

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
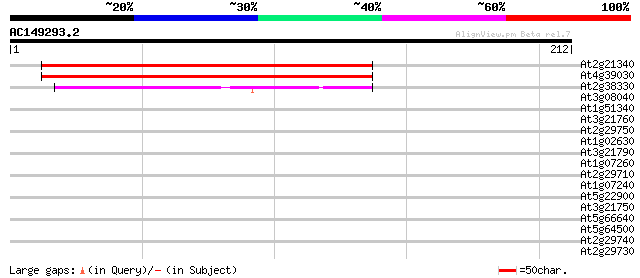
Query= AC149293.2 - phase: 2
(212 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g21340 unknown protein 204 2e-53
At4g39030 enhanced disease susceptibility 5 gene (EDS5) 179 1e-45
At2g38330 unknown protein 54 5e-08
At3g08040 putative integral membrane protein 35 0.029
At1g51340 putative protein 33 0.11
At3g21760 putative UDP-glucose glucosyltransferase 31 0.42
At2g29750 putative flavonol 3-O-glucosyltransferase 30 0.94
At1g02630 unknown protein 30 0.94
At3g21790 putative UDP-glucose glucosyltransferase 30 1.2
At1g07260 unknown protein 29 1.6
At2g29710 putative flavonol 3-O-glucosyltransferase 29 2.1
At1g07240 unknown protein 28 3.6
At5g22900 Na+/H+ antiporter-like protein 28 4.6
At3g21750 putative UDP-glucose glucosyltransferase 28 4.6
At5g66640 putative protein 27 6.1
At5g64500 unknown protein 27 7.9
At2g29740 putative flavonol 3-O-glucosyltransferase 27 7.9
At2g29730 putative flavonol 3-O-glucosyltransferase 27 7.9
>At2g21340 unknown protein
Length = 414
Score = 204 bits (520), Expect = 2e-53
Identities = 99/125 (79%), Positives = 110/125 (87%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L GWVAQSASLGMKDSWGPLKALA AS INGVGD+VLCT+LGYGIAGAAWATM SQVVAA
Sbjct: 193 LIGWVAQSASLGMKDSWGPLKALAVASAINGVGDVVLCTFLGYGIAGAAWATMVSQVVAA 252
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM LN KGY+AF+ +PS E +TI GLAAPVF+TMMSKV FY+LL+YFATSMGT+
Sbjct: 253 YMMMDALNKKGYSAFSFCVPSPSELLTIFGLAAPVFITMMSKVLFYTLLVYFATSMGTNI 312
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 313 IAAHQ 317
>At4g39030 enhanced disease susceptibility 5 gene (EDS5)
Length = 543
Score = 179 bits (453), Expect = 1e-45
Identities = 86/125 (68%), Positives = 106/125 (84%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L G VAQSASLGMK+SWGPLKALAAA++ING+GD +LC +LG GIAGAAWAT ASQ+V+A
Sbjct: 235 LVGLVAQSASLGMKNSWGPLKALAAATIINGLGDTILCLFLGQGIAGAAWATTASQIVSA 294
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM +LN +GYNA++ +IPS +E I LAAPVF+++ SK+AFYS +IY ATSMGTH
Sbjct: 295 YMMMDSLNKEGYNAYSFAIPSPQELWKISALAAPVFISIFSKIAFYSFIIYCATSMGTHV 354
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 355 LAAHQ 359
>At2g38330 unknown protein
Length = 521
Score = 54.3 bits (129), Expect = 5e-08
Identities = 44/122 (36%), Positives = 62/122 (50%), Gaps = 6/122 (4%)
Query: 18 AQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMMR 77
AQ A G KD+ PL A+ A +V+N V D +L LG+GI+GAA AT+ S+ + A++++
Sbjct: 228 AQGAFRGFKDTTTPLYAVVAGNVLNAVLDPILIFVLGFGISGAAAATVISEYLIAFILLW 287
Query: 78 TLNMKGYNAFALS--IPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAA 135
