

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149205.17 + phase: 0 /pseudo
(626 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
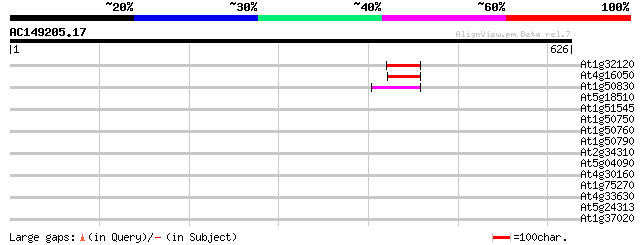
Score E
Sequences producing significant alignments: (bits) Value
At1g32120 hypothetical protein 51 2e-06
At4g16050 hypothetical protein 48 1e-05
At1g50830 hypothetical protein 46 7e-05
At5g18510 putative protein 42 0.001
At1g51545 unknown protein 42 0.001
At1g50750 hypothetical protein 39 0.007
At1g50760 hypothetical protein 39 0.011
At1g50790 hypothetical protein 38 0.019
At2g34310 unknown protein 33 0.37
At5g04090 putative protein 32 0.81
At4g30160 putative villin 32 1.1
At1g75270 GSH-dependent dehydroascorbate reductase 1, putative 32 1.4
At4g33630 unknown protein 30 4.0
At5g24313 unknown protein 29 6.9
At1g37020 hypothetical protein 29 6.9
>At1g32120 hypothetical protein
Length = 1206
Score = 51.2 bits (121), Expect = 2e-06
Identities = 19/38 (50%), Positives = 27/38 (71%)
Query: 421 IAWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSV 458
+AWK+Y RP++D LYFP++ EA +T Y +WW SV
Sbjct: 427 LAWKDYIRPIADGMLYFPARLHEADVTVGYIRWWKLSV 464
>At4g16050 hypothetical protein
Length = 900
Score = 48.1 bits (113), Expect = 1e-05
Identities = 19/37 (51%), Positives = 24/37 (64%)
Query: 422 AWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSV 458
AW +Y++PL LYFPS+ A +T RY WW KSV
Sbjct: 644 AWDDYNKPLVGLKLYFPSRVATASVTTRYRDWWAKSV 680
>At1g50830 hypothetical protein
Length = 768
Score = 45.8 bits (107), Expect = 7e-05
Identities = 19/55 (34%), Positives = 31/55 (55%)
Query: 404 NSDLIKMFLVNQTDPKAIAWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSV 458
+ DL + + + AW +Y++ L +LY PS+ + +TARY WW+KSV
Sbjct: 400 DQDLPGLATCQRNSTEKEAWNDYNKSLIGLNLYMPSRLDQGSVTARYRVWWLKSV 454
>At5g18510 putative protein
Length = 702
Score = 42.0 bits (97), Expect = 0.001
Identities = 17/36 (47%), Positives = 21/36 (58%)
Query: 422 AWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKS 457
AW +YS L +LY PS+ +TARY WW KS
Sbjct: 394 AWDDYSNSLEGLNLYLPSQLDRGYVTARYQDWWFKS 429
>At1g51545 unknown protein
Length = 629
Score = 41.6 bits (96), Expect = 0.001
Identities = 15/37 (40%), Positives = 21/37 (56%)
Query: 422 AWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSV 458
AW Y++ L LY PS+ +T RY WW+KS+
Sbjct: 326 AWDGYNKSLDGLMLYIPSRVATTSVTERYRDWWLKSI 362
>At1g50750 hypothetical protein
Length = 816
Score = 39.3 bits (90), Expect = 0.007
Identities = 15/46 (32%), Positives = 29/46 (62%), Gaps = 1/46 (2%)
Query: 413 VNQTD-PKAIAWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKS 457
VNQ + + AW +Y++P++D +L+ PS+ +T + +WW +S
Sbjct: 321 VNQNNLSQEAAWNDYNKPINDLALFIPSRSAIPRVTPTFCEWWRRS 366
>At1g50760 hypothetical protein
Length = 649
Score = 38.5 bits (88), Expect = 0.011
Identities = 12/36 (33%), Positives = 24/36 (66%)
Query: 422 AWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKS 457
AW +Y++P+ D +L+ PS+ +T+ + +WW K+
Sbjct: 393 AWNDYNKPIDDLALHIPSRSAIPRVTSTFCEWWRKT 428
>At1g50790 hypothetical protein
Length = 812
Score = 37.7 bits (86), Expect = 0.019
Identities = 34/129 (26%), Positives = 50/129 (38%), Gaps = 11/129 (8%)
Query: 422 AWKNYSRPLSDRSLYFPSKFFEAGITARYAKWWMKSVLRSQ-------GFVTNFEPRKRS 474
AW +Y +PL D LY PS+ + T + W K V+ S G T+F P S
Sbjct: 405 AWNDYYKPLDDLELYIPSRSAISCDTPMFCDSWRKRVVGSAKTLRTSIGDDTSFVP-SGS 463
Query: 475 ASSSKCEPSAKIPPEFPSHKLVGCTVTIGKSSDDGSNTSQGDKIVDDDAPSGSIPKLLKT 534
