

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149204.7 + phase: 0
(600 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
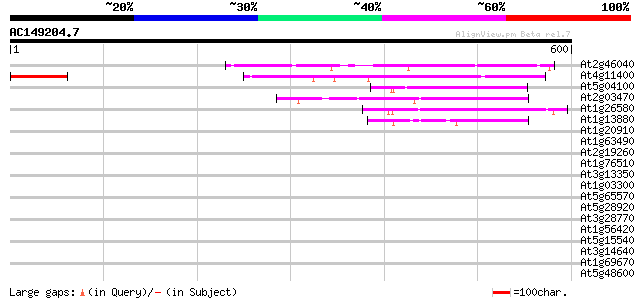
Score E
Sequences producing significant alignments: (bits) Value
At2g46040 unknown protein 240 1e-63
At4g11400 putative protein 219 3e-57
At5g04100 putative protein 145 6e-35
At2g03470 putative MYB family transcription factor 124 1e-28
At1g26580 hypothetical protein 112 5e-25
At1g13880 unknown protein 111 1e-24
At1g20910 unknown protein 39 0.011
At1g63490 RB-binding protein -like 35 0.12
At2g19260 unknown protein 34 0.27
At1g76510 putative DNA-binding protein 34 0.27
At3g13350 unknown protein 32 0.77
At1g03300 unknown protein 32 1.3
At5g65570 putative protein 30 2.9
At5g28920 putative protein 30 3.8
At3g28770 hypothetical protein 30 5.0
At1g56420 hypothetical protein 30 5.0
At5g15540 putative protein 29 6.6
At3g14640 putative cytochrome P450 29 6.6
At1g69670 putative cullin 29 6.6
At5g48600 chromosome condensation protein 29 8.6
>At2g46040 unknown protein
Length = 562
Score = 240 bits (613), Expect = 1e-63
Identities = 145/366 (39%), Positives = 204/366 (55%), Gaps = 48/366 (13%)
Query: 231 DDVLMLDPSSVNRESFGRKRKRDDLMSEMQSWVIRTAKNPCDPVLGSMPEKSKWKSYGNQ 290
+ +++L+ SV +E +++ + E W+ AK+PCDP LG +P++S+W SYG++
Sbjct: 227 EQIMILE--SVTQECSSPGKRKRECPLETLKWLSDVAKDPCDPSLGIVPDRSEWVSYGSE 284
Query: 291 EIWKKVLLFREAAFLKKDFGSDCEKLSWLAQRMHPSMYDVNLGVNYNLRQRI-------- 342
E WK++LLFR + + + S CEK Q+MHP +YD + G +YNLR+R+
Sbjct: 285 EPWKQLLLFRAS---RTNNDSACEKTWQKVQKMHPCLYDDSAGASYNLRERLSYEDYKRG 341
Query: 343 KRDNGVLVGKSASILQRTPDSEAKEKLLDSCVPESILDAPPTVNIPLGPNHQAEVPEWTG 402
K NG +G S D E + L +G QA+VPEWTG
Sbjct: 342 KTGNGSDIGSS--------DEEDRPCAL------------------VGSKFQAKVPEWTG 375
Query: 403 TTHKSDSKWLGTQIWPLQIVKSK---LLEGEPVGKGRQDSCRCQVQGSVECVRFHIAEKS 459
T +SDSKWLGT+IWPL ++K L+E + +GKGRQD C C GS+ECV+FHI K
Sbjct: 376 ITPESDSKWLGTRIWPLTKEQTKANLLIERDRIGKGRQDPCGCHNPGSIECVKFHITAKR 435
Query: 460 AKLKLEIGVAFYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPK 519
KLKLE+G AFY W D GE + WT E KK K ++ S+P SL F P
Sbjct: 436 DKLKLELGPAFYMWCFDVMGECTLQYWTDLELKKIKSLM-SSPPSLSPAFIHQAKMILPS 494
Query: 520 KSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDEESEFTPLKGSFGHQTDKSNSF---T 576
KSR +VSY++NV LLQ R Q+R TP +IDSD ++ + + ++N+F
Sbjct: 495 KSRGKIVSYFYNVTLLQYRASQSRITPHDIDSDTDQIFLRATENE--NPAPEANTFQKPA 552
