E ++VI FL
Sbjct: 188 RQGNLDLSSYHTRKQTLWEELTSLQAIP-------------RTVEDLLIERETNRVIDFL 234
Query: 178 TGLNDSFSVVKTQVLLIDPLPSINKVYSMVIQEESNIIPPTSLASNEDSSILVNASDARK 237
GLND + V++Q+L+ LPS+++V++M+ Q+E S SS+ ++ + +
Sbjct: 235 MGLNDCYDAVRSQILMKKTLPSLSEVFNMIDQDEIQRSARISTTPGMTSSVFAVSNQSSQ 294
Query: 238 PFLRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPTASSNAVTSEHAVDS 297
L +G + K CT+C R H + CY KH +P + K +N T VD
Sbjct: 295 SVL----NGDTYQKKERPVCTYCSRPGHVEDTCYKKHGYPTSFKSKQIAN--TELQIVDP 348
Query: 298 HT 299
T
Sbjct: 349 FT 350
>At1g57640
Length = 1444
Score = 160 bits (405), Expect = 5e-39
Identities = 141/605 (23%), Positives = 254/605 (41%), Gaps = 118/605 (19%)
Query: 1 SIYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRT 60
S Y + ++ +V P+L NY W K AL ++ KF F+DG++P P +
Sbjct: 19 SPYDLTAADNSGAVISHPILKTNNYEEWACGFKTALRSRKKFGFLDGTIPQPLDGSPDLE 78
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
W N L++SW+ ++ ++ I + A D+W ++++RFS + + +++ +
Sbjct: 79 DWLTINALLVSWMKMTIDSELLTNISHRDVARDLWEQIRKRFSVSNGPKNQKMKADLATC 138
Query: 121 KQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGL 180
KQ +V Y+ + +W+ +NS+RP+ +C C C C +R +D V Q+L GL
Sbjct: 139 KQEGMTVEGYYGKLNKIWDNINSYRPLRICKCGR-CICNLGTDQEKYREDDMVHQYLYGL 197
Query: 181 NDS-FSVVKTQVLLIDPLPSINKVYSMVIQEESNIIPPTSLASNEDSSILVNASDARKPF 239
N++ F +++ + PLP + +VY++V QEE + +S D + R
Sbjct: 198 NETKFHTIRSSLTSRVPLPGLEEVYNIVRQEEDMVNNRSSNEERTDVTAFAVQMRPRSEV 257
Query: 240 LRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFP-----------NAN-------- 280
+ K + S+ N + CT C R H+ E C++ +P N+N
Sbjct: 258 ISEKFAN-SEKLQNKKLCTHCNRGGHSPENCFVLIGYPEWWGDRPRGKSNSNGSTSRGRG 316
Query: 281 -----------KPTASSNAVTSEHAVDSHTSSEGTSSSSQ--TGLTQEQYVHLVSLLQQS 327
+PT + +T H + T S +GLT EQ+ +V LL
Sbjct: 317 RFGPGFNGGQPRPTYVNVVMTGPFPSSEHVNRVITDSDRDAVSGLTDEQWRGVVKLL--- 373
Query: 328 SLVPSATPPNPASTNHVATSFPSSIDFTSGINTIFSCSLHVPSDHWLIDSGANEHICSSL 387
+A + S H S S+ FTS W++D+GA+ H+ +L
Sbjct: 374 ----NAGRSDNKSNAHETQSGTCSL-FTS----------------WILDTGASHHMTGNL 412
Query: 388 HLFHSYYRIKPICVNLPNGSSVIVQYAGTVVFSPHFHITHVLYSPSFKVYLISVSKICQS 447
L + P+ + L +G+ + GTV H + V Y
Sbjct: 413 ELLSDMRSMSPVLIILADGNKRVAVSEGTVRLGSHLILKSVFY----------------- 455
Query: 448 LPYHVHFLLNTYVIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQAHFISSAVSPTNKMSC 507
++++++ +I +G + D + H +++A T+ +
Sbjct: 456 -------------VKELESD-LISVGQMMD-------------ENHCVNAAAVHTSVKA- 487
