

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149471.21 - phase: 0
(281 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
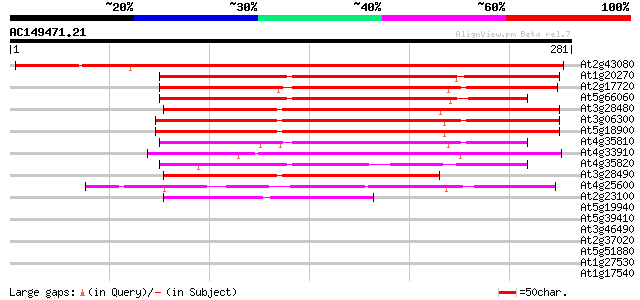
Score E
Sequences producing significant alignments: (bits) Value
At2g43080 unknown protein 405 e-113
At1g20270 unknown protein 220 6e-58
At2g17720 similar to prolyl 4-hydroxylase alpha subunit 207 5e-54
At5g66060 prolyl 4-hydroxylase, alpha subunit-like protein 183 1e-46
At3g28480 prolyl 4-hydroxylase like protein 182 2e-46
At3g06300 unknown protein 182 2e-46
At5g18900 unknown protein 175 2e-44
At4g35810 putative protein 169 2e-42
At4g33910 unknown protein 145 2e-35
At4g35820 putative protein 141 3e-34
At3g28490 prolyl 4-hydroxylase, putative 125 2e-29
At4g25600 unknown protein 92 2e-19
At2g23100 hypothetical protein 59 3e-09
At5g19940 unknown protein 33 0.13
At5g39410 unknown protein 30 1.1
At3g46490 putative protein 30 1.5
At2g37020 translin-like protein 30 1.5
At5g51880 unknown protein 29 3.2
At1g27530 unknown protein 29 3.2
At1g17540 protein kinase, putative 29 3.2
>At2g43080 unknown protein
Length = 283
Score = 405 bits (1042), Expect = e-113
Identities = 195/278 (70%), Positives = 234/278 (84%), Gaps = 5/278 (1%)
Query: 4 PAMKIVFGLLTFVTIGMIIGALSQLAFIRRLELEEPFTTTTTTRSLLPRGYTYWNN---- 59
PAMKIVFGLLTFVT+GM+IG+L QLAFI RLE + T + R L + Y +
Sbjct: 3 PAMKIVFGLLTFVTVGMVIGSLLQLAFINRLE-DSYGTGFPSLRGLRGQNTRYLRDVSRW 61
Query: 60 NNDKEAQILRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGK 119
NDK+A++LR+G VKPEV+SWSPRII+LH+FLS EEC+YL+ +A PRL++STVVD TGK
Sbjct: 62 ANDKDAELLRIGNVKPEVVSWSPRIIVLHDFLSPEECEYLKAIARPRLQVSTVVDVKTGK 121
Query: 120 GIKSDVRTSSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHH 179
G+KSDVRTSSGMFL+H ER YP+I AIEKRI+V+SQ+P ENGEL+QVLRYE Q+Y+PHH
Sbjct: 122 GVKSDVRTSSGMFLTHVERSYPIIQAIEKRIAVFSQVPAENGELIQVLRYEPQQFYKPHH 181
Query: 180 DYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGN 239
DYF+DTFNLKRGGQR+ATMLMYL D+VEGGET+FP AG +C+CGGK+ KG+ VKP KG+
Sbjct: 182 DYFADTFNLKRGGQRVATMLMYLTDDVEGGETYFPLAGDGDCTCGGKIMKGISVKPTKGD 241
Query: 240 AVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMRQ 277
AVLFWSMGLDGQSDP S+HGGC VL+GEKWSATKWMRQ
Sbjct: 242 AVLFWSMGLDGQSDPRSIHGGCEVLSGEKWSATKWMRQ 279
>At1g20270 unknown protein
Length = 287
Score = 220 bits (561), Expect = 6e-58
Identities = 118/209 (56%), Positives = 138/209 (65%), Gaps = 14/209 (6%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSH 135
EVLSW PR + HNFLS EEC+YL +A P + STVVD+ TGK S VRTSSG FL
Sbjct: 77 EVLSWEPRAFVYHNFLSKEECEYLISLAKPHMVKSTVVDSETGKSKDSRVRTSSGTFLRR 136
Query: 136 EERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRI 195
