

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149129.7 + phase: 0
(726 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
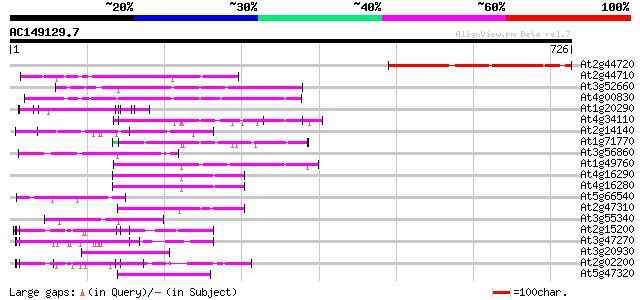
Score E
Sequences producing significant alignments: (bits) Value
At2g44720 unknown protein 259 4e-69
At2g44710 putative heterogeneous nuclear ribonucleoprotein 186 3e-47
At3g52660 putative RNA binding protein 130 3e-30
At4g00830 unknown protein 122 5e-28
At1g20290 hypothetical protein 74 4e-13
At4g34110 poly(A)-binding protein 61 2e-09
At2g14140 putative transposase of FARE2.8 (cds1) 59 7e-09
At1g71770 polyadenylate-binding protein 5 59 7e-09
At3g56860 unknown protein 59 1e-08
At1g49760 Putative Poly-A Binding Protein (F14J22.3) 59 1e-08
At4g16290 FCA delta protein 58 2e-08
At4g16280 FCA gamma protein 58 2e-08
At5g66540 unknown protein 57 3e-08
At2g47310 putative FCA-related protein 57 3e-08
At3g55340 putative protein 57 4e-08
At2g15200 putative protein on transposon FARE2.7 (cds2) 55 2e-07
At3g47270 putative protein 54 2e-07
At3g20930 unknown protein 54 2e-07
At2g02200 putative protein on transposon FARE2.10 (cds2) 54 3e-07
At5g47320 40S ribosomal protein S19 54 4e-07
>At2g44720 unknown protein
Length = 239
Score = 259 bits (662), Expect = 4e-69
Identities = 143/238 (60%), Positives = 166/238 (69%), Gaps = 16/238 (6%)
Query: 491 YPKSSMKRDYGRPVDLPPPRSRVSADYGSQVVSQRGPSYRD-YPARDSGYPDLHRDTSRT 549
YPK+S+KRDY R +LPPPRSR + Y S++ +R SYRD YP R SGY DL R +SR+
Sbjct: 16 YPKASLKRDYDRRDELPPPRSRPAVSYSSRLSPERHLSYRDDYPPRGSGYSDLPRSSSRS 75
Query: 550 APRRGYLDDGYDRRLERPPSPSPRLSYREGRPRDYDALSGSKRPYAAIDDISPRYADAGA 609
RR ++DD Y R ERPPS Y EGRPR Y+ L GSKRPYAA+DD+ PRYAD
Sbjct: 76 EIRRPFVDDLYSPRFERPPS------YSEGRPRAYEPLPGSKRPYAALDDLPPRYADVDV 129
Query: 610 RQSRSRLDYDYGGSASRYREAYGDRLERSSLGYSGSRSSLSNQNSHGAYSSRQDPSYGRG 669
R SR RLDYD G S+Y E+YGDR+ RSSLGY SR+S+SN +S G YSSRQ YG G
Sbjct: 130 RHSRPRLDYDVG--PSQYGESYGDRIPRSSLGYGSSRNSMSNHDSRGPYSSRQGMDYGGG 187
Query: 670 SLGGSD-GGIYSSSYGGDYLSRGSDVGGSSYSSMYSSRGMNGSSSYMSGSGGGSRSYY 726
S GSD GG+YSSSYGGD R D GGSSYSS+YSSRG+ GSS SGGG SYY
Sbjct: 188 SYSGSDVGGMYSSSYGGDLPRR--DGGGSSYSSIYSSRGLGGSSY----SGGGPGSYY 239
>At2g44710 putative heterogeneous nuclear ribonucleoprotein
Length = 300
Score = 186 bits (473), Expect = 3e-47
Identities = 116/299 (38%), Positives = 172/299 (56%), Gaps = 30/299 (10%)
Query: 14 EVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQ---EAGDEVEEEPE 70
EV+ES+D++ KDE LD +DN EY+ EEY G ++++ER + Q++ + E G+E+EEE E
Sbjct: 12 EVEESVDDFGKDERLDLDDNEPEYEAEEY-GGEEFEERELGQEDHELVNEEGEELEEEIE 70
Query: 71 ENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVI 130
V+EE G+ DE + E +++ +DDD D+HA EE H DV+EEE +V+
Sbjct: 71 --VEEEAGEFADEIGDGAE----DLDSEDDD----DDHAIEEVKHGETVDVEEEEHHDVL 120
Query: 131 KETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETV 190
E +K+KE E+FVG LDK A+E DL+KVF VGE+TEVR+ NPQTK++KG A LRF TV
Sbjct: 121 HERRKRKEFEIFVGSLDKGASEEDLKKVFGHVGEVTEVRILKNPQTKKSKGSAFLRFATV 180
Query: 191 EHVKRALAELKNP--VINGKQ-----------CGITITTCQDSDTLYLDNICKSWKKEAL 237
E KRA+ ELKN +I G+ I T+++D + SW +E +
Sbjct: 181 EQAKRAVKELKNTELLIVGESIISNLSILLIVISFMIFLVLQVKTIFIDGLLPSWNEERV 240
Query: 238 KEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDK 296
++ LK YG + L + F+ F +H + K + +++ G DK
Sbjct: 241 RDLLKPYGK---LEKVELARNMPSARRKDFGFVTFDTHEAAVSCAKFINNSELGEGEDK 296
>At3g52660 putative RNA binding protein
Length = 471
Score = 130 bits (326), Expect = 3e-30
Identities = 97/335 (28%), Positives = 167/335 (48%), Gaps = 32/335 (9%)
Query: 60 EAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLA 119
EA D +E E ++D GD EE+ E E YEEVE ++++ +E EE
Sbjct: 10 EAHDSMESEERVDLD---GDNDPEEILEEEVEYEEVE--EEEIEEIEEEIEEE------V 58
Query: 120 DVKEEEQVEVIKETQKQKEL--------------EVFVGGLDKEATEHDLRKVFSKVGEI 165
+V+EEE+ E T++++E EV++GG+ +ATE DL+ +GE+
Sbjct: 59 EVEEEEEEEDAVATEEEEEKKRHVELLALPPHGSEVYLGGIPTDATEGDLKGFCGSIGEV 118
Query: 166 TEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYL 225
TEVR+ + KG+A + F + + A+ L N GK+ I +T Q L+L
Sbjct: 119 TEVRIMREKDSGDGKGYAFVTFRSKDLAAEAIDTLNNTDFRGKR--IKCSTTQAKHRLFL 176
Query: 226 DNICKSWKKEALKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRL 285
N+ ++W + +K+ G + + L ++ N G N G AF+E+ +H+ ++ Y +
Sbjct: 177 GNVPRNWMESDIKKAANRIG-PGVQIVELPKEPQNMGRNRGFAFIEYYNHACAE--YSKQ 233
Query: 286 QKTDVVFGVDKPA-EVSFANSFIDLGDDIMA-QVKTVFIDLLPPSWDEDYVRALLKKYGA 343
+ ++ F +D A VS+A S G D A QVK ++I LP ++ ++AL + +G
Sbjct: 234 KMSNPSFKLDDNAPTVSWAESRSGGGGDSSASQVKALYIKNLPRDITQERLKALFEHHGK 293
Query: 344 VEKVELAKNMPGARRKNYGFVTFGTHAAAVECAES 378
+ KV + PG YGFV + + + ++
Sbjct: 294 ILKVVIPPAKPGKEDSRYGFVHYAERTSVMRALKN 328
>At4g00830 unknown protein
Length = 495
Score = 122 bits (307), Expect = 5e-28
Identities = 94/362 (25%), Positives = 189/362 (51%), Gaps = 19/362 (5%)
Query: 20 DEYEKDEHLDFED-NYHEYDPE-EYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEE 77
D + D+ +DFE+ +Y E + E E ++Y+E E D+ + G++ EE E E+
Sbjct: 3 DARDNDDRVDFEEGSYSEMEDEVEEEQVEEYEEEEEEDDDDDDVGNQNAEEREV---EDY 59
Query: 78 GDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQK 137
