

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
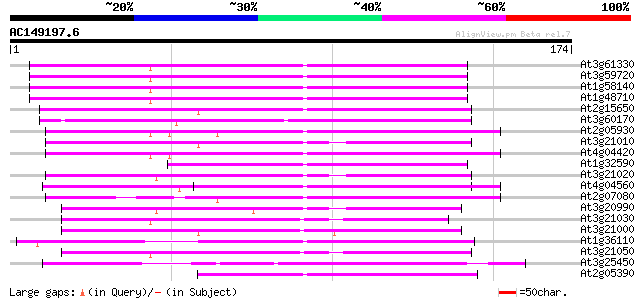
Query= AC149197.6 + phase: 0 /pseudo
(174 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g61330 copia-type polyprotein 100 7e-22
At3g59720 copia-type reverse transcriptase-like protein 100 7e-22
At1g58140 hypothetical protein 100 7e-22
At1g48710 hypothetical protein 100 7e-22
At2g15650 putative retroelement pol polyprotein 81 3e-16
At3g60170 putative protein 81 3e-16
At2g05930 copia-like retroelement pol polyprotein 62 2e-10
At3g21010 unknown protein 60 6e-10
At4g04420 putative transposon protein 59 1e-09
At1g32590 hypothetical protein, 5' partial 59 2e-09
At3g21020 unknown protein 54 3e-08
At4g04560 putative transposon protein 53 1e-07
At2g07080 putative gag-protease polyprotein 51 3e-07
At3g20990 unknown protein 49 1e-06
At3g21030 hypothetical protein 47 4e-06
At3g21000 unknown protein 47 5e-06
At1g36110 hypothetical protein 47 5e-06
At3g21050 hypothetical protein 45 2e-05
At3g25450 hypothetical protein 45 2e-05
At2g05390 putative retroelement pol polyprotein 43 8e-05
>At3g61330 copia-type polyprotein
Length = 1352
Score = 99.8 bits (247), Expect = 7e-22
Identities = 53/138 (38%), Positives = 78/138 (56%), Gaps = 3/138 (2%)
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSV--PALASNATDMQ*TANRDAKKKD 64
P +PV NYD W ++MK I DV E+V P + + Q RD++K+D
Sbjct: 7 PFQVPVLTKSNYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRD 66
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ ++Q + + FEK+V SAKE WE L Y G ++KKVRLQ LR ++E L M
Sbjct: 67 KKALCLIYQGLDEDTFEKVVEATSAKE-AWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 125 EEEETISQYFDKFVNLSN 142
+E E +S YF + + ++N
Sbjct: 126 KEGELVSDYFSRVLTVTN 143
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 99.8 bits (247), Expect = 7e-22
Identities = 53/138 (38%), Positives = 78/138 (56%), Gaps = 3/138 (2%)
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSV--PALASNATDMQ*TANRDAKKKD 64
P +PV NYD W ++MK I DV E+V P + + Q RD++K+D
Sbjct: 7 PFQVPVLTKSNYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRD 66
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ ++Q + + FEK+V SAKE WE L Y G ++KKVRLQ LR ++E L M
Sbjct: 67 KKALCLIYQGLDEDTFEKVVEATSAKE-AWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 125 EEEETISQYFDKFVNLSN 142
+E E +S YF + + ++N
Sbjct: 126 KEGELVSDYFSRVLTVTN 143
>At1g58140 hypothetical protein
Length = 1320
Score = 99.8 bits (247), Expect = 7e-22
Identities = 53/138 (38%), Positives = 78/138 (56%), Gaps = 3/138 (2%)
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSV--PALASNATDMQ*TANRDAKKKD 64
P +PV NYD W ++MK I DV E+V P + + Q RD++K+D
Sbjct: 7 PFQVPVLTKSNYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRD 66
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ ++Q + + FEK+V SAKE WE L Y G ++KKVRLQ LR ++E L M
Sbjct: 67 KKALCLIYQGLDEDTFEKVVEATSAKE-AWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 125 EEEETISQYFDKFVNLSN 142
