

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149038.3 + phase: 0
(1557 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
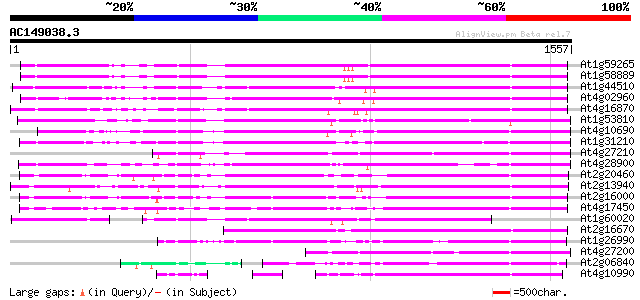
At1g59265 polyprotein, putative 930 0.0
At1g58889 polyprotein, putative 928 0.0
At1g44510 polyprotein, putative 923 0.0
At4g02960 putative polyprotein of LTR transposon 918 0.0
At4g16870 retrotransposon like protein 909 0.0
At1g53810 867 0.0
At4g10690 retrotransposon like protein 860 0.0
At1g31210 putative reverse transcriptase 834 0.0
At4g27210 putative protein 768 0.0
At4g28900 putative protein 714 0.0
At2g20460 putative retroelement pol polyprotein 704 0.0
At2g13940 putative retroelement pol polyprotein 697 0.0
At2g16000 putative retroelement pol polyprotein 681 0.0
At4g17450 retrotransposon like protein 628 e-180
At1g60020 hypothetical protein 626 e-179
At2g16670 putative retroelement pol polyprotein 616 e-176
At1g26990 polyprotein, putative 584 e-166
At4g27200 putative protein 518 e-146
At2g06840 putative retroelement pol polyprotein 460 e-129
At4g10990 putative retrotransposon polyprotein 453 e-127
>At1g59265 polyprotein, putative
Length = 1466
Score = 930 bits (2403), Expect = 0.0
Identities = 590/1568 (37%), Positives = 826/1568 (52%), Gaps = 181/1568 (11%)
Query: 31 KLDETNYLQWKQQVEGVLRGTKMVRHVV-SPQIPPVFLNDAAREAGTENPAYTEWEEQDS 89
KL TNYL W +QV + G ++ + S +PP + A A NP YT W+ QD
Sbjct: 25 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTTMPPATIGTDA--APRVNPDYTRWKRQDK 82
Query: 90 LLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGTRSI 149
L+ + +L IS S+ R + Q+W+ + QLR++L+ TKGT++I
Sbjct: 83 LIYSAVLGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQLRTQLKQWTKGTKTI 142
Query: 150 AEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELESQ 209
+++ + + + L +G P+ H + +E VLE LPEE+ P++ + ++ +L E+ +
Sbjct: 143 DDYMQGLVTRFDQLALLGKPMDHDEQVERVLENLPEEYKPVIDQIAAKDTPPTLTEIHER 202
Query: 210 LLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNNS 269
LL E++ A V + ++H + N +N + Y ++ N + P S
Sbjct: 203 LLNHESKILAVSSATVIPITANAVSHRNTTTTNNNNNGNRNNRYDNRNNNNNSKPW-QQS 261
Query: 270 SQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVPP 329
S F+PN ++ + + G +CQIC GH A C
Sbjct: 262 STNFHPN-----NNQSKPYLG----------------KCQICGVQGHSAKRC-------- 292
Query: 330 QYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRP 389
+ ++ V + P +P P+ P+
Sbjct: 293 ---------------------------SQLQHFLSSVNSQQPP--SPFTPWQPR------ 317
Query: 390 PAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLA 449
N + S +N W DSGATHH+T D NNL ++G D + + +G +
Sbjct: 318 --------ANLALGSPYSSNNWLLDSGATHHITSDFNNLSLHQPYTGGDDVMVADGSTIP 369
Query: 450 INSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQ 509
I+ GS S S+ P L L+N+L+VP+I KNL+SV + C N V EF VK
Sbjct: 370 ISHTGSTSLSTKSRP---LNLHNILYVPNIHKNLISVYRLCNANGVSVEFFPASFQVKDL 426
Query: 510 DSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCN 569
++ LL+G D+ LY++ VS AS SS AT
Sbjct: 427 NTGVPLLQGKTKDE-LYEWPIASSQPVSLFASPSSKAT---------------------- 463
Query: 570 NGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDF-C 628
