

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
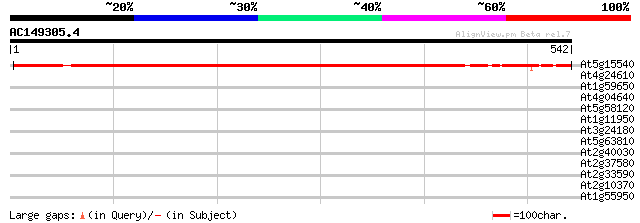
Query= AC149305.4 - phase: 0 /partial
(542 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g15540 putative protein 611 e-175
At4g24610 putative protein 37 0.028
At1g59650 unknown protein 34 0.18
At4g04640 ATPase gamma-subunit (AtpC1) 30 2.6
At5g58120 resistance protein - like 30 3.4
At1g11950 putative DNA-binding protein 30 4.5
At3g24180 unknown protein 29 5.8
At5g63810 beta-galactosidase (emb|CAB64746.1) 28 9.9
At2g40030 putative DNA-directed RNA polymerase II subunit 28 9.9
At2g37580 putative RING zinc finger protein 28 9.9
At2g33590 putative cinnamoyl-CoA reductase 28 9.9
At2g10370 putative replication protein A1 28 9.9
At1g55950 hypothetical protein 28 9.9
>At5g15540 putative protein
Length = 1755
Score = 611 bits (1576), Expect = e-175
Identities = 324/545 (59%), Positives = 402/545 (73%), Gaps = 23/545 (4%)
Query: 4 QLLESIIFIIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAG 63
QLLESIIFIIDSVLPL+RKLP S+ E+LEQDLK MI+RHSFL VVHAC+ S+LAG
Sbjct: 1067 QLLESIIFIIDSVLPLIRKLPLSVTEDLEQDLKHMIVRHSFLTVVHACV------SKLAG 1120
Query: 64 KGAAVIEHLIQVFFKCLDTEAVVNKQLVGRSLFCLGLLIRYGNCLLASSGNKLVDVKRSL 123
KG +++EHL+Q FFK L+ + N Q+ GRSLFCLGLLIR+GN L+++SG K ++ L
Sbjct: 1121 KGVSIVEHLLQFFFKRLEAQGSDNTQIAGRSLFCLGLLIRHGNSLISTSGGKNFNLSGCL 1180
Query: 124 NLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADDRLKIQALQ 183
NLF ++L ED ALK RSLQALG++LIARPEYMLE DIGKI+E TL+ A+ R+K+QALQ
Sbjct: 1181 NLFKRHLRTEDIALKVRSLQALGFILIARPEYMLEEDIGKIIETTLADEANGRMKMQALQ 1240
Query: 184 NMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYWNNILGRC 243
NM+EYLLDAE ++ +E+ + G +VPVAAGAGDTNICGGI+QL+W+ ILGRC
Sbjct: 1241 NMYEYLLDAEKQLGSEKASDNTVNSVEQGGHNVPVAAGAGDTNICGGIVQLFWDKILGRC 1300
Query: 244 VDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLMNMHEKYP 303
+DF+ Q+RQ++LKIVEVVLRQGLVHPITCVPYLIALETDP E+N K+AHHLLMNMHEKYP
Sbjct: 1301 LDFDDQIRQTSLKIVEVVLRQGLVHPITCVPYLIALETDPQEANQKLAHHLLMNMHEKYP 1360
Query: 304 AFFESCLGDGLQMSFMFMQSIF-VSPDENVNHKSQSKIAVSGKGKPEADSLAQSRVGVSR 362
AFFES LGDGLQMSF+FMQSI V+ + N + + + + GK + +L Q+R+GVSR
Sbjct: 1361 AFFESRLGDGLQMSFIFMQSISQVTSEPNQSLQQKGSTNMLGKNDHASSTLTQARLGVSR 1420
Query: 363 IYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEPLYLIYT 422
IYKLIRGNR+SRNKFM+SIVRKFDNP WN VI+FL YCTE LALLPF +PDEPLYL+Y+
Sbjct: 1421 IYKLIRGNRVSRNKFMTSIVRKFDNPTWNGSVISFLKYCTETLALLPFTSPDEPLYLVYS 1480
Query: 423 INRVVQVRAGPLEANFKAWSSSLLQSEGQGTPHGNGMYQRATDETIHSTQGQSMDLNGPF 482
INRV+Q+RAG +E+N KA LL + T HGNG YQ+ + I MDLN
Sbjct: 1481 INRVMQIRAGAVESNLKA----LLHKDSAKTQHGNGAYQQ---DPIPGHMNM-MDLNTRI 1532
Query: 483 QQNVDVQPYVDDMTSVDLNG-----TNHQLPDYPLSHNGRLKVKPQAAGFADSFTFSKDD 537
Q+ T +DLNG + Q Y + HNG+ V + +D S DD
Sbjct: 1533 QEEPRHWNSYGHATLIDLNGSVYQDSRDQFTSYQV-HNGKADVHKMTS--SDPPELSTDD 1589
Query: 538 LEKVQ 542
