

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149050.9 - phase: 0 /pseudo
(157 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
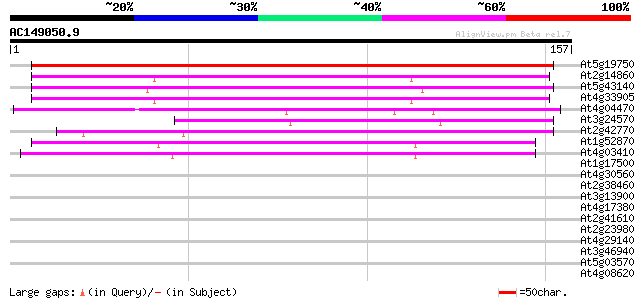
Sequences producing significant alignments: (bits) Value
At5g19750 unknown protein 220 3e-58
At2g14860 22 kDa peroxisomal membrane protein 77 5e-15
At5g43140 putative protein 74 2e-14
At4g33905 unknown protein 72 2e-13
At4g04470 PEROXISOMAL MEMBRANE PROTEIN PMP22 59 1e-09
At3g24570 unknown protein 56 7e-09
At2g42770 Unknown protein 55 1e-08
At1g52870 unknown protein 50 5e-07
At4g03410 unknown protein 46 9e-06
At1g17500 P-type ATPase, putative 30 0.53
At4g30560 cyclic nucleotide and calmodulin-regulated ion channel... 29 1.2
At2g38460 unknown protein 28 1.6
At3g13900 putative ATPase 28 2.7
At4g17380 putative protein 27 4.5
At2g41610 hypothetical protein 27 4.5
At2g23980 putative cyclic nucleotide and calmodulin-regulated io... 27 4.5
At4g29140 unknown protein 27 5.9
At3g46940 dUTP pyrophosphatase-like protein 27 5.9
At5g03570 transporter like protein 26 7.7
At4g08620 putative sulfate transporter 26 7.7
>At5g19750 unknown protein
Length = 288
Score = 220 bits (560), Expect = 3e-58
Identities = 102/146 (69%), Positives = 125/146 (84%)
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTLHF 66
Y ALL+ PV KA+T+A+LNL+GDLICQL I+K + D KRT F+FLGL LVGPTLHF
Sbjct: 118 YQALLSNSPVLTKAVTAALLNLVGDLICQLTINKTSSLDKKRTLTFTFLGLGLVGPTLHF 177
Query: 67 WYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVPKLKQEWFS 126
WYLYLS++VT G SGA++RL+LDQFVF+PIF+GVFLS++VTLEG+PS +PKL+QEW
Sbjct: 178 WYLYLSKVVTASGLSGAVIRLLLDQFVFAPIFVGVFLSAVVTLEGKPSNVIPKLQQEWTG 237
Query: 127 AVLANWQLWIPFQFLNFRFVPQQFQV 152
A++ANWQLWIPFQFLNFRFVPQ +QV
Sbjct: 238 AMIANWQLWIPFQFLNFRFVPQNYQV 263
>At2g14860 22 kDa peroxisomal membrane protein
Length = 252
Score = 76.6 bits (187), Expect = 5e-15
Identities = 42/147 (28%), Positives = 76/147 (51%), Gaps = 2/147 (1%)
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVID-KVQTPDLKRTFLFSFLGLVLVGPTLH 65
Y+ ++ +PV K++TS+++ + DL Q + ++ DL RT GL ++GPTLH
Sbjct: 77 YLGMVKSHPVVTKSVTSSLIYIAADLSSQTIAKTSSESYDLVRTARMGGYGLFVLGPTLH 136
Query: 66 FWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEG-RPSQAVPKLKQEW 124
+W+ ++S+L ++ + Q ++ PI +F S +L+G R S + +LK++
Sbjct: 137 YWFNFMSRLFPKQDLITTFKKMAMGQTIYGPIMTVIFFSLNASLQGERGSVILARLKRDL 196
