

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149135.13 - phase: 0
(316 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
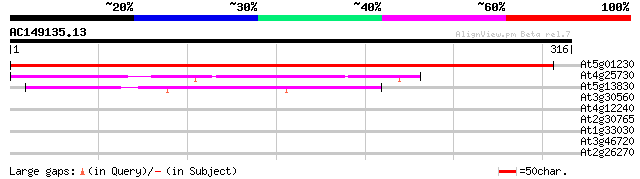
Score E
Sequences producing significant alignments: (bits) Value
At5g01230 cell division - like protein 575 e-165
At4g25730 unknown protein 131 5e-31
At5g13830 FtsJ (dbj|BAA83750.1) 67 1e-11
At3g30560 hypothetical protein 30 1.3
At4g12240 hypothetical proteins 30 1.7
At2g30765 hypothetical protein 30 2.2
At1g33030 putative protein 29 2.9
At3g46720 glucuronosyl transferase-like protein 29 3.8
At2g26270 hypothetical protein 28 8.5
>At5g01230 cell division - like protein
Length = 309
Score = 575 bits (1483), Expect = e-165
Identities = 283/307 (92%), Positives = 294/307 (95%), Gaps = 1/307 (0%)
Query: 1 MGKASRDKRDIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQVLS 60
MGKASRDKRDIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQVLS
Sbjct: 1 MGKASRDKRDIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQVLS 60
Query: 61 RKLYLPAKLAPDAKDENLPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRHFDGCKA 120
R+LYLPAK + ++KD +LPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRHFDGCKA
Sbjct: 61 RQLYLPAKSSAESKDGDLPLIVAIDLQPMAPIEGVIQVQGDITNARTAEVVIRHFDGCKA 120
Query: 121 NLVVCDGAPDVTGLHDMDEFVQSQLILAGLTIVTHVLKEGGKFIAKIFRGKDTSLLYCQL 180
+LVVCDGAPDVTGLHDMDEFVQSQLILAGLTIVTH+LKEGGKFIAKIFRGKDTSLLYCQL
Sbjct: 121 DLVVCDGAPDVTGLHDMDEFVQSQLILAGLTIVTHILKEGGKFIAKIFRGKDTSLLYCQL 180
Query: 181 KLFFPVVTFAKPKSSRNSSIEAFAVCENYSPPEGFNPKDLHRLLEKVGSPSGVDDTDCVS 240
KLFFP VTFAKPKSSRNSSIEAFAVCENYSPPEGFNP+DLHRLLEKVGSPSG D DC S
Sbjct: 181 KLFFPTVTFAKPKSSRNSSIEAFAVCENYSPPEGFNPRDLHRLLEKVGSPSGGSDLDCSS 240
Query: 241 GWLEGPNKVYIPFLACGDLTGYDSDRSYPLPKVA-GGTYQSLDPVQPPIAPPYKRALELK 299
GWLEGPNKVYIPFLACGDLTGYDSDRSYPLP+ A G +YQSLDP+QPPIAPPYKRALELK
Sbjct: 241 GWLEGPNKVYIPFLACGDLTGYDSDRSYPLPREADGSSYQSLDPIQPPIAPPYKRALELK 300
Query: 300 KASPQGF 306
KAS Q F
Sbjct: 301 KASAQSF 307
>At4g25730 unknown protein
Length = 821
Score = 131 bits (329), Expect = 5e-31
Identities = 83/240 (34%), Positives = 131/240 (54%), Gaps = 24/240 (10%)
Query: 1 MGKASRDKR-DIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQVL 59
MGK R D YYR AKE G+R+R+++KLLQ+D ++++ V+DLCAAPG W QV
Sbjct: 1 MGKVKGKHRLDKYYRLAKERGFRSRASYKLLQLDAKYSLLHSAHAVLDLCAAPGGWMQVA 60
Query: 60 SRKLYLPAKLAPDAKDENLPLIVAIDLQPMAPIEGVIQVQGDIT----NARTAEVVIRHF 115
K+ + + L++ IDL P+ P+ G + + DIT ++ +V+ +H
Sbjct: 61 VEKVPVGS------------LVLGIDLVPILPVRGCVTMTQDITRTECKSKIKQVMEQH- 107
Query: 116 DGCKA-NLVVCDGAPDVTGLHDMDEFVQSQLILAGLTIVTHVLKEGGKFIAKIFRGKD-T 173
G A NLV+ DG+P+V G + Q+ L++ + + T L G + K+FR +D
Sbjct: 108 -GVSAFNLVLHDGSPNVGGAWAQEAMSQNALVIDSVRLATEFLARNGNLVTKVFRSRDYN 166
Query: 174 SLLYCQLKLFFPVVTFAKPKSSRNSSIEAFAVCENYSPPEGFNPK--DLHRLLEKVGSPS 231
S+LYC +LF V F KP +SR++S E + V Y P +P+ D L ++ P+
Sbjct: 167 SVLYCLGRLFEKVEVF-KPPASRSASAETYLVGLKYLAPAKIDPRLLDYRHLFKESAEPT 225
>At5g13830 FtsJ (dbj|BAA83750.1)
Length = 224
Score = 67.0 bits (162), Expect = 1e-11
Identities = 52/225 (23%), Positives = 98/225 (43%), Gaps = 34/225 (15%)
Query: 10 DIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGSWSQVLSRKLYLPAKL 69
D +YR+A+ G+ ARSAFKLLQI +++ + + V+DL APG+W QV + L
Sbjct: 8 DFFYREAQRLGYVARSAFKLLQIQKQYKLIKPGSSVLDLGCAPGAWLQVACQSL------ 61
Query: 70 APDAKDENLPLIVAIDLQ----PMAPIEGVIQVQGDITNARTAEVVIRHFDGCKANLVVC 125
