

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149206.10 - phase: 0
(427 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
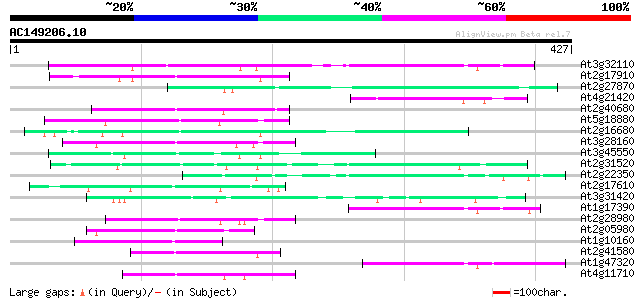
Score E
Sequences producing significant alignments: (bits) Value
At3g32110 non-LTR reverse transcriptase, putative 95 8e-20
At2g17910 putative non-LTR retroelement reverse transcriptase 73 3e-13
At2g27870 putative non-LTR retroelement reverse transcriptase 69 6e-12
At4g21420 hypothetical protein 68 8e-12
At2g40680 putative non-LTR retroelement reverse transcriptase 68 8e-12
At5g18880 SAE1-S9-protein - like 67 1e-11
At2g16680 putative non-LTR retroelement reverse transcriptase 67 2e-11
At3g28160 non-LTR reverse transcriptase, putative 66 4e-11
At3g45550 putative protein 65 7e-11
At2g31520 putative non-LTR retroelement reverse transcriptase 64 2e-10
At2g22350 putative non-LTR retroelement reverse transcriptase 64 2e-10
At2g17610 putative non-LTR retroelement reverse transcriptase 62 5e-10
At3g31420 hypothetical protein 62 6e-10
At1g17390 hypothetical protein 60 2e-09
At2g28980 putative non-LTR retroelement reverse transcriptase 60 2e-09
At2g05980 putative non-LTR retroelement reverse transcriptase 59 4e-09
At1g10160 putative reverse transcriptase 59 5e-09
At2g41580 putative non-LTR retroelement reverse transcriptase 59 7e-09
At1g47320 hypothetical protein 59 7e-09
At4g11710 putative protein 58 9e-09
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 94.7 bits (234), Expect = 8e-20
Identities = 102/390 (26%), Positives = 162/390 (41%), Gaps = 43/390 (11%)
Query: 30 WILSDFFAY-KDNALVEKIHQIALPLDETLDKLIWTDSVDGDLSNKLAFSFLPGH-GPTV 87
W +S + DN ++ + I + D+L W + DG + K AF+ L P
Sbjct: 1514 WDMSQIAPFVTDNKRLDLLAVIVDSVTGAHDRLAWGMTSDGRFTVKSAFAMLTNDDSPRQ 1573
Query: 88 HWAKM---LWNAYTPPTGAFITWRFLHNKLPTDDNLRKRGCYIVSICCCFCRKQAETSSH 144
+ + +W P W ++ + T+ RKR S C CR E+ H
Sbjct: 1574 DMSSLYGRVWKVQAPERVRVFLWLVVNQAIMTNSE-RKRRHLCDSDVCQVCRGGIESILH 1632
Query: 145 IFLQCPVTLQLWDWLLKATDQHLDFS-SILN--ISRMVQHVMNSA-------IVHIMWSI 194
+ CP +WD ++ Q F+ S+L S + Q +M + + W
Sbjct: 1633 VLRDCPAMSGIWDRIVPRRLQQSFFTMSLLEWLYSNLRQGLMTEGSDWSTMFAMAVWWGW 1692
Query: 195 WLECNNKYFDGVQKPMSTLFNTILAEVLRLSFMLDIVKGASSMQDFKLARLFSIPFKTNR 254
C+N + + F LA + +++ ++ + +L+ L R
Sbjct: 1693 KWRCSNIFGENKTCRDRVRFIKDLAVEVSIAYSREV--------ELRLSGL--------R 1736
Query: 255 VNPCREIIWVPPHGGCMKINCDGSVVGSPSCGSIGVIFRASQTMFCGAFAQNIGYATALE 314
VN + I W PP G KIN DG+ G+P S G + R S +CG FA NIG +A
Sbjct: 1737 VN--KPIRWTPPMEGWYKINTDGASRGNPGLASAGGVLRNSAGAWCGGFAVNIGRCSAPL 1794
