

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149127.6 - phase: 0
(421 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
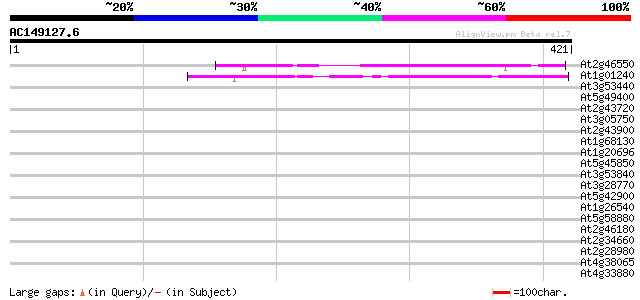
Score E
Sequences producing significant alignments: (bits) Value
At2g46550 unknown protein 168 6e-42
At1g01240 unknown protein 147 1e-35
At3g53440 unknown protein 39 0.004
At5g49400 putative protein 37 0.027
At2g43720 putative protein 33 0.30
At3g05750 hypothetical protein 32 0.50
At2g43900 putative inositol polyphosphate 5'-phosphatase 32 0.66
At1g68130 putative C2H2-type zinc finger protein 32 0.66
At1g20696 unknown protein 32 0.86
At5g45850 putative protein 31 1.1
At3g53840 protein kinase-like protein 31 1.1
At3g28770 hypothetical protein 31 1.1
At5g42900 unknown protein 31 1.5
At1g26540 unknown protein 31 1.5
At5g58880 putative protein 30 2.5
At2g46180 unknown protein 30 2.5
At2g34660 MRP-like ABC transporter 30 2.5
At2g28980 putative non-LTR retroelement reverse transcriptase 30 2.5
At4g38065 hypothetical protein 29 4.3
At4g33880 putative bHLH transcription factor (bHLH085) 29 4.3
>At2g46550 unknown protein
Length = 397
Score = 168 bits (425), Expect = 6e-42
Identities = 109/281 (38%), Positives = 149/281 (52%), Gaps = 55/281 (19%)
Query: 155 ATSFLISEHCKSTSSDFEPH--W--LGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDL 210
+T + S K + F+P W L +EK+ PWWRTT KD+LASLVA++S +YVENCDL
Sbjct: 149 STGYDESSEKKLSELSFDPSSPWNLLSSEKAAPWWRTTDKDELASLVAQRSLDYVENCDL 208
Query: 211 PEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRT 270
P P + ++ + PR D + +D +
Sbjct: 209 PTP--QKMKRSYYGSPRGFDSD------------------------------GLRDYSVS 236
Query: 271 FSSSESKDSDSSCNKNSKVNSES-AAKAELLKALCRSQTRAREAEKAAQEACDEKEHILS 329
+ + SSC + +SES +K+ELL+AL RSQTRAREAE A+EA EKEH++
Sbjct: 237 GQTIKGTSKGSSCKNRPEASSESDLSKSELLEALRRSQTRAREAENMAKEAYAEKEHLVK 296
Query: 330 LFFKQASQLFAYKQWLHVLQLENLCLQYKSKNQPLLNNLFP-------------YEGKKH 376
+ KQA++LF YKQWL +LQLE L LQ K+K NN P EG+K
Sbjct: 297 ILLKQAAELFGYKQWLQLLQLEALYLQIKNKEIDNKNNDDPGVSIPCWSNGKARKEGRKR 356
Query: 377 RKNRRKVKNSRRGIGKCIFAFAVGLALAGAGLLLGWTIGCM 417
R R K ++ +G A+G++L GAGLLLGWT+G M
Sbjct: 357 RSKRGKPNGAKYAVG-----LALGMSLVGAGLLLGWTVGWM 392
>At1g01240 unknown protein
Length = 331
Score = 147 bits (371), Expect = 1e-35
Identities = 104/301 (34%), Positives = 157/301 (51%), Gaps = 41/301 (13%)
Query: 134 NSDLYLTPKKKNEGEFWFSDDAT-SFLISEHCKST-----------SSDFEPHWLGAEKS 181
+SD K +G F ++ T SFL CK D+ A+ +
Sbjct: 56 SSDFSCDTKWWLKGSTGFDEEVTNSFLEDTKCKKLHEFVDLIGIREEEDYSFISKKADAT 115
Query: 182 QPWWR-TTGKDDLASLVAKKSFEY-VENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSL 239
PWWR TT KD+LA +VA KS ++ ++NCDLP P K + IH+ +
Sbjct: 116 TPWWRSTTDKDELALMVATKSVDHNIQNCDLPPPQ-KLHKSIHSSSGEK----------- 163
Query: 240 NQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAEL 299
K + S G S+ S+ESK++ + S S+ +K +L
Sbjct: 164 GFKTAVKSPWKQGVWKDRFERSLSYN------GSTESKNT----SPMSSPRSDDLSKGQL 213
Query: 300 LKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKS 359