K +I IEKRI+ Y+ IP ++GE +QVL YE Q Y PH+DYF D FN K GGQR+
Sbjct: 137 GRDK--IIKTIEKRIADYTFIPADHGEGLQVLHYEAGQKYEPHYDYFVDEFNTKNGGQRM 194
Query: 196 ATMLMYLGDNVEGGETHFPSAGSDECS---------CGGKLTKGLCVKPVKGNAVLFWSM 246
ATMLMYL D EGGET FP+A + S CG KGL VKP G+A+LFWSM
Sbjct: 195 ATMLMYLSDVEEGGETVFPAANMNFSSVPWYNELSECG---KKGLSVKPRMGDALLFWSM 251
Query: 247 GLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
D DP S+HGGCPV+ G KWS+TKWM
Sbjct: 252 RPDATLDPTSLHGGCPVIRGNKWSSTKWM 280
>At2g17720 similar to prolyl 4-hydroxylase alpha subunit
Length = 291
Score = 207 bits (527), Expect = 5e-54
Identities = 111/209 (53%), Positives = 139/209 (66%), Gaps = 16/209 (7%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFL-- 133
EV+SW PR ++ HNFL+ EEC++L +A P + STVVD TG S VRTSSG FL
Sbjct: 81 EVISWEPRAVVYHNFLTNEECEHLISLAKPSMVKSTVVDEKTGGSKDSRVRTSSGTFLRR 140
Query: 134 SHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
H+E ++ IEKRIS ++ IP+ENGE +QVL Y+ Q Y PH+DYF D FN K GGQ
Sbjct: 141 GHDE----VVEVIEKRISDFTFIPVENGEGLQVLHYQVGQKYEPHYDYFLDEFNTKNGGQ 196
Query: 194 RIATMLMYLGDNVEGGETHFPSAGS--------DECSCGGKLTKGLCVKPVKGNAVLFWS 245
RIAT+LMYL D +GGET FP+A +E S GK +GL V P K +A+LFW+
Sbjct: 197 RIATVLMYLSDVDDGGETVFPAARGNISAVPWWNELSKCGK--EGLSVLPKKRDALLFWN 254
Query: 246 MGLDGQSDPDSVHGGCPVLAGEKWSATKW 274
M D DP S+HGGCPV+ G KWS+TKW
Sbjct: 255 MRPDASLDPSSLHGGCPVVKGNKWSSTKW 283
>At5g66060 prolyl 4-hydroxylase, alpha subunit-like protein
Length = 267
Score = 183 bits (464), Expect = 1e-46
Identities = 101/195 (51%), Positives = 123/195 (62%), Gaps = 18/195 (9%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSH 135
E++SW PR + HNFL+ EEC YL +A P ++ STVVD TGK S VRTSSG FL+
Sbjct: 79 EIISWEPRASVYHNFLTKEECKYLIELAKPHMEKSTVVDEKTGKSTDSRVRTSSGTFLAR 138
Query: 136 EERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRI 195
K I IEKRIS ++ IP+E+GE +QVL YE Q Y PH+DYF D +N + GGQRI
Sbjct: 139 GRDK--TIREIEKRISDFTFIPVEHGEGLQVLHYEIGQKYEPHYDYFMDEYNTRNGGQRI 196
Query: 196 ATMLMYLGDNVEGGETHFPSAGSD-----------ECSCGGKLTKGLCVKPVKGNAVLFW 244
AT+LMYL D EGGET FP+A + EC G GL VKP G+A+LFW
Sbjct: 197 ATVLMYLSDVEEGGETVFPAAKGNYSAVPWWNELSECGKG-----GLSVKPKMGDALLFW 251
Query: 245 SMGLDGQSDPDSVHG 259
SM D DP S+HG
Sbjct: 252 SMTPDATLDPSSLHG 266
>At3g28480 prolyl 4-hydroxylase like protein
Length = 316
Score = 182 bits (462), Expect = 2e-46
Identities = 93/203 (45%), Positives = 136/203 (66%), Gaps = 7/203 (3%)
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEE 137
LSW+PR+ L FLS EECD+ +A +L+ S V D ++G+ ++S+VRTSSGMFLS +
Sbjct: 59 LSWTPRVFLYEGFLSDEECDHFIKLAKGKLEKSMVADNDSGESVESEVRTSSGMFLS--K 116
