

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149080.7 - phase: 0
(165 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
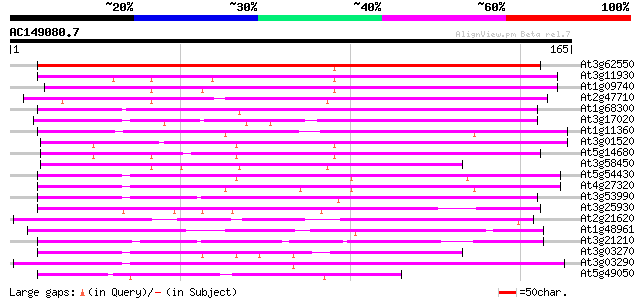
Score E
Sequences producing significant alignments: (bits) Value
At3g62550 unknown protein 158 1e-39
At3g11930 unknown protein 112 7e-26
At1g09740 putative ER6 protein 111 2e-25
At2g47710 unknown protein 104 2e-23
At1g68300 unknown protein 98 2e-21
At3g17020 unknown protein 81 3e-16
At1g11360 unknown protein 78 2e-15
At3g01520 unknown protein 75 2e-14
At5g14680 unknown protein 72 2e-13
At3g58450 putative protein 68 2e-12
At5g54430 unknown protein 66 1e-11
At4g27320 unknown protein 62 1e-10
At3g53990 unknown protein 62 2e-10
At3g25930 unknown protein 59 1e-09
At2g21620 RD2 protein (RD2) 54 3e-08
At1g48961 unknown protein 53 8e-08
At3g21210 unknown protein 52 1e-07
At3g03270 unknown protein 49 9e-07
At3g03290 hypothetical protein 45 2e-05
At5g49050 unknown protein 43 9e-05
>At3g62550 unknown protein
Length = 162
Score = 158 bits (399), Expect = 1e-39
Identities = 77/151 (50%), Positives = 107/151 (69%), Gaps = 3/151 (1%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDIT 68
R+I+VA+DE EES+ AL+W L NL S + LILLYVKPP VYS+ D G++ + D
Sbjct: 7 RKIVVAVDESEESMEALSWSLDNLFPYGSNNTLILLYVKPPLPVYSSLDAAGFIVTGDPV 66
Query: 69 ATMEKYSQQVADCVLEKAKIVCNDVQ---NVETRIENGDPRDVICQAVQKMGVDILVMGS 125
A ++KY ++ + V+ +++ V D + N+E R+ GD ++VIC AVQK+ VD+LVMG+
Sbjct: 67 AALKKYEYELVESVMARSRTVYQDYESDINIERRVGRGDAKEVICNAVQKLRVDMLVMGT 126
Query: 126 HGYGVIKRAFLGSVSNHCAQNVKCPVLIVKK 156
H YG KRA LGSVS +CA+ VKCPV+IVKK
Sbjct: 127 HDYGFFKRALLGSVSEYCAKRVKCPVVIVKK 157
>At3g11930 unknown protein
Length = 200
Score = 112 bits (281), Expect = 7e-26
Identities = 65/167 (38%), Positives = 100/167 (58%), Gaps = 14/167 (8%)
Query: 9 RRIMVAIDEGEESIYALTWCL---KNLVFQNSKDH-----LILLYVKPPRVVYSAFDG-- 58
+R++VAIDE + S YAL W + NL+ + L +++V+ P ++AF
Sbjct: 33 KRMVVAIDESDSSFYALQWVIDHFSNLLLTTAAAEAESGMLTVIHVQSPFNHFAAFPAGP 92
Query: 59 ---TGYLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQ 114
T SS + +++K Q+ + +L +A +C Q ET + G+ +++IC+AV+
Sbjct: 93 GGATAVYASSSMIESVKKAQQETSAALLSRALQMCRAKQIRTETLVLEGEAKEMICEAVE 152
Query: 115 KMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
KM VD+LV+GS G G IKRAFLGSVS++CA + CP+LIVK PK T
Sbjct: 153 KMHVDLLVVGSRGLGKIKRAFLGSVSDYCAHHANCPILIVKPPKEMT 199
>At1g09740 putative ER6 protein
Length = 171
Score = 111 bits (277), Expect = 2e-25
Identities = 61/161 (37%), Positives = 95/161 (58%), Gaps = 10/161 (6%)
