

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149134.1 - phase: 0
(555 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
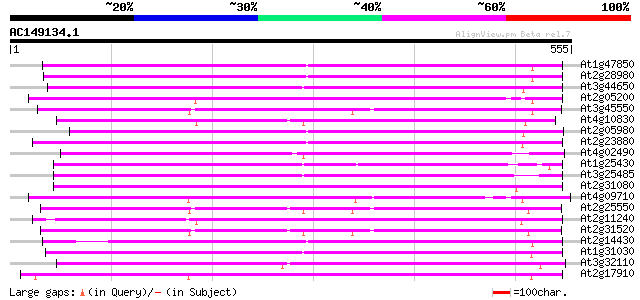
Score E
Sequences producing significant alignments: (bits) Value
At1g47850 hypothetical protein 322 4e-88
At2g28980 putative non-LTR retroelement reverse transcriptase 299 3e-81
At3g44650 putative protein 296 3e-80
At2g05200 putative non-LTR retroelement reverse transcriptase 295 6e-80
At3g45550 putative protein 293 2e-79
At4g10830 putative protein 292 3e-79
At2g05980 putative non-LTR retroelement reverse transcriptase 292 3e-79
At2g23880 putative non-LTR retroelement reverse transcriptase 291 6e-79
At4g02490 288 7e-78
At1g25430 hypothetical protein 286 2e-77
At3g25485 unknown protein 284 1e-76
At2g31080 putative non-LTR retroelement reverse transcriptase 282 4e-76
At4g09710 RNA-directed DNA polymerase -like protein 281 9e-76
At2g25550 putative non-LTR retroelement reverse transcriptase 281 9e-76
At2g11240 pseudogene 281 9e-76
At2g31520 putative non-LTR retroelement reverse transcriptase 280 2e-75
At2g14430 putative non-LTR retroelement reverse transcriptase 280 2e-75
At1g31030 F17F8.5 280 2e-75
At3g32110 non-LTR reverse transcriptase, putative 278 4e-75
At2g17910 putative non-LTR retroelement reverse transcriptase 270 2e-72
>At1g47850 hypothetical protein
Length = 799
Score = 322 bits (824), Expect = 4e-88
Identities = 180/520 (34%), Positives = 276/520 (52%), Gaps = 6/520 (1%)
Query: 33 ATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKN 92
AT +LT T EE + +F + S+ PGPD + + FF+ W+I +D A+ FF
Sbjct: 16 ATDQDMLTREVTSEENQKVLFAMPSNKFPGPDGYTSEFFKATWSITGQDFIAAIKSFFIK 75
Query: 93 GWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQ 152
G+LP NA + LIPK A + YR I+ N +K+I+K++A+RL +LP+ I + Q
Sbjct: 76 GFLPKGLNATILALIPKKDEATLMRDYRPISCCNVIYKVISKIIANRLKVMLPTFILQNQ 135
Query: 153 RGFVQGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFN 212
FV+ R + + + L +E + S A+KID++KAFD++ W FLL L+ F
Sbjct: 136 SAFVRERLLIENVLLATELVKDYHKDSISPRCAMKIDISKAFDSVQWQFLLNTLEALNFP 195
Query: 213 ELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILA 272
E FC+WIK + ++ + +NG GFF RG+RQG LSP LF I VLS I + A
Sbjct: 196 ENFCHWIKLCISTATFSVQVNGELAGFFGSKRGLRQGCALSPYLFVICMNVLSHMIDVAA 255
Query: 273 DKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKS 332
I + L CF DDLMVF + S+ + ++F +A SG +++ KS
Sbjct: 256 VHRNIGYHPKCKKLSLTHLCF-ADDLMVFIDGQQRSVEGVINIFKEFAGKSGLHISLEKS 314
Query: 333 FIFAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWK 392
++ G+++ NNI++ F G LP YLG P+ + + P+ DKV++K++ W
Sbjct: 315 TLYLAGVSELNRNNILSAFPFASGQLPVRYLGLPLLTKQMTTADYSPLLDKVRSKISSWT 374
Query: 393 ASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVA 452
A LS AGR+ L+ SV+ S+ MS Y P +KE+EK F+WSG K +
Sbjct: 375 ARSLSYAGRLALINSVIVSLSNFWMSAYRLPAGCIKEIEKLCSAFLWSGPELNPKKAKIT 434
Query: 453 WRKICADYEEGGLGVKSLICLNEATNLKICWNLM-QSDEQWANIIRSRVIRDHRCMNHHI 511
W +C +EGGLG+KSL+ N+ + LK+ W L+ + W N + + +IR + +
Sbjct: 435 WTSLCKLKQEGGLGIKSLLEANKVSCLKLIWRLVSRQSSLWVNWVWTYIIRKGSFWSAND 494
Query: 512 SSSI----WSGVKAEFHVIKDNCSWIVGDGKQINFWSDSW 547
SS+ W + V K C + G +FW D+W
Sbjct: 495 RSSLGSWMWKKLLKYRDVAKSMCKVEIKSGSSTSFWYDNW 534
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 299 bits (765), Expect = 3e-81
Identities = 176/519 (33%), Positives = 268/519 (50%), Gaps = 6/519 (1%)
Query: 34 TTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNG 93
T +LT T EE++ +F + ++ +PGPD + + FF+ W++ D A+ FF G
Sbjct: 741 TDQNILTREVTGEEIQKVLFAMPNNKSPGPDGYTSEFFKATWSLTGPDFIAAIQSFFVKG 800
Query: 94 WLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQR 153
+LP NA + LIPK A + YR I+ N +K+I+K+LA+RL +LPS I + Q
Sbjct: 801 FLPKGLNATILALIPKKDEAIEMKDYRPISCCNVLYKVISKILANRLKLLLPSFILQNQS 860
Query: 154 GFVQGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNE 213
FV+ R + + + L +E + +S A+KID++KAFD++ W FLL L+ F E
Sbjct: 861 AFVKERLLMENVLLATELVKDYHKESVTPRCAMKIDISKAFDSVQWQFLLNTLEALNFPE 920
Query: 214 LFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILAD 273
F +WIK + ++ + +NG GFF +RG+RQG LSP LF I VLS I A
Sbjct: 921 TFRHWIKLCISTATFSVQVNGELAGFFGSSRGLRQGCALSPYLFVICMNVLSHMIDEAAV 980
Query: 274 KGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSF 333
I L CF DDLMVF S+ + ++F +A SG +++ KS
Sbjct: 981 HRNIGYHPKCEKIGLTHLCF-ADDLMVFVDGHQWSIEGVINVFKEFAGRSGLQISLEKST 1039
Query: 334 IFAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKA 393
I+ G++ + ++ F G LP YLG P+ + + P+ + VK K++ W A
Sbjct: 1040 IYLAGVSASDRVQTLSSFPFANGQLPVRYLGLPLLTKQMTTADYSPLIEAVKTKISSWTA 1099
Query: 394 SLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAW 453
LS AGR+ L+ SV+ S+ MS Y P ++E+EK F+WSG V K +AW
Sbjct: 1100 RSLSYAGRLALLNSVIVSIANFWMSAYRLPAGCIREIEKLCSAFLWSGPVLNPKKAKIAW 1159
Query: 454 RKICADYEEGGLGVKSLICLNEATNLKICWNLMQSDEQ-WANIIRSRVIRDHRCMNHHIS 512
IC +EGGLG+KSL N+ + LK+ W L+ + W I + +IR + +
Sbjct: 1160 SSICQPKKEGGLGIKSLAEANKVSCLKLIWRLLSTQPSLWVTWIWTFIIRKGTFWSANER 1219
Query: 513 SSI----WSGVKAEFHVIKDNCSWIVGDGKQINFWSDSW 547
SS+ W + + K V +G +FW D W
Sbjct: 1220 SSLGSWMWKKLLKYRELAKSMHKVEVRNGSSTSFWYDHW 1258