GDT ++E+V+ EE+ DDD+ D E A ++ D ++ E+ +
Sbjct: 60 GDTKGGDMEDVQ---EEIAEDDDNHI-DIETADDDEKPPSPIDDEDREKYSHLLSLPPHG 115
Query: 138 ELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRAL 197
EVF+GGL ++ E DLR + ++GEI EVR+ + + +KG+A + F+T + ++A+
Sbjct: 116 S-EVFIGGLPRDVGEEDLRDLCEEIGEIFEVRLMKDRDSGDSKGYAFVAFKTKDVAQKAI 174
Query: 198 AELKNPVINGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLED 257
EL + GK I + + + L++ NI K+W ++ ++ ++ G +++ L++D
Sbjct: 175 EELHSKEFKGKT--IRCSLSETKNRLFIGNIPKNWTEDEFRKVIEDVG-PGVENIELIKD 231
Query: 258 DNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPA-EVSFAN-SFIDLGDDIMA 315
N N G AF+ + ++++ Y R + D F ++ A V++A+ A
Sbjct: 232 PTNTTRNRGFAFVLY--YNNACADYSRQKMIDSNFKLEGNAPTVTWADPKSSPEHSAAAA 289
Query: 316 QVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHAAAVEC 375
QVK +++ +P + + ++ L +++G V K+ G ++++GFV + ++A++
Sbjct: 290 QVKALYVKNIPENTSTEQLKELFQRHGEVTKIVTPPGKGG--KRDFGFVHYAERSSALKA 347
Query: 376 AE 377
+
Sbjct: 348 VK 349
Score = 36.6 bits (83), Expect = 0.051
Identities = 35/159 (22%), Positives = 66/159 (41%), Gaps = 14/159 (8%)
Query: 237 LKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDK 296
++++++ VE +++ +DD+++ N E + D+K + ++ D
Sbjct: 21 MEDEVEEEQVEEYEEEEEEDDDDDDVGNQNAEEREVEDYGDTKGGDMEDVQEEIAEDDDN 80
Query: 297 PAEVSFAN------SFIDLGD--------DIMAQVKTVFIDLLPPSWDEDYVRALLKKYG 342
++ A+ S ID D + VFI LP E+ +R L ++ G
Sbjct: 81 HIDIETADDDEKPPSPIDDEDREKYSHLLSLPPHGSEVFIGGLPRDVGEEDLRDLCEEIG 140
Query: 343 AVEKVELAKNMPGARRKNYGFVTFGTHAAAVECAESITS 381
+ +V L K+ K Y FV F T A + E + S
Sbjct: 141 EIFEVRLMKDRDSGDSKGYAFVAFKTKDVAQKAIEELHS 179
>At1g20290 hypothetical protein
Length = 497
Score = 73.6 bits (179), Expect = 4e-13
Identities = 39/125 (31%), Positives = 73/125 (58%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEEN 72
EE +E +E E++E + E+ E + EE +++ +E E++E +E +E EEE EE
Sbjct: 370 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 429
Query: 73 VDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKE 132
+EEE + +EE EE E EE E ++++ ++E EE E + +EE++ ++K+
Sbjct: 430 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKKYMIVKK 489
Query: 133 TQKQK 137
+K++
Sbjct: 490 KKKKR 494
Score = 72.8 bits (177), Expect = 6e-13
Identities = 41/125 (32%), Positives = 72/125 (56%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEEN 72
EE +E +E E++E + E+ E + EE +++ +E E++E +E +E EEE EE
Sbjct: 369 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 428
Query: 73 VDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKE 132
+EEE + +EE EE E EE E ++++ ++E EE E + +EEE+ +I +
Sbjct: 429 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKKYMIVK 488
Query: 133 TQKQK 137
+K+K
Sbjct: 489 KKKKK 493
Score = 70.9 bits (172), Expect = 2e-12
Identities = 41/130 (31%), Positives = 73/130 (55%)
Query: 12 GEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEE 71
G V+ +E E++E + E+ E + EE +++ +E E++E +E +E EEE EE
Sbjct: 358 GTIVQAEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 417
Query: 72 NVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIK 131
+EEE + +EE EE E EE E ++++ ++E EE E + +EEE+ E +
Sbjct: 418 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 477
Query: 132 ETQKQKELEV 141
E +++K + V
Sbjct: 478 EEEEKKYMIV 487
Score = 70.1 bits (170), Expect = 4e-12
Identities = 40/132 (30%), Positives = 73/132 (55%)
Query: 12 GEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEE 71
GE +E E++E + E+ E + EE +++ +E E++E +E +E EEE EE
Sbjct: 356 GEGTIVQAEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 415
Query: 72 NVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIK 131
+EEE + +EE EE E EE E ++++ ++E EE E + +EEE+ E +
Sbjct: 416 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 475
Query: 132 ETQKQKELEVFV 143
E +++++ + V
Sbjct: 476 EEEEEEKKYMIV 487
Score = 63.5 bits (153), Expect = 4e-10
Identities = 37/120 (30%), Positives = 67/120 (55%)
Query: 38 DPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEG 97
+ EE +++ +E E++E +E +E EEE EE +EEE + +EE EE E EE E
Sbjct: 364 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 423
Query: 98 DDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRK 157
++++ ++E EE E + +EEE+ E +E ++++E E ++E E + +K
Sbjct: 424 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKK 483
Score = 62.8 bits (151), Expect = 7e-10
Identities = 37/124 (29%), Positives = 68/124 (54%)
Query: 38 DPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEG 97
+ EE +++ +E E++E +E +E EEE EE +EEE + +EE EE E EE E
Sbjct: 365 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 424
Query: 98 DDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRK 157
++++ ++E EE E + +EEE+ E +E ++++E E ++E E + +
Sbjct: 425 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEKKY 484
Query: 158 VFSK 161
+ K
Sbjct: 485 MIVK 488
Score = 60.5 bits (145), Expect = 3e-09
Identities = 41/154 (26%), Positives = 80/154 (51%), Gaps = 4/154 (2%)
Query: 31 EDNYHEYDPEE-YVGDDDY---DERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVE 86
+ +++++ P +VG+ +E E++E +E +E EEE EE +EEE + +EE E