L+K+Q
Sbjct: 1590 LQKIQ 1594
>At4g24610 putative protein
Length = 1145
Score = 37.0 bits (84), Expect = 0.028
Identities = 16/40 (40%), Positives = 27/40 (67%)
Query: 406 ALLPFVAPDEPLYLIYTINRVVQVRAGPLEANFKAWSSSL 445
+++P+V PDE L+ ++ R++ V +EA FKAWSS +
Sbjct: 930 SVIPYVVPDELGILLNSMKRMLDVLRPNIEAKFKAWSSCI 969
>At1g59650 unknown protein
Length = 492
Score = 34.3 bits (77), Expect = 0.18
Identities = 15/54 (27%), Positives = 27/54 (49%)
Query: 454 PHGNGMYQRATDETIHSTQGQSMDLNGPFQQNVDVQPYVDDMTSVDLNGTNHQL 507
PH QR D+ + +G + D N PF++ + + V ++ + LNG +L
Sbjct: 344 PHFQESIQRLLDDEVEKVRGYTTDTNVPFRERLKILGRVANVDDLQLNGAEKKL 397
>At4g04640 ATPase gamma-subunit (AtpC1)
Length = 373
Score = 30.4 bits (67), Expect = 2.6
Identities = 15/48 (31%), Positives = 26/48 (53%), Gaps = 2/48 (4%)
Query: 183 QNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGG 230
+ + E L + +++T++VD VP VR + V + GD +CGG
Sbjct: 96 ETLVEVLYNINEQLQTDDVD--VPLTKVRPVKKVALVVVTGDRGLCGG 141
>At5g58120 resistance protein - like
Length = 1046
Score = 30.0 bits (66), Expect = 3.4
Identities = 36/136 (26%), Positives = 54/136 (39%), Gaps = 15/136 (11%)
Query: 31 LEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEHLIQVFFKCLDTEA--VVNK 88
L L Q+ILR+S + + CIK L + L G + L ++ LD EA +
Sbjct: 760 LPMSLTQLILRYSDIERIPDCIKALHQLFSLDLTGCRRLASLPELPGSLLDLEAEDCESL 819
Query: 89 QLVGRSLFCLGLLIRYGNCLLASSGNKLVDVKRSLNLFMKYL-------AGEDYALKARS 141
+ V L L+ + NC + ++R + K L A D+ K S
Sbjct: 820 ETVFSPLHTPRALLNFTNCFKLGGQARRAIIRRRSEIIGKALLPGREVPAEFDHRAKGNS 879
Query: 142 LQAL--GYVLIARPEY 155
L + GY RP Y
Sbjct: 880 LTIILNGY----RPSY 891
>At1g11950 putative DNA-binding protein
Length = 879
Score = 29.6 bits (65), Expect = 4.5
Identities = 18/54 (33%), Positives = 28/54 (51%), Gaps = 1/54 (1%)
Query: 14 DSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAA 67
DSVL L R LP + + +LEQ + + +S + C +C MS + K A+
Sbjct: 418 DSVLELKRILPVTWMSDLEQKAETFLASYSIKPPMSYC-RCSSDMSSMKRKAAS 470
>At3g24180 unknown protein
Length = 950
Score = 29.3 bits (64), Expect = 5.8
Identities = 18/52 (34%), Positives = 25/52 (47%), Gaps = 3/52 (5%)
Query: 192 AESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYWNNILGRC 243
A+ +T E DGK + +G S P + AGDT IC + W G+C
Sbjct: 323 AKDMWDTMEQDGKFDQENFNSGPSTP--SLAGDT-ICAAVSASAWVEAHGKC 371
>At5g63810 beta-galactosidase (emb|CAB64746.1)
Length = 741
Score = 28.5 bits (62), Expect = 9.9
Identities = 20/81 (24%), Positives = 33/81 (40%), Gaps = 3/81 (3%)
Query: 377 FMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPD-EPLYLIYTINR--VVQVRAGP 433
++ V + DN W ++ +F TY +L AP P+ L N + G
Sbjct: 139 YVPGTVFRADNEPWKHYMESFTTYIVNLLKQEKLFAPQGGPIILSQVENEYGYYEKDYGE 198
Query: 434 LEANFKAWSSSLLQSEGQGTP 454
+ WS+S+ S+ G P
Sbjct: 199 GGKRYAQWSASMAVSQNIGVP 219
>At2g40030 putative DNA-directed RNA polymerase II subunit
Length = 888
Score = 28.5 bits (62), Expect = 9.9