Query: 125 FSAVLANWQLWIPFQFLNFRFVPQQFQ 151
A+ W F+ FRF P Q
Sbjct: 197 LPALFNGVMYWPLCDFITFRFFPVHLQ 223
>At5g43140 putative protein
Length = 248
Score = 74.3 bits (181), Expect = 2e-14
Identities = 40/148 (27%), Positives = 77/148 (52%), Gaps = 2/148 (1%)
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLV-IDKVQTPDLKRTFLFSFLGLVLVGPTLH 65
Y+ L +P K++T++++ + DL Q++ ++ + DL RT + GL+ +GP+ H
Sbjct: 82 YLRKLESHPFMTKSITTSVIYMAADLTSQMITMEPTGSFDLIRTARMASFGLIFLGPSQH 141
Query: 66 FWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPS-QAVPKLKQEW 124
W+ YLS+++ ++++ Q +F P+ VF S L+G S + V +LK++
Sbjct: 142 LWFSYLSKILPKRDVLTTFKKIMMGQVLFGPVSNTVFYSYNAALQGENSEEIVARLKRDL 201
Query: 125 FSAVLANWQLWIPFQFLNFRFVPQQFQV 152
+ W F+ F++VP QV
Sbjct: 202 LPTLKNGLMYWPVCDFVTFKYVPVHLQV 229
>At4g33905 unknown protein
Length = 261
Score = 71.6 bits (174), Expect = 2e-13
Identities = 43/147 (29%), Positives = 72/147 (48%), Gaps = 2/147 (1%)
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVID-KVQTPDLKRTFLFSFLGLVLVGPTLH 65
Y+ ++ PV K++TS+++ + DL Q + V + DL RT GL+++GPTLH
Sbjct: 86 YLGMVKSRPVLTKSVTSSLIYIAADLSSQTIPQASVDSYDLVRTARMGGYGLLILGPTLH 145
Query: 66 FWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEG-RPSQAVPKLKQEW 124
+W+ +S L ++ + Q V+ P VF S L+G S+ V +LK++
Sbjct: 146 YWFNLMSSLFPKRDLITTFKKMAMGQTVYGPAMNVVFFSLNAALQGENGSEIVARLKRDL 205
Query: 125 FSAVLANWQLWIPFQFLNFRFVPQQFQ 151
+L W F+ F+F P Q
Sbjct: 206 LPTMLNGVMYWPLCDFITFKFCPVYLQ 232
>At4g04470 PEROXISOMAL MEMBRANE PROTEIN PMP22
Length = 190
Score = 58.5 bits (140), Expect = 1e-09
Identities = 40/156 (25%), Positives = 79/156 (50%), Gaps = 4/156 (2%)
Query: 2 TLYYRYMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVG 61
T RY++ L ++P+ KA+T+ +L+ + D++ Q + +Q L+R L +G
Sbjct: 9 TTLQRYLSQLQQHPLRTKAITAGVLSGVSDVVSQ-KLSGIQKIQLRRVLLKVIFAGGFLG 67
Query: 62 PTLHFWYLYLSQLVT-LPGTSGAILRLVLDQFVFSPIFLGVFLSSL-VTLEGRPSQAV-P 118
P HF++ YL + T +++L+Q SP+ +F+ V +E P V
Sbjct: 68 PAGHFFHTYLDKFFKGKKDTQTVAKKVILEQLTLSPLNHLLFMIYYGVVIERTPWTLVRE 127
Query: 119 KLKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQVRL 154
++K+ + + L W + ++N+++VP F+V L
Sbjct: 128 RIKKTYPTVQLTAWTFFPVVGWINYKYVPLHFRVIL 163
>At3g24570 unknown protein
Length = 235
Score = 56.2 bits (134), Expect = 7e-09
Identities = 34/113 (30%), Positives = 54/113 (47%), Gaps = 7/113 (6%)
Query: 47 KRTFLFSFLGLVLVGPTLHFWYLYLSQLVTL------PGTSGAILRLVLDQFVFSPIFLG 100