Query: 508 NSVSSNNNVSSIPSNAIWHFRLGHLSNQRLSMM-HSLYSSITIDNKAVCDICHFAKQRKL 566
P + +WH RLGH S++ ++++ L SS + VCD C AKQ +
Sbjct: 488 ------------PFD-LWHRRLGHASDKIVNLLPRELLSSGKEILENVCDTCMRAKQTRD 534
Query: 567 PYNLS 571
+ LS
Sbjct: 535 TFPLS 539
>At4g09480 unknown protein
Length = 301
Score = 156 bits (394), Expect = 9e-38
Identities = 70/210 (33%), Positives = 124/210 (58%), Gaps = 2/210 (0%)
Query: 3 YYVHPSEGPNSVTVTPL-LTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRTA 61
Y++H S+ + V+ +G + +W RS++ AL +NK FIDG++P PP + +
Sbjct: 26 YFLHSSDQAGLILVSDRPSSGSEFHSWRRSVRMALNVRNKLGFIDGTIPQPPSTHRDAGS 85
Query: 62 WERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLK 121
W RCN+++ +W++NSVS +I Q+++F A +W L RF + D RV + + +L+
Sbjct: 86 WSRCNDMVATWLMNSVSKKIGQSLLFMSTAESIWKNLLSRFKQDDAPRVFEIEQRLGSLQ 145
Query: 122 QGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLN 181
QG V Y+T + +LWEE ++ +P+CTC C C + + +V +FL GLN
Sbjct: 146 QGSMDVSTYYTELVTLWEEYKNYIELPLCTCG-RCECNASALWEKMQQRSRVTKFLMGLN 204
Query: 182 DSFSVVKTQVLLIDPLPSINKVYSMVIQEE 211
+++ + +L++ P+P+I V++MV Q+E
Sbjct: 205 EAYEATQRHILMLKPIPTIEDVFNMVAQDE 234
>At4g10980 putative protein
Length = 294
Score = 149 bits (376), Expect = 1e-35
Identities = 77/256 (30%), Positives = 142/256 (55%), Gaps = 7/256 (2%)
Query: 3 YYVHPSEGPNSVTVTPLL-TGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRTA 61
+++H S+ V V+ L T ++ +W RS+ AL +NK FIDG++ PP+D + A
Sbjct: 35 HHLHTSDHAGLVLVSERLNTASDFHSWRRSIWMALNVRNKLGFIDGTIVKPPLDHRDYGA 94
Query: 62 WERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLK 121
W RCN+ + +W++NSVS +I Q+++F A +W + RF + D RV + ++ ++
Sbjct: 95 WSRCNDTVSTWLMNSVSKKIGQSLLFIPTAEGIWKNMLSRFKQDDAPRVYDIEQRLSKIE 154
Query: 122 QGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLN 181
QG + Y+T +++LWEE ++ +P+CTC C C++ + V +FL GLN
Sbjct: 155 QGSMDISAYYTELQTLWEEHKNYVDLPVCTCG-RCECDAAVKWERLQQRSHVTKFLMGLN 213
Query: 182 DSFSVVKTQVLLIDPLPSINKVYSMVIQEE-SNIIPPTSLASN-EDSSILVNASDARKPF 239
+S+ + +L++ P+ +I + +++V Q+E I PT N ++ N+ D P+
Sbjct: 214 ESYEQTRRHILMLKPIRTIEEAFNIVTQDERQKAIRPTPKVDNVAFQTVSYNSGD---PY 270
Query: 240 LRGKSSGTSQSKNNSR 255
+ +G + N +R
Sbjct: 271 MENMENGFIAAYNTAR 286
>At1g47650 hypothetical protein
Length = 1409
Score = 146 bits (368), Expect = 1e-34
Identities = 83/283 (29%), Positives = 139/283 (48%), Gaps = 33/283 (11%)
Query: 1 SIYYVHPSEGPNSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLNRT 60
S +++H ++ P ++ L NY WN +M +L KNK FIDG++P P D N