+E E +S YF + + ++N
Sbjct: 126 KEGELVSDYFSRVLTVTN 143
>At1g48710 hypothetical protein
Length = 1352
Score = 99.8 bits (247), Expect = 7e-22
Identities = 53/138 (38%), Positives = 78/138 (56%), Gaps = 3/138 (2%)
Query: 7 PTHLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSV--PALASNATDMQ*TANRDAKKKD 64
P +PV NYD W ++MK I DV E+V P + + Q RD++K+D
Sbjct: 7 PFQVPVLTKSNYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRD 66
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ ++Q + + FEK+V SAKE WE L Y G ++KKVRLQ LR ++E L M
Sbjct: 67 KKALCLIYQGLDEDTFEKVVEATSAKE-AWEKLRTSYKGADQVKKVRLQTLRGEFEALQM 125
Query: 125 EEEETISQYFDKFVNLSN 142
+E E +S YF + + ++N
Sbjct: 126 KEGELVSDYFSRVLTVTN 143
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 81.3 bits (199), Expect = 3e-16
Identities = 46/139 (33%), Positives = 75/139 (53%), Gaps = 6/139 (4%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TAN-----RDAKKKD 64
+P+F+G+ YD W +KM I R + + VV + VP A + TA +A D
Sbjct: 9 IPIFDGEKYDFWSIKMATIFRTRKLWSVVEEGVPVEPVQAEETPETARAKTLREEAVTND 68
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
A+ + +V+++IF +I S+KE W+ L+ Y G +++ V+LQ LRR+YE L M
Sbjct: 69 TMALQILQTAVTDQIFSRIAAASSSKE-AWDVLKDEYQGSPQVRLVKLQSLRREYENLKM 127
Query: 125 EEEETISQYFDKFVNLSNQ 143
+ + I + DK + L Q
Sbjct: 128 YDNDNIKTFTDKLIVLEIQ 146
>At3g60170 putative protein
Length = 1339
Score = 80.9 bits (198), Expect = 3e-16
Identities = 44/137 (32%), Positives = 80/137 (58%), Gaps = 5/137 (3%)
Query: 10 LPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNAT---DMQ*TANRDAKKKDGK 66
+P F+G YD W + M+ R +++ +V + +PA+ T + Q +A +AK KD K
Sbjct: 12 IPRFDGY-YDFWSMTMENFLRSRELWRLVEEGIPAIVVGTTPVSEAQRSAVEEAKLKDLK 70
Query: 67 AMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEE 126
F+ Q++ EI E I+ +S + WE+++K Y G K+K+ +LQ LR+++E+L M+E
Sbjct: 71 VKNFLFQAIDREILETILD-KSTSKAIWESMKKKYQGSTKVKRAQLQALRKEFELLAMKE 129
Query: 127 EETISQYFDKFVNLSNQ 143
E I + + + + N+
Sbjct: 130 GEKIDTFLGRTLTVVNK 146
>At2g05930 copia-like retroelement pol polyprotein
Length = 916
Score = 61.6 bits (148), Expect = 2e-10
Identities = 44/152 (28%), Positives = 74/152 (47%), Gaps = 12/152 (7%)
Query: 12 VFEGKNYDQWIVKMKVICRFQDVLEVVNDSV----PALASN------ATDMQ*TANRDAK 61
+ E NY W VKM+ + R + S+ P + T+ Q +AK
Sbjct: 27 ILEKGNYGHWKVKMRALIRGLGKEAWIATSIGWKAPVIKGEDGEDVLKTEDQWNDAEEAK 86
Query: 62 KK-DGKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYE 120
+ +A+ + SV+ F++I + ESAKE W+ L K Y G +K+ R+ +L Q+E
Sbjct: 87 ATANSRALSLIFNSVNQNQFKQIQNCESAKE-AWDKLAKAYEGTSSVKRSRIDMLASQFE 145
Query: 121 VLTMEEEETISQYFDKFVNLSNQSQEW*KCYR 152
LTMEE E I ++ K +++++ K Y+
Sbjct: 146 NLTMEETENIEEFSGKISAIASEAHNLGKKYK 177
>At3g21010 unknown protein
Length = 438
Score = 60.1 bits (144), Expect = 6e-10
Identities = 40/139 (28%), Positives = 69/139 (48%), Gaps = 13/139 (9%)
Query: 12 VFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TAN-------RDAKKKD 64