KR + S G VGP HFWY L + + L T ++ +D +F P+ L
Sbjct: 78 KRVAITSMFGFGFVGPVGHFWYEGLDKFIKLKLRYVPKSTRFVAAKVAMDGLIFGPVDLL 137
Query: 101 VFLSSLVTLEGRPSQAVPK-LKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
VF + + G+ + V + LK+++ A+ W Q NFR+VP Q+Q+
Sbjct: 138 VFFTYMGFATGKNTAEVKEGLKRDFLPALALEGGAWPLLQIANFRYVPVQYQL 190
>At2g42770 Unknown protein
Length = 232
Score = 55.5 bits (132), Expect = 1e-08
Identities = 41/161 (25%), Positives = 68/161 (41%), Gaps = 22/161 (13%)
Query: 14 YPVPVK-ALTSAILNLIGDLICQLVIDKVQTPDLK---------------------RTFL 51
Y P+K A+T+ L GD I QL + LK R
Sbjct: 52 YRFPLKQAVTAGALTFTGDTIAQLSGRWKKRTALKQSSSELDEGELWNIFSEHDWIRALR 111
Query: 52 FSFLGLVLVGPTLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEG 111
S G +L GP + WY +L + P + +L+++L+Q + P + V + G
Sbjct: 112 MSSYGFLLYGPGSYAWYQFLDHSLPKPTATNLVLKVLLNQVILGPSVIAVIFAWNNLWLG 171
Query: 112 RPSQAVPKLKQEWFSAVLANWQLWIPFQFLNFRFVPQQFQV 152
+ S+ K +++ +L ++ W+P LNF VP Q +V
Sbjct: 172 KLSELGNKYQKDALPTLLYGFRFWVPVSILNFWVVPLQARV 212
>At1g52870 unknown protein
Length = 366
Score = 50.1 bits (118), Expect = 5e-07
Identities = 37/143 (25%), Positives = 61/143 (41%), Gaps = 2/143 (1%)
Query: 7 YMALLAKYPVPVKALTSAILNLIGDLICQLVIDK-VQTPDLKRTFLFSFLGLVLVGPTLH 65
Y L + PV K + S ++ +GD I Q K + D RT +G L G H
Sbjct: 173 YEEALKQNPVLAKMVISGVVYSVGDWIAQCYEGKPLFEIDRARTLRSGLVGFTLHGSLSH 232
Query: 66 FWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGR-PSQAVPKLKQEW 124
F+Y + +L +++ DQ V+S I+ ++ + L L P +LK +
Sbjct: 233 FYYQFCEELFPFQDWWVVPVKVAFDQTVWSAIWNSIYFTVLGFLRFESPISIFKELKATF 292
Query: 125 FSAVLANWQLWIPFQFLNFRFVP 147
+ A W+LW + + VP
Sbjct: 293 LPMLTAGWKLWPFAHLITYGLVP 315
>At4g03410 unknown protein
Length = 317
Score = 45.8 bits (107), Expect = 9e-06
Identities = 34/146 (23%), Positives = 60/146 (40%), Gaps = 2/146 (1%)
Query: 4 YYRYMALLAKYPVPVKALTSAILNLIGDLICQLVIDKVQTP-DLKRTFLFSFLGLVLVGP 62
++ Y +L PV K S I+ +GD I Q K D R +G L G
Sbjct: 128 WFAYEQILKTNPVLAKMAISGIVYSLGDWIAQCYEGKPLFEFDRTRVLRSGLVGFTLHGS 187
Query: 63 TLHFWYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGR-PSQAVPKLK 121
H++Y + L ++ DQ V+S I+ ++ + L L + P+ ++K
Sbjct: 188 LSHYYYQFCEALFPFQEWWVVPAKVAFDQTVWSAIWNSIYFTVLGLLRFQSPADIFSEIK 247
Query: 122 QEWFSAVLANWQLWIPFQFLNFRFVP 147
+ + A W+LW + + +P
Sbjct: 248 TTFLPMLTAGWKLWPLAHLVTYGVIP 273
>At1g17500 P-type ATPase, putative
Length = 1218