Sbjct: 63 SPFFLHSADHPGLNIISHRLDETNYGDWNVAMLISLDAKNKSGFIDGTLPRPLESDKNFR 122
Query: 61 AWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNL 120
W RC +++ SW++NSVSPQI ++I+ A D+W +L RF+ + R +L I +
Sbjct: 123 LWSRCKSMVKSWLLNSVSPQIYRSILRMNDASDIWRDLHSRFNVTNLPRTYNLTQEIQDF 182
Query: 121 KQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGL 180
+QG S+ +Y+T +K+LW++L S + PC C + + ++++ L GL
Sbjct: 183 RQGTLSLSEYYTRLKTLWDQLESTEELDD-----PCICGKAMRLQQKAEQAKIVKLLAGL 237
Query: 181 NDSFSVVKTQVLLIDPLPSINKVYSMVIQEESN------IIPPTSLASNEDSSILVNASD 234
NDS+++++ Q++ LPS+ +VY ++ Q+ S + PP + +E
Sbjct: 238 NDSYAIIRRQIIAKKALPSLAEVYHILDQDNSQQGFSNVVAPPAAFQVSE---------- 287
Query: 235 ARKPFLRGKSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFP 277
+ NN H E CY KH FP
Sbjct: 288 ------------AMMATNNDVNMLCSEWVGHIAERCYKKHGFP 318
Score = 82.8 bits (203), Expect = 1e-15
Identities = 58/197 (29%), Positives = 88/197 (44%), Gaps = 27/197 (13%)
Query: 374 LIDSGANEHICSSLHLFHSYYRIKPICVNLPNGSSVIVQYAGTVVFSPHFHITHVLYSPS 433
+IDSGA H+ LF + VNLP G +V + GT+ + + ++L+ P
Sbjct: 333 VIDSGAIHHVSHDKSLFLNLDTSVVSAVNLPAGPTVRISGVGTLRLNDDILLKNILFIPE 392
Query: 434 FKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKMIGLGNLCDGLYRLHPFAPASPQAH 493
F++ LIS+S + + V F + IQD +M+G G LY L P
Sbjct: 393 FRLNLISISSLTDDIGSRVIFDKTSCEIQDPIKARMLGQGRRIANLYVLDIEDPIV---- 448
Query: 494 FISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRLGHLSNQRLSMMHSLYSSITIDNKA 553
+V+++ ++WH RLGH S QRL ++ + NK
Sbjct: 449 ---------------------SVNAVVDISMWHRRLGHASLQRLDVISDSLGTTKPKNKG 487
Query: 554 --VCDICHFAKQRKLPY 568
C +CH AKQRKLP+
Sbjct: 488 SDYCHVCHLAKQRKLPF 504
>At4g16870 retrotransposon like protein
Length = 1474
Score = 139 bits (349), Expect = 2e-32
Identities = 150/642 (23%), Positives = 266/642 (41%), Gaps = 114/642 (17%)
Query: 12 NSVTVTPLLTGPNYLAWNRSMKRALGTKNKFVFIDGSV--PIPPMDDLN-------RTAW 62
N+ VT L T NYL W+ + L +DGS+ P P + N T W
Sbjct: 27 NTSNVTKL-TSNNYLMWSLQIHALLDGYELAGHLDGSIETPAPTLTTNNVVSANPQYTLW 85
Query: 63 ERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSINNLKQ 122
+R + LI S +I ++SP + + A +W L ++K + LR+ I LK+
Sbjct: 86 KRQDRLIFSALIGAISPPVQPLVSRATKASQIWKTLTNTYAKSSYDHIKQLRTQIKQLKK 145
Query: 123 GDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLTGLND 182
G K++ +Y S +L ++L + E+QV + L GL +
Sbjct: 146 GTKTIDEYVLSHTTLLDQLAI-------------------LGKPMEHEEQVERILEGLPE 186
Query: 183 SFSVVKTQVLLIDPLPSINKVYSMVIQEESNIIPPTSLASNEDSSILVNASDARKPFLRG 242