Query: 577 LLSPKK 582
LLSP K
Sbjct: 553 LLSPNK 558
>At4g11400 putative protein
Length = 573
Score = 219 bits (559), Expect = 3e-57
Identities = 133/346 (38%), Positives = 183/346 (52%), Gaps = 29/346 (8%)
Query: 251 KRDDLMSEMQSWVIRTAKNPCDPVLGSMPEKSKWKSYGNQEIWKKVLLFREAAFLKKDFG 310
KRDDL M W+ A +P DP +G +P SKWK Y + W +V + + +++D
Sbjct: 218 KRDDLPG-MLKWLALVATSPHDPAIGVIPHSSKWKQYNGNKCWLQVARAKNSLLVQRDNA 276
Query: 311 SDCEKLSWLAQRM---HPSMYDVNLGVNYNLRQRIKRDN-------GVLVGKSASILQRT 360
+ HPSMY+ + LR I+ N G S L ++
Sbjct: 277 ELRYRYHPFRGHQNIHHPSMYEDDRKSIGRLRYSIRPPNLSKHCSSSCCNGSSLVSLSKS 336
Query: 361 PDSEAKEKLLDSCVPESILDAP-----------PTVNIPLGPNHQAEVPEWTGTTHKSDS 409
++ ++ + + + P I +G HQA+V EWT + SDS
Sbjct: 337 RSTKCRKLTIIASERAGLTAGTSRARKRNKAEIPRRCIKVGHQHQAQVDEWTESGVDSDS 396
Query: 410 KWLGTQIWPLQIVKS--KLLEGEPVGKGRQDSCRCQVQGSVECVRFHIAEKSAKLKLEIG 467
KWLGT+IWP + ++ + L + VGKGR DSC C++ G VEC R HIAEK +LK E+G
Sbjct: 397 KWLGTRIWPPENSEALDQTLGNDLVGKGRPDSCSCELSGFVECTRLHIAEKRMELKRELG 456
Query: 468 VAFYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVS 527
F+ W ++ GEEV WT EEEK+FKD++ ++P S FW + FPKK RE LVS
Sbjct: 457 DDFFHWRFNQMGEEVCLRWTEEEEKRFKDMIIADPQS----FWTNAAKNFPKKKREELVS 512
Query: 528 YYFNVFLLQRRGYQNRHTPDNIDSDDEESEFTPLKGSFGHQTDKSN 573
YYFNVFL+ RR YQNR TP +IDSDD E F + GSFG S+
Sbjct: 513 YYFNVFLINRRRYQNRVTPKSIDSDD-EGAFGSVGGSFGRDAVTSS 557
Score = 55.1 bits (131), Expect = 1e-07
Identities = 22/61 (36%), Positives = 42/61 (68%)
Query: 1 MLGSEQTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKVGSSVKLVYSKYLSGL 60
++G + +DL+KLF++V+++ G+D V + LW++V E+ G + S+ L+Y KYL+ +
Sbjct: 50 VIGDGKNVDLFKLFVLVREREGFDTVSRKRLWEVVAEKLGFDCSLVPSLILIYLKYLNRM 109
Query: 61 E 61
E
Sbjct: 110 E 110
>At5g04100 putative protein
Length = 351
Score = 145 bits (366), Expect = 6e-35
Identities = 76/179 (42%), Positives = 104/179 (57%), Gaps = 12/179 (6%)
Query: 387 IPLGPNHQAEVPEWTGTTHKS-------DS---KWLGTQIWPLQIVKSKLLEGEPVGKGR 436
IP+GP QAE+P W T K DS +WLGT +WP +K K + + VG+GR
Sbjct: 166 IPIGPRFQAEIPVWIAPTKKGKFYGSPGDSNTLRWLGTGVWPTYSLK-KTVHSKKVGEGR 224
Query: 437 QDSCRCQVQGSVECVRFHIAEKSAKLKLEIGVAFYQWNLDKAGEE-VRRCWTAEEEKKFK 495
DSC C S C++ H E L+ EI AF W D+ GEE V + WTA+EE++F+
Sbjct: 225 SDSCSCASPRSTNCIKRHKKEAQELLEKEINRAFSTWEFDQMGEEIVLKSWTAKEERRFE 284
Query: 496 DVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDE 554