V + NY+ W K + + +VV + VP S ++ T RD +KD
Sbjct: 11 VSDDLNYEIWAPIAKTTLTEKGLWDVVENGVPPDPSKVPELAATIQAEEFSKWRDLAEKD 70
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ + S+++ F K + SAK+ W++LEK G ++ +L+ L +Q+E L+M
Sbjct: 71 MKALQILQSSLTDSAFRKTLSASSAKD-LWDSLEK-----GNNEQAKLRRLEKQFEELSM 124
Query: 125 EEEETISQYFDKFVNLSNQ 143
E E I+ Y D+ + + Q
Sbjct: 125 NEGEPINPYIDRVIEIVEQ 143
>At4g04420 putative transposon protein
Length = 1008
Score = 58.9 bits (141), Expect = 1e-09
Identities = 41/152 (26%), Positives = 70/152 (45%), Gaps = 12/152 (7%)
Query: 12 VFEGKNYDQWIVKMKVICRFQDVLEVVNDSV----PALASN-------ATDMQ*TANRDA 60
+ E NY W VKM+ + R + S+ P + D A
Sbjct: 15 MLEKGNYGHWKVKMRALIRGLGKEAWIATSIGWKAPVIKGEDGEDVLKTKDQWNDAEEAK 74
Query: 61 KKKDGKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYE 120
K + +A+ + V+ F++I + ESAKE W+ L K Y G +K+ R+ +L Q+E
Sbjct: 75 AKANSRALSLIFNFVNQNQFKRIQNCESAKE-AWDKLAKAYEGTSSVKRSRIDMLASQFE 133
Query: 121 VLTMEEEETISQYFDKFVNLSNQSQEW*KCYR 152
L+MEE E I ++ K +++++ K Y+
Sbjct: 134 NLSMEETENIEEFSGKISAIASEAHNLGKKYK 165
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 58.5 bits (140), Expect = 2e-09
Identities = 30/93 (32%), Positives = 55/93 (58%), Gaps = 1/93 (1%)
Query: 50 TDMQ*TANRDAKKKDGKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKK 109
T Q T + KD K ++ S+ I + I+ E++K+ WE++++ Y G+ +++
Sbjct: 11 TGAQRTELAEKTVKDHKVKNYLFASIDKTILKTILQKETSKD-LWESMKRKYQGNDRVQS 69
Query: 110 VRLQVLRRQYEVLTMEEEETISQYFDKFVNLSN 142
+LQ LRR +EVL M+ ETI+ YF + + ++N
Sbjct: 70 AQLQRLRRSFEVLEMKIGETITGYFSRVMEITN 102
>At3g21020 unknown protein
Length = 310
Score = 54.3 bits (129), Expect = 3e-08
Identities = 39/139 (28%), Positives = 68/139 (48%), Gaps = 13/139 (9%)
Query: 12 VFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPA-------LASNATDMQ*TANRDAKKKD 64
V + NY+ W K + + +VV + VP LA+ + + RD KD
Sbjct: 11 VSDDLNYEIWARIAKTTLTEKGLWDVVENGVPPDPSKVPELAAKIQPEELSKWRDLAVKD 70
Query: 65 GKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTM 124
KA+ + S+++ K ++ SAK+ W+ LEK G ++ +L+ L +Q+E L+M
Sbjct: 71 MKALQNLQSSLTDSALRKTLYASSAKD-VWDLLEK-----GNNEQAKLRRLEKQFEELSM 124
Query: 125 EEEETISQYFDKFVNLSNQ 143
+E E + Y DK ++ Q
Sbjct: 125 DEAEQTNSYLDKVRKIAEQ 143
>At4g04560 putative transposon protein
Length = 590
Score = 52.8 bits (125), Expect = 1e-07
Identities = 37/138 (26%), Positives = 59/138 (41%), Gaps = 28/138 (20%)
Query: 11 PVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATD-----MQ*TANRDAKKKDG 65
P+F +NY W +KMK I + + + E+V++ VP + Q T A KD
Sbjct: 8 PIFNKENYGFWRIKMKTIFQTKKLWEIVDEGVPKPPAEGDHSPEAVQQKTRCEAASLKDL 67
Query: 66 KAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTME 125
A+ + +VS+ IF +I SA + WE YE L M+
Sbjct: 68 TALQILQTAVSDSIFPRIAPASSALGKPWE-----------------------YENLKMK 104
Query: 126 EEETISQYFDKFVNLSNQ 143
E + I+ + K + + NQ
Sbjct: 105 ESDNINTFMTKLIEMGNQ 122
Score = 42.7 bits (99), Expect = 1e-04
Identities = 25/95 (26%), Positives = 49/95 (51%), Gaps = 1/95 (1%)