+ V Q+ D PSI +++ +I E+ ++ +L+S SS+ ++A+ A++
Sbjct: 187 DYKTVVDQIEGKDNTPSITEIHERLINHEAKLLSTAALSS---SSLPMSANVAQQ----- 238
Query: 243 KSSGTSQSKNNSRYCTFCRRNNHTVEYCYLKHDFPNANKPTASSNAVTSEHAVDSHTSSE 302
+ NN+R NN N NK N T+ ++ S
Sbjct: 239 ------RHHNNNR-------NN-------------NQNKNRTQGNTYTNNWQPSANNKSG 272
Query: 303 GTSSSSQTGLTQEQYVHLVSLLQQSSLVPSATPPNPASTNHVATSFPSSIDFTSGINTIF 362
G Q V S + P P+S++ +T P + +
Sbjct: 273 QRPFKPYLGKCQICNVQGHSARR----CPQLQAMQPSSSSSASTFTPWQPRANLAMGAPY 328
Query: 363 SCSLHVPSDHWLIDSGANEHICSSLHLFHSYYRIKPICVNLPNGSSVIVQYAGTVVFSPH 422
+ +++WL+DSGA HI S L+ + V + +G+S+ + G+ +
Sbjct: 329 T------ANNWLLDSGATHHITSDLNALALHQPYNGDDVMIADGTSLKITKTGSTFLPSN 382
Query: 423 FH---ITHVLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKMIGLGNLCDGL 479
+ VLY P + L+SV ++C + V F ++ ++D+ T ++ G D L
Sbjct: 383 ARDLTLNKVLYVPDIQKNLVSVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDEL 442
Query: 480 YRLHPFAPASPQAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRLGHLSNQRLSM 539
Y + +P+A + + SP +S WH RLGH S+ L+
Sbjct: 443 YE---WPVTNPKATALFTTPSPKTTLSS-----------------WHSRLGHPSSSILNT 482
Query: 540 MHSLYS---SITIDNKAVCDICHFAKQRKLPYNLSTLLLHLNLNYCILIFGVLFLLLVFM 596
+ S +S S++ NK C C K KLP+++S++ L Y IF +++
Sbjct: 483 LISKFSLPVSVSASNKLACSDCFINKSHKLPFSISSIKSTSPLEY---IFSDVWM----- 534
Query: 597 DTSIFSP----LLMILVDFYG--SFC*KINQKSQIMSKILSY 632
+ I SP ++LVD + ++ + QKSQ+ S +++
Sbjct: 535 -SPILSPDNYKYYLVLVDHHTRYTWLYPLQQKSQVKSTFIAF 575
>At1g21280 unknown protein (At1g21280)
Length = 237
Score = 127 bits (320), Expect = 4e-29
Identities = 68/216 (31%), Positives = 118/216 (54%), Gaps = 5/216 (2%)
Query: 1 SIYYVHPS-EGPNSVTVTPLLTGP-NYLAWNRSMKRALGTKNKFVFIDGSVPIPPMDDLN 58
S YY+ P P+ ++ L NY+AW + L KF FIDG++P P
Sbjct: 16 SPYYLPPDIHHPSDFSIQKLSKDEDNYVAWKIRFRSFLRVTKKFGFIDGTLPKPDPFSPL 75
Query: 59 RTAWERCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRIRVASLRSSIN 118
WE+CN +++ W++NS++ ++ +++++ E A +W +L+ F +++ LR +
Sbjct: 76 YQPWEQCNAMVMYWLMNSMTDKLLESVMYAETAHKMWEDLRRVFVPCVDLKIYQLRRRLA 135
Query: 119 NLKQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFRMEDQVIQFLT 178
L+QG SV +YF + +W EL+ + P+P C C C CE + A + R ++Q +FL
Sbjct: 136 TLRQGGDSVEEYFGKLSKVWMELSEYAPIPECKCG-GCNCECTKRAEEAREKEQRYEFLM 194
Query: 179 G--LNDSFSVVKTQVLLIDPLPSINKVYSMVIQEES 212
G LN F V T+++ P PS+++ ++MV ES
Sbjct: 195 GLKLNQGFEAVTTKIMFQKPPPSLHEAFAMVKDAES 230
>At2g15870 putative retroelement pol polyprotein
