

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149305.1 + phase: 0 /pseudo
(146 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
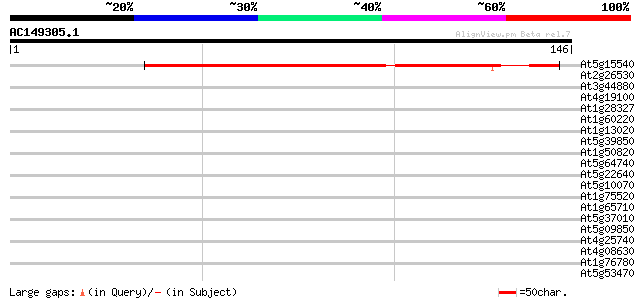
Score E
Sequences producing significant alignments: (bits) Value
At5g15540 putative protein 107 2e-24
At2g26530 AR781, similar to yeast pheromone receptor 33 0.070
At3g44880 lethal leaf-spot 1 homolog Lls1 31 0.20
At4g19100 putative protein 30 0.45
At1g28327 putative protein 30 0.59
At1g60220 unknown protein 29 0.78
At1g13020 eukaryotic initiation factor 4B (EIF4B5) 29 0.78
At5g39850 40S ribosomal protein S9-like 29 1.0
At1g50820 hypothetical protein 29 1.0
At5g64740 cellulose synthase 28 1.3
At5g22640 unknown protein 28 1.3
At5g10070 unknown protein 28 1.3
At1g75520 lateral root primordium protein LRP1 like 28 1.3
At1g65710 hypothetical protein 28 1.7
At5g37010 serine-rich protein - like 28 2.3
At5g09850 unknown protein 28 2.3
At4g25740 putative ribosomal protein S10 28 2.3
At4g08630 hypothetical protein 28 2.3
At1g76780 putative heat shock protein 28 2.3
At5g53470 unknown protein 27 2.9
>At5g15540 putative protein
Length = 1755
Score = 107 bits (268), Expect = 2e-24
Identities = 54/112 (48%), Positives = 78/112 (69%), Gaps = 13/112 (11%)
Query: 36 SEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKR 95
+EP KPGD SRQS+ F++ E++ LP++ Q+L+QRYQEFKNA++EDTVD+++Y+ N+KR
Sbjct: 1642 TEPLKPGDPLSRQSVAFDLSETRTDLPSTYQDLVQRYQEFKNAMREDTVDFTIYSTNVKR 1701
Query: 96 KRPTQTPRKVRKSGPPMVGGDNDEDDDED----WAGGSRNISFSGGRRSSLR 143
KRP TPRK +S V + D+DDD++ W GG GGR ++ R
Sbjct: 1702 KRP--TPRKTSRSAKKTVAYNEDDDDDDNDDRGWHGG-------GGRGAARR 1744
>At2g26530 AR781, similar to yeast pheromone receptor
Length = 317
Score = 32.7 bits (73), Expect = 0.070
Identities = 24/78 (30%), Positives = 34/78 (42%), Gaps = 12/78 (15%)
Query: 49 SIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKRKRPTQTPRKVRKS 108
S P + S PTSP+ + Y+EF+ A + D T + TPRK+
Sbjct: 10 SSPRQLSGCFLSAPTSPRRFNEFYREFEEAATRNFSD--RLTVPFDWEETPGTPRKI--- 64
Query: 109 GPPMVGGDNDEDDDEDWA 126
ND+DDD D+A
Sbjct: 65 -------TNDDDDDIDFA 75
>At3g44880 lethal leaf-spot 1 homolog Lls1
Length = 537
Score = 31.2 bits (69), Expect = 0.20
Identities = 22/84 (26%), Positives = 36/84 (42%), Gaps = 10/84 (11%)
Query: 37 EPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEF-KNALKEDTVDYSLYTANIKR 95
EPP+ +P + + +FS T ++L Y +N +D++ + +R
Sbjct: 214 EPPR---------LPDDFDKPEFSTVTIQRDLFYGYDTLMENVSDPSHIDFAHHKVTGRR 264
Query: 96 KRPTQTPRKVRKSGPPMVGGDNDE 119
R P KV SGP G ND+
Sbjct: 265 DRAKPLPFKVESSGPWGFQGANDD 288
>At4g19100 putative protein
Length = 214
Score = 30.0 bits (66), Expect = 0.45
Identities = 11/23 (47%), Positives = 16/23 (68%)
Query: 102 PRKVRKSGPPMVGGDNDEDDDED 124
P+K +KS P G +DEDDD++
Sbjct: 74 PKKTKKSKKPKPGNQSDEDDDDE 96
>At1g28327 putative protein
Length = 284
Score = 29.6 bits (65), Expect = 0.59
Identities = 19/49 (38%), Positives = 23/49 (46%), Gaps = 10/49 (20%)