Query: 58 RDAKKKDGKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRR 117
+ A + +A+ + S+ F ++ SAKE WE L+ + +K+ RL +L
Sbjct: 404 KTAANHNSQALSVIFGSLLRNKFTQVQGCLSAKE-VWEILQVSFECTNNVKRTRLDMLAS 462
Query: 118 QYEVLTMEEEETISQYFDKFVNLSNQSQEW*KCYR 152
++E LTME EE++ + K +++ ++ K Y+
Sbjct: 463 EFENLTMEAEESVDDFNGKLSSITQEAVVLGKTYK 497
>At2g07080 putative gag-protease polyprotein
Length = 627
Score = 51.2 bits (121), Expect = 3e-07
Identities = 40/142 (28%), Positives = 66/142 (46%), Gaps = 11/142 (7%)
Query: 12 VFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TANRDAKKK-DGKAMFF 70
+ + K Y W V M I R Q+ D A TA + K + +AM
Sbjct: 15 LLDTKRYGYWKVCMTQIIRGQE------DGFKITKPKANW---TAEEKLQSKFNARAMKA 65
Query: 71 MHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEETI 130
+ V + F+ I +SAK+ W+ L+K + G +K+ RL + Q+E L ME +E I
Sbjct: 66 IFNGVDEDEFKLIQGCKSAKQ-AWDTLQKSHEGTSSVKRTRLDHIATQFEYLKMEPDEKI 124
Query: 131 SQYFDKFVNLSNQSQEW*KCYR 152
++ K L+N+++ K Y+
Sbjct: 125 VKFSSKISALANEAEVMGKTYK 146
>At3g20990 unknown protein
Length = 379
Score = 48.9 bits (115), Expect = 1e-06
Identities = 35/132 (26%), Positives = 65/132 (48%), Gaps = 14/132 (10%)
Query: 17 NYDQWIVKMKVICRFQDVLEVVNDSVPA-------LASNATDMQ*TANRDAKKKDGKAMF 69
+Y+ W K +++ +VV + VP LA+ + + RD KD KA+
Sbjct: 16 DYEIWSSVTKATLTEKELWDVVENGVPPDPSKIPELATKIQPEELSRWRDLAVKDMKALQ 75
Query: 70 FMHQS-VSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEE 128
+ S +++ F K + SAK+ W++LEK G ++ +L+ L +Q+E + EE E
Sbjct: 76 ILQSSSLTDSAFRKTLSASSAKD-LWDSLEK-----GNNEQAKLRRLEKQFEEIKREENE 129
Query: 129 TISQYFDKFVNL 140
++ Y D+ +
Sbjct: 130 SLDSYIDRITEI 141
>At3g21030 hypothetical protein
Length = 411
Score = 47.4 bits (111), Expect = 4e-06
Identities = 36/127 (28%), Positives = 58/127 (45%), Gaps = 12/127 (9%)
Query: 17 NYDQWIVKMKVICRFQDVLEVVNDSV-------PALASNATDMQ*TANRDAKKKDGKAMF 69
+Y+ W +K V +VV + V P LA+ + R+ KD KA+
Sbjct: 14 DYEDWSPMVKTRLVENGVWDVVQNGVSPNPTTNPELAATIKAVDLAQWRNLVVKDNKALK 73
Query: 70 FMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEET 129
M S+ + +F K + SAK E W+ L+K K+ +L+ L +Q+E L M E E
Sbjct: 74 IMQSSLPDSVFRKTISIASAK-ELWDLLKK----GNDTKEAKLRRLEKQFENLMMYEGER 128
Query: 130 ISQYFDK 136
+ Y +
Sbjct: 129 MKLYLKR 135
>At3g21000 unknown protein
Length = 405
Score = 47.0 bits (110), Expect = 5e-06
Identities = 37/133 (27%), Positives = 65/133 (48%), Gaps = 10/133 (7%)
Query: 17 NYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TAN-------RDAKKKDGKAMF 69
+Y+ W K Q + +VV + VP S ++ T RD KD KA+
Sbjct: 16 DYEIWAPITKSTLIEQGLWDVVVNGVPQDPSKNPELAATIQPEELSKWRDFVVKDAKALQ 75
Query: 70 FMHQSVSNEIFEKIVHYESAKEETWEALEK--LYSGDGKLKKVRLQVLRRQYEVLTMEEE 127
+ S+++ +F K + SAK + W+ L K + +L++V ++ L +Q E L M ++
Sbjct: 76 ILQSSLTDSVFRKTLSASSAK-DVWDLLRKGNEQATIRRLEQVTIRRLEKQLEDLKMVDK 134
Query: 128 ETISQYFDKFVNL 140
E+ S Y DK + +
Sbjct: 135 ESGSSYLDKALEI 147
>At1g36110 hypothetical protein
Length = 745