+VK NP S FW+ FP+KS++ L+SYY+NVFL++R +NIDSDD+
Sbjct: 285 ALVKKNPLSSSDGFWEFASNAFPQKSKKDLLSYYYNVFLIKRMRLLKSSAANNIDSDDD 343
>At2g03470 putative MYB family transcription factor
Length = 450
Score = 124 bits (311), Expect = 1e-28
Identities = 88/282 (31%), Positives = 136/282 (48%), Gaps = 21/282 (7%)
Query: 286 SYGNQEIWKKVLLFREAAFLKK------DFGSDCEKLSWLAQRMHPSMYDVNLGVNYNLR 339
SY N + + + + A+ KK D CE +W H G+
Sbjct: 24 SYKNHNQFDEAIPYHHASMEKKTNVLVEDLIGLCENPTWTNDANHVDKGFETTGL----- 78
Query: 340 QRIKRDNGVLVGKSASILQRTPDSEAKEKLLDSCVPESILDAPPTVNIPLGPNHQAEVPE 399
+ D+ V + + ++ S+ K ++ V +++ PP + +G NHQA++PE
Sbjct: 79 --CQEDSQSGVTTQSDLSHQSSGSDFTWKPVED-VYTCLMNQPPRKQVLVGSNHQADIPE 135
Query: 400 WTGTTHKSDSKWLGTQIWPLQIVKSKLLEGEP-----VGKGRQDSCRCQVQGSVECVRFH 454
+ S+ + ++++ ++ G+GR++ C C +GS+ CVR H
Sbjct: 136 FVKEEILDQSEARTKEDLEGKLMRKCVIPMSDSDLCGTGQGRKE-CLCLDKGSIRCVRRH 194
Query: 455 IAEKSAKLKLEIGVA-FYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDI 513
I E L IG F + L + GEEV WT EEE F VV SNP S R FW +
Sbjct: 195 IIEARESLVETIGYERFMELGLCEMGEEVASLWTEEEEDLFHKVVYSNPFSAGRDFWKQL 254
Query: 514 FTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDEE 555
TFP ++ + LVSYYFNVF+L+RRG QNR ++DSDD+E
Sbjct: 255 KGTFPSRTMKELVSYYFNVFILRRRGIQNRFKALDVDSDDDE 296
>At1g26580 hypothetical protein
Length = 493
Score = 112 bits (281), Expect = 5e-25
Identities = 81/244 (33%), Positives = 124/244 (50%), Gaps = 27/244 (11%)
Query: 378 ILDAPPTVNIPLGPNHQAEVPEWTGT-------------THKSD----SKWLGTQIWPLQ 420
+LD +P+GP HQAE+PEW G+ H S K GT + P+
Sbjct: 126 LLDQRAKKQVPIGPGHQAEIPEWEGSQTGNIETSGMSVQNHISGCADGEKLFGTSVIPMP 185
Query: 421 IVKSKLLEGEPVGKGRQDSCRCQVQGSVECVRFHIAEKSAKLKLEIG-VAFYQWNLDKAG 479
+ + + VGKGR+ C C+ + SV CV HI E +L G F + L + G
Sbjct: 186 GLTTVAHIDDIVGKGRK-FCVCRDRDSVRCVCQHIKEAREELVKTFGNETFKELGLCEMG 244
Query: 480 EEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRG 539
E+ W+ E+ + F +VV SNP +L + FW + F ++++ +VS+YFNVF+L+RR
Sbjct: 245 EKGALKWSDEDAQLFHEVVYSNPVTLGQNFWRHLEAAFCSRTQKEIVSFYFNVFVLRRRA 304
Query: 540 YQNRHTPDNIDSDDEESE--FTPLKGSFGHQTDKSNSFTLLSP-----KKPQPTPHTKGR 592
QNR +IDSDD+E + G+ + D+ +S + SP KK P H +G
Sbjct: 305 IQNRAFILDIDSDDDEWHGCYGGSSGTRYVEEDEEDS-AIESPLHQGTKKVYPLHHEEGE 363
Query: 593 QLVN 596
+ V+
Sbjct: 364 EDVS 367
>At1g13880 unknown protein
Length = 424
Score = 111 bits (278), Expect = 1e-24
Identities = 67/186 (36%), Positives = 103/186 (55%), Gaps = 20/186 (10%)