Query: 11 IMVAIDEGEESIYALTWCLKNLVFQNSKDH--LILLYVKPPRVVYSA-------FDGTGY 61
++VA+D E S+ AL W L NL +S ++L+V+P V + F G
Sbjct: 10 VVVAVDGSEVSMEALRWALDNLKLSSSSSDSSFVVLHVQPSPSVAAGVSPGTIPFGGPSG 69
Query: 62 LFSSDITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKMGVDI 120
L TA +E++ +++ D +LE A +C + NV+T++ GDP+ IC+AV+ + D+
Sbjct: 70 LEVPAFTAAIEQHQKRITDTILEHASQICAEKSVNVKTQVVIGDPKYKICEAVENLHADL 129
Query: 121 LVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
LVMGS YG IKR FLGSVSN+C + CPV+I+K + ++
Sbjct: 130 LVMGSRAYGRIKRMFLGSVSNYCTNHAHCPVVIIKPKEDSS 170
>At2g47710 unknown protein
Length = 162
Score = 104 bits (260), Expect = 2e-23
Identities = 59/160 (36%), Positives = 90/160 (55%), Gaps = 9/160 (5%)
Query: 5 TENGRRIMVA-IDEGEESIYALTWCLKNLVFQNSKDH---LILLYVKPPRVVYSAFDGTG 60
T +G+ +MV +D+ E+S YAL W L + ++ L +++ KP V G G
Sbjct: 3 TGDGKSVMVVGVDDSEQSTYALEWTLDRFFAPYAPNYPFKLFIVHAKPNAVSAVGLAGPG 62
Query: 61 YLFSSDITATMEKYSQQVADCVLEKAKIVCND--VQNVETRIENGDPRDVICQAVQKMGV 118
++++ ++ + A V+EKAK +C V + GD R+++C+ V K
Sbjct: 63 ---TAEVVPYVDADLKHTAAKVVEKAKAICQSRSVHGAVIEVFEGDARNILCEVVDKHHA 119
Query: 119 DILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPK 158
ILV+GSHGYG IKRA LGS S++CA + C V+IVKKPK
Sbjct: 120 SILVVGSHGYGAIKRAVLGSTSDYCAHHAHCSVMIVKKPK 159
>At1g68300 unknown protein
Length = 160
Score = 97.8 bits (242), Expect = 2e-21
Identities = 57/149 (38%), Positives = 84/149 (56%), Gaps = 3/149 (2%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSD-- 66
+++MVAIDE E S AL W L L + D I+L+ P + S + Y +
Sbjct: 10 KQVMVAIDESECSKRALQWTLVYLK-DSLADSDIILFTAQPHLDLSCVYASSYGAAPIEL 68
Query: 67 ITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSH 126
I + E + + + E KI +E G+P++ IC+A +K+GVD+LV+GSH
Sbjct: 69 INSLQESHKNAGLNRLDEGTKICAETGVTPRKVLEFGNPKEAICEAAEKLGVDMLVVGSH 128
Query: 127 GYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
G G ++R FLGSVSN+C N KCPVL+V+
Sbjct: 129 GKGALQRTFLGSVSNYCVNNAKCPVLVVR 157
>At3g17020 unknown protein
Length = 163
Score = 80.9 bits (198), Expect = 3e-16
Identities = 58/161 (36%), Positives = 86/161 (53%), Gaps = 19/161 (11%)
Query: 8 GRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILL-------YVKPPRVVYSAFDGTG 60
GRRI VA+D + S AL+W + N+V DHLIL+ Y + ++ G+
Sbjct: 6 GRRIGVAVDFSDCSKKALSWAIDNVV--RDGDHLILITIAHDMNYEEGEMQLWETV-GSP 62
Query: 61 YLFSSDIT--ATMEKYS----QQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQ 114
++ S+ + A M+KY+ + D V A+ V +I GDPR+ IC A +
Sbjct: 63 FIPMSEFSDAAVMKKYALKPDAETLDIVNTAAR---KKTITVVMKIYWGDPREKICAAAE 119
Query: 115 KMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