Query: 93 IKRKRPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGRRSS 141
I R P + RKVRK DDDE W+GGS+ + G SS
Sbjct: 54 IFRPIPFKGMRKVRKH----------VDDDEGWSGGSKKVKVRGLTSSS 92
>At1g60220 unknown protein
Length = 604
Score = 29.3 bits (64), Expect = 0.78
Identities = 16/52 (30%), Positives = 25/52 (47%), Gaps = 7/52 (13%)
Query: 73 QEFKNALKEDTVDYSLYTANIKRKRPTQTPRKVRKSGPPMVGGDNDEDDDED 124
++F + LKE N K K P R + S ++ D+D+DDD+D
Sbjct: 248 KQFDSGLKESK-------GNKKSKEPYGKKRPMESSTYSLIDDDDDDDDDDD 292
>At1g13020 eukaryotic initiation factor 4B (EIF4B5)
Length = 549
Score = 29.3 bits (64), Expect = 0.78
Identities = 16/35 (45%), Positives = 17/35 (47%), Gaps = 6/35 (17%)
Query: 106 RKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGRRS 140
R SGPP + ED D W GG GGRRS
Sbjct: 108 RSSGPPGRMSRDREDSDGSWGGG------GGGRRS 136
>At5g39850 40S ribosomal protein S9-like
Length = 197
Score = 28.9 bits (63), Expect = 1.0
Identities = 17/41 (41%), Positives = 24/41 (58%), Gaps = 2/41 (4%)
Query: 84 VDYSLYTANIKRKRPTQTPRKVRKSGPPMV-GGDNDEDDDE 123
VD+SL T+ RP + R+ ++G GGD DEDD+E
Sbjct: 158 VDFSL-TSPFGGGRPGRVKRRNERAGAKKASGGDGDEDDEE 197
>At1g50820 hypothetical protein
Length = 528
Score = 28.9 bits (63), Expect = 1.0
Identities = 27/96 (28%), Positives = 41/96 (42%), Gaps = 11/96 (11%)
Query: 29 VSMMLDVSEPPKPGDVFSRQSIPFN--IGESQFSLPTSPQELIQRYQEFKNALKEDTVDY 86
+S + + EP + G V S +++ IG+ S TS ++ N ED +
Sbjct: 422 ISCVTPMCEPQRKGFVESDETLKSRTIIGDDT-SFVTSGSKM--------NRTSEDEMKI 472
Query: 87 SLYTANIKRKRPTQTPRKVRKSGPPMVGGDNDEDDD 122
+ + N +RK Q R V DNDEDDD
Sbjct: 473 AENSTNKRRKYMKQAHRGTMGLCQKQVPSDNDEDDD 508
>At5g64740 cellulose synthase
Length = 1084
Score = 28.5 bits (62), Expect = 1.3
Identities = 21/55 (38%), Positives = 25/55 (45%), Gaps = 7/55 (12%)
Query: 98 PTQTPRKVRKSGPPMVGGDNDEDD----DEDWAGGSRNISF---SGGRRSSLRNS 145
P R R G P V GD +EDD D ++ G+ I F S G S RNS
Sbjct: 82 PQCKTRFKRLKGSPRVEGDEEEDDIDDLDNEFEYGNNGIGFDQVSEGMSISRRNS 136
>At5g22640 unknown protein
Length = 871
Score = 28.5 bits (62), Expect = 1.3
Identities = 14/28 (50%), Positives = 18/28 (64%)
Query: 114 GGDNDEDDDEDWAGGSRNISFSGGRRSS 141
G D+D+DDD+D G S S GRR+S
Sbjct: 692 GDDDDDDDDDDDLGPSSFGSADKGRRNS 719
>At5g10070 unknown protein
Length = 264
Score = 28.5 bits (62), Expect = 1.3
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 4/60 (6%)
Query: 66 QELIQRYQEFKNALKEDTVDYSLYTANIK-RKRPTQTPRKVRKSGPPMVGGDNDEDDDED 124
+++++ Y + +N+ + +V S T + K+P V+K G GD E+DDED
Sbjct: 185 KDILKEYSKCENSAEIISVQNSWLTQQTQIPKQPPALEEHVKKDGE---SGDESEEDDED 241
>At1g75520 lateral root primordium protein LRP1 like
Length = 346
Score = 28.5 bits (62), Expect = 1.3
Identities = 13/33 (39%), Positives = 20/33 (60%), Gaps = 4/33 (12%)
Query: 97 RPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGGS 129
R TQ +++R++ GGDN++D D GGS
Sbjct: 184 RETQNAKRLREAS----GGDNNDDKDHSGGGGS 212
>At1g65710 hypothetical protein
Length = 455
Score = 28.1 bits (61), Expect = 1.7
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 5/51 (9%)