Score = 30.0 bits (66), Expect = 0.53
Identities = 20/89 (22%), Positives = 46/89 (51%), Gaps = 2/89 (2%)
Query: 23 SAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVLVGPTLHFWYLYLSQLVTL-PGTS 81
+A ++ +G + +I V F+++ VL+ ++ WYL+++ + P S
Sbjct: 1042 TADMDAVGTTMFTCIIWAVNVQIALTVSHFTWIQHVLIWGSIGLWYLFVALYGMMPPSLS 1101
Query: 82 GAILRLVLDQFVFSPIF-LGVFLSSLVTL 109
G I R++++ +PI+ + FL ++ T+
Sbjct: 1102 GNIYRILVEILAPAPIYWIATFLVTVTTV 1130
>At4g30560 cyclic nucleotide and calmodulin-regulated ion
channel-like protein
Length = 733
Score = 28.9 bits (63), Expect = 1.2
Identities = 22/71 (30%), Positives = 40/71 (55%), Gaps = 6/71 (8%)
Query: 35 QLVIDKVQTPD--LKRTFLFSFLGLVLVGPTLHFW-YLYLSQLVTLPGTSGAILRLVLDQ 91
+LVID Q L++ F+ FL VL P + W +LY+S+ ++ T A+ ++L Q
Sbjct: 190 ELVIDPAQIAKRYLQQYFIIDFLS-VLPLPQIVVWRFLYISKGASVLATKRALRSIILVQ 248
Query: 92 FVFSPIFLGVF 102
++ P F+ ++
Sbjct: 249 YI--PRFIRLY 257
>At2g38460 unknown protein
Length = 524
Score = 28.5 bits (62), Expect = 1.6
Identities = 15/51 (29%), Positives = 26/51 (50%), Gaps = 7/51 (13%)
Query: 67 WYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAV 117
W +Y +Q V LPG S A+L F+ + G +++ + EG P+ +
Sbjct: 306 WRIYFNQEVVLPGVSLALL-------FFTVLSFGTLMTATLQWEGIPTYII 349
>At3g13900 putative ATPase
Length = 1243
Score = 27.7 bits (60), Expect = 2.7
Identities = 17/59 (28%), Positives = 34/59 (56%), Gaps = 4/59 (6%)
Query: 52 FSFLGLVLVGPTLHFWYLYLSQLVTL-PGTSGAILRLVLDQFVFSPIFLGVFLSSLVTL 109
F+++ VL+ ++ WY++L+ L P SG I ++ + +PIF +L+SL+ +
Sbjct: 1088 FTWIQHVLIWGSIVTWYIFLALFGMLPPKVSGNIFHMLSETLAPAPIF---WLTSLLVI 1143
>At4g17380 putative protein
Length = 574
Score = 26.9 bits (58), Expect = 4.5
Identities = 18/67 (26%), Positives = 30/67 (43%)
Query: 84 ILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAVPKLKQEWFSAVLANWQLWIPFQFLNF 143
IL + + FV + IF+ + LV + S L+Q +LA ++P +F
Sbjct: 363 ILESIHNDFVSNSIFMSEATNMLVVMGPNMSGKSTYLQQVCLVVILAQIGCYVPARFATI 422
Query: 144 RFVPQQF 150
R V + F
Sbjct: 423 RVVDRIF 429
>At2g41610 hypothetical protein
Length = 304
Score = 26.9 bits (58), Expect = 4.5
Identities = 17/42 (40%), Positives = 25/42 (59%), Gaps = 2/42 (4%)
Query: 18 VKALTSAILNLIGDLICQLVIDKVQTPDLKRTFLFSFLGLVL 59
V+ LT +L LI D + V+ V+ P L R F+F FL L++
Sbjct: 242 VQGLT--LLVLIRDCVYLAVMSPVEEPILFRVFVFGFLLLLI 281
>At2g23980 putative cyclic nucleotide and calmodulin-regulated ion
channel protein