++ + LVMG+ G G +KR +GSVSNH NV CPV +VK
Sbjct: 120 QIPLSSLVMGNRGLGGLKRMIMGSVSNHVVNNVACPVTVVK 160
>At1g11360 unknown protein
Length = 242
Score = 77.8 bits (190), Expect = 2e-15
Identities = 51/173 (29%), Positives = 87/173 (49%), Gaps = 25/173 (14%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYL------ 62
R+I +A+D +ES YA+ W ++N + S D ++LL+V+P V+Y A G L
Sbjct: 38 RKIGIAVDLSDESAYAVQWAVQN--YLRSGDAVVLLHVQPTSVLYGADWGAMDLSPQWDP 95
Query: 63 --------FSSDITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQ 114
D K + VA ++E D+ +++ D ++ +C V+
Sbjct: 96 NNEESQRKLEDDFDIVTNKKASDVAQPLVEA------DIPFKIHIVKDHDMKERLCLEVE 149
Query: 115 KMGVDILVMGSHGYGVIKRAF---LGSVSNHCAQNVKCPVLIVKKPKSTTGGD 164
++G+ L+MGS G+G KR+ LGSVS++ + CPV++V+ P G D
Sbjct: 150 RLGLSTLIMGSRGFGATKRSSKGRLGSVSDYSVHHCACPVVVVRFPDDKDGED 202
>At3g01520 unknown protein
Length = 175
Score = 74.7 bits (182), Expect = 2e-14
Identities = 50/166 (30%), Positives = 85/166 (51%), Gaps = 12/166 (7%)
Query: 10 RIMVAIDEGEESIY---------ALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTG 60
++MVA++ Y A W L+ +V N+ D ILL + V FD
Sbjct: 7 KVMVAVNASTIKDYPNPSISCKRAFEWTLEKIVRSNTSDFKILL-LHVQVVDEDGFDDVD 65
Query: 61 YLFSS-DITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKMGV 118
+++S + M + ++ +LE C+++ E I+ GDP+DVICQ V+++
Sbjct: 66 SIYASPEDFRDMRQSNKAKGLHLLEFFVNKCHEIGVGCEAWIKTGDPKDVICQEVKRVRP 125
Query: 119 DILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTTGGD 164
D LV+GS G G ++ F+G+VS C ++ +CPV+ +K+ T D
Sbjct: 126 DFLVVGSRGLGRFQKVFVGTVSAFCVKHAECPVMTIKRNADETPSD 171
>At5g14680 unknown protein
Length = 175
Score = 71.6 bits (174), Expect = 2e-13
Identities = 49/159 (30%), Positives = 82/159 (50%), Gaps = 14/159 (8%)
Query: 10 RIMVAIDEGEESIY---------ALTWCLKNLVFQNSKDH-LILLYVKPPRVVYSAFDGT 59
R+MVA++E Y A W LK +V N+ L+LL+V+ FD
Sbjct: 7 RVMVAVNESTLKGYPHASISSKKAFEWTLKKIVRSNTSGFKLLLLHVQVQDE--DGFDDM 64
Query: 60 GYLFSS-DITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKMG 117
+++S D M + ++ +LE C+D+ E I GDP ++IC V+++
Sbjct: 65 DSIYASPDDFRQMRERNKAKGLHLLEFFVKKCHDIGVGCEAWIRKGDPTELICHEVRRVR 124
Query: 118 VDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKK 156
D LV+GS G G ++ F+G+VS C ++ +CPV+ +K+
Sbjct: 125 PDFLVVGSRGLGPFQKVFVGTVSEFCVKHAECPVITIKR 163
>At3g58450 putative protein
Length = 174
Score = 68.2 bits (165), Expect = 2e-12
Identities = 45/136 (33%), Positives = 74/136 (54%), Gaps = 12/136 (8%)
Query: 10 RIMVAIDEGEESIYALTWCLKNLVFQNSKDH--------LILLYVKPP--RVVYSAFDGT 59
++MVAIDE + S AL W + +L S + L LL+V P + +Y +