+ + E KI + +L +DGS Q I DD S+ T
Sbjct: 464 KMNRRSEDGRKIAEDSTKKRL---KFMKSARKNDGSTMGQKQVISDDADDDDSLTVAQIT 520
Query: 535 MSFEKSVED 543
++K +
Sbjct: 521 TLYKKKYSE 529
>At2g34310 unknown protein
Length = 470
Score = 33.5 bits (75), Expect = 0.37
Identities = 37/141 (26%), Positives = 59/141 (41%), Gaps = 5/141 (3%)
Query: 474 SASSSKCEPSAKIPPEFPSHKLVGCTVTIGKSSDDGSNTSQGDKIVDDDAPSGSIPKLL- 532
S S+ K ++PP S LVG GKS NT G+K+ A +G +
Sbjct: 119 SNSTDKNMADQELPPSVTSIVLVGRNNGNGKSFT--GNTLLGEKLFISKADAGGVTMEED 176
Query: 533 -KTMSFEKSVEDGLLAEKYVDGGAPLPAKDNTPTPLIFVDDCKHVLEDGNESEEARLSTD 591
K E+ +E LAE V K++ T + ++ +H L+ E+ E
Sbjct: 177 DKLKEKEREIESKSLAEAEVIAMKERSRKEHDHTMNMAHEEVEHALKQNTETHEKETQHH 236
Query: 592 TICQSENQT-ESYSYLSELIV 611
I S++ ES ++ LI+
Sbjct: 237 VIDVSDSTADESVFVVNPLIM 257
>At5g04090 putative protein
Length = 300
Score = 32.3 bits (72), Expect = 0.81
Identities = 18/45 (40%), Positives = 25/45 (55%), Gaps = 1/45 (2%)
Query: 500 VTIGKSSDDGSNTSQGDKIVDDDAPS-GSIPKLLKTMSFEKSVED 543
VTI S+D SN S D +VD DAP+ GS+ ++ + S D
Sbjct: 215 VTIASFSNDSSNQSLNDPLVDPDAPTFGSLGQIPQNFSLSDLTAD 259
>At4g30160 putative villin
Length = 974
Score = 32.0 bits (71), Expect = 1.1
Identities = 20/79 (25%), Positives = 43/79 (54%), Gaps = 1/79 (1%)
Query: 472 KRSASSSKCEPSAKIPPEFPS-HKLVGCTVTIGKSSDDGSNTSQGDKIVDDDAPSGSIPK 530
K SA +S+ KIPP+ PS K V + +S SN+ + ++ ++D GS+
Sbjct: 832 KSSAIASRSALFEKIPPQEPSIPKPVKASPKTPESPAPESNSKEQEEKKENDKEEGSMSS 891
Query: 531 LLKTMSFEKSVEDGLLAEK 549
+++++ ++ ++G+ E+
Sbjct: 892 RIESLTIQEDAKEGVEDEE 910
>At1g75270 GSH-dependent dehydroascorbate reductase 1, putative
Length = 213
Score = 31.6 bits (70), Expect = 1.4
Identities = 20/42 (47%), Positives = 24/42 (56%), Gaps = 4/42 (9%)
Query: 481 EPSAKIPPEFPS--HKLVGCTVTIGKSSD--DGSNTSQGDKI 518
EPS K PPEF S K+ G VT KS D DGS + D++
Sbjct: 86 EPSLKTPPEFASVGSKIFGAFVTFLKSKDANDGSEKALVDEL 127
>At4g33630 unknown protein
Length = 684
Score = 30.0 bits (66), Expect = 4.0
Identities = 21/61 (34%), Positives = 30/61 (48%), Gaps = 2/61 (3%)
Query: 474 SASSSKCEPSAKIPPEFPSHKLVGCTVTIGKSSDDGSNTSQGDKIVDDDAPSGSIPKLLK 533
SA+SS+ PS+K F H+L T SS GSN+S D+ +I +LK
Sbjct: 19 SATSSRLTPSSK--RSFYPHRLPDPTALCRCSSSSGSNSSSSSSSDDNPRWDSAIQDVLK 76
Query: 534 T 534
+
Sbjct: 77 S 77
>At5g24313 unknown protein
Length = 81
Score = 29.3 bits (64), Expect = 6.9
Identities = 16/50 (32%), Positives = 22/50 (44%)
Query: 517 KIVDDDAPSGSIPKLLKTMSFEKSVEDGLLAEKYVDGGAPLPAKDNTPTP 566
K++DD+ PS P + F S L+AE Y P P K + P
Sbjct: 27 KLLDDNVPSSYFPSMESRQVFLPSSHRSLVAEHYDRSPPPPPPKRSEVEP 76
>At1g37020 hypothetical protein
Length = 611
Score = 29.3 bits (64), Expect = 6.9
Identities = 16/36 (44%), Positives = 25/36 (69%), Gaps = 2/36 (5%)
Query: 504 KSSDDGSNTSQGDKIVDDDAPSGSIPKLLKTMSFEK 539
K+ +DG +T + +I+ D APS SIP+ L+ +S EK
Sbjct: 364 KNLNDGKSTVK--RIITDGAPSPSIPEHLQPVSDEK 397
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.350 0.153 0.557
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,032,037
Number of Sequences: 26719
Number of extensions: 521820
Number of successful extensions: 2438
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 2422
Number of HSP's gapped (non-prelim): 22
length of query: 626
length of database: 11,318,596
effective HSP length: 105
effective length of query: 521
effective length of database: 8,513,101
effective search space: 4435325621
effective search space used: 4435325621
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149205.17