LN N LS I GR + + T+ V F +L A G MA
Sbjct: 288 KLN---ENVVLLSPQIKVGRANQYLKSGGLLIGRTVALLVPF-TLATSLAAQNGPTQMAG 343
Query: 136 HQ 137
HQ
Sbjct: 344 HQ 345
>At3g08040 putative integral membrane protein
Length = 526
Score = 35.0 bits (79), Expect = 0.029
Identities = 49/152 (32%), Positives = 62/152 (40%), Gaps = 23/152 (15%)
Query: 7 PAGLLYLCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMA 66
PA LL L Q G KD+ PL A A VIN V D + L GI GAA A +
Sbjct: 224 PALLLSLA---MQGIFRGFKDTKTPLFATVVADVINIVLDPIFIFVLRLGIIGAAIAHVI 280
Query: 67 SQVVAAYMMMRTLNMK--------GYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFY 118
SQ ++ L K G F + +G +L LA + +T
Sbjct: 281 SQYFMTLILFVFLAKKVNLIPPNFGDLQFGRFLKNG-----LLLLARTIAVTFCQ----- 330
Query: 119 SLLIYFATSMGTHTMAAHQ--YGVNLSRKLLN 148
+L A +GT MAA Q V L+ LLN
Sbjct: 331 TLAAAMAARLGTTPMAAFQICLQVWLTSSLLN 362
>At1g51340 putative protein
Length = 509
Score = 33.1 bits (74), Expect = 0.11
Identities = 37/135 (27%), Positives = 57/135 (41%), Gaps = 5/135 (3%)
Query: 3 SRSIPAGLLYLCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAW 62
S PA LL L AQ G KD+ PL A V N + D + G+ GAA
Sbjct: 210 SLGAPAVLLSLA---AQGVFRGFKDTTTPLFATVIGDVTNIILDPIFIFVFRLGVTGAAT 266
Query: 63 ATMASQVVAAYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLI 122
A + SQ + +++ L M + F +S +F + + M +++ +L
Sbjct: 267 AHVISQYLMCGILLWKL-MGQVDIFNMS-TKHLQFCRFMKNGFLLLMRVIAVTFCVTLSA 324
Query: 123 YFATSMGTHTMAAHQ 137
A G+ +MAA Q
Sbjct: 325 SLAAREGSTSMAAFQ 339
>At3g21760 putative UDP-glucose glucosyltransferase
Length = 485
Score = 31.2 bits (69), Expect = 0.42
Identities = 20/73 (27%), Positives = 33/73 (44%), Gaps = 8/73 (10%)
Query: 15 GWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTY------LGYGIAGAAWATMASQ 68
G++ ++A +G W P A+ A I G + C + L +G+ A W A Q
Sbjct: 339 GFLERTAEIGKIVGWAPQSAILANPAIGGF--VSHCGWNSTLESLWFGVPMATWPLYAEQ 396
Query: 69 VVAAYMMMRTLNM 81
V A+ M+ L +
Sbjct: 397 QVNAFEMVEELGL 409
>At2g29750 putative flavonol 3-O-glucosyltransferase
Length = 481
Score = 30.0 bits (66), Expect = 0.94
Identities = 18/69 (26%), Positives = 29/69 (41%), Gaps = 9/69 (13%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
+CGW Q L K A ++ G + LG+G+ A W A Q + A
Sbjct: 348 VCGWAPQVEILAHK---------AVGGFVSHCGWNSILESLGFGVPIATWPMYAEQQLNA 398
Query: 73 YMMMRTLNM 81
+ M++ L +
Sbjct: 399 FTMVKELGL 407
>At1g02630 unknown protein
Length = 389
Score = 30.0 bits (66), Expect = 0.94
Identities = 13/49 (26%), Positives = 24/49 (48%)
Query: 15 GWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWA 63
G++A++ + SW P+ + ++ + VG + YL I A WA
Sbjct: 256 GFIAENLKSQLLQSWYPILLITVYNISDFVGKSLTALYLWQSIKSATWA 304