Query: 315 AEYSACMFAIEKAKELHLTNIWIETDSVNVIRAFHFNTGV----PWKMHIR-WHNCLLFC 369
AE + + A LT++ +E DS V+ TG+ P +R HN L
Sbjct: 1795 AELWGVYYGLYMAWAKQLTHLELEVDSEVVVG--FLKTGIGETHPLSFLVRLCHNFLSKD 1852
Query: 370 RSIRSLCTHVNREGNLVADALAKNGQGLAL 399
++R +HV RE N +AD LA + L+L
Sbjct: 1853 WTVR--ISHVYREANSLADGLANHAFSLSL 1880
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 73.2 bits (178), Expect = 3e-13
Identities = 56/204 (27%), Positives = 83/204 (40%), Gaps = 26/204 (12%)
Query: 31 ILSDFFAYKDNALVEKIHQIALPLDETLDKLIWTDSVDGDLSNKLAFSFLPG--HGPTVH 88
+L D F +KD ++ K PL D W S +G S K + FL H
Sbjct: 954 MLRDLFPWKDVEIILKQR----PLFFKEDSFCWLHSHNGLYSVKTGYEFLSKQVHHRLYQ 1009
Query: 89 WAKM----------LWNAYTPPTGAFITWRFLHNKLPTDDNLRKRGCYIVSICCCFCRKQ 138
AK+ +WN +T P W+ LH +P +D LR RG C C +
Sbjct: 1010 EAKVKPSVNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIR-SDDGCLMCDTE 1068
Query: 139 AETSSHIFLQCPVTLQLWDWL-LKATDQHLDFSSILNISRMVQHVMNSAIVH-------- 189
ET +HI +CP+ Q+W L + S N+SR++ + + H
Sbjct: 1069 NETINHILFECPLARQVWAITHLSSAGSEFSNSVYTNMSRLIDLTQQNDLPHHLRFVSPW 1128
Query: 190 IMWSIWLECNNKYFDGVQKPMSTL 213
I+W +W N F+G +TL
Sbjct: 1129 ILWFLWKNRNALLFEGKGSITTTL 1152
>At2g27870 putative non-LTR retroelement reverse transcriptase
Length = 314
Score = 68.6 bits (166), Expect = 6e-12
Identities = 82/309 (26%), Positives = 118/309 (37%), Gaps = 36/309 (11%)
Query: 121 RKRGCYIVSICCCFCRKQAETSSHIFLQCPVTLQLWDWLLKA-------TDQHLD--FSS 171
R+R S C C+ +T HI CP +W L+ A T L+ F++
Sbjct: 11 RRRRHLSDSDICQICKGAEKTIIHILRDCPAMEGIWIRLVPAGKRREFFTQSLLEWLFAN 70
Query: 172 ILNISRMVQHVMNSAIVHIMWSIWL-ECNNKYFDGVQKPMSTLFNTILAEVLRLSFMLDI 230
+ + + + ++ +W W C N + GVQ + S I
Sbjct: 71 LGDRRKTCESTWSTLFALSIWWAWKWRCGNIF--GVQDKCRDRVRFLKDLARETSMAHVI 128
Query: 231 VKGASSMQDFKLARLFSIPFKTNRVNPCREIIWVPPHGGCMKINCDGSVVGSPSCGSIGV 290
V+ S ++ RL I W P G K+N DG+ G+P S G
Sbjct: 129 VRTLSGGHGERVERL---------------IAWSKPEEGWWKLNTDGASRGNPGLASAGG 173
Query: 291 IFRASQTMFCGAFAQNIGYATALEAEYSACMFAIEKAKELHLTNIWIETDSVNVIRAFHF 350
+ R + + G FA NIG +A AE + + A E +T + IE DS V+
Sbjct: 174 VLRDEEGAWRGGFALNIGVCSAPLAELWGVYYGLYIAWERRVTRLEIEVDSEIVVGFLKI 233
Query: 351 NTGVPWKMHIRWHNCLLF-CRSIRSLCTHVNREGNLVADALAKNGQGLALYS-SHPLAFI 408
+ C F R R +HV RE N +AD GLA Y+ S PL F
Sbjct: 234 GINEVHPLSFLVRLCHDFISRDWRVRISHVYREANRLAD-------GLANYAFSLPLGFH 286
Query: 409 SSFYVRDCL 417
S V D L
Sbjct: 287 SLSLVPDSL 295
>At4g21420 hypothetical protein
Length = 229
Score = 68.2 bits (165), Expect = 8e-12
Identities = 49/146 (33%), Positives = 75/146 (50%), Gaps = 21/146 (14%)
Query: 260 EIIWVPPHGGCMKINCDGSVVGSPSCGSIGVIFRASQTMFCGAFAQNIGYATALEAEYSA 319