Sbjct: 342 DSDFYKFTPSMVHVGEGTIVQAEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 401
Query: 87 EVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQKELEVFVGGL 146
E E EE E ++++ ++E EE E + +EEE+ E +E ++++E E
Sbjct: 402 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 461
Query: 147 DKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNK 180
++E E + + + E + M V + K+ +
Sbjct: 462 EEEEEEEEEEEEEEEEEEEEKKYMIVKKKKKKRR 495
>At4g34110 poly(A)-binding protein
Length = 629
Score = 61.2 bits (147), Expect = 2e-09
Identities = 62/294 (21%), Positives = 119/294 (40%), Gaps = 37/294 (12%)
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
+F+ LD+ L FS G I ++ V+ + ++KG+ +++ E ++A+ +L
Sbjct: 126 IFIKNLDESIDHKALHDTFSSFGNIVSCKVAVD-SSGQSKGYGFVQYANEESAQKAIEKL 184
Query: 201 KNPVINGKQCGI-TITTCQDSDT---------LYLDNICKSWKKEALKEKLKHYGVESFK 250
++N KQ + Q+ D+ +Y+ N+ +S + LK YG K
Sbjct: 185 NGMLLNDKQVYVGPFLRRQERDSTANKTKFTNVYVKNLAESTTDDDLKNAFGEYG----K 240
Query: 251 DLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQ------KTDVVFGVDKPAE----- 299
+ + + EG + G F+ F + D+ A + L K V K +E
Sbjct: 241 ITSAVVMKDGEGKSKGFGFVNFENADDAARAVESLNGHKFDDKEWYVGRAQKKSERETEL 300
Query: 300 -VSFANSFIDLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARR 358
V + + + D Q +++ L PS ++ ++ + +G V ++ ++ P
Sbjct: 301 RVRYEQNLKEAADKF--QSSNLYVKNLDPSISDEKLKEIFSPFGTVTSSKVMRD-PNGTS 357
Query: 359 KNYGFVTFGTHAAAVECAESITSAGLGE-------GNKKAKVRARLSRPLPKVR 405
K GFV F T A E ++ + +K R RL +VR
Sbjct: 358 KGSGFVAFATPEEATEAMSQLSGKMIESKPLYVAIAQRKEDRRVRLQAQFSQVR 411
Score = 56.6 bits (135), Expect = 5e-08
Identities = 53/249 (21%), Positives = 105/249 (41%), Gaps = 21/249 (8%)
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
++VG LD T+ L F ++G + VR+ + T+R+ G+ + F + RA+ EL
Sbjct: 38 LYVGDLDFNVTDSQLFDAFGQMGTVVTVRVCRDLVTRRSLGYGYVNFTNPQDAARAIQEL 97
Query: 201 KNPVINGKQCGITITTCQDS------DTLYLDNICKSWKKEALKEKLKHYGVESFKDLTL 254
+ GK + + S +++ N+ +S +AL + +G + +
Sbjct: 98 NYIPLYGKPIRVMYSHRDPSVRRSGAGNIFIKNLDESIDHKALHDTFSSFG--NIVSCKV 155
Query: 255 LEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPAEVSFANSFIDLGDDIM 314
D + + G F +++ + A K ++K + + DK + F+ +
Sbjct: 156 AVDSSGQSKGYG-----FVQYANEESAQKAIEKLNGMLLNDKQV---YVGPFLRRQERDS 207
Query: 315 AQVKT----VFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFGTHA 370
KT V++ L S +D ++ +YG + + K+ G + K +GFV F
Sbjct: 208 TANKTKFTNVYVKNLAESTTDDDLKNAFGEYGKITSAVVMKDGEG-KSKGFGFVNFENAD 266
Query: 371 AAVECAESI 379
A ES+
Sbjct: 267 DAARAVESL 275
Score = 49.7 bits (117), Expect = 6e-06
Identities = 55/217 (25%), Positives = 89/217 (40%), Gaps = 32/217 (14%)
Query: 135 KQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVK 194
K K V+V L + T+ DL+ F + G+IT + + + K +KGF + FE +
Sbjct: 211 KTKFTNVYVKNLAESTTDDDLKNAFGEYGKITSAVVMKDGEGK-SKGFGFVNFENADDAA 269
Query: 195 RALAELKNPVINGKQCGI-------------------TITTCQD---SDTLYLDNICKSW 232
RA+ L + K+ + + D S LY+ N+ S
Sbjct: 270 RAVESLNGHKFDDKEWYVGRAQKKSERETELRVRYEQNLKEAADKFQSSNLYVKNLDPSI 329
Query: 233 KKEALKEKLKHYG-VESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVV 291
E LKE +G V S K ++ D N GT+ G F+ F++ ++ +A +L +
Sbjct: 330 SDEKLKEIFSPFGTVTSSK---VMRDPN--GTSKGSGFVAFATPEEATEAMSQLSGKMI- 383
Query: 292 FGVDKPAEVSFANSFIDLGDDIMAQVKTVFIDLLPPS 328
KP V+ A D + AQ V + PS
Sbjct: 384 --ESKPLYVAIAQRKEDRRVRLQAQFSQVRPVAMQPS 418
>At2g14140 putative transposase of FARE2.8 (cds1)
Length = 783
Score = 59.3 bits (142), Expect = 7e-09
Identities = 71/246 (28%), Positives = 107/246 (42%), Gaps = 29/246 (11%)
Query: 37 YDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEE-V 95
Y+ EY GD+ G E+ E + GDE E E EE EEEG +EE E+VEY +E
Sbjct: 390 YENFEYRGDE-----GTEKQEIPKQGDE-EMEGEEEKQEEEGK--EEEEEKVEYRGDEGT 441
Query: 96 EGDDDDVAGDDEHASEENVHEHLADVKEEEQVEV--IKETQKQ-------KELEVFVGGL 146
E + GD+E EE E KEEE+VE +ET+KQ +E+E
Sbjct: 442 EKQEIPKQGDEEMEGEEEKQEEEGKEKEEEKVEYRGDEETEKQEIPKQGDEEMEGEEEKQ 501
Query: 147 DKEATEHDLRKVFSKVGEIT-EVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVI 205
++E E + KV K T V T + + + R E E E +
Sbjct: 502 EEEGKEEEEEKVEYKDHHSTCNVEETEKQENPKQGDEEMEREEGKEEKVEEHDEYNDAA- 560
Query: 206 NGKQCGITITTCQDSDTLYLDNICKSWKKEA--------LKEKLKHYGVESFKDLTLLED 257
++ I ++ +D+DT + + K+E ++E +H E + +L D
Sbjct: 561 -DQEAYINLSDDEDNDTAPTEKESQPQKEETTEVPKEENVEEHDEHDETEDQEAYVILSD 619
Query: 258 DNNEGT 263
D + GT
Sbjct: 620 DEDNGT 625
Score = 55.5 bits (132), Expect = 1e-07
Identities = 43/158 (27%), Positives = 77/158 (48%), Gaps = 20/158 (12%)
Query: 8 IADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEE 67
I G+E E +E +++E + E+ EY +E + ++G E+ EG+E ++ EE
Sbjct: 406 IPKQGDEEMEGEEEKQEEEGKEEEEEKVEYRGDEGTEKQEIPKQGDEEMEGEE--EKQEE 463
Query: 68 EPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENV-----------HE 116
E +E +E+ GDEE E+ E + D+++ G++E EE H
Sbjct: 464 EGKEKEEEKVEYRGDEETEKQEIPKQ----GDEEMEGEEEKQEEEGKEEEEEKVEYKDHH 519
Query: 117 HLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHD 154
+V+E E+ E K+ ++ E E G +++ EHD
Sbjct: 520 STCNVEETEKQENPKQGDEEMERE---EGKEEKVEEHD 554