Query: 138 RKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIAT 197
R+ ++ +E +++ ++ +P ENGE MQ+L YE Q Y PH DYF D NL+ GG RIAT
Sbjct: 117 RQDDIVSNVEAKLAAWTFLPEENGESMQILHYENGQKYEPHFDYFHDQANLELGGHRIAT 176
Query: 198 MLMYLGDNVEGGETHFP-----SAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQS 252
+LMYL + +GGET FP + + S +G VKP KG+A+LF+++ + +
Sbjct: 177 VLMYLSNVEKGGETVFPMWKGKATQLKDDSWTECAKQGYAVKPRKGDALLFFNLHPNATT 236
Query: 253 DPDSVHGGCPVLAGEKWSATKWM 275
D +S+HG CPV+ GEKWSAT+W+
Sbjct: 237 DSNSLHGSCPVVEGEKWSATRWI 259
>At3g06300 unknown protein
Length = 299
Score = 182 bits (461), Expect = 2e-46
Identities = 100/212 (47%), Positives = 133/212 (62%), Gaps = 14/212 (6%)
Query: 74 KPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFL 133
K + +S PR + FL+ ECD+L +A L+ S V D + G+ SDVRTSSG F+
Sbjct: 37 KVKQVSSKPRAFVYEGFLTDLECDHLISLAKENLQRSAVADNDNGESQVSDVRTSSGTFI 96
Query: 134 SHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
S + K P++ IE ++S ++ +P ENGE +QVLRYE Q Y H DYF D N+ RGG
Sbjct: 97 S--KGKDPIVSGIEDKLSTWTFLPKENGEDLQVLRYEHGQKYDAHFDYFHDKVNIARGGH 154
Query: 194 RIATMLMYLGDNVEGGETHFPSA----------GSDECSCGGKLTKGLCVKPVKGNAVLF 243
RIAT+L+YL + +GGET FP A D+ S K KG+ VKP KGNA+LF
Sbjct: 155 RIATVLLYLSNVTKGGETVFPDAQEFSRRSLSENKDDLSDCAK--KGIAVKPKKGNALLF 212
Query: 244 WSMGLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
+++ D DP S+HGGCPV+ GEKWSATKW+
Sbjct: 213 FNLQQDAIPDPFSLHGGCPVIEGEKWSATKWI 244
>At5g18900 unknown protein
Length = 298
Score = 175 bits (444), Expect = 2e-44
Identities = 95/210 (45%), Positives = 132/210 (62%), Gaps = 10/210 (4%)
Query: 74 KPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFL 133
K + +S PR + FL+ ECD++ +A LK S V D ++G+ S+VRTSSG F+
Sbjct: 36 KVKQVSSKPRAFVYEGFLTELECDHMVSLAKASLKRSAVADNDSGESKFSEVRTSSGTFI 95
Query: 134 SHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
S + K P++ IE +IS ++ +P ENGE +QVLRYE Q Y H DYF D N+ RGG
Sbjct: 96 S--KGKDPIVSGIEDKISTWTFLPKENGEDIQVLRYEHGQKYDAHFDYFHDKVNIVRGGH 153
Query: 194 RIATMLMYLGDNVEGGETHFPSA--------GSDECSCGGKLTKGLCVKPVKGNAVLFWS 245
R+AT+LMYL + +GGET FP A ++ +G+ VKP KG+A+LF++
Sbjct: 154 RMATILMYLSNVTKGGETVFPDAEIPSRRVLSENKEDLSDCAKRGIAVKPRKGDALLFFN 213
Query: 246 MGLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
+ D DP S+HGGCPV+ GEKWSATKW+
Sbjct: 214 LHPDAIPDPLSLHGGCPVIEGEKWSATKWI 243
>At4g35810 putative protein
Length = 307
Score = 169 bits (428), Expect = 2e-42
Identities = 104/225 (46%), Positives = 127/225 (56%), Gaps = 47/225 (20%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSD----------- 124
EV+SW PR + HNFL+ EEC++L +A P + S VVD TGK I S
Sbjct: 81 EVISWEPRAFVYHNFLTNEECEHLISLAKPSMMKSKVVDVKTGKSIDSRFCTLTSVVVFT 140