+++W P G +K+N DGS G SIG +FR + F ++++IG AT+ AE++A
Sbjct: 60 KVVWKKPENGRIKLNFDGSR-GREGQASIGGVFRNHKAEFLLGYSESIGEATSTMAEFAA 118
Query: 320 CMFAIEKAKELHLTNIWIETDSVNV---------IRAFHFNTGVPWKMHI--RWHNCLLF 368
+E A E LT++W+E D+ + +R N V + + +NC+L
Sbjct: 119 LKRGLELALENGLTDLWLEGDAKIIMDIISRRGRLRCEKTNKHVNYIKVVMPELNNCVL- 177
Query: 369 CRSIRSLCTHVNREGNLVADALAKNG 394
+HV REGN VAD LAK G
Sbjct: 178 --------SHVYREGNRVADKLAKLG 195
>At2g40680 putative non-LTR retroelement reverse transcriptase
Length = 296
Score = 68.2 bits (165), Expect = 8e-12
Identities = 42/160 (26%), Positives = 72/160 (44%), Gaps = 11/160 (6%)
Query: 63 WTDSVDGDLSNKLAFSFLPGHGPTVHWAKMLWNAYTPPTGAFITWRFLHNKLPTDDNLRK 122
W + S+ + L P WAK +W + P FITW +H++L T +R
Sbjct: 102 WKTGFKHNFSSNETWKMLRVEKPICRWAKEIWFSEATPKFFFITWLAIHDRLTTGARMRS 161
Query: 123 RGCYIVSICCCFCRKQAETSSHIFLQCPVTLQLWDWLLK---------ATDQHLDFSSIL 173
V C FC + ET +H+F QCP + Q+W+ L+K + D+ + +
Sbjct: 162 WNTQ-VDTTCKFCAEPVETRNHLFFQCPYSTQVWEKLMKGLLQGHYTSSWDRIVTLLTSS 220
Query: 174 NISRMVQHVMNSAIVHIMWSIWLECNNKYFDGVQKPMSTL 213
++ R ++ + +IW E NN+ G + P++ L
Sbjct: 221 SLGRKHLLLLRYTFQIKVHTIWRERNNRR-HGEENPITPL 259
>At5g18880 SAE1-S9-protein - like
Length = 295
Score = 67.4 bits (163), Expect = 1e-11
Identities = 50/203 (24%), Positives = 87/203 (42%), Gaps = 21/203 (10%)
Query: 27 NGEWILSDFFAYKDNALVEKIHQIALPLDET-LDKLIWTDSVDGDL---SNKLAFSFLPG 82
NG+W L + + + +P + D +W ++ L S++ + +
Sbjct: 61 NGDWFLPAARSDNSQLFLAALTMAPVPHESRGQDSFLWRNAAGSYLPSFSSRDTWEQIRV 120
Query: 83 HGPTVHWAKMLWNAYTPPTGAFITWRFLHNKLPTDDNLRKRGCYIVSICCCFCRKQAETS 142
H PTV WAK++W P + ITW +LPT D LR G I S C ET
Sbjct: 121 HSPTVPWAKVVWFKEYIPRFSLITWMSFLERLPTRDRLRGWGMNIPS-SWVLCSNGDETH 179
Query: 143 SHIFLQCPVTLQLWDW------------LLKATDQHLDFSSILNISRMVQHVMNSAIVHI 190
+H+F +C +L +W++ L A+ L + + +++ ++ SA+ H
Sbjct: 180 AHLFFECSFSLAIWEFFASKFRPSPPFGLPAASSWILQLPLRSHSTTILKLLLQSAVYH- 238
Query: 191 MWSIWLECNNKYFDGVQKPMSTL 213
+W E N + F + S+L
Sbjct: 239 ---VWKERNARIFTSISSSASSL 258
>At2g16680 putative non-LTR retroelement reverse transcriptase
Length = 1319
Score = 67.0 bits (162), Expect = 2e-11
Identities = 80/365 (21%), Positives = 135/365 (36%), Gaps = 53/365 (14%)
Query: 12 HNISFSQMKVANYL--VNGEWIL---SDFFAYKDNALVEKIHQIALPLDETLDKLIWTDS 66
H++ MKV++ + + W L ++ F KD L+ +HQ PL + D W +
Sbjct: 904 HSLMNIHMKVSHLIDPLTRNWNLKKLTELFHEKDVQLI--MHQ--RPLISSEDSYCWAGT 959
Query: 67 VDG--------DLSNKLAFSFLPGHG---PTVH-WAKMLWNAYTPPTGAFITWRFLHNKL 114
+G + S++ F L P+V+ +W+ T P W+ L L
Sbjct: 960 NNGLYTVKSGYERSSRETFKNLFKEADVYPSVNPLFDKVWSLETVPKIKVFMWKALKGAL 1019