Sbjct: 31 KVMVAIDESKNSFDALEWAVDHLRVVISAEPETGQEGGLLTLLHVHPTYLQYIYPSGGTA 90
Query: 60 GYLFSSD-ITATMEKYSQQVADCVLEKAKIVCND-VQNVETRIENGDPRDVICQAVQKMG 117
++++D + M K ++ + +A +C + ET I GDP+++ICQAV++
Sbjct: 91 SAVYATDSVPEPMRKAREESTTNLFTRALEICRGKMVKTETMILEGDPKEMICQAVEQTH 150
Query: 118 VDILVMGSHGYGVIKR 133
VD+LV+GS G G+IKR
Sbjct: 151 VDLLVVGSRGLGMIKR 166
>At5g54430 unknown protein
Length = 250
Score = 65.9 bits (159), Expect = 1e-11
Identities = 44/165 (26%), Positives = 83/165 (49%), Gaps = 13/165 (7%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSS--D 66
R+I VA+D EES +A+ W + + + D ++LL+V P V++ A G L + D
Sbjct: 48 RKIGVAVDLSEESSFAVRWAVDHYI--RPGDAVVLLHVSPTSVLFGADWGPLPLKTQIED 105
Query: 67 ITATMEKYSQQVADCVLEKAKIVCNDVQNVETR-----IENGDPRDVICQAVQKMGVDIL 121
A + + K + ++ + +++ D R+ +C ++++G+ +
Sbjct: 106 PNAQPQPSQEDFDAFTSTKVADLAKPLKELGFPYKIHIVKDHDMRERLCLEIERLGLSAV 165
Query: 122 VMGSHGYGVIKR----AFLGSVSNHCAQNVKCPVLIVKKPKSTTG 162
+MGS G+G K+ LGSVS++C + CPV++V+ P G
Sbjct: 166 IMGSRGFGAEKKRGSDGKLGSVSDYCVHHCVCPVVVVRYPDDRDG 210
>At4g27320 unknown protein
Length = 260
Score = 62.4 bits (150), Expect = 1e-10
Identities = 45/169 (26%), Positives = 82/169 (47%), Gaps = 17/169 (10%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYL------ 62
R+I VA+D EES +A+ W + + + D +++L+V P V++ A G L
Sbjct: 45 RKIGVAVDLSEESAFAVRWAVDHYI--RPGDAVVILHVSPTSVLFGADWGPLPLQTPPPP 102
Query: 63 FSSDITATMEKYSQQVADCVLE-KAKIVCNDVQNVETR-----IENGDPRDVICQAVQKM 116
++ K SQ+ D K + ++ +++ D R+ +C +++
Sbjct: 103 SAATDPGAQPKPSQEDFDAFTSSKVADLAKPLKEAGFPHKIHIVKDHDMRERLCLETERL 162
Query: 117 GVDILVMGSHGYGVIKRAF---LGSVSNHCAQNVKCPVLIVKKPKSTTG 162
+ ++MGS G+G KR LGSVS++C + CPV++V+ P G
Sbjct: 163 NLSAVIMGSRGFGAEKRGSDGKLGSVSDYCVHHCVCPVVVVRYPDDRDG 211
>At3g53990 unknown protein
Length = 160
Score = 61.6 bits (148), Expect = 2e-10
Identities = 44/156 (28%), Positives = 75/156 (47%), Gaps = 12/156 (7%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDIT 68
R I +A+D E S AL W ++NL + D + +++ P S + + S +
Sbjct: 5 RNIGIAMDFSESSKNALKWAIENLA--DKGDTIYIIHTLPLSGDESR-NSLWFKSGSPLI 61
Query: 69 ATMEKYSQQVADCVLEKAKIVCNDVQN---------VETRIENGDPRDVICQAVQKMGVD 119
E ++ + K I C D+ + V T++ GD R+ + AV+ + +D
Sbjct: 62 PLAEFREPEIMEKYGVKTDIACLDMLDTGSRQKEVHVVTKLYWGDAREKLVDAVKDLKLD 121
Query: 120 ILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
+VMGS G ++R +GSVS+ Q+ CPV +VK
Sbjct: 122 SIVMGSRGLSALQRIIMGSVSSFVIQHAPCPVTVVK 157
>At3g25930 unknown protein
Length = 154
Score = 58.9 bits (141), Expect = 1e-09