Score = 49.3 bits (116), Expect = 8e-06
Identities = 44/165 (26%), Positives = 71/165 (42%), Gaps = 25/165 (15%)
Query: 8 IADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEE 67
I G+E E +E +++E + E+ EY +E + ++G E+ EG+E E
Sbjct: 446 IPKQGDEEMEGEEEKQEEEGKEKEEEKVEYRGDEETEKQEIPKQGDEEMEGEE------E 499
Query: 68 EPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDD----EHASEENVHEH------ 117
+ EE EEE + + + EE E ++ GD+ E EE V EH
Sbjct: 500 KQEEEGKEEEEEKVEYKDHHSTCNVEETEKQENPKQGDEEMEREEGKEEKVEEHDEYNDA 559
Query: 118 --------LADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHD 154
L+D ++ + KE+Q QKE E ++ EHD
Sbjct: 560 ADQEAYINLSDDEDNDTAPTEKESQPQKE-ETTEVPKEENVEEHD 603
Score = 41.6 bits (96), Expect = 0.002
Identities = 45/154 (29%), Positives = 68/154 (43%), Gaps = 19/154 (12%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEEN 72
EE KE +E EK E+ D + + E+ DE +E++EG+E ++VEE E N
Sbjct: 503 EEGKE--EEEEKVEYKDHHSTCNVEETEKQENPKQGDEE-MEREEGKE--EKVEEHDEYN 557
Query: 73 -----------VDEEEGDTGDEEVEEVEYVYEEVE-GDDDDVAGDDEHASEENVHEH--L 118
D+E+ DT E E E E +++V DEH E+ + L
Sbjct: 558 DAADQEAYINLSDDEDNDTAPTEKESQPQKEETTEVPKEENVEEHDEHDETEDQEAYVIL 617
Query: 119 ADVKEEEQVEVIKETQKQKELEVFVGGLDKEATE 152
+D ++ KE+Q QK V G K+ E
Sbjct: 618 SDDEDNGTAPTEKESQPQKVETTEVPGETKKDDE 651
Score = 40.4 bits (93), Expect = 0.004
Identities = 39/134 (29%), Positives = 64/134 (47%), Gaps = 18/134 (13%)
Query: 10 DNGEEVKESIDEYE-KDEHLDFEDNYHEY-DPEEYVG--DDDYDERGIEQDEGQEAGDEV 65
+N ++ E ++ E K+E ++ D Y++ D E Y+ DD+ ++ + E Q +E
Sbjct: 531 ENPKQGDEEMEREEGKEEKVEEHDEYNDAADQEAYINLSDDEDNDTAPTEKESQPQKEET 590
Query: 66 EEEP-EENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEE 124
E P EENV EE D DE ++ YV + D+D+ E S+ ++
Sbjct: 591 TEVPKEENV--EEHDEHDETEDQEAYVI--LSDDEDNGTAPTEKESQP---------QKV 637
Query: 125 EQVEVIKETQKQKE 138
E EV ET+K E
Sbjct: 638 ETTEVPGETKKDDE 651
>At1g71770 polyadenylate-binding protein 5
Length = 668
Score = 59.3 bits (142), Expect = 7e-09
Identities = 60/265 (22%), Positives = 105/265 (38%), Gaps = 24/265 (9%)
Query: 134 QKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHV 193
Q ++VG LD E L +F++V + +R+ T R+ G+A + F E
Sbjct: 40 QTHPNSSLYVGDLDPSVNESHLLDLFNQVAPVHNLRVC-RDLTHRSLGYAYVNFANPEDA 98
Query: 194 KRALAELKNPVINGKQCGITITTCQDSDTL------YLDNICKSWKKEALKEKLKHYGVE 247
RA+ L I + I ++ S L ++ N+ S +AL E +G
Sbjct: 99 SRAMESLNYAPIRDRPIRIMLSNRDPSTRLSGKGNVFIKNLDASIDNKALYETFSSFGT- 157
Query: 248 SFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPAEVSFANSFI 307
L+ + G + G F++F ++ A +L G+ + F F+
Sbjct: 158 ---ILSCKVAMDVVGRSKGYGFVQFEKEETAQAAIDKLN------GMLLNDKQVFVGHFV 208
Query: 308 DLGDDIMAQ------VKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNY 361
D ++ V++ LP +D ++ KYG + + K+ G R ++
Sbjct: 209 RRQDRARSESGAVPSFTNVYVKNLPKEITDDELKKTFGKYGDISSAVVMKDQSGNSR-SF 267
Query: 362 GFVTFGTHAAAVECAESITSAGLGE 386
GFV F + AA E + LGE
Sbjct: 268 GFVNFVSPEAAAVAVEKMNGISLGE 292
Score = 45.1 bits (105), Expect = 1e-04
Identities = 55/269 (20%), Positives = 110/269 (40%), Gaps = 32/269 (11%)
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
VF+ LD L + FS G I ++ ++ R+KG+ ++FE E + A+ +L
Sbjct: 134 VFIKNLDASIDNKALYETFSSFGTILSCKVAMDV-VGRSKGYGFVQFEKEETAQAAIDKL 192
Query: 201 KNPVINGKQCGI-TITTCQDS-----------DTLYLDNICKSWKKEALKEKLKHYGVES 248
++N KQ + QD +Y+ N+ K + LK+ YG
Sbjct: 193 NGMLLNDKQVFVGHFVRRQDRARSESGAVPSFTNVYVKNLPKEITDDELKKTFGKYG--D 250
Query: 249 FKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQ----KTDVVF------GVDKPA 298
+++D + + G F+ F S + A +++ DV++ D+
Sbjct: 251 ISSAVVMKDQSGNSRSFG--FVNFVSPEAAAVAVEKMNGISLGEDVLYVGRAQKKSDREE 308
Query: 299 EV--SFANSFIDLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGA 356
E+ F I + + Q +++ L S +++ ++ + +YG V ++ N G
Sbjct: 309 ELRRKFEQERISRFEKL--QGSNLYLKNLDDSVNDEKLKEMFSEYGNVTSCKVMMNSQGL 366
Query: 357 RRKNYGFVTFGTHAAAVECAESITSAGLG 385
R +GFV + A+ + + +G
Sbjct: 367 SR-GFGFVAYSNPEEALLAMKEMNGKMIG 394
>At3g56860 unknown protein
Length = 478
Score = 58.9 bits (141), Expect = 1e-08
Identities = 48/223 (21%), Positives = 99/223 (43%), Gaps = 34/223 (15%)
Query: 12 GEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEE 71
GEE E+ + +K + E+ ++ GD +E+ E +E +E EEE E+
Sbjct: 9 GEESNEAEEPSQKLKQTPEEEQQLVIKNQDNQGD-------VEEVEYEEVEEEQEEEVED 61
Query: 72 NVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIK 131
+ DE++GD +++ + +E +G+ E +E + + L +E+ + ++K
Sbjct: 62 DDDEDDGDENEDQTDG-----NRIEAAATSGSGNQEDDDDEPIQDLLEPFSKEQVLSLLK 116
Query: 132 ETQKQK----------------ELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQ 175
E ++ ++FV GL + L + F + GEI + + +
Sbjct: 117 EAAEKHVDVANRIREVADEDPVHRKIFVHGLGWDTKTETLIEAFKQYGEIEDCKAVFDKI 176
Query: 176 TKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQ 218
+ ++KG+ + +++ + A LK P K+ G +T CQ
Sbjct: 177 SGKSKGYGFILYKSRSGARNA---LKQP---QKKIGSRMTACQ 213
Score = 40.4 bits (93), Expect = 0.004
Identities = 25/69 (36%), Positives = 36/69 (51%), Gaps = 4/69 (5%)
Query: 131 KETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETV 190
+ TQK+ ++V + E L FSK GEI E + ++ T R KGF L +++