Query: 115 PTDDNLRKRGCYIVSICCCFCRKQAETSSHIFLQCPVTLQLWDW-LLKATDQHLDFSSIL 173
+D LR RG C FC+++ ET +H+ QCP Q+W L++A S
Sbjct: 1020 AVEDRLRSRGIRTAD-GCLFCKEEIETINHLLFQCPFARQVWALSLIQAPATGFGTSIFS 1078
Query: 174 NISRMVQHVMNSAIVH--------IMWSIWLECNNKYFDGVQKPMSTLFNTILAEVLRLS 225
NI+ ++Q+ N I ++W IW N F G S + E
Sbjct: 1079 NINHVIQNSQNFGIPRHMRTVSPWLLWEIWKNRNKTLFQGTGLTSSEIVAKAYEECNLWI 1138
Query: 226 FMLDIVKGASSMQDFKLARLFSIPFKTNRVNPCREIIWVPPHGGCMKINCDGSVVGSPSC 285
+ G S + K W PP G +K N +
Sbjct: 1139 NAQEKSSGGVSPSEHK---------------------WNPPPAGELKCNIGVAWSRQKQL 1177
Query: 286 GSIGVIFRASQTMFCGAFAQNIGYATAL-EAEYSACMFAIEKAKELHLTNIWIETDSVNV 344
+ + R S ++ +L +A+ + +A+E H + S ++
Sbjct: 1178 AGVSWVLRDSMGQVLLHSRRSYSQVYSLFDAKIKSWDWALESMDHFHFDKVTFAATSHDI 1237
Query: 345 IRAFH 349
I+A H
Sbjct: 1238 IKALH 1242
>At3g28160 non-LTR reverse transcriptase, putative
Length = 630
Score = 65.9 bits (159), Expect = 4e-11
Identities = 50/192 (26%), Positives = 77/192 (40%), Gaps = 19/192 (9%)
Query: 41 NALVEKIHQIALPLDETLDKLIWT---DSVDGDLSNKLAFSFLPGHGPTVHWAKMLWNAY 97
N + I + + D+ +W DS G S+ + + W + +W
Sbjct: 228 NQIENHIELVRQARSQESDRSLWKQKEDSFKGSFSSPKTWQQIRTISNECEWYRGVWFPS 287
Query: 98 TPPTGAFITWRFLHNKLPTDDNLRKRGCYIVSICCCFCRKQAETSSHIFLQCPVTLQLWD 157
+ P +F+TW HN+L T D L K C FC ++ ET H+F CP + Q+W
Sbjct: 288 STPKYSFVTWLAFHNRLATGDRLYKWNSE-ARATCVFCDEELETRDHLFFSCPYSSQIWI 346
Query: 158 WLLKATDQHLDFSS-------ILNISRMVQHVMN-----SAIVHIMWSIWLECNNKYFDG 205
L K + SS +L+ S+ HV A++H S+W E N +
Sbjct: 347 ALAKGLLNGRNVSSWSLITPHLLDSSQPYLHVFTLRYTFQALIH---SLWRERNGRRHGE 403
Query: 206 VQKPMSTLFNTI 217
P S L I
Sbjct: 404 PAIPASKLTKLI 415
>At3g45550 putative protein
Length = 851
Score = 65.1 bits (157), Expect = 7e-11
Identities = 68/276 (24%), Positives = 103/276 (36%), Gaps = 49/276 (17%)
Query: 30 WILSDFFAYKDNALVEKIHQIALPLDETLDKLIWTDSVDGDLSNKLAFSFLPGHGPT--- 86
W + + D + + I +I L D+LIW+ + GD + + + +L H P+
Sbjct: 593 WDNAKLQTFVDQSDHDYIKRIYLSTRSKTDRLIWSYNSTGDYTVRSGY-WLSTHDPSNTI 651
Query: 87 ---------VHWAKMLWNAYTPPTGAFITWRFLHNKLPTDDNLRKRGCYIVSICCCFCRK 137
V +WN P WR L LPT D L RG I C CR+
Sbjct: 652 PTMAKPHGSVDLKTKIWNLPIMPKLKHFLWRILSKALPTTDRLTTRGMRI-DPGCPRCRR 710
Query: 138 QAETSSHIFLQCPVTLQLWDWLLKATDQHLDFSSIL------NISRMVQHVMNSAIVH-- 189
+ E+ +H CP W + +D L SSIL NIS ++ + N+ I
Sbjct: 711 ENESINHALFTCPFATMAW----RLSDTPLYRSSILSNNIEDNISNILLLLQNTTITDSQ 766
Query: 190 ------IMWSIWLECNNKYFDGVQKPMSTLFNTILAEVLRLSFMLDIVKGASSMQDFKLA 243
++W IW NN F+ LR S + +V+ + ++ A
Sbjct: 767 KLIPFWLLWRIWKARNNVVFNN----------------LRESPSITVVRAKAETNEWLNA 810
Query: 244 RLFSIPFK-TNRVNPCREIIWVPPHGGCMKINCDGS 278