Length = 1264
Score = 119 bits (297), Expect = 2e-26
Identities = 71/241 (29%), Positives = 121/241 (49%), Gaps = 26/241 (10%)
Query: 54 MDDLNRTAWE-----RCNNLILSWIINSVSPQIAQTIVFHEYAIDVWIELQERFSKVDRI 108
+D+ N W CN ++ SW++NS+SPQI ++I+ D+W +L RF+ +
Sbjct: 82 LDETNYGDWNVAMLISCNTMVKSWLLNSMSPQIYRSILRMNDVSDIWRDLNSRFNMTNLP 141
Query: 109 RVASLRSSINNLKQGDKSVLDYFTSIKSLWEELNSHRPMPMCTCPYPCRCESMRAARDFR 168
R +L I +L+Q S+ +Y+T +K+LW++LNS + PC C +
Sbjct: 142 RTYNLTQEIQDLRQSTLSLSEYYTRLKTLWDQLNSTEEL-----DDPCTCGKALRLQQKA 196
Query: 169 MEDQVIQFLTGLNDSFSVVKTQVLLIDPLPSINKVYSMVIQEESN------IIPPTSLAS 222
++++FL GLN+S+++++ QV+ LPS+ +VY +V Q+ S + PP +
Sbjct: 197 ERAKIVKFLAGLNESYAIIRRQVIAKKILPSLAEVYHIVDQDNSQQGFSNVVAPPVAFQV 256
Query: 223 NEDSSILVNASDARKPFLRGKSSGTSQSKNNSR-YCTFCRRNNHTVEYCYLKHDFPNANK 281
+E + + N D +++ N R C+F R H E CY KH FP
Sbjct: 257 SEVT--VANIIDPTICYVQ-------NCPNKGRPMCSFYNRVGHIAERCYKKHGFPPGFT 307
Query: 282 P 282
P
Sbjct: 308 P 308
Score = 58.9 bits (141), Expect = 2e-08
Identities = 50/191 (26%), Positives = 79/191 (41%), Gaps = 41/191 (21%)
Query: 427 HVLYSPSFKVYLISVSKICQSLPYHVHFLLNTYVIQDVKTQKMIGLGNLCDGLYRLHPFA 486
+VL+ P F++ LIS+S + + V F + + IQD+ +M+G G LY +
Sbjct: 344 NVLFIPEFRLNLISISSLTDDIGSRVIFDQHAFEIQDLIKGRMLGHGRRVANLYVM---- 399
Query: 487 PASPQAHFISSAVSPTNKMSCNSVSSNNNVSSIPSNAIWHFRLGHLSNQRLSMMHSLYSS 546
+ +N +V+++ + WH RL H S QRL ++ +
Sbjct: 400 ---------------------DVEDTNVSVNAVVDISTWHNRLRHASLQRLDVISESLGT 438
Query: 547 ITIDNKA--VCDICHFAKQRKLPYNLSTLLLHLNLNYCILIFGVLFLLLVFMDTSIFSPL 604
NK C +CH AK RKL + N C IF +L + I+ P
Sbjct: 439 TKHKNKGSDYCHVCHLAKHRKLSFPSQN-------NVCNEIFEMLHI-------DIWGPF 484
Query: 605 LMILVDFYGSF 615
+ VD Y F
Sbjct: 485 SVETVDGYQYF 495
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.356 0.156 0.555
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,822,824
Number of Sequences: 26719
Number of extensions: 1081669
Number of successful extensions: 7195
Number of sequences better than 10.0: 98
Number of HSP's better than 10.0 without gapping: 57
Number of HSP's successfully gapped in prelim test: 41
Number of HSP's that attempted gapping in prelim test: 6821
Number of HSP's gapped (non-prelim): 230
length of query: 1441
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1329
effective length of database: 8,326,068
effective search space: 11065344372
effective search space used: 11065344372
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC149491.1