Identities = 45/159 (28%), Positives = 78/159 (48%), Gaps = 22/159 (13%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNL--VFQNSKDHLILLYVK----PPRVVYSA--FDGTG 60
+ +M+ IDE S L W L+N ++SK ++ + PP V+ S+ F
Sbjct: 2 KNVMLIIDESNASYDLLIWALENQKDTIESSKVYIFAKQPQNSFTPPTVLSSSVGFAQIF 61
Query: 61 YLFS--SDITATMEKYSQQVADCVLEKAKIVC-NDVQNVETRIENGDPRDVICQAVQKMG 117
Y FS S++ ++ + ++A +LEKAK +C N ET ++GDP+D+I + +Q+
Sbjct: 62 YPFSPNSELIRLAQEKNMKIALGILEKAKKICLNHGIKAETFTDDGDPKDLIRKIIQERN 121
Query: 118 VDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKK 156
++++V C QN C +L+VKK
Sbjct: 122 INLIVTSDQ-----------QSLKKCTQNTDCSLLVVKK 149
>At2g21620 RD2 protein (RD2)
Length = 187
Score = 54.3 bits (129), Expect = 3e-08
Identities = 42/154 (27%), Positives = 67/154 (43%), Gaps = 21/154 (13%)
Query: 2 AGITENGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGY 61
+G GR ++VA+D G S +A W L + HL+ V S + Y
Sbjct: 33 SGERRRGRDVIVAVDHGPNSKHAFDWALVHFCRLADTLHLV-------HAVSSVKNDVVY 85
Query: 62 LFSSDITATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDIL 121
S A MEK + + + K+ R+ GD VIC+ +K+ +
Sbjct: 86 ETSQ---ALMEKLAVEAYQVAMVKSV----------ARVVEGDAGKVICKEAEKVKPAAV 132
Query: 122 VMGSHGYGVIKRAFLGSVSNHCAQNVK-CPVLIV 154
++G+ G +++ GSVS +C N K PV+IV
Sbjct: 133 IVGTRGRSLVRSVLQGSVSEYCFHNCKSAPVIIV 166
>At1g48961 unknown protein
Length = 219
Score = 52.8 bits (125), Expect = 8e-08
Identities = 40/155 (25%), Positives = 70/155 (44%), Gaps = 20/155 (12%)
Query: 6 ENGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSS 65
E+ RRI+V +++ + + AL W L NL+ Q L+ +Y PPR S
Sbjct: 2 EDVRRIVVVVEDKQAARTALQWALHNLLRQGDVIVLLHVYSPPPRKK-----------KS 50
Query: 66 DITATMEKYSQQVADCVLEKAKIVCNDVQNVETRI---ENGDPRDVICQAVQKMGVDILV 122
+ ++ +A E +C+ N T I E D +I Q V+++G +L+
Sbjct: 51 TAARLLRRHGYNLALSFRE----ICDSFFNTNTEIIVREGDDDGRMIAQVVKEIGASMLL 106
Query: 123 MGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKP 157
+G H + R + + A+N C V+ +K+P
Sbjct: 107 VGLHQNSFLYRWAISGID--VARNFNCKVMAIKQP 139
>At3g21210 unknown protein
Length = 686
Score = 52.4 bits (124), Expect = 1e-07
Identities = 40/149 (26%), Positives = 70/149 (46%), Gaps = 15/149 (10%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDIT 68
R+I +A++ EES + + W + N + Q D +I+L+V P ++ A D Y +
Sbjct: 15 RKIGIAVELSEESAFTVRWAVDNYIRQG--DDIIILHVSPTAGLFGA-DWGYYPLQTQPP 71
Query: 69 ATMEKYSQQVADCVLEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSHGY 128
T +VAD L K + T +++ D R+ +C Q++ + L+MG
Sbjct: 72 YTTASIFSKVAD--LGKPLKEAGFPHTIHT-VKDYDKRERLCLETQRLNLTALIMGFGD- 127
Query: 129 GVIKRAFLGSVSNHCAQNVKCPVLIVKKP 157
GSVS+ C + C V++V+ P
Sbjct: 128 --------GSVSDFCVHHCVCQVVVVRYP 148