P + R WV P +K N D S
Sbjct: 811 TQTQGPRRLPKRTTAAGNTTWVKPQMPYIKCNFDAS 846
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 63.9 bits (154), Expect = 2e-10
Identities = 86/391 (21%), Positives = 150/391 (37%), Gaps = 53/391 (13%)
Query: 32 LSDFFAYKDNALVEKIHQIALPLDETLDKLIWTDSVDGDLSNKLAFSFL----------- 80
+S F D+ IH+I L + DK+IW + G+ + + + L
Sbjct: 1133 ISQFVDQSDHGF---IHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTHDPSTNIPAI 1189
Query: 81 -PGHGPTVHWAKMLWNAYTPPTGAFITWRFLHNKLPTDDNLRKRGCYIVSICCCFCRKQA 139
P HG ++ +WN P WR L L T + L RG I I C C ++
Sbjct: 1190 NPPHG-SIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPI-CPRCHREN 1247
Query: 140 ETSSHIFLQCPVTLQLWDWLLKAT---DQHLDFSSILNISRMVQHVMNSAI--------V 188
E+ +H CP W WL ++ +Q + NIS ++ V ++ + V
Sbjct: 1248 ESINHALFTCPFATMAW-WLSDSSLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPV 1306
Query: 189 HIMWSIWLECNNKYFDGVQKPMSTLFNTILAEVLRLSFMLDIVKGASSMQDFKLARLFSI 248
++W IW NN F+ ++ S T+L+ L+ +
Sbjct: 1307 WLIWRIWKARNNVVFNKFRESPS---KTVLSAKAETHDWLNATQSHK-----------KT 1352
Query: 249 PFKTNRVNPCREIIWVPPHGGCMKINCDGSVVGSPSCGSIGVIFRASQTMFCGAFAQNIG 308
P T ++ +I W P +K N D + G I R + +
Sbjct: 1353 PSPTRQIAE-NKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLA 1411
Query: 309 Y-ATALEAEYSACMFAIEKAKELHLTNIWIETDS---VNVIRAFHFNTGVPWKMHIRWHN 364
+ + LEAE A + A+++ T +++E D +N+I F++ + + +
Sbjct: 1412 HTSNPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLINGISFHSSLANHL----ED 1467
Query: 365 CLLFCRSIRSL-CTHVNREGNLVADALAKNG 394
+ S+ + R+GN +A LAK G
Sbjct: 1468 ISFWANKFASIQFGFIRRKGNKLAHVLAKYG 1498
>At2g22350 putative non-LTR retroelement reverse transcriptase
Length = 321
Score = 63.9 bits (154), Expect = 2e-10
Identities = 75/307 (24%), Positives = 118/307 (38%), Gaps = 39/307 (12%)
Query: 132 CCFCRKQAETSSHIFLQCPVTLQLWDWLLKATDQHLDFSSILNISRMVQHVMNSA----- 186
C C+ ET H+ CP +W L+ DQ F + + + +++
Sbjct: 35 CQICQGGEETILHVLRDCPAMAGIWSRLVPR-DQIRQFFTASLLEWIYKNLRERGSWPTV 93
Query: 187 -IVHIMWSIWLECNNKYFDGVQKPMSTLFNTILAEVLRLSFMLDIVKGASSMQDFKLARL 245
++ + W C N F G K R+ F+ D+ + + F
Sbjct: 94 FVMAVWWGWKWRCGN-IFGGNGKCRD-----------RVKFIKDLAEEVAIANAFVKGN- 140
Query: 246 FSIPFKTNRVNPCREIIWVPPHGGCMKINCDGSVVGSPSCGSIGVIFRASQTMFCGAFAQ 305
+ +RV R + WV P G +K+N DG+ G+P + G + R + G FA
Sbjct: 141 ---EVRVSRVE--RLVSWVSPEDGWVKLNTDGASRGNPGFATAGGVLRDHNGAWIGGFAV 195
Query: 306 NIGYATALEAEYSACMFAIEKAKELHLTNIWIETDSVNVIRAFHFNTGVPWKMHIRWHNC 365
NIG +A AE + + A + +E DS V+ TG+ + +
Sbjct: 196 NIGVCSAPLAELWGVYYGLFIAWGRGARRVELEVDSKMVVG--FLTTGIADSHPLSF--L 251
Query: 366 LLFCRSIRS-----LCTHVNREGNLVADALAKN----GQGLALYSSHPLAFISSFYVRDC 416