Sbjct: 241 EHTQKK----IYVSNVGAELDPQKLLMFFSKFGEIEEGPLGLDKYTGRPKGFCLFVYKSS 296
Query: 191 EHVKRALAE 199
E KRAL E
Sbjct: 297 ESAKRALEE 305
>At1g49760 Putative Poly-A Binding Protein (F14J22.3)
Length = 671
Score = 58.9 bits (141), Expect = 1e-08
Identities = 55/279 (19%), Positives = 117/279 (41%), Gaps = 22/279 (7%)
Query: 135 KQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVK 194
+Q ++VG LD T+ L + F++ G++ VR+ + T+R+ G+ + + T +
Sbjct: 41 QQGTTSLYVGDLDATVTDSQLFEAFTQAGQVVSVRVCRDMTTRRSLGYGYVNYATPQDAS 100
Query: 195 RALAELKNPVINGKQCGITITTCQDS------DTLYLDNICKSWKKEALKEKLKHYGVES 248
RAL EL +NG+ + + S +++ N+ KS +AL E +G
Sbjct: 101 RALNELNFMALNGRAIRVMYSVRDPSLRKSGVGNIFIKNLDKSIDHKALHETFSAFG--- 157
Query: 249 FKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPAEVS-FANSFI 307
L+ + G + G F+++ + ++ A + K + + DK V F +
Sbjct: 158 -PILSCKVAVDPSGQSKGYGFVQYDTDEAAQGA---IDKLNGMLLNDKQVYVGPFVHKLQ 213
Query: 308 DLGDDIMAQVKTVFIDLLPPSWDEDYVRALLKKYGAVEKVELAKNMPGARRKNYGFVTFG 367
+ V++ L S ++ + + ++G + ++ G + K +GFV F
Sbjct: 214 RDPSGEKVKFTNVYVKNLSESLSDEELNKVFGEFGVTTSCVIMRDGEG-KSKGFGFVNFE 272
Query: 368 THAAAVECAESITSAG-------LGEGNKKAKVRARLSR 399
A +++ +G+ KK++ L +
Sbjct: 273 NSDDAARAVDALNGKTFDDKEWFVGKAQKKSERETELKQ 311
Score = 40.4 bits (93), Expect = 0.004
Identities = 45/217 (20%), Positives = 87/217 (39%), Gaps = 30/217 (13%)
Query: 134 QKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHV 193
+K K V+V L + ++ +L KVF + G +T + + ++KGF + FE +
Sbjct: 219 EKVKFTNVYVKNLSESLSDEELNKVFGEFG-VTTSCVIMRDGEGKSKGFGFVNFENSDDA 277
Query: 194 KRALAELKNPVINGKQCGI-------------------TITTCQD---SDTLYLDNICKS 231
RA+ L + K+ + ++ D LY+ N+ +S
Sbjct: 278 ARAVDALNGKTFDDKEWFVGKAQKKSERETELKQKFEQSLKEAADKSQGSNLYVKNLDES 337
Query: 232 WKKEALKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVV 291
+ L+E +G + ++ D + G + G F+ FS+ ++ A + +
Sbjct: 338 VTDDKLREHFAPFG--TITSCKVMRDPS--GVSRGSGFVAFSTPEEATRAITEMNGKMI- 392
Query: 292 FGVDKPAEVSFANSFIDLGDDIMAQVKTVFIDLLPPS 328
V KP V+ A D + AQ + +PP+
Sbjct: 393 --VTKPLYVALAQRKEDRKARLQAQFSQMRPVNMPPA 427
Score = 40.4 bits (93), Expect = 0.004
Identities = 29/128 (22%), Positives = 57/128 (43%), Gaps = 8/128 (6%)
Query: 79 DTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQKE 138
+ D+ V+ + + D + G + SE ++K++ + + + K +
Sbjct: 272 ENSDDAARAVDALNGKTFDDKEWFVGKAQKKSERET-----ELKQKFEQSLKEAADKSQG 326
Query: 139 LEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQ-TKRNKGFALLRFETVEHVKRAL 197
++V LD+ T+ LR+ F+ G IT ++ +P R GF + F T E RA+
Sbjct: 327 SNLYVKNLDESVTDDKLREHFAPFGTITSCKVMRDPSGVSRGSGF--VAFSTPEEATRAI 384
Query: 198 AELKNPVI 205
E+ +I
Sbjct: 385 TEMNGKMI 392
>At4g16290 FCA delta protein
Length = 533
Score = 57.8 bits (138), Expect = 2e-08
Identities = 41/178 (23%), Positives = 84/178 (47%), Gaps = 9/178 (5%)
Query: 133 TQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEH 192
+ + +++FVG + + ATE ++R F + G + EV + + +T + +G +++ T +
Sbjct: 114 SDRSSTVKLFVGSVPRTATEEEIRPYFEQHGNVLEVALIKDKRTGQQQGCCFVKYATSKD 173
Query: 193 VKRALAELKNPVINGKQCGITITTCQDSD-----TLYLDNICKSWKKEALKEKLKHYGVE 247
RA+ L N + G D + TL S K+A +++++ ++
Sbjct: 174 ADRAIRALHNQITLPGGTGPVQVRYADGERERIGTLEFKLFVGSLNKQATEKEVEEIFLQ 233
Query: 248 --SFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPAEVSFA 303
+D+ L+ D+ + GC F+++SS + A L T + G ++P V FA
Sbjct: 234 FGHVEDVYLMRDEYRQSR--GCGFVKYSSKETAMAAIDGLNGTYTMRGCNQPLIVRFA 289
>At4g16280 FCA gamma protein
Length = 747
Score = 57.8 bits (138), Expect = 2e-08
Identities = 41/178 (23%), Positives = 84/178 (47%), Gaps = 9/178 (5%)
Query: 133 TQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEH 192
+ + +++FVG + + ATE ++R F + G + EV + + +T + +G +++ T +
Sbjct: 114 SDRSSTVKLFVGSVPRTATEEEIRPYFEQHGNVLEVALIKDKRTGQQQGCCFVKYATSKD 173
Query: 193 VKRALAELKNPVINGKQCGITITTCQDSD-----TLYLDNICKSWKKEALKEKLKHYGVE 247
RA+ L N + G D + TL S K+A +++++ ++
Sbjct: 174 ADRAIRALHNQITLPGGTGPVQVRYADGERERIGTLEFKLFVGSLNKQATEKEVEEIFLQ 233
Query: 248 --SFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPAEVSFA 303
+D+ L+ D+ + GC F+++SS + A L T + G ++P V FA
Sbjct: 234 FGHVEDVYLMRDEYRQSR--GCGFVKYSSKETAMAAIDGLNGTYTMRGCNQPLIVRFA 289
>At5g66540 unknown protein
Length = 524
Score = 57.4 bits (137), Expect = 3e-08
Identities = 46/155 (29%), Positives = 78/155 (49%), Gaps = 20/155 (12%)
Query: 9 ADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDD-DYDERGIE------QDEGQEA 61
A N EE+++ K H ED+ E D + + DD D +++ IE +DE +E
Sbjct: 89 AKNPEEIRKLGKLALKVSH---EDDIDEMDMDGFDSDDVDDEDKEIESNDSEGEDEEEEE 145
Query: 62 GDEVEEEPEENVDEEEGDTGDEEVE-------EVEYVYEEVEGDDDDVAGDDEHASEENV 114
DE EEE EE +EEE D +E +E E+E EE E ++ + ++ +
Sbjct: 146 EDEEEEEEEEEEEEEEKDGDNEGIEDKFFKIKELEEFLEEGEAEEYGIDHKNKKGVAQRK 205
Query: 115 HEHLADVKEEEQVEVIKETQKQKELEVFVGGLDKE 149
++L+D ++EE + + ++ E + F GG D+E
Sbjct: 206 KQNLSDDEDEEDDD---DEEEDVEFDAFAGGDDEE 237
>At2g47310 putative FCA-related protein
Length = 324
Score = 57.4 bits (137), Expect = 3e-08
Identities = 45/184 (24%), Positives = 80/184 (43%), Gaps = 23/184 (12%)
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