>At3g21790 putative UDP-glucose glucosyltransferase
Length = 495
Score = 29.6 bits (65), Expect = 1.2
Identities = 19/73 (26%), Positives = 32/73 (43%), Gaps = 8/73 (10%)
Query: 15 GWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTY------LGYGIAGAAWATMASQ 68
G+ ++ +G W P A+ A I G + C + L +G+ AAW A Q
Sbjct: 336 GFFDRTKDIGKVIGWAPQVAVLANPAIGGF--VTHCGWNSTLESLWFGVPTAAWPLYAEQ 393
Query: 69 VVAAYMMMRTLNM 81
A++M+ L +
Sbjct: 394 KFNAFLMVEELGL 406
>At1g07260 unknown protein
Length = 476
Score = 29.3 bits (64), Expect = 1.6
Identities = 18/71 (25%), Positives = 33/71 (46%), Gaps = 4/71 (5%)
Query: 15 GWVAQSASLGMKDSWGPLKALAAASVINGV----GDIVLCTYLGYGIAGAAWATMASQVV 70
G++ ++AS G+ W P + A + G G + L +G+ A W A Q +
Sbjct: 334 GFLDRTASKGLVCDWAPQVEVLAHKALGGFVSHCGWNSVLESLWFGVPIATWPMYAEQQL 393
Query: 71 AAYMMMRTLNM 81
A+ M++ L +
Sbjct: 394 NAFSMVKELGL 404
>At2g29710 putative flavonol 3-O-glucosyltransferase
Length = 467
Score = 28.9 bits (63), Expect = 2.1
Identities = 30/131 (22%), Positives = 52/131 (38%), Gaps = 10/131 (7%)
Query: 15 GWVAQSASLGMKDSWGPLKALAAASVINGV----GDIVLCTYLGYGIAGAAWATMASQVV 70
G++ + + GM W P + A + G G + L +G+ W A Q +
Sbjct: 324 GFMDRVSGRGMICGWSPQVEILAHKAVGGFVSHCGWNSIVESLWFGVPIVTWPMYAEQQL 383
Query: 71 AAYMMMRTLNMK-----GYNAFALSIPSGREFITILGLAAPVFMTMMSK-VAFYSLLIYF 124
A++M++ L + Y+ + I S E T + ++ K V S +I
Sbjct: 384 NAFLMVKELKLAVELKLDYSVHSGEIVSANEIETAISCVMNKDNNVVRKRVMDISQMIQR 443
Query: 125 ATSMGTHTMAA 135
AT G + AA
Sbjct: 444 ATKNGGSSFAA 454
>At1g07240 unknown protein
Length = 480
Score = 28.1 bits (61), Expect = 3.6
Identities = 19/71 (26%), Positives = 32/71 (44%), Gaps = 4/71 (5%)
Query: 15 GWVAQSASLGMKDSWGP----LKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVV 70
G+V ++ G+ SW P L A ++ G + L YG+ A W A Q +
Sbjct: 334 GFVDRTMGRGIVCSWAPQVDILAHKATGGFVSHCGWNSVQESLWYGVPIATWPMYAEQQL 393
Query: 71 AAYMMMRTLNM 81
A+ M++ L +
Sbjct: 394 NAFEMVKELGL 404
>At5g22900 Na+/H+ antiporter-like protein
Length = 822
Score = 27.7 bits (60), Expect = 4.6
Identities = 18/56 (32%), Positives = 31/56 (55%), Gaps = 1/56 (1%)
Query: 93 SGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAAHQYGVNLSRKLLN 148
+GR+ ITI GL++ + T++ V F+ L T HT+ + +Y V S + L+
Sbjct: 146 TGRKAITI-GLSSVLLSTLVCSVIFFGNLRDVGTKNSDHTLNSLEYVVIYSIQCLS 200
>At3g21750 putative UDP-glucose glucosyltransferase
Length = 473
Score = 27.7 bits (60), Expect = 4.6
Identities = 18/71 (25%), Positives = 32/71 (44%), Gaps = 4/71 (5%)
Query: 15 GWVAQSASLGMKDSWGP----LKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVV 70
G++ ++ +G SW P L + A + + G + L +G+ AAW A Q