Query: 125 --------------------VRTSSGMFLS--HEERKYPMIHAIEKRISVYSQIPIENGE 162
VRTSSG FL+ H+E ++ IE RIS ++ IP ENGE
Sbjct: 141 FQLNLERFENSKFANPSLCRVRTSSGTFLNRGHDE----IVEEIENRISDFTFIPPENGE 196
Query: 163 LMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGS---- 218
+QVL YE Q Y PHHDYF D FN+++GGQRIAT+LMYL D EGGET FP+A
Sbjct: 197 GLQVLHYEVGQRYEPHHDYFFDEFNVRKGGQRIATVLMYLSDVDEGGETVFPAAKGNVSD 256
Query: 219 ----DECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSVHG 259
DE S GK +GL V P K +A+LFWSM D DP S+HG
Sbjct: 257 VPWWDELSQCGK--EGLSVLPKKRDALLFWSMKPDASLDPSSLHG 299
>At4g33910 unknown protein
Length = 288
Score = 145 bits (366), Expect = 2e-35
Identities = 86/211 (40%), Positives = 115/211 (53%), Gaps = 5/211 (2%)
Query: 70 LGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVV--DANTGKGIKSDVRT 127
+G + +VLSW PR I NF + E+C + A LK S + T + K RT
Sbjct: 73 IGSIPFQVLSWRPRAIYFPNFATAEQCQAIIERAKVNLKPSALALRKGETAENTKG-TRT 131
Query: 128 SSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFN 187
SSG F+S E + +E++I+ + IP +GE +LRYE Q Y H+D F+ T
Sbjct: 132 SSGTFISASEESTGALDFVERKIARATMIPRSHGESFNILRYELGQKYDSHYDVFNPTEY 191
Query: 188 LKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECSCG--GKLTKGLCVKPVKGNAVLFWS 245
+ QRIA+ L+YL D EGGET FP G K GL VKP KG+ +LF+S
Sbjct: 192 GPQSSQRIASFLLYLSDVEEGGETMFPFENGSNMGIGYDYKQCIGLKVKPRKGDGLLFYS 251
Query: 246 MGLDGQSDPDSVHGGCPVLAGEKWSATKWMR 276
+ +G D S+HG CPV GEKW ATKW+R
Sbjct: 252 VFPNGTIDQTSLHGSCPVTKGEKWVATKWIR 282
>At4g35820 putative protein
Length = 272
Score = 141 bits (356), Expect = 3e-34
Identities = 88/192 (45%), Positives = 111/192 (56%), Gaps = 26/192 (13%)
Query: 76 EVLSWSPRIILLHNFLSY--------EECDYLRGVALPRLKISTVVDANTGKGIKSDVRT 127
EV++ PR + HNFL+ EECD+L +A P + S V +A TG G +S RT
Sbjct: 89 EVITKEPRAFVYHNFLALFFKICKTNEECDHLISLAKPSMARSKVRNALTGLGEESSSRT 148
Query: 128 SSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFN 187
SSG F+ K ++ IEKRIS ++ IP ENGE +QV+ YE Q + PH D
Sbjct: 149 SSGTFIRSGHDK--IVKEIEKRISEFTFIPQENGETLQVINYEVGQKFEPHFD------- 199
Query: 188 LKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMG 247
G QRIAT+LMYL D +GGET FP A G K KG+ V+P KG+A+LFWSM
Sbjct: 200 ---GFQRIATVLMYLSDVDKGGETVFPEAK------GIKSKKGVSVRPKKGDALLFWSMR 250
Query: 248 LDGQSDPDSVHG 259
DG DP S HG
Sbjct: 251 PDGSRDPSSKHG 262
>At3g28490 prolyl 4-hydroxylase, putative
Length = 213
Score = 125 bits (315), Expect = 2e-29
Identities = 67/139 (48%), Positives = 95/139 (68%), Gaps = 3/139 (2%)
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVV-DANTGKGIKSDVRTSSGMFLSHE 136
LSW+PR L FLS EECD+L +A +L+ S VV D ++G+ S+VRTSSGMFL+