Query: 383 PTVNIPLGPNHQAEVPEWTGTTHKSDS---------KWLGTQIWPLQIVKSKLLEGEPVG 433
P +P+G ++QA++PE S + G + P+ ++++ + +G
Sbjct: 124 PRKTVPIGSDYQADIPECVKEEANDQSGQGVGYDEEQVTGKCVIPMPDCETEVCK---IG 180
Query: 434 KGRQDSCRCQVQGSVECVRFHIAEKSAKLKLEIGVAFYQWNLD----KAGEEVRRCWTAE 489
KGR++ C C +GS+ CV+ HI E L IG Y LD + GEEV T +
Sbjct: 181 KGRKE-CICLDKGSIRCVQQHIMENREDLFATIG---YDRCLDIGLCEMGEEVAARLTED 236
Query: 490 EEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNI 549
EE F ++V SNP S+DR FW + + FP ++ + +VSYYFNVF+L+RR QNR ++
Sbjct: 237 EEDLFHEIVYSNPVSMDRDFWKHLKSAFPSRTMKEIVSYYFNVFILRRRAIQNRSKSLDV 296
Query: 550 DSDDEE 555
DSDD+E
Sbjct: 297 DSDDDE 302
>At1g20910 unknown protein
Length = 398
Score = 38.5 bits (88), Expect = 0.011
Identities = 25/76 (32%), Positives = 37/76 (47%), Gaps = 3/76 (3%)
Query: 6 QTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYG---LGVKVGSSVKLVYSKYLSGLET 62
Q L++ KL+ V + GGY+VV N+LW VGE + V + + Y K L E
Sbjct: 135 QPLNILKLWRAVVNLGGYEVVTTNKLWRQVGESFNPPKTCTTVSYTFRNFYEKALLEYEK 194
Query: 63 PLKNVVDGEFPKRDLV 78
L+N + P L+
Sbjct: 195 CLRNNGELNLPGSTLI 210
>At1g63490 RB-binding protein -like
Length = 1458
Score = 35.0 bits (79), Expect = 0.12
Identities = 22/60 (36%), Positives = 29/60 (47%), Gaps = 4/60 (6%)
Query: 6 QTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKVGSSVKLV----YSKYLSGLE 61
+ LDL KLF VK GGY+ V K + W V + G K+ K V Y ++L E
Sbjct: 125 EELDLCKLFNAVKRFGGYEKVVKGKKWGEVYQFMSSGEKISKCAKHVLCQLYKEHLHDFE 184
>At2g19260 unknown protein
Length = 631
Score = 33.9 bits (76), Expect = 0.27
Identities = 19/59 (32%), Positives = 32/59 (54%), Gaps = 4/59 (6%)
Query: 382 PPTVNIPLGPNHQAEVPEWTGTTHKSDSKWLGTQIWPLQIVKSKLLEGEPVGKGRQDSC 440
P + I +G QA+VP+W+G T SD+ ++G PL+I +S+ + K + C
Sbjct: 482 PFVIGIRIGKMFQADVPDWSGPT-MSDTSFVGE---PLEIGQSEYMHDLKKAKNSKKQC 536
>At1g76510 putative DNA-binding protein
Length = 434
Score = 33.9 bits (76), Expect = 0.27
Identities = 16/34 (47%), Positives = 21/34 (61%)
Query: 6 QTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEY 39
Q L+ KL+ V GGYDVV ++LW VGE +
Sbjct: 171 QPLNCLKLWRAVIKLGGYDVVTTSKLWRQVGESF 204
>At3g13350 unknown protein
Length = 319
Score = 32.3 bits (72), Expect = 0.77
Identities = 19/58 (32%), Positives = 31/58 (52%), Gaps = 3/58 (5%)
Query: 7 TLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKVGSS---VKLVYSKYLSGLE 61