>At3g44650 putative protein
Length = 762
Score = 296 bits (757), Expect = 3e-80
Identities = 158/515 (30%), Positives = 268/515 (51%), Gaps = 6/515 (1%)
Query: 38 LLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPN 97
+LT + + EE+K +F + +D +PGPD F + FF+ W I+ + A+ FF G+LP
Sbjct: 1 MLTRVVSAEEIKKVLFSMPNDKSPGPDGFTSEFFKESWEILGPEFILAIQSFFALGFLPK 60
Query: 98 NFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQ 157
N+ + LIPK + + YR I+ N +K+I+K+LA+RL +LP I+ Q FV+
Sbjct: 61 GVNSTILALIPKKLESKEMKDYRPISCCNVMYKVISKILANRLKLLLPQFIAGNQSSFVK 120
Query: 158 GRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCN 217
R + + + L ++ + S A+KID++KA D++ W FL+ L F E+F +
Sbjct: 121 DRLLIENVLLATDLVKDYHKDSISERCAIKIDISKASDSVQWSFLINTLTAMHFPEMFIH 180
Query: 218 WIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGLI 277
WI+ + + + +NG GFF +RG+RQG LSP LF I +VLS+ + + G I
Sbjct: 181 WIRLCITTPSFSVQVNGELAGFFQSSRGLRQGCALSPYLFVICMDVLSKLLDKVVGIGRI 240
Query: 278 DLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFIFAG 337
+ L H + DDLM+ + S+ + +F ++ SG +++ KS IF+
Sbjct: 241 GYHPHCKRMGLT-HLSFADDLMILTDGQCRSIEGIIEVFDLFSKWSGLKISMEKSTIFSA 299
Query: 338 GITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLS 397
G++ T + F VG LP YLG P+ + + + P+ ++++ ++ W + LS
Sbjct: 300 GLSSTSRAQLHTHFPFEVGELPIRYLGLPLVTKRLSSVDYAPLIEQIRKRIGSWSSRFLS 359
Query: 398 IAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKIC 457
AGR L+ S++ S +S + P ++E+EK +F+WSG K ++W ++C
Sbjct: 360 FAGRFNLISSIIWSSCNFWLSAFQLPRACIQEIEKLCSSFLWSGTNLNSKKAKISWNQVC 419
Query: 458 ADYEEGGLGVKSLICLNEATNLKICWNLM-QSDEQWANIIRSRVIRDHRCM----NHHIS 512
EGGLG++SL N+ LK+ W ++ D W + +++ N ++
Sbjct: 420 KPKSEGGLGLRSLKEANDVCCLKLVWRIISHGDSLWVKWVEHNLLKREIFWIVKENANLG 479
Query: 513 SSIWSGVKAEFHVIKDNCSWIVGDGKQINFWSDSW 547
S IW + V K C VG+G+ +FW D W
Sbjct: 480 SWIWKKILKYRGVAKRFCKAEVGNGESTSFWFDDW 514
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 295 bits (754), Expect = 6e-80
Identities = 179/543 (32%), Positives = 273/543 (49%), Gaps = 21/543 (3%)
Query: 19 DHLLVEEAIPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIV 78
D +V+EAI +V N LT +P EEVK+AVF +N+ APGPD F A F+ YW+I+
Sbjct: 299 DFSVVDEAIEPMVSQGDNDFLTRIPNDEEVKDAVFSINASKAPGPDGFTAGFYHSYWHII 358
Query: 79 KKDVYEAVLDFFKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLAD 138
DV + FF + P N I LIPK V YR IAL N +KI+ K++
Sbjct: 359 STDVGREIRLFFTSKNFPRRMNETHIRLIPKDLGPRKVADYRPIALCNIFYKIVAKIMTK 418
Query: 139 RLAKILPSIISKEQRGFVQGRNIRDCIALTSEAINVLDNKSFGG--NLALKIDVTKAFDT 196
R+ ILP +IS+ Q FV GR I D + +T E ++ L S ++A+K D++KA+D
Sbjct: 419 RMQLILPKLISENQSAFVPGRVISDNVLITHEVLHFLRTSSAKKHCSMAVKTDMSKAYDR 478
Query: 197 LNWDFLLLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLL 256
+ WDFL VL+ FGF+ ++ +W+ + S +NG G RG+RQGDPLSP L
Sbjct: 479 VEWDFLKKVLQRFGFHSIWIDWVLECVTSVSYSFLINGTPQGKVVPTRGLRQGDPLSPCL 538
Query: 257 FCIVEEVLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLF 316
F + EVLS + + + S N H + DD M F K+ S L +
Sbjct: 539 FILCTEVLSGLCTRAQRLRQLPGVRVSINGPRVNHLLFADDTMFFSKSDPESCNKLSEIL 598
Query: 317 TRYADCSGQIMNIRKSFI-FAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGI 375
+RY SGQ +N KS + F+ + + IL YLG P G+ K
Sbjct: 599 SRYGKASGQSINFHKSSVTFSSKTPRSVKGQVKRILKIRKEGGTGKYLGLPEHFGRRKRD 658
Query: 376 HFQPIADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIK 435
F I DK++ K W + LS AG+ ++K+V+ SM +++MS + P + ++++ +
Sbjct: 659 IFGAIIDKIRQKSHSWASRFLSQAGKQVMLKAVLASMPLYSMSCFKLPSALCRKIQSLLT 718
Query: 436 NFIWSGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQSDEQWANI 495
F W RK VAW K+ GGLG + + N++ K+ W L+ S E
Sbjct: 719 RFWWDTKPDVRKTSWVAWSKLTNPKNAGGLGFRDIERCNDSLLAKLGWRLLNSPES---- 774
Query: 496 IRSRVIRDHRCMNHHISSSI-----------WSGVKAEFHVIKDNCSWIVGDGKQINFWS 544
+ SR++ C H SS + W + A ++K+ W++ +G++++ W+
Sbjct: 775 LLSRILLGKYC---HSSSFMECKLPSQPSHGWRSIIAGREILKEGLGWLITNGEKVSIWN 831
Query: 545 DSW 547
D W
Sbjct: 832 DPW 834
>At3g45550 putative protein
Length = 851
Score = 293 bits (750), Expect = 2e-79
Identities = 178/532 (33%), Positives = 279/532 (51%), Gaps = 19/532 (3%)
Query: 28 PKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVL 87
P V N LT E+ A+ + D APGPD A F++ W+IV DV + V
Sbjct: 34 PSSVTNIINSELTQDFRDSEIFEAICQIGDDKAPGPDGLTARFYKQCWDIVGNDVIKEVK 93
Query: 88 DFFKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSI 147
FF++ + + N +I +IPK N ++ YR IAL N +K+I+K + +RL L SI
Sbjct: 94 LFFESSHMKTSVNHTNICMIPKIQNPQTLSDYRPIALCNVLYKVISKCMVNRLKAHLNSI 153
Query: 148 ISKEQRGFVQGRNIRDCIALTSEAINVLD-----NKSFGGNLALKIDVTKAFDTLNWDFL 202
+S Q F+ GR I D + + E ++ L +K++ +A+K DV+KA+D + WDFL
Sbjct: 154 VSDSQAAFIPGRIINDNVMIAHEIMHSLKVRKRVSKTY---MAVKTDVSKAYDRVEWDFL 210
Query: 203 LLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEE 262
++ FGF + + WI + S + +NG+ HG+ + RG+RQGDPLSP LF + +
Sbjct: 211 ETTMRLFGFCDKWIGWIMAAVKSVHYSVLINGSPHGYISPTRGIRQGDPLSPYLFILCGD 270
Query: 263 VLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADC 322
+LS I + A G I + H + DD + FC+A + + LK +F Y
Sbjct: 271 ILSHLIKVKASSGDIRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYY 330
Query: 323 SGQIMNIRKSFIFAG----GITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQ 378