Sbjct: 35 LSWTPRAFLYKGFLSDEECDHLIKLAKGKLEKSMVVADVDSGESEDSEVRTSSGMFLT-- 92
Query: 137 ERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIA 196
+R+ ++ +E +++ ++ +P ENGE +Q+L YE Q Y PH DYF D L+ GG RIA
Sbjct: 93 KRQDDIVANVEAKLAAWTFLPEENGEALQILHYENGQKYDPHFDYFYDKKALELGGHRIA 152
Query: 197 TMLMYLGDNVEGGETHFPS 215
T+LMYL + +GGET FP+
Sbjct: 153 TVLMYLSNVTKGGETVFPN 171
>At4g25600 unknown protein
Length = 291
Score = 92.4 bits (228), Expect = 2e-19
Identities = 72/242 (29%), Positives = 119/242 (48%), Gaps = 34/242 (14%)
Query: 39 PFTTTTTTRSLLPRGYTYWNNNNDKEAQ-ILRLGYVKPE---VLSWSPRIILLHNFLSYE 94
PF + + + L + T + ++D +A +L +V P LSW PR+ L FLS E
Sbjct: 21 PFCSGGSRKELRDKEIT--SKSDDTQASYVLGSKFVDPTRVLQLSWLPRVFLYRGFLSEE 78
Query: 95 ECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEERKYPMIHAIEKRISVYS 154
ECD+L IS + + +D +T P++ IE+++S ++
Sbjct: 79 ECDHL---------ISLRKETTEVYSVDADGKTQLD----------PVVAGIEEKVSAWT 119
Query: 155 QIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFP 214
+P ENG ++V Y + + DYF + + +AT+++YL + +GGE FP
Sbjct: 120 FLPGENGGSIKVRSYTSEKSGKKL-DYFGEEPSSVLHESLLATVVLYLSNTTQGGELLFP 178
Query: 215 SAG---SDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSA 271
++ + C GG + ++PVKGNA+LF++ L+ D S H CPV+ GE A
Sbjct: 179 NSEMKPKNSCLEGGNI-----LRPVKGNAILFFTRLLNASLDGKSTHLRCPVVKGELLVA 233
Query: 272 TK 273
TK
Sbjct: 234 TK 235
>At2g23100 hypothetical protein
Length = 1036
Score = 58.9 bits (141), Expect = 3e-09
Identities = 32/105 (30%), Positives = 59/105 (55%), Gaps = 3/105 (2%)
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEE 137
LSW+PR+ L NF + ++C+ + +A P+LK ST+ K + R+ + +E
Sbjct: 800 LSWNPRVFYLPNFATKQQCEAVIDMAKPKLKPSTLALRKETKHFQMQYRS---LHQHTDE 856
Query: 138 RKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYF 182
+ ++ AIE++I++ ++ P + E +LRY+ Q Y H+D F
Sbjct: 857 DESGVLAAIEEKIALATRFPKDYYESFNILRYQLGQKYDSHYDAF 901
>At5g19940 unknown protein
Length = 239
Score = 33.5 bits (75), Expect = 0.13
Identities = 33/137 (24%), Positives = 55/137 (40%), Gaps = 5/137 (3%)
Query: 147 EKRISVYSQ----IPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIATMLMYL 202
E R ++S +P++ +VLR + + L+ G + + L+
Sbjct: 16 EPRTQIHSSRRLNLPLQYSIPYKVLRSRSRRLGLVVSSVSAPNVELRTGPDDLISTLLSK 75
Query: 203 GDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSVHGGCP 262
N +GG T P + G+L K CVK N ++F + S P S GG
Sbjct: 76 VANSDGGVTLSPEQHKEVAQVAGELQK-YCVKEPVKNPLIFGDWEVVYCSRPTSPGGGYR 134
Query: 263 VLAGEKWSATKWMRQSV 279
+ G + TK M Q++
Sbjct: 135 SVIGRLFFKTKEMIQAI 151
>At5g39410 unknown protein
Length = 454
Score = 30.4 bits (67), Expect = 1.1
Identities = 14/33 (42%), Positives = 23/33 (69%)