+++V + K ATE+D+R+VF K G +TE+ + + T + ++++ VE A+A
Sbjct: 111 KLYVAPISKTATEYDIRQVFEKYGNVTEIILPKDKMTGERAAYCFIKYKKVEEGNAAIAA 170
Query: 200 LKNP-VINGKQCGITITTCQ------------------DSDTLYLDNICKSWKKEALKEK 240
L G+ + + + D+ LY+ + K K + E
Sbjct: 171 LTEQFTFPGEMLPVKVRFAEAERERIGKCRCFAPVQLPDNPKLYVRCLNKQTTKMEVNEV 230
Query: 241 LKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVDKPAEV 300
YG+ +D+ + DD G AF++FS + A K L + G D+P V
Sbjct: 231 FSRYGI--IEDIYMALDDMK--ICRGYAFVQFSCKEMALAAIKALNGLFTIRGSDQPLIV 286
Query: 301 SFAN 304
FA+
Sbjct: 287 RFAD 290
>At3g55340 putative protein
Length = 597
Score = 57.0 bits (136), Expect = 4e-08
Identities = 42/162 (25%), Positives = 77/162 (46%), Gaps = 12/162 (7%)
Query: 45 DDDYDERGIEQDEGQEAGD-----EVEEEPEENVDEEEGDTGDEEVEEVEYVYEEVEGDD 99
DD+ E + E D + + + VDE EGD G +E ++ + + +
Sbjct: 63 DDEIRENEVGNGGSSEKTDTKIKKKRKRDDAVEVDELEGDEGTKEEQKPQKKKNKKKKKK 122
Query: 100 DDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQKEL---EVFVGGLDKEATEHDLR 156
V + A E NV E + + E++EV + +++ + +++VGG+ ++TE ++R
Sbjct: 123 RKVNKTPKKAEEGNVEEKV----KVEEIEVNTDNKEEDGVVPNKLYVGGIPYQSTEDEIR 178
Query: 157 KVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALA 198
F G I +V + P+ G A + F+T + KRALA
Sbjct: 179 SYFRSCGVIIKVDCKMRPEDGAFSGIAFITFDTEDGAKRALA 220
Score = 37.7 bits (86), Expect = 0.023
Identities = 28/77 (36%), Positives = 40/77 (51%), Gaps = 6/77 (7%)
Query: 141 VFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAEL 200
V++G L + TE D+RK+FS I VR+ N +T KG+A + F+ V AL +L
Sbjct: 264 VYIGNLAWDTTERDIRKLFSDC-VINSVRLGKNKETGEFKGYAHVDFKDSVSVAIAL-KL 321
Query: 201 KNPVINGKQCGITITTC 217
VI CG + C
Sbjct: 322 DQQVI----CGRPVKIC 334
>At2g15200 putative protein on transposon FARE2.7 (cds2)
Length = 650
Score = 54.7 bits (130), Expect = 2e-07
Identities = 46/139 (33%), Positives = 69/139 (49%), Gaps = 16/139 (11%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHE-YDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEE 71
EE KE ++ E++E E+ E + EY GD+ G E+ E + GDE E EE
Sbjct: 419 EEQKEEDEKKEQEEEKQEEEGKEEELEKVEYRGDE-----GKEKQEIPKQGDEEMEGEEE 473
Query: 72 NVDEEEGDTGDEEVEEVEYVYEE-------VEGDDDDVAGDDEHASEENVHEHLADVKEE 124
EEEG +EE E+VEY +E + D+++ G++E EE E V +E
Sbjct: 474 K-QEEEGK--EEEEEKVEYRGDEGKEKQEIPKQGDEEMEGEEEKQEEEGKEEEEEKVLKE 530
Query: 125 EQVEVIKETQKQKELEVFV 143
E VE E + ++ E +V
Sbjct: 531 ENVEEHDEHDETEDQEAYV 549
Score = 47.0 bits (110), Expect = 4e-05
Identities = 43/151 (28%), Positives = 72/151 (47%), Gaps = 7/151 (4%)
Query: 10 DNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEG--QEAGDEVEE 67
+ G+E + EY DE + ++ + D E ++ +E G E++E + GDE +E
Sbjct: 437 EEGKEEELEKVEYRGDEGKEKQEIPKQGDEEMEGEEEKQEEEGKEEEEEKVEYRGDEGKE 496
Query: 68 EPEENVDEEEGDTGDEEVEEVEYVYEEVEG--DDDDVAGDDEHASEENVHEH--LADVKE 123
+ E +E G+EE +E E EE E +++V DEH E+ + L+D ++
Sbjct: 497 KQEIPKQGDEEMEGEEEKQEEEGKEEEEEKVLKEENVEEHDEHDETEDQEAYVILSDDED 556
Query: 124 EEQVEVIKETQKQKELEVFVGGLDKEATEHD 154
KE+Q QKE E ++ EHD
Sbjct: 557 NGTTPTEKESQPQKE-ETTEVPKEENVEEHD 586
Score = 46.6 bits (109), Expect = 5e-05
Identities = 37/144 (25%), Positives = 64/144 (43%), Gaps = 13/144 (9%)
Query: 8 IADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEE 67
I G+E E +E +++E + E+ EY +E + ++G E+ EG+E E+
Sbjct: 460 IPKQGDEEMEGEEEKQEEEGKEEEEEKVEYRGDEGKEKQEIPKQGDEEMEGEE-----EK 514
Query: 68 EPEENVDEEEGDTGDEEVEEVEYVYEEVE--------GDDDDVAGDDEHASEENVHEHLA 119
+ EE +EEE EE E ++E E DD+D + E
Sbjct: 515 QEEEGKEEEEEKVLKEENVEEHDEHDETEDQEAYVILSDDEDNGTTPTEKESQPQKEETT 574
Query: 120 DVKEEEQVEVIKETQKQKELEVFV 143
+V +EE VE + + ++ E +V
Sbjct: 575 EVPKEENVEEHDKHDETEDQEAYV 598
Score = 44.7 bits (104), Expect = 2e-04
Identities = 67/260 (25%), Positives = 107/260 (40%), Gaps = 38/260 (14%)
Query: 5 IYLIADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDE 64
+ L+ +N E +K+ I E +L E Y EE E+DE +
Sbjct: 386 VKLVYENCEVLKKKIGRLEG--YLGIERVPASYTREEQK----------EEDEKK----- 428
Query: 65 VEEEPEENVDEEEGDTGDEEVEEVEYVYEE-VEGDDDDVAGDDEHASEENVHEHLADVKE 123
E EE EEEG +EE+E+VEY +E E + GD+E EE E +E
Sbjct: 429 ---EQEEEKQEEEGK--EEELEKVEYRGDEGKEKQEIPKQGDEEMEGEEEKQEEEGKEEE 483
Query: 124 EEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFA 183
EE+VE + K+K+ E+ G ++ E + ++ K E +V N +
Sbjct: 484 EEKVEYRGDEGKEKQ-EIPKQGDEEMEGEEEKQEEEGKEEEEEKVLKEENVE-------- 534
Query: 184 LLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKH 243
E EH + E + + + G T T + + + K+E ++E KH
Sbjct: 535 ----EHDEHDETEDQEAYVILSDDEDNGTTPT--EKESQPQKEETTEVPKEENVEEHDKH 588
Query: 244 YGVESFKDLTLLEDDNNEGT 263
E + +L DD + GT
Sbjct: 589 DETEDQEAYVILSDDEDNGT 608
Score = 42.7 bits (99), Expect = 7e-04
Identities = 43/131 (32%), Positives = 63/131 (47%), Gaps = 18/131 (13%)
Query: 13 EEVKESIDEYE-KDEHLDFEDNYHEY-DPEEYVGDDDYDERGIE--QDEGQEAGDEVEEE 68
EE KE +E K+E+++ D + E D E YV D ++ G + E Q +E E
Sbjct: 517 EEGKEEEEEKVLKEENVEEHDEHDETEDQEAYVILSDDEDNGTTPTEKESQPQKEETTEV 576
Query: 69 P-EENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQV 127
P EENV EE D DE ++ YV + D+D+ E S+ ++EE
Sbjct: 577 PKEENV--EEHDKHDETEDQEAYVI--LSDDEDNGTAPTEKESQR---------QKEETT 623