SGQ +N++KS I G G T TR+ ++NI G YLG P G+ K F
Sbjct: 331 SGQKINVQKSLITFGSRVYGSTQTRLKTLLNIPNQGGGG---KYLGLPEQFGRKKKEMFN 387
Query: 379 PIADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFI 438
I D+VK + A W A LS AG+ L+KSV +M V+ MS + P I+ E+E + NF
Sbjct: 388 YIIDRVKERTASWSAKFLSPAGKEILLKSVALAMPVYAMSCFKLPQGIVSEIESLLMNFW 447
Query: 439 WSGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQ-SDEQWANIIR 497
W KR + VAW+++ +EGGLG + L N+A K W ++Q + +A +++
Sbjct: 448 WEKASNKRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRIIQYPNSLFARVMK 507
Query: 498 SRVIRDHRCMNHHISSSI---WSGVKAEFHVIKDNCSWIVGDGKQINFWSDS 546
+R +D+ ++ S WS + + +++ +++GDGK I D+
Sbjct: 508 ARYFKDNSIIDAKTRSQQSYGWSSLLSGIALLRKGTRYVIGDGKTIRLGIDN 559
>At4g10830 putative protein
Length = 1294
Score = 292 bits (748), Expect = 3e-79
Identities = 171/503 (33%), Positives = 269/503 (52%), Gaps = 11/503 (2%)
Query: 47 EVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANSIIL 106
E+ NA+ + D APGPD A F++ W IV DV + V FF+ ++ + N +I +
Sbjct: 794 EIYNAICHIGDDKAPGPDGLTARFYKSCWEIVGPDVIKEVKIFFRTSYMKQSINHTNICM 853
Query: 107 IPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDCIA 166
IPK N +++ YR IAL N +KII+K L +RL L +I+S Q F+ GR + D +
Sbjct: 854 IPKITNPETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGRLVNDNVM 913
Query: 167 LTSEAINVLDNKSFGGN--LALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILH 224
+ E ++ L + +A+K DV+KA+D + W+FL ++ FGF+E + WI +
Sbjct: 914 IAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIKWIMGAVK 973
Query: 225 SSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAASR 284
S + +NG HG RG+RQGDPLSP LF + ++L+ I +G D+
Sbjct: 974 SVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAEG--DIRGIRI 1031
Query: 285 NNCLP--FHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFI-FAGGITD 341
N +P H + DD + FC++ + + LK +F Y SGQ +N+ KS I F +
Sbjct: 1032 GNGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFGSRVHG 1091
Query: 342 TRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGR 401
T N + NILG YLG P G+ K F I ++VK + + W A LS AG+
Sbjct: 1092 TTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYLSPAGK 1151
Query: 402 IQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYE 461
++KSV SM V+ MS + P+ I+ E+E + NF W + KR++ +AW+++ +
Sbjct: 1152 EIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRLQYSKK 1211
Query: 462 EGGLGVKSLICLNEATNLKICWNLMQS-DEQWANIIRSRVIRDHRCMN---HHISSSIWS 517
EGGLG + L N+A K W ++ + + +A I+++R R+ ++ S W+
Sbjct: 1212 EGGLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKARYFREDSILDAKRQRYQSYGWT 1271
Query: 518 GVKAEFHVIKDNCSWIVGDGKQI 540
+ A VIK +IVGDGK +
Sbjct: 1272 SMLAGLDVIKKGSRFIVGDGKTV 1294
>At2g05980 putative non-LTR retroelement reverse transcriptase
Length = 1352
Score = 292 bits (748), Expect = 3e-79
Identities = 163/494 (32%), Positives = 257/494 (51%), Gaps = 6/494 (1%)
Query: 60 APGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANSIILIPKTPNADSVDQY 119
+PGPD + FF+ W ++ +D+ A+ FF G+LP N + LI K + Y
Sbjct: 617 SPGPDGYTVEFFKTAWPVLGRDLVIAIQSFFLKGFLPKGINTTILALISKKHEVSGMKDY 676
Query: 120 RTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDCIALTSEAINVLDNKS 179
R I+ N +KI++K++A+RL +ILP+ I+ Q F++ R + + + L SE + +S
Sbjct: 677 RPISCCNVLYKIVSKLMANRLKEILPASIAPNQSAFIKDRLMMENLLLASELVKDYHKES 736
Query: 180 FGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGF 239
ALKID++KAFD + W FL+ VLK E+F +WI+ + ++ + +NG GF
Sbjct: 737 ISSRSALKIDISKAFDFVQWPFLINVLKAIHLPEMFIHWIELCIGTASFSVQVNGELSGF 796
Query: 240 FNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLM 299
F RG+RQG LSP L+ I VLS + A + I RN L CF DD+M
Sbjct: 797 FRSERGLRQGCSLSPYLYVICMNVLSCMLDKAAVEKKISYHPRCRNMNLTHLCF-ADDIM 855
Query: 300 VFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFIFAGGITDTRMNNIVNILGFNVGSLP 359
VF S+ ++F ++A S +++ KS IF GI+ +I+ F +G+LP
Sbjct: 856 VFSDGTSKSIQGTLAIFEKFAAMSWLKISLEKSTIFMAGISPNAKTSILQQFPFELGTLP 915
Query: 360 FTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSI 419
YLG P+ + + P+ +K++A++ W LS AGR+QL+KSV+ S+ +S+
Sbjct: 916 VKYLGLPLLTKRMTQSDYLPLVEKIRARITSWTNRFLSFAGRLQLIKSVLSSITNFWLSV 975
Query: 420 YSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNL 479
+ P L+E+EK F+WSG K +AW ++C EEGGLG+K L NE + L
Sbjct: 976 FRLPKACLQEIEKMFSAFLWSGPDLNTKKAKIAWSEVCKLKEEGGLGLKPLKEANEVSLL 1035
Query: 480 KICWNLMQS-DEQWANIIRSRVIRDHRCM----NHHISSSIWSGVKAEFHVIKDNCSWIV 534
K+ W ++ + D W + +IR N + S +W + + + V
Sbjct: 1036 KLIWRILSARDSLWVKWVNKHLIRKETFWSVKENTGLGSWLWRKILKQRDKARLFHRMEV 1095
Query: 535 GDGKQINFWSDSWC 548
G +FW D WC
Sbjct: 1096 RSGTFTSFWHDHWC 1109
>At2g23880 putative non-LTR retroelement reverse transcriptase
Length = 1216
Score = 291 bits (745), Expect = 6e-79
Identities = 166/530 (31%), Positives = 267/530 (50%), Gaps = 6/530 (1%)
Query: 23 VEEAIPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDV 82
+E +P +RLLT + T EE+K +F + D +PGPD + + F++ W I+ +V
Sbjct: 154 LENLLPFRCSEDDHRLLTRVVTGEEIKKVIFSMPKDKSPGPDGYTSEFYKASWEIIGDEV 213
Query: 83 YEAVLDFFKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAK 142
A+ FF G+LP N+ + LIPK A + YR I+ N +K I+K+LA+RL +