L+AL SQTRAREAE+AA+EAC EK+ ++++ KQASQ+ AYKQWL +L++E L LQ K
Sbjct: 214 LEALRHSQTRAREAERAAREACAEKDRVITILLKQASQMLAYKQWLKLLEMEALYLQMKK 273
Query: 360 KNQPLLNNLFPYEGKKHRKNRRKVKNSRRG-IGKCIFAFAVGLALAGAGLLLGWTIGCMF 418
+ + +G +K +++ + ++G G+ + AFA+G +L GAGLLLGWT+G +
Sbjct: 274 EEE----QEEQVKGMNLKKRKQRGEKKKKGETGRYMMAFALGFSLIGAGLLLGWTVGWLL 329
Query: 419 P 419
P
Sbjct: 330 P 330
>At3g53440 unknown protein
Length = 512
Score = 39.3 bits (90), Expect = 0.004
Identities = 37/127 (29%), Positives = 61/127 (47%), Gaps = 23/127 (18%)
Query: 213 PNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFS 272
P+++ ++ + + E +KE+ + +S +S S +STT++S C+ + S R FS
Sbjct: 82 PSLRRSLRLSSRECPEVNKEKPKTAS--------TSSSRTKSSTTVSSSCALRRSPR-FS 132
Query: 273 S--SESKD-SDSSCNKNS-----------KVNSESAAKAELLKALCRSQTRAREAEKAAQ 318
S ES D S SS K S ++SE + LK+ RS++R R K
Sbjct: 133 SGGGESVDQSSSSIGKKSGNSVLSSCSTKSISSEKVKGGDGLKSKDRSRSRVRPKTKKMF 192
Query: 319 EACDEKE 325
CD+ E
Sbjct: 193 SGCDDNE 199
>At5g49400 putative protein
Length = 280
Score = 36.6 bits (83), Expect = 0.027
Identities = 30/130 (23%), Positives = 48/130 (36%), Gaps = 16/130 (12%)
Query: 210 LPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEM----------------CSSDSNGC 253
L P ++ + L + D EE + + N K E+ SDS
Sbjct: 134 LKNPKLRTKPSVDDLDGSDDDDEEERPDATNGKAEVEKRSKKSKRKHRSKSDSESDSEAS 193
Query: 254 TSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREA 313
T + G S + S SSS+S+D K K + + E + S + + E+
Sbjct: 194 VFETDSDGSSGESSSEYSSSSDSEDERRRRRKAKKSKKKQKQRKERRRRYSSSSSESSES 253
Query: 314 EKAAQEACDE 323
E A+ DE
Sbjct: 254 ESASDSDSDE 263
>At2g43720 putative protein
Length = 147
Score = 33.1 bits (74), Expect = 0.30
Identities = 26/107 (24%), Positives = 47/107 (43%), Gaps = 8/107 (7%)
Query: 231 KEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSC-----NK 285
K+ N+VSS +Q L D T C+++ DRT + +E SC N
Sbjct: 16 KKVNEVSSASQSLLSPVQDHINFTLQKAYFKCAYECFDRTRTHAEISRCAESCSVPITNA 75
Query: 286 NSKVNSESAAKAELLK---ALCRSQTRAREAEKAAQEACDEKEHILS 329
+ ++E + E L +C+ + + +K EA ++ EH ++
Sbjct: 76 QNYFDNEMSVFQERLNRSLVVCQDKFEVAKQQKTRSEAVNDLEHCVN 122
>At3g05750 hypothetical protein
Length = 798
Score = 32.3 bits (72), Expect = 0.50
Identities = 48/184 (26%), Positives = 71/184 (38%), Gaps = 29/184 (15%)
Query: 121 NSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSD-DATSFLISEHCKSTSSDFEPHWLGAE 179
NSD R K E + ++ + K + D D SF S K SSD
Sbjct: 439 NSDKRIKKGEKVIKCNITVDGGLKTGDDDRKKDMDVISFTFSSPIKGLSSD--------- 489
Query: 180 KSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSL 239
SQ + + +D ++L K D N +K+ L T K E+ SSL
Sbjct: 490 -SQYFLKKNDQDAESALCFNK-------IDSDSLNFLLEKKLREL----TSKMESSCSSL 537
Query: 240 NQKLEMCSSDSNGCTSTTLT-------SGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSE 292
Q+ E S + + T + +G S +SD +SSS K + +VNS
Sbjct: 538 TQEEESSGSITKDWVNGTRSLPSDDQDNGLSESESDSDYSSSFYKKKIFQAEDDEEVNSF 597
Query: 293 SAAK 296
S A+
Sbjct: 598 STAE 601
>At2g43900 putative inositol polyphosphate 5'-phosphatase
Length = 1305
Score = 32.0 bits (71), Expect = 0.66