Query: 95 RKRPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGRRSSLRNS 145
R+R +TP + R+ G + + GGSR +S S +RS +RN+
Sbjct: 166 RERRRRTPSRERERS-----GSRERGNVGGGGGGSRRVSRSPAKRSEIRNA 211
>At5g37010 serine-rich protein - like
Length = 637
Score = 27.7 bits (60), Expect = 2.3
Identities = 16/48 (33%), Positives = 23/48 (47%), Gaps = 1/48 (2%)
Query: 95 RKRPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGRRSSL 142
R+R +TP + R G + + GGSR +S S GRRS +
Sbjct: 178 RERRRRTPSRERDDSKSNRSGSRERGSSGN-GGGSRRVSRSPGRRSEI 224
>At5g09850 unknown protein
Length = 353
Score = 27.7 bits (60), Expect = 2.3
Identities = 19/69 (27%), Positives = 34/69 (48%), Gaps = 2/69 (2%)
Query: 36 SEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKR 95
++P +P S+Q+IP + + T+ + L Q Y++ +NA K+ T+ K
Sbjct: 277 AKPSRPSP--SQQTIPRDKEHKEVDFDTARKRLQQNYRQAENAKKQRTIQVMDIHDIPKP 334
Query: 96 KRPTQTPRK 104
K+ PRK
Sbjct: 335 KKGGFFPRK 343
>At4g25740 putative ribosomal protein S10
Length = 177
Score = 27.7 bits (60), Expect = 2.3
Identities = 21/68 (30%), Positives = 27/68 (38%), Gaps = 1/68 (1%)
Query: 79 LKEDTVDYSLYTANIKRKRPTQTPRKVRKSGPPMVGGDNDEDDDED-WAGGSRNISFSGG 137
L D V +L + RP P R+ GPP GD D D + GG R GG
Sbjct: 82 LPSDVVPATLKKSAKPGGRPFGGPPGDRQRGPPRSDGDRPRFGDRDGYRGGPRGGDEKGG 141
Query: 138 RRSSLRNS 145
+ + S
Sbjct: 142 APADFQPS 149
>At4g08630 hypothetical protein
Length = 779
Score = 27.7 bits (60), Expect = 2.3
Identities = 14/46 (30%), Positives = 23/46 (49%), Gaps = 3/46 (6%)
Query: 93 IKRKRPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGGSRNISFSGGR 138
+ + P + P V + P G++D D DE + G +I +GGR
Sbjct: 256 LTKNPPLRHPHAVAQ---PPSNGESDVDYDESYTSGMPSIGLAGGR 298
>At1g76780 putative heat shock protein
Length = 1871
Score = 27.7 bits (60), Expect = 2.3
Identities = 20/72 (27%), Positives = 33/72 (45%), Gaps = 6/72 (8%)
Query: 66 QELIQ-RYQEFKNALKEDTVDYSLYT-----ANIKRKRPTQTPRKVRKSGPPMVGGDNDE 119
+EL++ + K +K+ DY L + ++ + RK+ KS +G DE
Sbjct: 1172 EELVETEISDHKEKVKKKDEDYILRSQDTGKVDLGERERRSKQRKIHKSVEDEIGDQEDE 1231
Query: 120 DDDEDWAGGSRN 131
D +E A SRN
Sbjct: 1232 DAEEAAAVVSRN 1243
>At5g53470 unknown protein
Length = 338
Score = 27.3 bits (59), Expect = 2.9
Identities = 9/13 (69%), Positives = 10/13 (76%)
Query: 115 GDNDEDDDEDWAG 127
GD DE DD+DW G
Sbjct: 76 GDEDESDDDDWEG 88
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.329 0.143 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,628,278
Number of Sequences: 26719
Number of extensions: 160103
Number of successful extensions: 1304
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 28
Number of HSP's that attempted gapping in prelim test: 1260
Number of HSP's gapped (non-prelim): 62
length of query: 146
length of database: 11,318,596
effective HSP length: 90
effective length of query: 56
effective length of database: 8,913,886
effective search space: 499177616
effective search space used: 499177616
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC149305.1