Sbjct: 214 IIAIQSFFAKGFLPKGVNSTILALIPKKKEAREIKDYRPISCCNVLYKAISKILANRLKR 273
Query: 143 ILPSIISKEQRGFVQGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFL 202
ILP I Q FV+ R + + + L +E + S A+KID++KAFD+L W FL
Sbjct: 274 ILPKFIVGNQSAFVKDRLLIENVLLATELVKDYHKDSISTRCAMKIDISKAFDSLQWSFL 333
Query: 203 LLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEE 262
VL F F +WI + ++ I +NG G+F RG+RQG LSP LF I +
Sbjct: 334 THVLAAMNFPGEFIHWISLCMSTASFSIQVNGELAGYFRSARGLRQGCSLSPYLFVISMD 393
Query: 263 VLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADC 322
VLSR + A + L CF DDLM+ K+ S+ + + ++A
Sbjct: 394 VLSRMLDKAAGAREFGYHPRCKTLGLTHLCF-ADDLMILTDGKIRSVDGIVKVLNQFAAK 452
Query: 323 SGQIMNIRKSFIFAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIAD 382
G + + K+ ++ G++D + + F VG LP YLG P+ + + P+ D
Sbjct: 453 LGLKICMEKTTLYLAGVSDHSRQLMSSRYSFGVGKLPVRYLGLPLVTKRLTTSDYSPLID 512
Query: 383 KVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGD 442
+++ ++ W + LS AGR+ L+ SV+ S+ M+ + P + + E+ + +WSG
Sbjct: 513 QIRRRIGMWTSRYLSFAGRLSLINSVLWSITNFWMNAFRLPRECINEINRISSALLWSGP 572
Query: 443 VTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQ-SDEQWANIIRSRVI 501
K V+W +IC +EGGLG++SL N+ ++LK+ W L+ D W R ++
Sbjct: 573 ELNPKKAKVSWDEICKPKKEGGLGLQSLREANKVSSLKLIWRLLSCQDSLWVKWTRMNLL 632
Query: 502 RDHRC----MNHHISSSIWSGVKAEFHVIKDNCSWIVGDGKQINFWSDSW 547
+ + + S IW + V K C V +G +FW D+W
Sbjct: 633 KKESFWSIGTHSTLGSWIWRRLLKHREVAKSFCKIEVNNGVNTSFWFDNW 682
>At4g02490
Length = 657
Score = 288 bits (736), Expect = 7e-78
Identities = 163/503 (32%), Positives = 258/503 (50%), Gaps = 24/503 (4%)
Query: 51 AVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANSIILIPKT 110
A+F + + APGPD F FF W IVK+ A+ +FF G L FNA +I LIPK
Sbjct: 2 ALFSMPRNKAPGPDGFPVEFFLEAWPIVKESFIAAIKEFFLTGHLLKGFNATAITLIPKV 61
Query: 111 PNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDCIALTSE 170
P AD ++ +R +A +K+I ++++ RL + + Q GF++GR + + + L SE
Sbjct: 62 PGADQLNLFRPVACCTTIYKVITRLISKRLNLFIDQAVQSNQVGFIKGRLLCENVLLASE 121
Query: 171 AINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHSSKMFI 230
++ + L++D+TKA+D +NW+FL+ +LK +F NWI + + I
Sbjct: 122 LVDNFQAEGDTSRGCLQVDLTKAYDNVNWEFLINILKALNLPPIFINWIWVCISTPSYSI 181
Query: 231 SMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAASRNNCLP- 289
+ NG GFF +G+RQGDP+S LF +V ++L+RS+ + A +G L CL
Sbjct: 182 AYNGELIGFFVGKKGIRQGDPMSSHLFVLVMDILARSLDLGAVEGRFVL----HPKCLAP 237
Query: 290 --FHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFIFAGGITDTRMNNI 347
H + DD++VFC +SSL+ + + + SG +N++K+ + G R +
Sbjct: 238 MITHLSFADDILVFCDGSLSSLVAILDILDVFKKGSGLGINLQKTALLLDGGNFERNRIM 297
Query: 348 VNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGRIQLVKS 407
LG + GSLP YLG P+ K K +QP+ D++ ++ W A LS AGR+QL+KS
Sbjct: 298 AASLGVSQGSLPVRYLGVPLMSQKMKKHDYQPLVDRINSRFTSWTARHLSFAGRLQLLKS 357
Query: 408 VVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYEEGGLGV 467
V+ S + SI+ P + L ++E+ F+WSG + ++W +C+ E GGLG+
Sbjct: 358 VIYSTINFWASIFILPNQCLHKLEQMCNAFLWSGAPNSAREAKISWDIVCSSKESGGLGL 417
Query: 468 KSLICLNEATNLKICWNLM-QSDEQWANIIRSRVIRDHRCMNHHISSSIWSGVKAEFHVI 526
K L N+ LK+ W L S W + +R +W + V
Sbjct: 418 KRLSSWNKVLALKLIWLLFTASGSLWVSWVR----------------WVWRKLCKLREVA 461
Query: 527 KDNCSWIVGDGKQINFWSDSWCG 549
+ VG G FW D+W G
Sbjct: 462 RPFVICEVGSGITARFWQDNWTG 484
>At1g25430 hypothetical protein
Length = 1213
Score = 286 bits (732), Expect = 2e-77
Identities = 159/521 (30%), Positives = 269/521 (51%), Gaps = 32/521 (6%)
Query: 44 TKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANS 103
+ E+++ A+F L + + GPD F A FF W+IV +V +A+ +FF +G L +NA +
Sbjct: 448 SNEDIRAALFSLPRNKSCGPDGFTAEFFIDSWSIVGAEVTDAIKEFFSSGCLLKQWNATT 507
Query: 104 IILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRD 163
I+LIPK N +R I+ +N +K+I ++L DRL ++L +IS Q F+ GR++ +
Sbjct: 508 IVLIPKIVNPTCTSDFRPISCLNTLYKVIARLLTDRLQRLLSGVISSAQSAFLPGRSLAE 567
Query: 164 CIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTIL 223
+ L ++ ++ + + LK+D+ KAFD++ W+F++ L+ E F NWI +
Sbjct: 568 NVLLATDLVHGYNWSNISPRGMLKVDLKKAFDSVRWEFVIAALRALAIPEKFINWISQCI 627
Query: 224 HSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAAS 283
+ +S+NG GFF +G+RQGDPLSP LF + E S + + GLI +
Sbjct: 628 STPTFTVSINGGNGGFFKSTKGLRQGDPLSPYLFVLAMEAFSNLLHSRYESGLIHYHPKA 687
Query: 284 RNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFIFAGGITDTR 343
N + H + DD+M+F SL + +A SG +N KS ++ G+
Sbjct: 688 SNLSIS-HLMFADDVMIFFDGGSFSLHGICETLDDFASWSGLKVNKDKSHLYLAGLNQLE 746
Query: 344 MNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGRIQ 403
+N GF +G+LP YLG P+ K + ++P+ +K+ A+ W LS AGRIQ
Sbjct: 747 -SNANAAYGFPIGTLPIRYLGLPLMNRKLRIAEYEPLLEKITARFRSWVNKCLSFAGRIQ 805
Query: 404 LVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYEEG 463
L+ SV+ + MS + P +K +E F+WSG++ + K + V+W +C EG
Sbjct: 806 LISSVIFGSINFWMSTFLLPKGCIKRIESLCSRFLWSGNIEQAKGIKVSWAALCLPKSEG 865
Query: 464 GLGVKSLICLNEATNLKICWNL-MQSDEQWANIIRSRVIRDHRCMNHHISSSIWSGVKAE 522
GLG++ L+ N+ ++++ W L + D WA D + ++H S W+ +
Sbjct: 866 GLGLRRLLEWNKTLSMRLIWRLFVAKDSLWA---------DWQHLHHLSRGSFWAVEGGQ 916
Query: 523 FHVIKDNCSW----------------IVGDGKQINFWSDSW 547