Query: 7 KIVFGLLTFVTIGMIIGALSQLAFIRRLELEEP 39
K +FG+ +VT+G+ +G LS+ +F R L L+ P
Sbjct: 303 KSLFGIFRYVTLGVSLGLLSKFSFGRWLLLKFP 335
>At3g46490 putative protein
Length = 306
Score = 30.0 bits (66), Expect = 1.5
Identities = 17/36 (47%), Positives = 21/36 (58%), Gaps = 2/36 (5%)
Query: 156 IPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRG 191
+P+E E M+VLR EK + Y P HD D N RG
Sbjct: 69 LPLE--EKMKVLRNEKYRGYAPFHDSLLDPENQVRG 102
>At2g37020 translin-like protein
Length = 238
Score = 30.0 bits (66), Expect = 1.5
Identities = 18/57 (31%), Positives = 27/57 (46%), Gaps = 2/57 (3%)
Query: 129 SGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDT 185
+ + L H+ R P + IEK + G L ++L QYYR H D+ S+T
Sbjct: 49 ANLLLVHQSRPIPEV--IEKAKEKIVDLKQYYGRLAEILEECPGQYYRYHGDWRSET 103
>At5g51880 unknown protein
Length = 227
Score = 28.9 bits (63), Expect = 3.2
Identities = 27/126 (21%), Positives = 54/126 (42%), Gaps = 9/126 (7%)
Query: 150 ISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIATMLMYLGDNVEGG 209
I + ++ + ++ RY Q++ H D +D L+ G + T+L+YL N
Sbjct: 96 IKIRRKVAVGLNPNIRFYRYSAGQHFGRHIDESAD---LEDGNRTYYTLLIYLSGNSTKS 152
Query: 210 ETHFPSAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQS------DPDSVHGGCPV 263
++ S+ +++ S L G V N+++ ++G + D +H G V
Sbjct: 153 KSKSSSSKTNDSSSAEPLVGGETVFYGSRNSIVAEVAPVEGMALFHIHGDKCMLHEGRNV 212
Query: 264 LAGEKW 269
G K+
Sbjct: 213 SKGVKY 218
>At1g27530 unknown protein
Length = 174
Score = 28.9 bits (63), Expect = 3.2
Identities = 14/41 (34%), Positives = 20/41 (48%), Gaps = 2/41 (4%)
Query: 54 YTYWNNNNDKEAQILRLGYVKPEVLSWSPRIILLHNFLSYE 94
YT N +ND + R+ PE W+ + +HN L YE
Sbjct: 44 YTQMNKSNDNDW--FRISASNPEGTRWTGKCWYVHNLLKYE 82
>At1g17540 protein kinase, putative
Length = 740
Score = 28.9 bits (63), Expect = 3.2
Identities = 20/76 (26%), Positives = 30/76 (39%), Gaps = 3/76 (3%)
Query: 157 PIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSA 216
P NGEL + K YR S+ F + A+ LG + GG +H +
Sbjct: 209 PALNGELSPPTPHYKPSLYRSSMSELSNEFPSDHSAESNASFYSILGRSTYGGSSHSSMS 268
Query: 217 ---GSDECSCGGKLTK 229
+E GG +T+
Sbjct: 269 EITDGEESLSGGNITE 284
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,821,126
Number of Sequences: 26719
Number of extensions: 303765
Number of successful extensions: 595
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 552
Number of HSP's gapped (non-prelim): 22
length of query: 281
length of database: 11,318,596
effective HSP length: 98
effective length of query: 183
effective length of database: 8,700,134
effective search space: 1592124522
effective search space used: 1592124522
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC149471.21