L C S +HV RE N +AD LA GL L S P +SS + D
Sbjct: 252 LRLCYDFLSKGWIVRISHVYREANRLADGLANYAFSLSLGLHLLESRP-DVVSSILLDDV 310
Query: 417 LGLPFSR 423
G+ + R
Sbjct: 311 AGVSYPR 317
>At2g17610 putative non-LTR retroelement reverse transcriptase
Length = 773
Score = 62.4 bits (150), Expect = 5e-10
Identities = 57/233 (24%), Positives = 94/233 (39%), Gaps = 48/233 (20%)
Query: 16 FSQMKVANYLVNGEWILSDFFAYKDNALVEKIHQIALPLDETL--------DKLIWTDSV 67
+SQ+KV++ L+ G W ++ L + IHQ +P + D + W +
Sbjct: 411 YSQLKVSDLLIEGRW--------NEDLLCKLIHQNDIPHIRAIRPSITGANDAITWIYTH 462
Query: 68 DGDLSNKLAFSFLPGHGPTVHWA-------------KMLWNAYTPPTGAFITWRFLHNKL 114
DG+ S K + L H + +W PP WR HN L
Sbjct: 463 DGNYSVKSGYHLLRKLSQQQHASLPSPNEVSAQTVFTNIWKQNAPPKIKHFWWRSAHNAL 522
Query: 115 PTDDNLRKRGCYIVSICCCFCRKQAETSSHIFLQCPVTLQLWDWL---------LKATDQ 165
PT NL++R I C C + +E +H+ QC V+ ++W+ L +
Sbjct: 523 PTAGNLKRRR-LITDDTCQRCGEASEDVNHLLFQCRVSKEIWEQAHIKLCPGDSLMSNSF 581
Query: 166 HLDFSSILNISRMVQHVMNSAIVHIMWSIW-----LECNNKYF---DGVQKPM 210
+ + SI +++ + + S I W IW L NNK + D +QK +
Sbjct: 582 NQNLESIQKLNQSARKDV-SLFPFIGWRIWKMRNDLIFNNKRWSIPDSIQKAL 633
>At3g31420 hypothetical protein
Length = 1491
Score = 62.0 bits (149), Expect = 6e-10
Identities = 87/364 (23%), Positives = 133/364 (35%), Gaps = 57/364 (15%)
Query: 59 DKLIWTDSVDGDLSNKLAF---SFLPG----HGPT--VHWAKMLWNAYTPPTGAFITWRF 109
D + W + DG + K A+ + LPG H P + ++LW T P W+
Sbjct: 1125 DLIGWHYTKDGMYTVKSAYWLATHLPGTTGTHPPPGDIKLKQLLWKTKTAPKIKHFCWKI 1184
Query: 110 LHNKLPTDDNLRKRGCYIVSICCCFCRKQAETSSHIFLQCPVTLQLW-----------DW 158
L + T + LR R SIC CR + ETS H+F +C W D
Sbjct: 1185 LSGAIATGEMLRYRHINKQSICKRCCRDE-ETSQHLFFECDYAKATWRGAGLPNLIFQDS 1243
Query: 159 LLKATDQHLDFSSILNISRMVQHVMNSAIVHIMWSIWLECNNKYFDGVQKPMSTLFNTIL 218
++ ++ F ++ + + I+W +W N F P
Sbjct: 1244 IVTLEEK---FRAMFTFNPSTTNYWRQLPFWILWRLWRSRNILTFQQKHIPWEVTVQLAK 1300
Query: 219 AEVLRLSFMLDIVKGASSMQDFKLARLFSIPFKTNRVNPCREIIWVPPHGGCMKINCDGS 278
+ L + D V+ + L+R S + NR W P G K N DGS
Sbjct: 1301 QDALEWQDIEDRVQVIN-----PLSRRHSNRYAANR--------WTRPVCGWKKCNYDGS 1347
Query: 279 ---VVGSPSCGSIGVIFRASQTMFCGAFAQNIGYAT--ALEAEYSACMFAIEKAKELHLT 333
++ S + G + R F G Q G T ALE+ A + A++ T
Sbjct: 1348 YSTIINSKA----GWVIRDEHGQFIGG-GQAKGKHTNNALESALQALIIAMQSCWSHGHT 1402
Query: 334 NIWIETDSVNVIRAFHFNTG----VPWKMHIR-WHNCLLFCRSIRSLCTHVNREGNLVAD 388
+ E D++ V + + W I+ W CR + +NR N AD
Sbjct: 1403 KVCFEGDNIEVYQILNEGKARFDVYNWIRDIQAWKRRFQECRFL-----WINRRNNKPAD 1457
Query: 389 ALAK 392
LAK
Sbjct: 1458 TLAK 1461
>At1g17390 hypothetical protein
Length = 322
Score = 60.5 bits (145), Expect = 2e-09