D+ +W VG+G + ++W D+W
Sbjct: 917 ----SDSWTWKRLLSLRPLAHQFLVCKVGNGLKADYWYDNW 953
>At3g25485 unknown protein
Length = 979
Score = 284 bits (726), Expect = 1e-76
Identities = 162/505 (32%), Positives = 258/505 (51%), Gaps = 26/505 (5%)
Query: 44 TKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANS 103
T EE+K +F ++SD +PGPD F F++ W I+ + AV FF+ G+LP N+
Sbjct: 370 TGEEIKTVLFFMSSDKSPGPDGFTTEFYKATWEIIGAEFIVAVKSFFEKGFLPKGVNSTI 429
Query: 104 IILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRD 163
+ LIPK + YR I+ N +K+I+K++A RL +ILP I+ Q FV+ R +
Sbjct: 430 LALIPKKFETKEMKDYRPISCCNVIYKVISKLIAKRLKEILPQFIAGNQSAFVKDRLLIQ 489
Query: 164 CIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTIL 223
+ L +E + +S A+KID++KAFD++ W FL VL + F++ F +WI +
Sbjct: 490 NLLLATEIVKDYHKESVSDRCAIKIDISKAFDSVQWSFLENVLHSLNFSQEFIHWIMLCI 549
Query: 224 HSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAAS 283
++ + +NG GFFN +RG+RQG LSP LF I +VLS+ + A
Sbjct: 550 TTASFSVQVNGELVGFFNSSRGLRQGCSLSPYLFVIAMDVLSKMLDRAAGFKKFGYHPRC 609
Query: 284 RNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFIFAGGITDTR 343
+ L H + DDLMV K+ S+ + S+F +A SG +++ KS I+ G +
Sbjct: 610 QTIGLT-HLTFADDLMVLSDGKVRSVEGIVSVFDEFAKKSGLKISMEKSTIYLAGSANMY 668
Query: 344 MNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGRIQ 403
I F VGSLP YLG P+ + ++P+ +++K K+ W A LS AGR+
Sbjct: 669 RREIEERFKFAVGSLPVRYLGLPLVTKRFSSADYRPLVEQIKRKIGTWTARYLSYAGRLN 728
Query: 404 LVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYEEG 463
L+ SV+ S+ ++ + P + ++E++K F+WSG K V W IC EG
Sbjct: 729 LISSVIWSICNFWIAAFRLPRECIREIDKLCSAFLWSGPELNSKRAKVNWEAICKPKREG 788
Query: 464 GLGVKSLICLNEATNLKICWN-LMQSDEQWANIIRSRVIRDHRCMNHHISSSIWSGVKAE 522
GLG++SL N+ + LK+ W + + D W +++ R
Sbjct: 789 GLGLRSLKEANDVSCLKLIWRWVSRGDSLWRKLLKYR----------------------- 825
Query: 523 FHVIKDNCSWIVGDGKQINFWSDSW 547
K C + +G + +FW D+W
Sbjct: 826 -DTAKQFCKVDIRNGGRTSFWFDNW 849
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 282 bits (721), Expect = 4e-76
Identities = 170/513 (33%), Positives = 262/513 (50%), Gaps = 9/513 (1%)
Query: 44 TKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANS 103
TK EV +AV + APGPD + F+Q W V V VL+FF+ G LP + N
Sbjct: 293 TKAEVVSAVKSMGRFKAPGPDGYQPVFYQQCWETVGPSVTRFVLEFFETGVLPASTNDAL 352
Query: 104 IILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRD 163
++LI K + + Q+R ++L N FKII K++ RL ++ +I Q F+ GR D
Sbjct: 353 LVLIAKVAKPERIQQFRPVSLCNVLFKIITKMMVTRLKNVISKLIGPAQASFIPGRLSID 412
Query: 164 CIALTSEAINVLDNKSFG-GNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTI 222
I L EA++ + K G + LK+D+ KA+D + WDFL L+ G +E + + I
Sbjct: 413 NIVLVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRVRWDFLQETLEAAGLSEGWTSRIMAG 472
Query: 223 LHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAA 282
+ M + NG + F RG+RQGDPLSP LF + E L I K IA
Sbjct: 473 VTDPSMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIEASVGKREWKPIAV 532
Query: 283 SRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFIFAGGITDT 342
S H + DDL++F +A ++ + +++ + R+ + SGQ +++ KS IF
Sbjct: 533 SCGGSKLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSHNVSR 592
Query: 343 RMNNIVNI-LGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGR 401
M +++ G YLG PI + + F + ++V A+LA WK LS+AGR
Sbjct: 593 EMEQLISEESGIGCTKELGKYLGMPILQKRMNKETFGEVLERVSARLAGWKGRSLSLAGR 652
Query: 402 IQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYE 461
I L K+V+ S+ VH MS P+ L ++++ + F+W + K+K ++WRKIC
Sbjct: 653 ITLTKAVLSSIPVHVMSAILLPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSWRKICKPKA 712
Query: 462 EGGLGVKSLICLNEATNLKICWNLMQSDEQ-WANIIRSRV----IRDHRCMNHHIS-SSI 515
EGG+G++S +N+A K+ W L+Q E WA ++R + ++D + SS
Sbjct: 713 EGGIGLRSARDMNKALVAKVGWRLLQDKESLWARVVRKKYKVGGVQDTSWLKPQPRWSST 772
Query: 516 WSGVKAEF-HVIKDNCSWIVGDGKQINFWSDSW 547
W V V+ W+ GDG I FW D W
Sbjct: 773 WRSVAVGLREVVVKGVGWVPGDGCTIRFWLDRW 805
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 281 bits (718), Expect = 9e-76
Identities = 171/550 (31%), Positives = 271/550 (49%), Gaps = 25/550 (4%)
Query: 19 DHLLVEEAIPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIV 78
D +V+EA+ ++ + N L + + E+K A+F +++D APGPD F A FF YW+I+
Sbjct: 345 DLQVVQEALSPIISSHCNEELIKISSLLEIKEALFSISADKAPGPDGFSASFFHAYWDII 404
Query: 79 KKDVYEAVLDFFKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLAD 138
+ DV + FF + L N + LIPK V YR IAL N ++KI+ K+L
Sbjct: 405 EADVSRDIRSFFVDSCLSPRLNETHVTLIPKISAPRKVSDYRPIALCNVQYKIVAKILTR 464
Query: 139 RLAKILPSIISKEQRGFVQGRNIRDCIALTSEAINVL--DNKSFGGNLALKIDVTKAFDT 196
RL L +IS Q FV GR I D + +T E ++ L ++A+K D++KA+D
Sbjct: 465 RLQPWLSELISLHQSAFVPGRAIADNVLITHEILHFLRVSGAKKYCSMAIKTDMSKAYDR 524
Query: 197 LNWDFLLLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLL 256
+ W+FL VL GF++ + W+ + + +NG+ G +RG+RQGDPLSP L
Sbjct: 525 IKWNFLQEVLMRLGFHDKWIRWVMQCVCTVSYSFLINGSPQGSVVPSRGLRQGDPLSPYL 584
Query: 257 FCIVEEVLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLF 316
F + EVLS +KG++ I +R + H + DD M FCK + L ++
Sbjct: 585 FILCTEVLSGLCRKAQEKGVMVGIRVARGSPQVNHLLFADDTMFFCKTNPTCCGALSNIL 644