Query: 128 EVIKETQKQKE 138
EV +ET+K E
Sbjct: 624 EVPRETKKDDE 634
>At3g47270 putative protein
Length = 671
Score = 54.3 bits (129), Expect = 2e-07
Identities = 62/262 (23%), Positives = 109/262 (40%), Gaps = 52/262 (19%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHE-YDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEE 71
EE KE ++ E++E E+ E + EY GD+ +++ I + +E E E++ EE
Sbjct: 293 EEQKEEDEKKEQEEEKQEEEGKEEELEKVEYRGDERTEKQEIPKQGDEEMEGEEEKQKEE 352
Query: 72 NVDEEE------GDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENV----HEHLADV 121
+EEE GD G E+ E + EE+EG+++ + + EE V H +V
Sbjct: 353 GKEEEEEKVEYRGDEGTEKQEIPKQGDEEMEGEEEKQEEEGKEEEEEKVEYRDHHSTCNV 412
Query: 122 KEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKG 181
+E E+ E K+ ++ E E G ++ EHD + +T+ K
Sbjct: 413 EETEKQENPKQGDEEMEREE---GKEENVEEHD-----------------EHDETEDQKA 452
Query: 182 FALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKL 241
+ +L + NG + Q +T + K+E ++E
Sbjct: 453 YVIL---------------SDDEDNGTAPTEKESQPQKEETTEVP------KEENVEEHD 491
Query: 242 KHYGVESFKDLTLLEDDNNEGT 263
+H E + +L DD + GT
Sbjct: 492 EHDETEDQEAYVILLDDEDNGT 513
Score = 53.5 bits (127), Expect = 4e-07
Identities = 56/197 (28%), Positives = 84/197 (42%), Gaps = 36/197 (18%)
Query: 8 IADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVE- 66
I G+E E +E +K+E + E+ EY +E + ++G E+ EG+E E E
Sbjct: 334 IPKQGDEEMEGEEEKQKEEGKEEEEEKVEYRGDEGTEKQEIPKQGDEEMEGEEEKQEEEG 393
Query: 67 -EEPEENVD---------------EEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHA- 109
EE EE V+ +E GDEE+E E E VE D+ +D+ A
Sbjct: 394 KEEEEEKVEYRDHHSTCNVEETEKQENPKQGDEEMEREEGKEENVEEHDEHDETEDQKAY 453
Query: 110 -----SEEN-----------VHEHLADVKEEEQVEVIKETQKQKELEVFVGGLDKE--AT 151
E+N E +V +EE VE E + ++ E +V LD E T
Sbjct: 454 VILSDDEDNGTAPTEKESQPQKEETTEVPKEENVEEHDEHDETEDQEAYVILLDDEDNGT 513
Query: 152 EHDLRKVFSKVGEITEV 168
++ + EITEV
Sbjct: 514 APTEKESQPQKEEITEV 530
Score = 49.3 bits (116), Expect = 8e-06
Identities = 56/185 (30%), Positives = 77/185 (41%), Gaps = 44/185 (23%)
Query: 10 DNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQ---------DEGQE 60
+ G+E + EY DE + ++ + D E ++ E G E+ DEG E
Sbjct: 311 EEGKEEELEKVEYRGDERTEKQEIPKQGDEEMEGEEEKQKEEGKEEEEEKVEYRGDEGTE 370
Query: 61 ------AGDEVEEEPEENVDEEEGDTGDEEVEEVEY-------VYEEVEGDDDDVAGDD- 106
GDE E E EE EEEG +EE E+VEY EE E ++ GD+
Sbjct: 371 KQEIPKQGDE-EMEGEEEKQEEEGK--EEEEEKVEYRDHHSTCNVEETEKQENPKQGDEE 427
Query: 107 ---EHASEENVHEH--------------LADVKEEEQVEVIKETQKQKELEVFVGGLDKE 149
E EENV EH L+D ++ KE+Q QKE E ++
Sbjct: 428 MEREEGKEENVEEHDEHDETEDQKAYVILSDDEDNGTAPTEKESQPQKE-ETTEVPKEEN 486
Query: 150 ATEHD 154
EHD
Sbjct: 487 VEEHD 491
Score = 39.3 bits (90), Expect = 0.008
Identities = 40/134 (29%), Positives = 65/134 (47%), Gaps = 18/134 (13%)
Query: 10 DNGEEVKESIDEYE-KDEHLDFEDNYHEY-DPEEYV--GDDDYDERGIEQDEGQEAGDEV 65
+N ++ E ++ E K+E+++ D + E D + YV DD+ + + E Q +E
Sbjct: 419 ENPKQGDEEMEREEGKEENVEEHDEHDETEDQKAYVILSDDEDNGTAPTEKESQPQKEET 478
Query: 66 EEEP-EENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEE 124
E P EENV EE D DE ++ YV + D+D+ E S+ ++E
Sbjct: 479 TEVPKEENV--EEHDEHDETEDQEAYVI--LLDDEDNGTAPTEKESQP---------QKE 525
Query: 125 EQVEVIKETQKQKE 138
E EV +ET+K E
Sbjct: 526 EITEVPRETKKDDE 539
>At3g20930 unknown protein
Length = 374
Score = 54.3 bits (129), Expect = 2e-07
Identities = 32/114 (28%), Positives = 59/114 (51%)
Query: 94 EVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEH 153
E+ G +A +++ E ++ D ++ + + E+ K ++F+ GL +E
Sbjct: 237 ELAGVPGVLAVVPDNSFESLNKDYEGDSTQDSRDQDDSESPPVKTKKLFITGLSFYTSEK 296
Query: 154 DLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAELKNPVING 207
LR F GE+ EV++ ++ +KR+KG+A L + T E AL E+ +ING
Sbjct: 297 TLRAAFEGFGELVEVKIIMDKISKRSKGYAFLEYTTEEAAGTALKEMNGKIING 350
>At2g02200 putative protein on transposon FARE2.10 (cds2)
Length = 586
Score = 53.9 bits (128), Expect = 3e-07
Identities = 47/135 (34%), Positives = 68/135 (49%), Gaps = 18/135 (13%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHE-YDPEEYVGDDDYDERGIEQDEGQEAGDEVEEEPEE 71
EE KE ++ E++E E+ E + EY GD+ G E+ E + GDE E EE
Sbjct: 204 EEQKEEDEKKEQEEEKQEEEGKEEKLEKIEYRGDE-----GTEKQEIPKQGDEEMEGEEE 258
Query: 72 NVDEEEGDTGDEEVEEVEYVYEE-VEGDDDDVAGDDEHASEENVHEHLADVKEEEQVEV- 129
EEEG +EE E+VEY +E E + GD+E EE + +EEE+VE
Sbjct: 259 K-QEEEGK--EEEEEKVEYRGDEGTEKQEIPKQGDEEMEGEEEKQKEEGKEEEEEKVEYR 315
Query: 130 -------IKETQKQK 137
++ET+KQ+
Sbjct: 316 DHHSTCNVEETEKQE 330
Score = 50.1 bits (118), Expect = 4e-06
Identities = 52/185 (28%), Positives = 83/185 (44%), Gaps = 24/185 (12%)
Query: 10 DNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQ-----EAGDE 64
+ G+E K EY DE + ++ + D E ++ +E G E++E + + G E
Sbjct: 222 EEGKEEKLEKIEYRGDEGTEKQEIPKQGDEEMEGEEEKQEEEGKEEEEEKVEYRGDEGTE 281
Query: 65 VEEEPEENVDEEEGDT-------GDEEVEEVEY-------VYEEVEGDDDDVAGDDEHAS 110
+E P++ +E EG+ +EE E+VEY EE E ++ GD+E
Sbjct: 282 KQEIPKQGDEEMEGEEEKQKEEGKEEEEEKVEYRDHHSTCNVEETEKQENPKQGDEEMER 341
Query: 111 EENVHEHLADVKEEEQVEVIKETQKQKELEVFVGGLDKE--ATEHDLRKVFSKVGEITEV 168