Identities = 35/155 (22%), Positives = 60/155 (38%), Gaps = 12/155 (7%)
Query: 143 KKNEGEFWFSDDATSFLISEHCKSTSSDFEPHWLGAEKSQPWWRTTGKDDLASLVAKKSF 202
KKNEG+ S S S SD + + ++KS D S +KKS
Sbjct: 1114 KKNEGDSNSKSSKKSDGDSNSKSSKKSDGDSNSKSSKKSD--------GDSNSKSSKKS- 1164
Query: 203 EYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGC 262
+ + + K ++ +++D + N SS + CS T +
Sbjct: 1165 ---DGDSNSKSSKKSDGDSNSKSSKKSDGDSNSKSSKKSDGDSCSKSQKKSDGDTNSKSQ 1221
Query: 263 SFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKA 297
D D + S + D DSS + K + +S++K+
Sbjct: 1222 KKGDGDSSSKSHKKNDGDSSSKSHKKNDGDSSSKS 1256
Score = 29.6 bits (65), Expect = 3.3
Identities = 16/72 (22%), Positives = 31/72 (42%)
Query: 227 RETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKN 286
+++D + N SS + S S + + D D SS+ D DS+ +
Sbjct: 1138 KKSDGDSNSKSSKKSDGDSNSKSSKKSDGDSNSKSSKKSDGDSNSKSSKKSDGDSNSKSS 1197
Query: 287 SKVNSESAAKAE 298
K + +S +K++
Sbjct: 1198 KKSDGDSCSKSQ 1209
>At1g68130 putative C2H2-type zinc finger protein
Length = 419
Score = 32.0 bits (71), Expect = 0.66
Identities = 40/164 (24%), Positives = 64/164 (38%), Gaps = 11/164 (6%)
Query: 180 KSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSL 239
+SQP + + A ++ EN + + +I P H LQ R++ E Q S+L
Sbjct: 200 RSQPSNHRLHEQQQHTTNATQTASTAENNENGDLSIGPILPGHPLQRRQSPPSEQQPSTL 259
Query: 240 ------NQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSES 293
N +E+ S C T S +T S E K S K
Sbjct: 260 LYPFVTNGSIELQLLPSRNCADETSLSLSIGTMDQKTMSEVEKK----SYEKGETSLERE 315
Query: 294 AAKAELLKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQ 337
A+ E + + ++ EA++ Q A E H LF ++AS+
Sbjct: 316 EARRETKRQIEIAELEFAEAKRIRQHARAEL-HKAHLFREEASR 358
>At1g20696 unknown protein
Length = 141
Score = 31.6 bits (70), Expect = 0.86
Identities = 28/102 (27%), Positives = 43/102 (41%), Gaps = 9/102 (8%)
Query: 141 PKKKNEGEFWFSDDATSFLISEHCKSTSSDFEPHWLGAEKSQPWWRTTGKDDLASLVAKK 200
PK+ + F F +D EH K+ S G EK W++ + A VAK
Sbjct: 35 PKRPSSAFFVFMEDFRVTYKEEHPKNKSVAAVGK-AGGEK----WKSLSDSEKAPYVAKA 89
Query: 201 SFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQK 242
VE E N+K + K P+E ++ + VS +N +
Sbjct: 90 DKRKVEY----EKNMKAYNKKLEEGPKEDEESDKSVSEVNDE 127
>At5g45850 putative protein
Length = 444
Score = 31.2 bits (69), Expect = 1.1
Identities = 35/136 (25%), Positives = 62/136 (44%), Gaps = 11/136 (8%)
Query: 188 TGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCS 247
T + L L+ +SFE+V EP+ PF+ T + K + S L + +EM S
Sbjct: 50 TENEILIRLLQDQSFEHVM-----EPSSVPFKWEQTPGKPKDKKNLVEESDLIKAMEMVS 104
Query: 248 SDSN---GCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELLKALC 304
S ++ C+S+ ++ + D DR SS+ S+D D + S AE +
Sbjct: 105 STASFSVNCSSSGVSEFENNGDGDR--SSNVSRD-DVILEYRDLIMSRFLPAAEAISLKM 161
Query: 305 RSQTRAREAEKAAQEA 320
+ + +AEK +++
Sbjct: 162 KKEASRVKAEKKKKQS 177
>At3g53840 protein kinase-like protein
Length = 640
Score = 31.2 bits (69), Expect = 1.1
Identities = 41/190 (21%), Positives = 73/190 (37%), Gaps = 17/190 (8%)