+ ++V +D++ P V + D+ N ++ ++++
Sbjct: 62 ---GPLRSGGIVVGMDIKKVKVPPQCDSRVQTIAADVLNFPRQKIRELSPQQLGFSVILS 118
Query: 126 DGAPDVTGLHDMDEFVQSQLILAGLTIVT---------------------HVLKEGGKFI 164
D V+G+ D + ++L + L + VL+ GG +
Sbjct: 119 DMCHSVSGITTRDAALSAELGMRALDLAVGQAAISQSPNDDDGGPNESRPGVLRHGGHLV 178
Query: 165 AKIFRGKDTSLLYCQLKLFFPVVTFAKPKSSRNSSIEAFAVCENY 209
K+ +D K F ++ +PK++R SS E + +C+ +
Sbjct: 179 IKLLESEDAQDFARICKPIFNKASWLRPKATRPSSREIYLICQGF 223
>At3g30560 hypothetical protein
Length = 1473
Score = 30.4 bits (67), Expect = 1.3
Identities = 28/95 (29%), Positives = 41/95 (42%), Gaps = 8/95 (8%)
Query: 175 LLYCQLKLFFPVVTFAKPKSSRNSSIEAFAVCE-NYSPPEGFNPKDLHRLLEKVGSPSGV 233
LLY Q+ F+ K K+ R + F V N+ PP+ + L L+ + P G
Sbjct: 824 LLYEQIPNFY--TWNGKDKNFRRRKMPGFVVGRINHVPPKIDDAYHLRILINNIRGPKGF 881
Query: 234 DDTDCVSGWLEGPNKVYIPFLACGDLTGYDSDRSY 268
DD + G + +K Y AC L D D+ Y
Sbjct: 882 DDIKTIEGVV---HKTYRD--ACYALGLLDDDKDY 911
>At4g12240 hypothetical proteins
Length = 364
Score = 30.0 bits (66), Expect = 1.7
Identities = 14/41 (34%), Positives = 21/41 (51%)
Query: 262 YDSDRSYPLPKVAGGTYQSLDPVQPPIAPPYKRALELKKAS 302
+ S P PK+ G + LD P PPY A++L+ A+
Sbjct: 27 FSSSSINPKPKIRVGVWWDLDNKPPASFPPYDAAVKLRTAA 67
>At2g30765 hypothetical protein
Length = 116
Score = 29.6 bits (65), Expect = 2.2
Identities = 18/69 (26%), Positives = 36/69 (52%), Gaps = 2/69 (2%)
Query: 36 FNIFEGVKR-VVDLCAAPGSWSQVLSRKLYLPAKLAPDAKDENLPLIVAIDLQPMAPIEG 94
F ++ R V +CAAP ++ ++L A + +KDE++ LI ++ + ++
Sbjct: 8 FTLYRATNRQFVAVCAAPSAFFFFFCEHVFLFADVETSSKDESV-LITKVETETSNEVKV 66
Query: 95 VIQVQGDIT 103
+QV G+ T
Sbjct: 67 HVQVSGEKT 75
>At1g33030 putative protein
Length = 352
Score = 29.3 bits (64), Expect = 2.9
Identities = 25/104 (24%), Positives = 49/104 (47%), Gaps = 11/104 (10%)
Query: 10 DIYYRKAKEEGWRARSAFKLLQIDEEFNIFEGVKRVVDLCAAPGS-WSQVLSRKLYLPAK 68
D +R+ + + + + + + +N F+GVK +VD+ GS S+++S+ ++
Sbjct: 155 DSRFREVFQSSMKGFNEVFIEEFLKNYNGFDGVKSLVDVGGGDGSLLSRIISKHTHIIKA 214
Query: 69 LAPDAKDENLPLIVAIDLQPMAPIEGVIQVQGDI-TNARTAEVV 111
+ D LP ++ L P G+ V GD+ TN E +
Sbjct: 215 INFD-----LPTVINTSL----PSPGIEHVAGDMFTNTPKGEAI 249
>At3g46720 glucuronosyl transferase-like protein
Length = 447
Score = 28.9 bits (63), Expect = 3.8
Identities = 17/61 (27%), Positives = 27/61 (43%), Gaps = 10/61 (16%)
Query: 68 KLAPDAKDENLPLIVAIDLQP----------MAPIEGVIQVQGDITNARTAEVVIRHFDG 117
K D KD + +V +L P M P+E +++ ++ N RTA VI +
Sbjct: 153 KFLIDMKDPEVQNMVVENLHPLKYKDLPTSGMGPLERFLEICAEVVNKRTASAVIINTSS 212
Query: 118 C 118
C
Sbjct: 213 C 213
>At2g26270 hypothetical protein
Length = 470
Score = 27.7 bits (60), Expect = 8.5
Identities = 14/34 (41%), Positives = 22/34 (64%), Gaps = 4/34 (11%)
Query: 95 VIQVQGDITNARTAEVVIRHFDGCKANLVVCDGA 128
V++V G+ + A+V+ F GC A+ VVC+GA
Sbjct: 150 VLKVAGE----QGAKVIDSWFIGCNASFVVCEGA 179
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,605,956
Number of Sequences: 26719
Number of extensions: 345890
Number of successful extensions: 718
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 709
Number of HSP's gapped (non-prelim): 10
length of query: 316
length of database: 11,318,596
effective HSP length: 99
effective length of query: 217
effective length of database: 8,673,415
effective search space: 1882131055
effective search space used: 1882131055
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC149135.13