Identities = 50/155 (32%), Positives = 71/155 (45%), Gaps = 13/155 (8%)
Query: 259 REIIWVPPHGGCMKINCDGSVVGSPSCGSIGVIFRASQTMFCGAFAQNIGYATALEAEYS 318
R I W PP G K+N DG+ G+P + G + R +C F+ NIG +A AE
Sbjct: 100 RLIAWSPPRVGWFKLNTDGASRGNPRLATAGGVVRDGDGNWCYGFSLNIGICSAPLAELW 159
Query: 319 ACMFAIEKAKELHLTNIWIETDSVNVIRAFHFNTGV----PWKMHIR-WHNCLLFCRSIR 373
+ + A E +T + +E DS V+ TG+ P +R H L S+R
Sbjct: 160 GAYYGLNIAWERGVTQLEMEIDSEMVVG--FLRTGIDDSHPLSFLVRLCHGLLSKDWSVR 217
Query: 374 SLCTHVNREGNLVADALAKNG----QGLALYSSHP 404
+HV RE N +AD LA G L++S P
Sbjct: 218 --ISHVYREANRLADGLANYAFFLPLGFHLFNSTP 250
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 60.1 bits (144), Expect = 2e-09
Identities = 39/157 (24%), Positives = 71/157 (44%), Gaps = 18/157 (11%)
Query: 74 KLAFSFLPGHGPTVHWAKMLWNAYTPPTGAFITWRFLHNKLPTDDNLRKRGCYIVSICCC 133
K+ ++ + H P +W K +W Y+ P +F+ W + N+L T D ++ + + C
Sbjct: 1339 KVTWNNVRTHQPQQNWYKGVWFPYSTPKYSFLLWLTVQNRLSTGDRIKAWNSGQL-VTCT 1397
Query: 134 FCRKQAETSSHIFLQCPVTLQLWDWL---LKATDQHLDFSSIL------NISR----MVQ 180
C ET H+F C T +W+ L L +T+ D++ + N+ R + +
Sbjct: 1398 LCNNAEETRDHLFFSCQYTSYVWEALTQRLLSTNYSRDWNRLFTLLCTSNLPRDHLFLFR 1457
Query: 181 HVMNSAIVHIMWSIWLECNNKYFDGVQKPMSTLFNTI 217
+V ++I H IW E N + + P + L I
Sbjct: 1458 YVFQASIYH----IWRERNARRHGEISSPTNRLIKLI 1490
>At2g05980 putative non-LTR retroelement reverse transcriptase
Length = 1352
Score = 59.3 bits (142), Expect = 4e-09
Identities = 34/131 (25%), Positives = 61/131 (45%), Gaps = 7/131 (5%)
Query: 59 DKLIWT---DSVDGDLSNKLAFSFLPGHGPTVHWAKMLWNAYTPPTGAFITWRFLHNKLP 115
D+ +W D+ S+ + + W + +W + + P +F+TW HN+L
Sbjct: 1170 DRSLWKQKEDTFKSSFSSSKTWQQIRSISLRCDWYRGVWFSASTPKYSFVTWLAFHNRLT 1229
Query: 116 TDDNLRKRGCYIVSICCCFCRKQAETSSHIFLQCPVTLQLWDWLLKATDQHLDFSSILNI 175
T D + K C FC ++ ET H+F CP + +W L K L+ +ILN
Sbjct: 1230 TSDKICKWNSG-ARYDCVFCGEELETRDHLFFSCPYSSHVWFSLTKGL---LNGRNILNW 1285
Query: 176 SRMVQHVMNSA 186
+ + H+++S+
Sbjct: 1286 NLITPHLLDSS 1296
>At1g10160 putative reverse transcriptase
Length = 1118
Score = 58.9 bits (141), Expect = 5e-09
Identities = 33/113 (29%), Positives = 52/113 (45%), Gaps = 1/113 (0%)
Query: 50 IALPLDETLDKLIWTDSVDGDLSNKLAFSFLPGHGPTVHWAKMLWNAYTPPTGAFITWRF 109
I P D+T I + S +S+L +V W K +W P AFI W
Sbjct: 904 IDCPHDDTYLWKIGHHAPSNRFSTADTWSYLQPSSTSVLWHKAVWFKDHVPKQAFICWVV 963
Query: 110 LHNKLPTDDNLRKRGCYIVSICCCFCRKQAETSSHIFLQCPVTLQLWDWLLKA 162
HN+L T D LR+ G + + C C E+ H+F +C + ++W + ++A
Sbjct: 964 AHNRLHTRDRLRRWG-FSIPPTCVLCNDLDESREHLFFRCQFSSEIWSFFMRA 1015
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 58.5 bits (140), Expect = 7e-09