EE E V +EE VE E + ++ E +V D E T ++ S+ E TEV
Sbjct: 342 EEGKEE---KVLKEENVEEHDEHDETEDQEAYVILSDDEDNGTAPTEKESESQKEETTEV 398
Query: 169 RMTVN 173
N
Sbjct: 399 AKEEN 403
Score = 46.6 bits (109), Expect = 5e-05
Identities = 39/144 (27%), Positives = 63/144 (43%), Gaps = 16/144 (11%)
Query: 8 IADNGEEVKESIDEYEKDEHLDFEDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEE 67
I G+E E +E +K+E + E+ EY D + +E+ E QE + +E
Sbjct: 285 IPKQGDEEMEGEEEKQKEEGKEEEEEKVEYR-------DHHSTCNVEETEKQENPKQGDE 337
Query: 68 EPEENVDEEEGDTGDEEVEEVEYVYEEVE--------GDDDDVAGDDEHASEENVHEHLA 119
E E +EE +E VEE + ++E E DD+D E+ E
Sbjct: 338 EMEREEGKEEKVLKEENVEEHD-EHDETEDQEAYVILSDDEDNGTAPTEKESESQKEETT 396
Query: 120 DVKEEEQVEVIKETQKQKELEVFV 143
+V +EE VE E + ++ E +V
Sbjct: 397 EVAKEENVEEHDEHDETEDQEAYV 420
Score = 45.1 bits (105), Expect = 1e-04
Identities = 66/257 (25%), Positives = 109/257 (41%), Gaps = 36/257 (14%)
Query: 57 EGQEAGDEVEEEPEENVDEEEGDTGDEEVEEVEYVYEE-VEGDDDDVAGDDEHASEENVH 115
E Q+ DE +E+ EE EEEG +E++E++EY +E E + GD+E EE
Sbjct: 204 EEQKEEDEKKEQEEEK-QEEEGK--EEKLEKIEYRGDEGTEKQEIPKQGDEEMEGEEEKQ 260
Query: 116 EHLADVKEEEQVEVIKETQKQKELEVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQ 175
E +EEE+VE + +K+ E+ G ++ E + +K K E +V +
Sbjct: 261 EEEGKEEEEEKVEYRGDEGTEKQ-EIPKQGDEEMEGEEEKQKEEGKEEEEEKVEYRDHHS 319
Query: 176 TKRNKGFALLRFETVEHVKRALAELKNPVINGKQCGITITTCQDSDTLYLDNICKSWKKE 235
T + E E+ K+ E++ GK+ K K+E
Sbjct: 320 TCN-----VEETEKQENPKQGDEEMERE--EGKE-------------------EKVLKEE 353
Query: 236 ALKEKLKHYGVESFKDLTLLEDDNNEGTNCGCAFLEFSSHSDSKDAYKRLQKTDVVFGVD 295
++E +H E + +L DD + GT A E S S K+ + K + V D
Sbjct: 354 NVEEHDEHDETEDQEAYVILSDDEDNGT----APTEKESES-QKEETTEVAKEENVEEHD 408
Query: 296 KPAEVSFANSFIDLGDD 312
+ E +++ L DD
Sbjct: 409 EHDETEDQEAYVILSDD 425
Score = 39.7 bits (91), Expect = 0.006
Identities = 37/148 (25%), Positives = 68/148 (45%), Gaps = 21/148 (14%)
Query: 12 GEEVKESID-EYEKDEHLDFEDNYHEYDPEEYV-------GDDDYD-----ERGIEQDEG 58
GEE K+ + + E++E +++ D++ + EE GD++ + E + ++E
Sbjct: 295 GEEEKQKEEGKEEEEEKVEYRDHHSTCNVEETEKQENPKQGDEEMEREEGKEEKVLKEEN 354
Query: 59 QEAGDEVEEEPEENV-----DEEEGDTGDEEVEEVEYVYEEVE-GDDDDVAGDDEHASEE 112
E DE +E ++ D+E+ T E E E E +++V DEH E
Sbjct: 355 VEEHDEHDETEDQEAYVILSDDEDNGTAPTEKESESQKEETTEVAKEENVEEHDEHDETE 414
Query: 113 NVHEH--LADVKEEEQVEVIKETQKQKE 138
+ + L+D ++ KE+Q QKE
Sbjct: 415 DQEAYVILSDDEDNGTAPTEKESQPQKE 442
Score = 38.9 bits (89), Expect = 0.010
Identities = 39/130 (30%), Positives = 61/130 (46%), Gaps = 19/130 (14%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEY-DPEEYV---GDDDYDERGIEQDEGQEAGDEVEEE 68
EE KE ++ K+E+++ D + E D E YV D+D E++ + + E
Sbjct: 342 EEGKE--EKVLKEENVEEHDEHDETEDQEAYVILSDDEDNGTAPTEKESESQKEETTEVA 399
Query: 69 PEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENVHEHLADVKEEEQVE 128
EENV EE D DE ++ YV + D+D+ E S+ ++EE E
Sbjct: 400 KEENV--EEHDEHDETEDQEAYVI--LSDDEDNGTAPTEKESQP---------QKEEIKE 446
Query: 129 VIKETQKQKE 138
V +ET+K E
Sbjct: 447 VPRETKKDDE 456
Score = 34.3 bits (77), Expect = 0.25
Identities = 24/84 (28%), Positives = 40/84 (47%), Gaps = 9/84 (10%)
Query: 13 EEVKESIDEYEKDEHLDFEDNYHEY-DPEEYV--GDDDYDERGIEQDEGQEAGDEVEEEP 69
E KE E K+E+++ D + E D E YV DD+ + + E Q +E++E P
Sbjct: 389 ESQKEETTEVAKEENVEEHDEHDETEDQEAYVILSDDEDNGTAPTEKESQPQKEEIKEVP 448
Query: 70 EENVDEEEG------DTGDEEVEE 87
E ++E T +EE+ +
Sbjct: 449 RETKKDDEDVNQTPLSTQEEEITQ 472
Score = 33.1 bits (74), Expect = 0.56
Identities = 26/107 (24%), Positives = 44/107 (40%), Gaps = 16/107 (14%)
Query: 10 DNGEEVKESIDEYEKDEHLDF--EDNYHEYDPEEYVGDDDYDERGIEQDEGQEAGDEVEE 67
DNG E E +K+E + E+N E+D + D + + ++ A E E
Sbjct: 378 DNGTAPTEKESESQKEETTEVAKEENVEEHDEHDETEDQEAYVILSDDEDNGTAPTEKES 437
Query: 68 EPEENVDEEEGDTGDEEVEEVEYVYEEVEGDDDDVAGDDEHASEENV 114
+P++ EE++ V E + DD+DV EE +
Sbjct: 438 QPQK--------------EEIKEVPRETKKDDEDVNQTPLSTQEEEI 470
>At5g47320 40S ribosomal protein S19
Length = 212
Score = 53.5 bits (127), Expect = 4e-07
Identities = 29/120 (24%), Positives = 55/120 (45%)
Query: 140 EVFVGGLDKEATEHDLRKVFSKVGEITEVRMTVNPQTKRNKGFALLRFETVEHVKRALAE 199
++++GGL EH L+ FS +TE R+ N T R++G+ + F + + A++
Sbjct: 32 KLYIGGLSPGTDEHSLKDAFSSFNGVTEARVMTNKVTGRSRGYGFVNFISEDSANSAISA 91
Query: 200 LKNPVINGKQCGITITTCQDSDTLYLDNICKSWKKEALKEKLKHYGVESFKDLTLLEDDN 259
+ +NG + + S L LD + +K+ K + + F D L++ N
Sbjct: 92 MNGQELNGFNISVNVAKDWPSLPLSLDESIEEAEKKENKMMSRSVWKDPFVDAFLMKKKN 151
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,466,153
Number of Sequences: 26719
Number of extensions: 975571
Number of successful extensions: 11650
Number of sequences better than 10.0: 850
Number of HSP's better than 10.0 without gapping: 370
Number of HSP's successfully gapped in prelim test: 505
Number of HSP's that attempted gapping in prelim test: 5082
Number of HSP's gapped (non-prelim): 3507
length of query: 726
length of database: 11,318,596
effective HSP length: 107
effective length of query: 619
effective length of database: 8,459,663
effective search space: 5236531397
effective search space used: 5236531397
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC149129.7