Query: 317 TRYADCSGQIMNIRKSFIFAGGIT--DTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKG 374
+Y SGQ +N+ KS I T D + +++ N G + YLG P G+ K
Sbjct: 645 KKYELASGQSINLAKSAITFSSKTPQDIKRRVKLSLRIDNEGGIG-KYLGLPEHFGRRKR 703
Query: 375 IHFQPIADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWI 434
F I D+++ + W LS AG+ L+K+V+ SM + M + P + K+++ +
Sbjct: 704 DIFSSIVDRIRQRSHSWSIRFLSSAGKQILLKAVLSSMPSYAMMCFKLPASLCKQIQSVL 763
Query: 435 KNFIWSGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQSDEQWAN 494
F W KRKM V+W K+ EGGLG + + K+ W +++
Sbjct: 764 TRFWWDSKPDKRKMAWVSWDKLTLPINEGGLGFREI-------EAKLSWRILKEPHS--- 813
Query: 495 IIRSRVIRDHRC---------MNHHISSSIWSGVKAEFHVIKDNCSWIVGDGKQINFWSD 545
+ SRV+ C + +S W G+ A +++ W +G G IN W++
Sbjct: 814 -LLSRVLLGKYCNTSSFMDCSASPSFASHGWRGILAGRDLLRKGLGWSIGQGDSINVWTE 872
Query: 546 SWCGQVLAQT 555
+W QT
Sbjct: 873 AWLSPSSPQT 882
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 281 bits (718), Expect = 9e-76
Identities = 178/531 (33%), Positives = 278/531 (51%), Gaps = 23/531 (4%)
Query: 31 VDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFF 90
V T N LT + E+ +A+ + D APGPD A F++ W+IV DV V FF
Sbjct: 798 VTNTVNLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFF 857
Query: 91 KNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISK 150
+ ++ + N +I +IPK N ++ YR IAL N +K+I+K L +RL L SI+S
Sbjct: 858 ETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSD 917
Query: 151 EQRGFVQGRNIRDCIALTSEAINVLD-----NKSFGGNLALKIDVTKAFDTLNWDFLLLV 205
Q F+ GR I D + + E ++ L +K++ +A+K DV+KA+D + WDFL
Sbjct: 918 SQAAFIPGRIINDNVMIAHEVMHSLKVRKRVSKTY---MAVKTDVSKAYDRVEWDFLETT 974
Query: 206 LKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLS 265
++ FGF + WI + S + +NG+ HG+ RG+RQGDPLSP LF + ++LS
Sbjct: 975 MRLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILS 1034
Query: 266 RSISILADKGLIDLIAASRNNCLP--FHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCS 323
I+ A G DL N P H + DD + FC+A + + LK +F Y S
Sbjct: 1035 HLINGRASSG--DLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYS 1092
Query: 324 GQIMNIRKSFIFAG----GITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQP 379
GQ +N++KS I G G T +R+ I+ I G YLG P G+ K F+
Sbjct: 1093 GQKINVQKSMITFGSRVYGSTQSRLKQILEIPNQGGGG---KYLGLPEQFGRKKKEMFEY 1149
Query: 380 IADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIW 439
I D+VK + + W A LS AG+ ++KSV +M V+ MS + P I+ E+E + NF W
Sbjct: 1150 IIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWW 1209
Query: 440 SGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQ-SDEQWANIIRS 498
+R + VAW+++ +EGGLG + L N+A K W L+Q + +A ++++
Sbjct: 1210 EKASNQRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKA 1269
Query: 499 RVIRDHRCMNHHI---SSSIWSGVKAEFHVIKDNCSWIVGDGKQINFWSDS 546
R +D ++ + S W+ + ++K ++GDG+ I D+
Sbjct: 1270 RYFKDVSILDAKVRKQQSYGWASLLDGIALLKKGTRHLIGDGQNIRIGLDN 1320
>At2g11240 pseudogene
Length = 1044
Score = 281 bits (718), Expect = 9e-76
Identities = 172/534 (32%), Positives = 265/534 (49%), Gaps = 19/534 (3%)
Query: 23 VEEAIPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDV 82
++EAI + N EE+K+A F +++D APGPD F A FFQ W V ++
Sbjct: 242 IKEAIKPFISPEQN--------PEEIKSACFSIHADKAPGPDGFSASFFQSNWMTVGPNI 293
Query: 83 YEAVLDFFKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAK 142
+ FF + L N I LIPK + + YR IAL +KII+K+L+ RL
Sbjct: 294 VLEIQSFFSSSTLQPTINKTHITLIPKIQSLKRMVDYRPIALCTVFYKIISKLLSRRLQP 353
Query: 143 ILPSIISKEQRGFVQGRNIRDCIALTSEAINVLDNKSFGGN----LALKIDVTKAFDTLN 198
IL IIS+ Q FV R D + +T EA++ L KS G +A+K +++KA+D +
Sbjct: 354 ILQEIISENQSAFVPKRASNDNVLITHEALHYL--KSLGAEKRCFMAVKTNMSKAYDRIE 411
Query: 199 WDFLLLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFC 258
WDF+ LV++ GF++ + +WI + + +NG+ G RG+RQGDPLSP LF
Sbjct: 412 WDFIKLVMQEMGFHQTWISWILQCITTVSYSFLLNGSAQGAVTPERGLRQGDPLSPFLFI 471
Query: 259 IVEEVLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTR 318
I EVLS G + + S+ N H + DD + FC++ + S + +
Sbjct: 472 ICSEVLSGLCRKAQLDGSLLGLRVSKGNPRVNHLLFADDTIFFCRSDLKSCKTFLCILKK 531
Query: 319 YADCSGQIMNIRKSFI-FAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHF 377
Y + SGQ++N KS I F+ D ILG + YLG P G+ K F
Sbjct: 532 YEEASGQMINKSKSAITFSRKTPDHIKTEAQQILGIQLVGGLGKYLGLPKMFGRKKRDLF 591
Query: 378 QPIADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNF 437
I D+++ + W + LS AG+ ++KSV+ SM +TMS + + + K ++ + +F
Sbjct: 592 NQIVDRIRQRSLSWSSRFLSTAGKTTMLKSVLASMPTYTMSCFKLLVSLCKRIQSALTHF 651
Query: 438 IWSGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQSDE-QWANII 496
W K+KM +AW K+ + +EGGLG K + N+A K+ W ++QS I+
Sbjct: 652 WWDSSADKKKMCWIAWSKMAKNKKEGGLGFKDITNFNDALLAKLSWRIVQSPSCVLVRIL 711
Query: 497 RSRVIRDHR---CMNHHISSSIWSGVKAEFHVIKDNCSWIVGDGKQINFWSDSW 547
+ R C SS W G+ +IK ++G G W++ W
Sbjct: 712 LGKYCRTSSFLDCSVTAASSHGWRGICTGKDLIKSQLGKVIGSGLDTLVWNEPW 765
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 280 bits (715), Expect = 2e-75
Identities = 177/531 (33%), Positives = 278/531 (52%), Gaps = 23/531 (4%)
Query: 31 VDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFF 90
V T N LT + E+ +A+ + D APGPD A F++ W+IV DV V FF
Sbjct: 572 VTNTVNLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFF 631