Query: 150 WFSDDATSFLISEHCKSTSSDFEPHWLGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCD 209
+FS +F +C ST + P G + P +R D+ SL F+ +
Sbjct: 18 YFSSTTQAFKRCPNCGSTRVPY-PLSTGLDCGDPGYRIRC-DNYGSLW----FDTLNGST 71
Query: 210 LPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSN-----GCTSTTLTSGCSF 264
P I P + L+P E+N+ S++ K D N C++T + C+
Sbjct: 72 NPIKTIDPSGQRFVLRP--PGFEQNKCVSVDIKYHGIQLDLNLPFNVSCSNTVIIMNCTK 129
Query: 265 QDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAR----EAEKAAQEA 320
D S + +S C+K N E+ + + C +T A + +A +
Sbjct: 130 DGLDAYSSQGFNCSDNSLCHKFLNANLEARGNCRGVTSCCWYKTGASVNTYKVYRARPDM 189
Query: 321 CDEKEHILSL 330
C + ++L
Sbjct: 190 CSAYQSFMNL 199
>At3g28770 hypothetical protein
Length = 2081
Score = 31.2 bits (69), Expect = 1.1
Identities = 26/99 (26%), Positives = 48/99 (48%), Gaps = 10/99 (10%)
Query: 231 KEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSK-- 288
KEE++ +N +L+ + TT + ++ ++ + E K+S+ S +KN +
Sbjct: 959 KEEDKKEYVNNELKKQEDNKK---ETTKSENSKLKEENK--DNKEKKESEDSASKNREKK 1013
Query: 289 --VNSESAAKAELLKALCRSQTRAREAEKAAQEACDEKE 325
+S K E K +SQ + RE EK ++E +KE
Sbjct: 1014 EYEEKKSKTKEEAKKEKKKSQDKKRE-EKDSEERKSKKE 1051
>At5g42900 unknown protein
Length = 246
Score = 30.8 bits (68), Expect = 1.5
Identities = 61/234 (26%), Positives = 94/234 (40%), Gaps = 39/234 (16%)
Query: 74 FSTRLHGDNVSIGSDQTVK-NFDAVSFVGNAATAAIEQQWN--VYPKYMKNSDTRTSKIE 130
FS+R D+ SD + S + +A +E +W + Y+K+ +E
Sbjct: 9 FSSRRFSDDSDDSSDDASSVEGETTSSMYSAGKEYMETEWTNEKHSLYLKS-------ME 61
Query: 131 ASLNSDLY--LTPKKKNEGEFWFSDDATSFLISEHCKSTSSDF----EPHWLGAEKSQPW 184
AS LY L KNE ++T F K + F + W QP
Sbjct: 62 ASFVDQLYNSLGALGKNENV----SESTRF--GSGRKPSQEQFKVLHDGFWQKINVKQPE 115
Query: 185 WRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQ-VSSLNQKL 243
R G+ S E++ + P IK ++ + Q TD+ ENQ VSS N K
Sbjct: 116 HRINGRH------GGNSHEFLRS-----PWIKHYKPLVKTQIPVTDEPENQVVSSSNGKK 164
Query: 244 EMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSS-CNKNSKVNSESAAK 296
+CSS G S+ +D D+ S E++ SD + N+ K + S+ K
Sbjct: 165 GICSS---GSASSLKQLSSHSRDHDQ-ISVGEAEVSDQNFVNEGIKGENGSSKK 214
>At1g26540 unknown protein
Length = 695
Score = 30.8 bits (68), Expect = 1.5
Identities = 24/76 (31%), Positives = 38/76 (49%), Gaps = 1/76 (1%)
Query: 227 RETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKN 286
R+ +E+NQ S+LN+ E C+ G T+ T D D+ SS + + S +++
Sbjct: 427 RKRKREQNQNSNLNETDETCNVSKAGVNGTSDTIRVDDVD-DQPLSSWINIPTVLSSDQS 485
Query: 287 SKVNSESAAKAELLKA 302
S V SAA E +A
Sbjct: 486 SNVVDNSAADVEETQA 501
>At5g58880 putative protein
Length = 1088
Score = 30.0 bits (66), Expect = 2.5
Identities = 29/100 (29%), Positives = 43/100 (43%), Gaps = 3/100 (3%)
Query: 207 NCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQD 266
N + +I RK L+ D E + LN K D S + +
Sbjct: 352 NSPVSASDISTTRKRLDLENEYIDHTEQNLP-LNGKEATIEDDDKSVVSRK-SEEKEVEM 409
Query: 267 SDRTFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRS 306
+D T S+ E D DSSC++ S+ KAEL +A+C+S
Sbjct: 410 NDETDSNKEECD-DSSCSEESESELCRLNKAELREAICQS 448
>At2g46180 unknown protein