Identities = 33/149 (22%), Positives = 56/149 (37%), Gaps = 17/149 (11%)
Query: 77 FKCLDTEAVVNKQLVGRSLFCLGLLIRYGNCLLASSGNKLVDVKRSLNLFMK-------- 128
F+ +DT Q+ G + + +RYG ++ S + ++ LF++
Sbjct: 221 FRVVDTYL----QVRGTAKAARNIDMRYGVSKISDSSSSKAWTEKMRTLFIRKGSGFSSR 276
Query: 129 -YLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADDRLKIQALQNMFE 187
+ G+ Y R + +G + E + G L + DD+L + Q
Sbjct: 277 SVITGDAY----RHVNEVGIPIEIAQRITFEERVSVHNRGYLQKLVDDKLCLSYTQGSTT 332
Query: 188 YLLDAESKMETEEVDGKVPGHSVRAGQSV 216
Y L SK TE G+V V G V
Sbjct: 333 YSLRDGSKGHTELKPGQVVHRRVMDGDVV 361
>At2g37580 putative RING zinc finger protein
Length = 630
Score = 28.5 bits (62), Expect = 9.9
Identities = 24/78 (30%), Positives = 31/78 (38%), Gaps = 13/78 (16%)
Query: 409 PFVAPDE-----PLYLIYTINRVVQVRA--------GPLEANFKAWSSSLLQSEGQGTPH 455
PF AP PL LI+ + V V A GP ++ + SSS S TPH
Sbjct: 49 PFPAPPRSIDLTPLKLIFVVIAFVAVPALVYALFFNGPCSSSRRNSSSSRTSSSSDDTPH 108
Query: 456 GNGMYQRATDETIHSTQG 473
T+ T+ S G
Sbjct: 109 ATVDTPPITETTVTSESG 126
>At2g33590 putative cinnamoyl-CoA reductase
Length = 321
Score = 28.5 bits (62), Expect = 9.9
Identities = 13/45 (28%), Positives = 22/45 (48%)
Query: 348 PEADSLAQSRVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNK 392
PE + +A + G + K + R ++SS+ F NP W+K
Sbjct: 96 PEVELIAPAVDGTLNVLKACIEANVKRVVYVSSVAAAFMNPMWSK 140
>At2g10370 putative replication protein A1
Length = 267
Score = 28.5 bits (62), Expect = 9.9
Identities = 21/86 (24%), Positives = 36/86 (41%), Gaps = 3/86 (3%)
Query: 313 GLQMSFMFMQSIFVSPDENVN---HKSQSKIAVSGKGKPEADSLAQSRVGVSRIYKLIRG 369
G ++ FM M ++S + + S S I V K E ++ Q V + + ++ R
Sbjct: 158 GFKLPFMIMNKKYISSTSIFSFFPNISCSYILVVSKDTLECEAYGQIAVDLEKRFRAFRK 217
Query: 370 NRISRNKFMSSIVRKFDNPKWNKFVI 395
N + +VR + W K VI
Sbjct: 218 NNVVIALAWWKVVRYYQGCNWFKIVI 243
>At1g55950 hypothetical protein
Length = 203
Score = 28.5 bits (62), Expect = 9.9
Identities = 28/100 (28%), Positives = 43/100 (43%), Gaps = 11/100 (11%)
Query: 179 IQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTN----------IC 228
+Q L+N + KMETEE K G ++ A + VAA D++ IC
Sbjct: 38 LQPLENPPTESYGDDEKMETEEEGEKYLGKNLPASAFIIVAAERHDSDSYSETELEKKIC 97
Query: 229 GGIIQLYWNNILGRCVDFNTQVRQSALKIVE-VVLRQGLV 267
G ++ N+ D + Q L + +VL QG+V
Sbjct: 98 GSVVVAKMNDDKMVSTDTKKKYFQRILSDDDKIVLLQGMV 137
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,830,557
Number of Sequences: 26719
Number of extensions: 499828
Number of successful extensions: 1161
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 1150
Number of HSP's gapped (non-prelim): 17
length of query: 542
length of database: 11,318,596
effective HSP length: 104
effective length of query: 438
effective length of database: 8,539,820
effective search space: 3740441160
effective search space used: 3740441160
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC149305.4