Length = 747
Score = 26.9 bits (58), Expect = 4.5
Identities = 20/70 (28%), Positives = 37/70 (52%), Gaps = 4/70 (5%)
Query: 35 QLVIDKVQTPD--LKRTFLFSFLGLVLVGPTLHFWYLYLSQLVTLPGTSGAILRLVLDQF 92
+LVID Q L++ F+ L ++ V + + +LY S+ + T A+ +VL Q+
Sbjct: 190 ELVIDPAQIAKRYLQQYFIIDLLSVLPVPQIIVWRFLYTSRGANVLATKQALRYIVLVQY 249
Query: 93 VFSPIFLGVF 102
+ P FL ++
Sbjct: 250 I--PRFLRMY 257
>At4g29140 unknown protein
Length = 532
Score = 26.6 bits (57), Expect = 5.9
Identities = 23/68 (33%), Positives = 32/68 (46%), Gaps = 8/68 (11%)
Query: 42 QTPD---LKRTFLFSFLGLVLVGPTLHFWYLYLSQL-----VTLPGTSGAILRLVLDQFV 93
Q PD L +T+L L +L LH +YL VTL SGA+ L + F+
Sbjct: 168 QDPDIAKLAQTYLIFSLPDLLTNTLLHPIRIYLRAQGIIHPVTLASLSGAVFHLPANLFL 227
Query: 94 FSPIFLGV 101
S + LG+
Sbjct: 228 VSYLRLGL 235
>At3g46940 dUTP pyrophosphatase-like protein
Length = 166
Score = 26.6 bits (57), Expect = 5.9
Identities = 10/17 (58%), Positives = 14/17 (81%)
Query: 30 GDLICQLVIDKVQTPDL 46
GD I QL+I+K+ TPD+
Sbjct: 129 GDRIAQLIIEKIVTPDV 145
>At5g03570 transporter like protein
Length = 498
Score = 26.2 bits (56), Expect = 7.7
Identities = 14/51 (27%), Positives = 26/51 (50%), Gaps = 7/51 (13%)
Query: 67 WYLYLSQLVTLPGTSGAILRLVLDQFVFSPIFLGVFLSSLVTLEGRPSQAV 117
W YL+Q + LPG S A+L F+ + G +++ + +G P+ +
Sbjct: 302 WRNYLNQEIVLPGVSLALL-------FFTVLSFGTLMTATLEWKGIPTYII 345
>At4g08620 putative sulfate transporter
Length = 649
Score = 26.2 bits (56), Expect = 7.7
Identities = 20/74 (27%), Positives = 34/74 (45%), Gaps = 6/74 (8%)
Query: 24 AILNLIGDLICQLVIDKVQTPDLKRTFLFS---FLGLVLVG---PTLHFWYLYLSQLVTL 77
A+++L+ +CQ VID + P+ +F+ F G+ G L F +LS +
Sbjct: 143 AVVSLLVGTLCQAVIDPKKNPEDYLRLVFTATFFAGIFQAGLGFLRLGFLIDFLSHAAVV 202
Query: 78 PGTSGAILRLVLDQ 91
GA + + L Q
Sbjct: 203 GFMGGAAITIALQQ 216
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.332 0.146 0.456
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,242,747
Number of Sequences: 26719
Number of extensions: 123758
Number of successful extensions: 386
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 368
Number of HSP's gapped (non-prelim): 22
length of query: 157
length of database: 11,318,596
effective HSP length: 91
effective length of query: 66
effective length of database: 8,887,167
effective search space: 586553022
effective search space used: 586553022
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC149050.9