Query: 91 KNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISK 150
+ ++ + N +I +IPK N ++ YR IAL N +K+I+K L +RL L SI+S
Sbjct: 632 ETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSD 691
Query: 151 EQRGFVQGRNIRDCIALTSEAINVLD-----NKSFGGNLALKIDVTKAFDTLNWDFLLLV 205
Q F+ GR I D + + E ++ L +K++ +A+K DV+KA+D + WDFL
Sbjct: 692 SQAAFIPGRIINDNVMIAHEVMHSLKVRKRVSKTY---MAVKTDVSKAYDRVEWDFLETT 748
Query: 206 LKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLS 265
++ FGF + WI + S + +NG+ HG+ RG+RQGDPLSP LF + ++LS
Sbjct: 749 MRLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILS 808
Query: 266 RSISILADKGLIDLIAASRNNCLP--FHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCS 323
I+ A G DL N P H + DD + FC+A + + LK +F Y S
Sbjct: 809 HLINGRASSG--DLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYS 866
Query: 324 GQIMNIRKSFIFAG----GITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQP 379
GQ +N++KS I G G T +++ I+ I G YLG P G+ K F+
Sbjct: 867 GQKINVQKSMITFGSRVYGSTQSKLKQILEIPNQGGGG---KYLGLPEQFGRKKKEMFEY 923
Query: 380 IADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIW 439
I D+VK + + W A LS AG+ ++KSV +M V+ MS + P I+ E+E + NF W
Sbjct: 924 IIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWW 983
Query: 440 SGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICWNLMQ-SDEQWANIIRS 498
+R + VAW+++ +EGGLG + L N+A K W L+Q + +A ++++
Sbjct: 984 EKASNQRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKA 1043
Query: 499 RVIRDHRCMNHHI---SSSIWSGVKAEFHVIKDNCSWIVGDGKQINFWSDS 546
R +D ++ + S W+ + ++K ++GDG+ I D+
Sbjct: 1044 RYFKDVSILDAKVRKQQSYGWASLLDGIALLKKGTRHLIGDGQNIRIGLDN 1094
>At2g14430 putative non-LTR retroelement reverse transcriptase
Length = 1277
Score = 280 bits (715), Expect = 2e-75
Identities = 167/520 (32%), Positives = 259/520 (49%), Gaps = 36/520 (6%)
Query: 33 ATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKN 92
AT +LT T EE++ +F + S+ +PGPD +
Sbjct: 615 ATDQDMLTREVTSEEIQKVLFAMPSNKSPGPDGY-------------------------- 648
Query: 93 GWLPNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQ 152
NA + LIPK A + YR I+ N +K+I+K++A+RL +LP+ I + Q
Sbjct: 649 ----TRLNATILALIPKKDEATLMRDYRPISCCNVIYKVISKIIANRLKVMLPTFILQNQ 704
Query: 153 RGFVQGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFN 212
FV+ R + + + L +E + S A+KID++KAFD++ W FLL L+ F
Sbjct: 705 SAFVRERLLMENVLLATELVKDYHKDSISPRCAMKIDISKAFDSVQWQFLLNTLEALKFP 764
Query: 213 ELFCNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILA 272
E F +WIK + ++ + +N Q GFF RG+RQG LSP LF I VLS I + A
Sbjct: 765 EKFRHWIKLCISTATFSVQVNSEQAGFFGSKRGLRQGCALSPYLFVICMNVLSHMIDVAA 824
Query: 273 DKGLIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKS 332
I + L CF DDLMVF + S+ + ++F +A SG +++ KS
Sbjct: 825 VHRNIGYHPKCKKLSLTHLCF-ADDLMVFIDGQQRSVEGVINIFKDFAGKSGLHISLEKS 883
Query: 333 FIFAGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWK 392
++ +++ NNI++ F G LP YLG P+ + + P+ DKV++K++ W
Sbjct: 884 TLYLAEVSELNRNNILSAFPFASGQLPVRYLGFPLLTKQMTTADYSPLLDKVRSKISSWT 943
Query: 393 ASLLSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVA 452
A LS AGR+ L+ SV+ S+ MS Y P +KE+EK F+WSG K +
Sbjct: 944 ARSLSYAGRLALINSVIVSLSNFWMSAYRLPAGCIKEIEKLCSAFLWSGPELNPKKAKIT 1003
Query: 453 WRKICADYEEGGLGVKSLICLNEATNLKICWNLM-QSDEQWANIIRSRVIRDHRCMNHHI 511
W +C +EGGLG+KSL+ N+ + LK+ W L+ + W N + + +IR + +
Sbjct: 1004 WTSLCKLKQEGGLGIKSLLEANKVSCLKLIWRLVSRQSSLWVNWVWTYIIRKGSFWSAND 1063
Query: 512 SSSI----WSGVKAEFHVIKDNCSWIVGDGKQINFWSDSW 547
SS+ W + V K C + G +FW D+W
Sbjct: 1064 RSSLGSWMWKKLLNYRDVAKSMCKVEIKSGSSTSFWYDNW 1103
>At1g31030 F17F8.5
Length = 872
Score = 280 bits (715), Expect = 2e-75
Identities = 156/517 (30%), Positives = 264/517 (50%), Gaps = 6/517 (1%)
Query: 36 NRLLTMLPTKEEVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWL 95
N +LT + EE+K +F + D +PGPD + + F++ W+I+ ++ V FF+ G+L
Sbjct: 93 NEMLTREVSSEEIKTVLFSMPKDKSPGPDGYTSEFYKATWDIIGQEFTLPVQSFFQKGFL 152
Query: 96 PNNFNANSIILIPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGF 155
P N+ + LIPK A + YR I+ N +K+I+K++A+RL +LP I++ Q F
Sbjct: 153 PKGINSIILALIPKKLAAKEMRDYRPISCCNVLYKVISKIIANRLKLLLPRFIAENQSAF 212
Query: 156 VQGRNIRDCIALTSEAINVLDNKSFGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELF 215
V+ R + + + L +E + S A+KID++KAFD++ W FL L F+ F
Sbjct: 213 VKDRLLIENLLLATELVKDYHKDSISARCAIKIDISKAFDSVQWSFLTNTLVAMNFSPTF 272
Query: 216 CNWIKTILHSSKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSISILADKG 275
+WI + ++ + +NG G+F RG+RQG LSP LF I +VLS+ + A
Sbjct: 273 IHWINLCITTASFSVQVNGDLVGYFQSKRGLRQGCSLSPYLFVICMDVLSKMLDKAAGVR 332
Query: 276 LIDLIAASRNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFIF 335
+ L H + DDLMV K S+ + +F + SG +++ KS ++
Sbjct: 333 KFGFHPKCQRLGLT-HLSFADDLMVLSDGKTRSIEGILEVFDEFCKRSGLRISLEKSTLY 391
Query: 336 AGGITDTRMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASL 395
G++ I F+VG LP YLG P+ + + P+ +++K ++A W
Sbjct: 392 MAGVSPIIKQEIAAKFLFDVGQLPVRYLGLPLVTKRLTSADYSPLLEQIKKRIATWTFRF 451
Query: 396 LSIAGRIQLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRK 455
S AGR L+KSV+ S+ ++ + P + ++E++K +F+WSG ++W