TLDL++LF+ V +GG + V K+ W V + + S+ ++ Y K+L LE
Sbjct: 70 TLDLHRLFIEVTSRGGIERVVKDRKWKEVIGAFSFPTTITSASFVLRKYYLKFLFQLE 127
>At1g03300 unknown protein
Length = 670
Score = 31.6 bits (70), Expect = 1.3
Identities = 27/120 (22%), Positives = 50/120 (41%), Gaps = 14/120 (11%)
Query: 179 QDCKTNETSLRKLVNQNAAMEIMDDVSDVAN-SMPGLSDGSKS-CANDDANDSGDDVLML 236
++ K T ++ N+A +MD++ N S +S+ KS C NDD +D
Sbjct: 397 ENSKDGSTKRKREEKHNSASSVMDEIDGTCNGSESEISNTGKSICNNDDVDD-------- 448
Query: 237 DPSSVNRESFGRKRKRDDLMSEMQSWVIRTAKNPCDPVLGSMPEKSKWKSYGNQEIWKKV 296
P S + + ++ + ++A+ P +P WKSY E++K +
Sbjct: 449 QPLSTELPYYQSLSVVNSFAADAEETPAKSART-ISPFAKKLP---FWKSYETDELYKSL 504
>At5g65570 putative protein
Length = 738
Score = 30.4 bits (67), Expect = 2.9
Identities = 32/96 (33%), Positives = 37/96 (38%), Gaps = 3/96 (3%)
Query: 59 GLETPLKNVVDGEFPKRDLVGDRVKFGERLTELQAELVLDDYGEGDVGDEVKSVYGCRKK 118
GL L V F LV VKFG+ +A+LVLD E DV + G +K
Sbjct: 189 GLAVILGLEVSNVFVGSALVDMYVKFGKTR---EAKLVLDRVEEKDVVLITALIVGYSQK 245
Query: 119 LCDTNRVKVVNPELNASELEKVYEYIDGRKSCGTNK 154
DT VK L Y Y SCG K
Sbjct: 246 GEDTEAVKAFQSMLVEKVQPNEYTYASVLISCGNLK 281
>At5g28920 putative protein
Length = 510
Score = 30.0 bits (66), Expect = 3.8
Identities = 23/91 (25%), Positives = 45/91 (49%), Gaps = 7/91 (7%)
Query: 143 YIDGRKSCGTNKMKDTNLVSNMAKKVESEGLVDVLMQDCKTNETSLRKLVNQNAAMEIMD 202
Y RKS + + N + + +K+ +GL+ + N+TSL N +++++ D
Sbjct: 206 YTKKRKSMQIDTNRLENNQARLDRKLACQGLIITKLTRAVKNDTSL----NLASSLKLHD 261
Query: 203 -DVSDVANSMPGLSDGSKSCANDDANDSGDD 232
D +D+ + L+ + + NDD + S DD
Sbjct: 262 LDTNDITSE--PLAPATPATPNDDESASSDD 290
>At3g28770 hypothetical protein
Length = 2081
Score = 29.6 bits (65), Expect = 5.0
Identities = 23/88 (26%), Positives = 39/88 (44%), Gaps = 5/88 (5%)
Query: 145 DGRKSCGTNKMKDTNLVSNMAKKVESEGLVDVLMQDCKTNETSLRKLVNQNAAMEIMDDV 204
+G K GTN T S +K VE G+ D ++D N +N + D+V
Sbjct: 1732 EGSKDGGTNS---TGKDSKDSKSVEINGVKDDSLKDDSKNGDINE--INNGKEDSVKDNV 1786
Query: 205 SDVANSMPGLSDGSKSCANDDANDSGDD 232
+++ + L++ + S N D D+ D
Sbjct: 1787 TEIQGNDNSLTNSTSSEPNGDKLDTNKD 1814
>At1g56420 hypothetical protein
Length = 183
Score = 29.6 bits (65), Expect = 5.0
Identities = 14/33 (42%), Positives = 23/33 (69%), Gaps = 1/33 (3%)
Query: 31 LWDLVGEEYGLGVKVGSSVKLVYSKYLSGLETP 63
+WDL+ E+ GLG ++ +V+ V+ +LSG E P
Sbjct: 86 VWDLIIEKDGLGKEINETVERVFC-HLSGQEPP 117
>At5g15540 putative protein
Length = 1755