Sbjct: 327 GFLDRTVEIGKIISWAPQVDVLNSPAIGAFVTHCGWNSILESLWFGVPMAAWPIYAEQQF 386
Query: 71 AAYMMMRTLNM 81
A+ M+ L +
Sbjct: 387 NAFHMVDELGL 397
>At5g66640 putative protein
Length = 451
Score = 27.3 bits (59), Expect = 6.1
Identities = 13/38 (34%), Positives = 20/38 (52%), Gaps = 5/38 (13%)
Query: 147 LNHLCLNCYMELIGICQRPGCF*DLLQ*SELHLDCY*E 184
+N CL C+ C RP ++L+ + H+DCY E
Sbjct: 93 VNPRCLCCFH-----CHRPFVMHEILKKGKFHIDCYKE 125
>At5g64500 unknown protein
Length = 484
Score = 26.9 bits (58), Expect = 7.9
Identities = 36/132 (27%), Positives = 57/132 (42%), Gaps = 25/132 (18%)
Query: 13 LCGWVAQSASLGMKDSWGP-----LKALAAASVINGVGDIVLCTYLGYGIAGAAWATMAS 67
+ G++A + LG WGP + + A +I G G V+C GI G T++
Sbjct: 278 ILGYIAYNFVLGAYSYWGPKAGYNIYKMENADMIFG-GVTVVC-----GIVG----TLSG 327
Query: 68 QVVAAYMMMRTLNMKGYNAFALSIPSGREFI-TILGLAAPVFMTMMSKVAFYSL--LIYF 124
V+ YM N A + S FI I AA F +M + +A +++ L+ F
Sbjct: 328 GVILDYMDATISN-------AFKVLSVSTFIGAIFCFAAFCFKSMYAFLALFAVGELLVF 380
Query: 125 ATSMGTHTMAAH 136
AT + + H
Sbjct: 381 ATQGPVNFIVLH 392
>At2g29740 putative flavonol 3-O-glucosyltransferase
Length = 474
Score = 26.9 bits (58), Expect = 7.9
Identities = 17/71 (23%), Positives = 31/71 (42%), Gaps = 4/71 (5%)
Query: 15 GWVAQSASLGMKDSWGPLKALAAASVINGV----GDIVLCTYLGYGIAGAAWATMASQVV 70
G++ + LG+ W P + A I G G + L +G+ A W A Q +
Sbjct: 337 GFMNRVMGLGLVCGWAPQVEILAHKAIGGFVSHCGWNSILESLRFGVPIATWPMYAEQQL 396
Query: 71 AAYMMMRTLNM 81
A+ +++ L +
Sbjct: 397 NAFTIVKELGL 407
>At2g29730 putative flavonol 3-O-glucosyltransferase
Length = 467
Score = 26.9 bits (58), Expect = 7.9
Identities = 29/129 (22%), Positives = 48/129 (36%), Gaps = 15/129 (11%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
+CGW Q L K G + S++ L +G+ W A Q + A
Sbjct: 335 ICGWSPQVEILAHKAVGGFVSHCGWNSIVES---------LWFGVPIVTWPMYAEQQLNA 385
Query: 73 YMMMRTLNMK-----GYNAFALSIPSGREFITILGLAAPVFMTMMSK-VAFYSLLIYFAT 126
++M++ L + Y + I + E T + ++ K V S +I AT
Sbjct: 386 FLMVKELKLAVELKLDYRVHSDEIVNANEIETAIRYVMDTDNNVVRKRVMDISQMIQRAT 445
Query: 127 SMGTHTMAA 135
G + AA
Sbjct: 446 KNGGSSFAA 454
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.338 0.146 0.479
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,092,625
Number of Sequences: 26719
Number of extensions: 141992
Number of successful extensions: 476
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 468
Number of HSP's gapped (non-prelim): 20
length of query: 212
length of database: 11,318,596
effective HSP length: 95
effective length of query: 117
effective length of database: 8,780,291
effective search space: 1027294047
effective search space used: 1027294047
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC149293.2