Identities = 32/123 (26%), Positives = 57/123 (46%), Gaps = 10/123 (8%)
Query: 93 LWNAYTPPTGAFITWRFLHNKLPTDDNLRKRGCYIVSICCCFCRKQAETSSHIFLQCPVT 152
+WN + P W+ L + +D LR RG ++ C C ++ ET +HI QCP+
Sbjct: 775 IWNLNSAPKIKVFLWKVLKGAVAVEDRLRTRGV-LIEDGCSMCPEKNETLNHILFQCPLA 833
Query: 153 LQLWDWLLKATDQHLDFSSIL-NISRMVQHVMNSAI--------VHIMWSIWLECNNKYF 203
Q+W + H SI N++ ++ + N+ + I+W +W N + F
Sbjct: 834 RQVWALTPMQSPNHGFGDSIFTNVNHVIGNCHNTELSPHLRYVSPWIIWILWKNRNKRLF 893
Query: 204 DGV 206
+G+
Sbjct: 894 EGI 896
>At1g47320 hypothetical protein
Length = 259
Score = 58.5 bits (140), Expect = 7e-09
Identities = 47/159 (29%), Positives = 73/159 (45%), Gaps = 7/159 (4%)
Query: 269 GCMKINCDGSVVGSPSCGSIGVIFRASQTMFCGAFAQNIGYATALEAEYSACMFAIEKAK 328
GC+KIN DG+ G+P + G + + ++ +CG F+ NIG ++A AE + + A
Sbjct: 99 GCLKINTDGASRGNPGLATAGGVLQDNEGRWCGGFSLNIGRSSAPMAELWGAYYGLYLAW 158
Query: 329 ELHLTNIWIETDSVNVIRAFHFNTGV----PWKMHIRWHNCLLFCRSIRSLCTHVNREGN 384
E ++I +E DS V+ TG+ P +R + + + R HV RE N
Sbjct: 159 ERKSSHIELEVDSEIVVG--FLKTGISDHHPLSFLVRLCHGFI-SKDWRVRIFHVYREAN 215
Query: 385 LVADALAKNGQGLALYSSHPLAFISSFYVRDCLGLPFSR 423
AD LA L L + F+ D LG+ R
Sbjct: 216 RFADGLANYVFSLPLGFHFVPNVVDLFWKEDNLGVTCPR 254
>At4g11710 putative protein
Length = 473
Score = 58.2 bits (139), Expect = 9e-09
Identities = 39/140 (27%), Positives = 63/140 (44%), Gaps = 10/140 (7%)
Query: 87 VHWAKMLWNAYTPPTGAFITWRFLHNKLPTDDNLRKRGCYIVSICCCFCRKQAETSSHIF 146
V W K +W A P AF W +HN+L T D + V C C K E+ H+F
Sbjct: 303 VAWYKGVWFAQAIPKHAFCMWLAVHNRLSTGDRMTLWNMG-VDATCILCNKALESRDHLF 361
Query: 147 LQCPVTLQLWDWLLKA---TDQHLDFSSIL-NISR----MVQHVMNSAIVHI-MWSIWLE 197
CP ++W+ L K T + D+ +I+ N+SR + + I+ + ++++W E
Sbjct: 362 FSCPFATEIWEPLAKTIYNTCFYTDWQTIINNVSRNWPDRIAGFLARCILQVTIYTLWRE 421
Query: 198 CNNKYFDGVQKPMSTLFNTI 217
N + S L + I
Sbjct: 422 RNERKHGASPNSSSRLISWI 441
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.139 0.458
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,108,458
Number of Sequences: 26719
Number of extensions: 428079
Number of successful extensions: 1302
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 69
Number of HSP's successfully gapped in prelim test: 28
Number of HSP's that attempted gapping in prelim test: 1124
Number of HSP's gapped (non-prelim): 119
length of query: 427
length of database: 11,318,596
effective HSP length: 102
effective length of query: 325
effective length of database: 8,593,258
effective search space: 2792808850
effective search space used: 2792808850
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149206.10