Score = 29.3 bits (64), Expect = 6.6
Identities = 25/117 (21%), Positives = 53/117 (44%), Gaps = 8/117 (6%)
Query: 141 YEYIDGRKSCGTNKMKDTNLVSNMAKKVESEGLVDVLMQDCKTNETSLRKLVNQNAAMEI 200
+EY+ +C D + +K+ SE V V MQ + +T L + + +
Sbjct: 121 FEYVTPGPTC------DPLFTNEGPQKIISEPSVPVKMQ--RQTDTHLARSIEPEPVKRV 172
Query: 201 MDDVSDVANSMPGLSDGSKSCANDDANDSGDDVLMLDPSSVNRESFGRKRKRDDLMS 257
+ +S + ++S + A DS + + ++ S +++ G+K+++DDL S
Sbjct: 173 LRPNHVEDHSWQHETLTNQSPKDVTAYDSRPETITMNELSASKKPKGKKKRKDDLSS 229
>At3g14640 putative cytochrome P450
Length = 514
Score = 29.3 bits (64), Expect = 6.6
Identities = 22/55 (40%), Positives = 26/55 (47%), Gaps = 3/55 (5%)
Query: 45 VGSSVKLVYSKYLSGLET---PLKNVVDGEFPKRDLVGDRVKFGERLTELQAELV 96
VG KLV K S E P + G+ R G K G+R+ ELQAELV
Sbjct: 179 VGEWSKLVSDKGSSSCEVDVWPWLVSMTGDVISRTAFGSSYKEGQRIFELQAELV 233
>At1g69670 putative cullin
Length = 732
Score = 29.3 bits (64), Expect = 6.6
Identities = 12/48 (25%), Positives = 28/48 (58%), Gaps = 3/48 (6%)
Query: 136 ELEKVYEYIDGRKSCGTNKMKDTNLVSNMAKK---VESEGLVDVLMQD 180
E+E+V Y+D + + + +++N ++ +E+ GLV++L+ D
Sbjct: 235 EVERVVNYLDAKSEAKITSVVEREMIANHVQRLVHMENSGLVNMLLND 282
>At5g48600 chromosome condensation protein
Length = 1241
Score = 28.9 bits (63), Expect = 8.6
Identities = 33/164 (20%), Positives = 68/164 (41%), Gaps = 8/164 (4%)
Query: 43 VKVGSSVKLVYSKYLSGLETPLKNVVDGEFPKRDLVGDRVKFGERLTELQ-----AELVL 97
V++ ++ KL+ K G+E + E K +L ++ E+Q + ++
Sbjct: 867 VQIETNQKLI-KKLTKGIEEATREKERLEGEKENLHVTFKDITQKAFEIQETYKKTQQLI 925
Query: 98 DDYGEGDVGDEVKSVYGCRKKLCDTNRVKVVNPELNASELEKVYEYIDGRKSCGTNKMKD 157
D++ DV KS Y KK D + V+ E +++K Y ++ R+ K+ D
Sbjct: 926 DEHK--DVLTGAKSDYENLKKSVDELKASRVDAEFKVQDMKKKYNELEMREKGYKKKLND 983
Query: 158 TNLVSNMAKKVESEGLVDVLMQDCKTNETSLRKLVNQNAAMEIM 201
+ + + LVD + +L + + A+E++
Sbjct: 984 LQIAFTKHMEQIQKDLVDPDKLQATLMDNNLNEACDLKRALEMV 1027
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,693,347
Number of Sequences: 26719
Number of extensions: 676420
Number of successful extensions: 1523
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 1485
Number of HSP's gapped (non-prelim): 43
length of query: 600
length of database: 11,318,596
effective HSP length: 105
effective length of query: 495
effective length of database: 8,513,101
effective search space: 4213984995
effective search space used: 4213984995
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149204.7