Length = 725
Score = 30.0 bits (66), Expect = 2.5
Identities = 28/99 (28%), Positives = 42/99 (42%), Gaps = 8/99 (8%)
Query: 230 DKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKV 289
+KE N++ +LE S TS L S +D R SS + + + + K
Sbjct: 292 EKENNELKLKRSELEAALEASQKSTSRKLFPK-STEDLSRHLSSLDEEKAGTFPGKEDME 350
Query: 290 NSESAAKAELLKALCRSQTRAREAEKAAQEACDEKEHIL 328
S + EL +A RE +KA QE K+H+L
Sbjct: 351 KSLQRLEKELEEA-------RREKDKARQELKRLKQHLL 382
>At2g34660 MRP-like ABC transporter
Length = 1623
Score = 30.0 bits (66), Expect = 2.5
Identities = 35/146 (23%), Positives = 61/146 (40%), Gaps = 8/146 (5%)
Query: 266 DSDRT--FSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREAE---KAAQEA 320
DS R FSS E+ S+ + + V S AA AE L++L RA++ + ++
Sbjct: 1450 DSGRVQEFSSPENLLSNEGSSFSKMVQSTGAANAEYLRSLVLDNKRAKDDSHHLQGQRKW 1509
Query: 321 CDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKSKNQPLLNNLFPYEGKKHRKNR 380
+ F A+ L + L L++E+ K N ++ EGK ++
Sbjct: 1510 LASSRWAAAAQFALAASLTSSHNDLQSLEIEDDSSILKRTNDAVVTLRSVLEGKHDKEIA 1569
Query: 381 RKVKN---SRRGIGKCIFAFAVGLAL 403
++ SR G ++ GLA+
Sbjct: 1570 ESLEEHNISREGWLSSLYRMVEGLAV 1595
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 30.0 bits (66), Expect = 2.5
Identities = 19/57 (33%), Positives = 29/57 (50%), Gaps = 4/57 (7%)
Query: 335 ASQLFAYKQWLHVLQLENLCLQYKSK----NQPLLNNLFPYEGKKHRKNRRKVKNSR 387
A +L AY W H+ +LE L+ KSK N NN + ++ + RK R ++ R
Sbjct: 634 AEELKAYTDWTHLSELEEGFLKQKSKLHWMNVGDGNNSYFHKAAQVRKMRNSIREIR 690
>At4g38065 hypothetical protein
Length = 1050
Score = 29.3 bits (64), Expect = 4.3
Identities = 21/79 (26%), Positives = 36/79 (44%), Gaps = 7/79 (8%)
Query: 233 ENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSE 292
E ++SSL+QKLE S++S C T S + +++ + + K+ +
Sbjct: 906 EGEISSLSQKLE-TSNESVSCFRQEATK------SRAELETKQTELKEVTTQMQEKLRTS 958
Query: 293 SAAKAELLKALCRSQTRAR 311
A K EL+K + T R
Sbjct: 959 EAEKTELVKEVASLSTEKR 977
>At4g33880 putative bHLH transcription factor (bHLH085)
Length = 352
Score = 29.3 bits (64), Expect = 4.3
Identities = 20/73 (27%), Positives = 30/73 (40%), Gaps = 2/73 (2%)
Query: 228 ETDKEENQVSSLNQKLEMCSSDSNGCTS--TTLTSGCSFQDSDRTFSSSESKDSDSSCNK 285
E ++EE + +K S N T+ T S C+ QD SSS+ D + N
Sbjct: 203 EGEEEEGETKLKKRKNGAMMSRQNSSTTFCTEEESNCADQDGGGEDSSSKEDDPSKALNL 262
Query: 286 NSKVNSESAAKAE 298
N K + A +
Sbjct: 263 NGKTRASRGAATD 275
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.127 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,518,874
Number of Sequences: 26719
Number of extensions: 406766
Number of successful extensions: 1658
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 38
Number of HSP's that attempted gapping in prelim test: 1618
Number of HSP's gapped (non-prelim): 69
length of query: 421
length of database: 11,318,596
effective HSP length: 102
effective length of query: 319
effective length of database: 8,593,258
effective search space: 2741249302
effective search space used: 2741249302
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149127.6