Sbjct: 452 FSFAGRFNLIKSVLWSICNFWLAAFRLPRQCIREIDKLCSSFLWSGSEMSSHKAKISWDI 511
Query: 456 ICADYEEGGLGVKSLICLNEATNLKICWNLM-QSDEQWANIIRSRVIRDHRCMNHHISSS 514
+C EGGLG+++L N+ + LK+ W ++ S+ W + +IR + S+S
Sbjct: 512 VCKPKAEGGLGLRNLKEANDVSCLKLVWRIISNSNSLWTKWVAEYLIRKKSIWSLKQSTS 571
Query: 515 ----IWSGVKAEFHVIKDNCSWIVGDGKQINFWSDSW 547
IW + V K VG+G+ +FW D W
Sbjct: 572 MGSWIWRKILKIRDVAKSFSRVEVGNGESASFWYDHW 608
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 278 bits (712), Expect = 4e-75
Identities = 169/515 (32%), Positives = 258/515 (49%), Gaps = 13/515 (2%)
Query: 47 EVKNAVFDLNSDDAPGPDVFGACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANSIIL 106
EV+ A+ + APGPD F F+Q W +V + V + V+DFF +G P N ++L
Sbjct: 972 EVEGAIRSMGKYKAPGPDGFQPVFYQQGWEVVGESVTKFVMDFFSSGSFPQETNDVLVVL 1031
Query: 107 IPKTPNADSVDQYRTIALVNFKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDCIA 166
I K + + Q+R I+L N FK I KV+ RL ++ +I Q F+ GR D I
Sbjct: 1032 IAKVLKPEKITQFRPISLCNVLFKTITKVMVGRLKGVINKLIGPAQTSFIPGRLSTDNIV 1091
Query: 167 LTSEAINVLDNKS-FGGNLALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHS 225
+ E ++ + K G + LK+D+ KA+D + WD L LK G + WI +
Sbjct: 1092 VVQEVVHSMRRKKGVKGWMLLKLDLEKAYDRIRWDLLEDTLKAAGLPGTWVQWIMKCVEG 1151
Query: 226 SKMFISMNGAQHGFFNCNRGVRQGDPLSPLLFCIVEEVLSRSI--SILADKGLIDLIAAS 283
M + NG + F RG+RQGDPLSP LF + E L I SI A K I S
Sbjct: 1152 PSMRLLWNGEKTDAFKPLRGLRQGDPLSPYLFVLCIERLCHLIESSIAAKKW--KPIKIS 1209
Query: 284 RNNCLPFHCFYVDDLMVFCKAKMSSLIVLKSLFTRYADCSGQIMNIRKSFI-FAGGITDT 342
++ H + DDL++F +A + + VL+ + ++ SGQ +++ KS I F+ +
Sbjct: 1210 QSGPRLSHICFADDLILFAEASIDQIRVLRGVLEKFCGASGQKVSLEKSKIYFSKNVLRD 1269
Query: 343 RMNNIVNILGFNVGSLPFTYLGAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGRI 402
I G YLG PI + + F + + ++LA WK +LS AGR+
Sbjct: 1270 LGKRISEESGMKATKDLGKYLGVPILQKRINKETFGEVIKRFSSRLAGWKGRMLSFAGRL 1329
Query: 403 QLVKSVVQSMLVHTMSIYSWPIKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYEE 462
L K+V+ S+LVHTMS P L ++K + F+W + K+K VAW ++C E
Sbjct: 1330 TLTKAVLTSILVHTMSTIKLPQSTLDGLDKVSRAFLWGSTLEKKKQHLVAWTRVCLPRRE 1389
Query: 463 GGLGVKSLICLNEATNLKICWNLMQSDEQ-WANIIRSRV-IRDHRCMNHHISSSIWSGVK 520
GGLG++S +N+A K+ W ++ WA ++RS+ + D N ++ S WS
Sbjct: 1390 GGLGIRSATAMNKALIAKVGWRVLNDGSSLWAQVVRSKYKVGDVHDRNWTVAKSNWSSTW 1449
Query: 521 AEF-----HVIKDNCSWIVGDGKQINFWSDSWCGQ 550
VI W++GDG+QI FW+D W +
Sbjct: 1450 RSVGVGLRDVIWREQHWVIGDGRQIRFWTDRWLSE 1484
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 270 bits (690), Expect = 2e-72
Identities = 165/548 (30%), Positives = 273/548 (49%), Gaps = 11/548 (2%)
Query: 11 FLLTIFLQDHLLVE----EAIPKLVDATTNRLLTMLPTKEEVKNAVFDLNSDDAPGPDVF 66
F +F ++L E + V + N L T+ EV NAVF +N + APGPD F
Sbjct: 370 FFENLFTSTYILTHNNHLEGLQAKVTSEMNHNLIQEVTELEVYNAVFSINKESAPGPDGF 429
Query: 67 GACFFQIYWNIVKKDVYEAVLDFFKNGWLPNNFNANSIILIPKTPNADSVDQYRTIALVN 126
A FFQ +W++VK + + FF+ G LP ++N I LIPK + + R I+L +
Sbjct: 430 TALFFQQHWDLVKHQILTEIFGFFETGVLPQDWNHTHICLIPKITSPQRMSDLRPISLCS 489
Query: 127 FKFKIINKVLADRLAKILPSIISKEQRGFVQGRNIRDCIALTSEAINVL--DNKSFGGNL 184
+KII+K+L RL K LP+I+S Q FV R I D I + E I+ L +++ ++
Sbjct: 490 VLYKIISKILTQRLKKHLPAIVSTTQSAFVPQRLISDNILVAHEMIHSLRTNDRISKEHM 549
Query: 185 ALKIDVTKAFDTLNWDFLLLVLKTFGFNELFCNWIKTILHSSKMFISMNGAQHGFFNCNR 244
A K D++KA+D + W FL ++ GFN + +WI + S + +NG +G R
Sbjct: 550 AFKTDMSKAYDRVEWPFLETMMTALGFNNKWISWIMNCVTSVSYSVLINGQPYGHIIPTR 609
Query: 245 GVRQGDPLSPLLFCIVEEVLSRSISILADKGLIDLIAASRNNCLPFHCFYVDDLMVFCKA 304
G+RQGDPLSP LF + E L ++ G I I H + DD ++ CKA
Sbjct: 610 GIRQGDPLSPALFVLCTEALIHILNKAEQAGKITGIQFQDKKVSVNHLLFADDTLLMCKA 669
Query: 305 KMSSLIVLKSLFTRYADCSGQIMNIRKSFIFAGGITDTRMNN-IVNILGFNVGSLPFTYL 363
L ++Y SGQ++N+ KS I G D ++ + I + G ++ YL
Sbjct: 670 TKQECEELMQCLSQYGQLSGQMINLNKSAITFGKNVDIQIKDWIKSRSGISLEGGTGKYL 729
Query: 364 GAPIFKGKPKGIHFQPIADKVKAKLAKWKASLLSIAGRIQLVKSVVQSMLVHTMSIYSWP 423
G P K F I +K++++L W A LS G+ L+KS+ ++ V+ MS + P
Sbjct: 730 GLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTLSQGGKEVLLKSIALALPVYVMSCFKLP 789
Query: 424 IKILKEMEKWIKNFIWSGDVTKRKMVTVAWRKICADYEEGGLGVKSLICLNEATNLKICW 483
+ +++ + +F W+ KRK+ ++W+++ ++GG G K L C N+A K W
Sbjct: 790 KNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLPKDQGGFGFKDLQCFNQALLAKQAW 849
Query: 484 NLMQ-SDEQWANIIRSRVIRDHRCMNHHISSS---IWSGVKAEFHVIKDNCSWIVGDGKQ 539
++Q ++ + +SR + ++ S W + ++ ++G+G++
Sbjct: 850 RVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYAWRSILFGRELLMQGLRTVIGNGQK 909
Query: 540 INFWSDSW 547
W+D W
Sbjct: 910 TFVWTDKW 917
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.140 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,481,810
Number of Sequences: 26719
Number of extensions: 528691
Number of successful extensions: 1756
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 64
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1518
Number of HSP's gapped (non-prelim): 105
length of query: 555
length of database: 11,318,596
effective HSP length: 104
effective length of query: 451
effective length of database: 8,539,820
effective search space: 3851458820
effective search space used: 3851458820
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC149134.1