Score = 47.0 bits (110), Expect = 5e-06
Identities = 32/144 (22%), Positives = 63/144 (43%), Gaps = 19/144 (13%)
Query: 3 LESFP--THLPVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TANRDA 60
LE+ P T P+ NY W ++M++ + E + + D
Sbjct: 10 LENVPSSTQCPMLTSTNYTVWAIRMQIARSVNKMWETIEPGI----------------DD 53
Query: 61 KKKDGKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYE 120
K+ A + QS+ + ++ +AK W++++ Y G ++K+ RLQ L ++E
Sbjct: 54 GYKNTMARGLIFQSIPESLTLQVGTLATAKL-VWDSIKTRYVGADRVKEARLQTLMAEFE 112
Query: 121 VLTMEEEETISQYFDKFVNLSNQS 144
+ M+E E I + + L+ +S
Sbjct: 113 KMKMKESEKIDVFAGRLAELATRS 136
>At3g21050 hypothetical protein
Length = 420
Score = 45.4 bits (106), Expect = 2e-05
Identities = 36/134 (26%), Positives = 60/134 (43%), Gaps = 12/134 (8%)
Query: 17 NYDQWIVKMKVICRFQDVLEVVNDSV-------PALASNATDMQ*TANRDAKKKDGKAMF 69
+Y+ W K V +VV + V P LA+ + R+ KD KA+
Sbjct: 14 DYEHWSPIAKKRLVENGVWDVVQNGVSPNPTKIPQLAATIQAVDLAQWRNRVIKDNKALK 73
Query: 70 FMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEET 129
M S+ + +F K + SAK E W+ L+K K+ +L+ L +Q+E L M E E
Sbjct: 74 IMQSSLPDSVFRKTISIASAK-ELWDLLKK----GNDTKEAKLRRLEKQFEKLMMYEGEP 128
Query: 130 ISQYFDKFVNLSNQ 143
+ Y + ++ +
Sbjct: 129 MDLYLKRVEEITER 142
>At3g25450 hypothetical protein
Length = 1343
Score = 45.1 bits (105), Expect = 2e-05
Identities = 34/150 (22%), Positives = 61/150 (40%), Gaps = 23/150 (15%)
Query: 11 PVFEGKNYDQWIVKMKVICRFQDVLEVVNDSVPALASNATDMQ*TANRDAKKKDGKAMFF 70
P+ NY W ++M+ + R + + + D +K D A
Sbjct: 23 PMLNSVNYTVWTMRMEAVLRVHKLWGTIEPG---------------SADEEKND-MARAL 66
Query: 71 MHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQYEVLTMEEEETI 130
+ QS+ + + V + WEA++ G ++K+ RLQ L +++ L M++ ETI
Sbjct: 67 LFQSIPESLILQ-VGKQKTSSAVWEAIKSRNLGAERVKEARLQTLMAEFDKLKMKDSETI 125
Query: 131 SQYFDKFVNLSNQSQEW*KCYRLDED*ESS 160
Y + ++ K L ED E S
Sbjct: 126 DDYVGRISEITT------KAAALGEDIEES 149
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 43.1 bits (100), Expect = 8e-05
Identities = 22/87 (25%), Positives = 48/87 (54%), Gaps = 1/87 (1%)
Query: 59 DAKKKDGKAMFFMHQSVSNEIFEKIVHYESAKEETWEALEKLYSGDGKLKKVRLQVLRRQ 118
D +K+ A + QSV ++ ++++K WEA++ G ++K+ +LQ L +
Sbjct: 19 DDMEKNDMARALLFQSVPESTILQVGKHKTSKA-MWEAIKTRNLGAERVKEAKLQTLMAE 77
Query: 119 YEVLTMEEEETISQYFDKFVNLSNQSQ 145
++ L M++ ETI ++ + +S +S+
Sbjct: 78 FDRLNMKDNETIDEFVGRISEISTKSE 104
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.332 0.140 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,596,112
Number of Sequences: 26719
Number of extensions: 136999
Number of successful extensions: 859
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 791
Number of HSP's gapped (non-prelim): 64
length of query: 174
length of database: 11,318,596
effective HSP length: 92
effective length of query: 82
effective length of database: 8,860,448
effective search space: 726556736
effective search space used: 726556736
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149197.6