>At3g03270 unknown protein
Length = 201
Score = 49.3 bits (116), Expect = 9e-07
Identities = 40/138 (28%), Positives = 68/138 (48%), Gaps = 20/138 (14%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSA---FDGTGYLFSS 65
R + V +D S AL W +NL+ D +IL++V+P ++ F+ TG
Sbjct: 5 RTVGVGMDYSPTSKLALRWAAENLL--EDGDTVILIHVQPQNADHTRKILFEETGSPLIP 62
Query: 66 -----DITATME---KYSQQVADCV--LEKAKIVCNDVQNVETRIENGDPRDVICQAVQK 115
++ + + Y +V D + L +AK V V ++ GDPR+ +C AV+
Sbjct: 63 LEEFREVNLSKQYGLAYDPEVLDVLDTLSRAKKV-----KVVAKVYWGDPREKLCDAVEN 117
Query: 116 MGVDILVMGSHGYGVIKR 133
+ +D +V+GS G G +KR
Sbjct: 118 LKLDSIVLGSRGLGSLKR 135
>At3g03290 hypothetical protein
Length = 274
Score = 45.1 bits (105), Expect = 2e-05
Identities = 40/163 (24%), Positives = 67/163 (40%), Gaps = 3/163 (1%)
Query: 2 AGITENGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGY 61
A ITE G R+MV +D+ S AL W LK+ + S+D+L LLY P +
Sbjct: 99 AEITEAGNRVMVVVDKVIASTGALEWALKHTL--QSQDYLFLLYFSKPFRKGKRKNRKSE 156
Query: 62 LFSSDITATMEKYSQQVADCV-LEKAKIVCNDVQNVETRIENGDPRDVICQAVQKMGVDI 120
+ + ++ T++K Q + +E ++ + + E +E + V V K
Sbjct: 157 VKTDELVHTLKKLCQTKRPGIEVEIRRLQGKEKEKGEKIVEEAKEQQVSLLVVGKEKKPP 216
Query: 121 LVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTTGG 163
+ +G KR +C + C + VK GG
Sbjct: 217 VWRLLKRWGWKKRRGRAGTLKYCLEKASCMTIAVKPKNRKLGG 259
>At5g49050 unknown protein
Length = 150
Score = 42.7 bits (99), Expect = 9e-05
Identities = 31/113 (27%), Positives = 57/113 (50%), Gaps = 10/113 (8%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVF----QNSKDHLILLYVKPPRVVYSAFDGTGYLFS 64
R ++V +D+ S +AL L +L F N + L++L+ +P + G G +
Sbjct: 39 RVVVVGVDDSAHSYHALETAL-DLFFIPFKANPRFKLVVLHARPTATFFLGVAGPGTV-- 95
Query: 65 SDITATMEKYSQQVADCVLEKAKIVCN--DVQNVETRIENGDPRDVICQAVQK 115
DI +E+ + AD V +K VC+ V+ + GDPR+++ +AV++
Sbjct: 96 -DIIPMVEEDLNKTADLVKKKCAEVCSAKSVEISSLEVIEGDPRNIMLEAVER 147
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.137 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,786,508
Number of Sequences: 26719
Number of extensions: 148977
Number of successful extensions: 383
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 30
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 327
Number of HSP's gapped (non-prelim): 53
length of query: 165
length of database: 11,318,596
effective HSP length: 92
effective length of query: 73
effective length of database: 8,860,448
effective search space: 646812704
effective search space used: 646812704
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149080.7