H+S WH RLGHP ++ SV+ + + N S C
Sbjct: 464 --------HSS---------------WHARLGHPAPSILNSVISNYSLSVLNPSHKFLSC 500
Query: 629 SACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIF 688
S C + KS+++P S +P E I+ D+W + + SH Y Y++I VD ++RYTW++
Sbjct: 501 SDCLINKSNKVPFSQSTINSTRPLEYIYSDVWS-SPILSHDNYRYYVIFVDHFTRYTWLY 559
Query: 689 PLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTH 748
PLK KS TF FK ++E ++ I + +D GGEF ++ + GI+H + PHT
Sbjct: 560 PLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEFVALWEYFSQHGISHLTSPPHTP 619
Query: 749 HQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLH 808
NG ERKHRHIVETGLTLL++A +P YW +AF A YLINRLP+P+L +SPF L
Sbjct: 620 EHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQLESPFQKLF 679
Query: 809 LQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFIS 867
P+Y L+ FGC+C+P+ RPYN HKL+ +S++CVFLGYS + Y CL T R++IS
Sbjct: 680 GTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHLQTSRLYIS 739
Query: 868 KDVIFNEYKFPYSELFTS-----GQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTP 922
+ V F+E FP+S + Q S HT LP+ L C ++TP
Sbjct: 740 RHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRTPVLPAPSCSDPHHAATP 799
Query: 923 TTTLHT-------------ASPHSSFPES------NQSNHHHSIQ------DTHAS---S 954
++ +S SSFP S Q+ + Q TH+S S
Sbjct: 800 PSSPSAPFRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQTQTHSSQNTS 859
Query: 955 HSNHHNISPGPIFN--PTPISTHPPSPSP-----SSHSHNTYHSISVEPVTSQPSTQ-AE 1006
+N N SP + TP + SPSP SS + T SI + P P Q
Sbjct: 860 QNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHP--PPPLAQIVN 917
Query: 1007 PHRIHPNNTHSMATRAKHGIVQKRKHPTL---LLTHIEPTGYRQAMKQPQWLQAMQLEHE 1063
+ P NTHSM TRAK GI++ +L L EP QA+K +W AM E
Sbjct: 918 NNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSLAAESEPRTAIQALKDERWRNAMGSEIN 977
Query: 1064 ALMKNNTWTLVPLPADR-QAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKET 1122
A + N+TW LVP P VGC+W+F K N DGS+N+YKARLVAKG++Q PG DY ET
Sbjct: 978 AQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARLVAKGYNQRPGLDYAET 1037
Query: 1123 FSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PSLVC 1181
FSPV+K ++R VL +AV W I+QLDVNNAFL G L ++VYM+QPPGF D P+ VC
Sbjct: 1038 FSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDKDRPNYVC 1097
Query: 1182 KLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILIT 1241
KL KALYGLKQAPRAW+ L++ LL +GF +S D SLF+L + +MLVYVDDILIT
Sbjct: 1098 KLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVYVDDILIT 1157
Query: 1242 GSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMA 1301
G+ +L+ + L+ FS+KD +L YFLGIE G L LSQ++YI DLL + NM
Sbjct: 1158 GNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRVPTG-LHLSQRRYILDLLARTNMI 1216
Query: 1302 NANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAP 1361
A + +PMA S KL+ Y ++DPT +R IVG LQY+ TRP+ISY+VN++ QF+ P
Sbjct: 1217 TAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRLSQFMHMP 1276
Query: 1362 LEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACIL 1421
E+H +A+KRILRYL GT +HG+ + LSL A+ DADWA D DD ST+G +
Sbjct: 1277 TEEHLQALKRILRYLAGTPNHGIFLKKG---NTLSLHAYSDADWAGDKDDYVSTNGYIVY 1333
Query: 1422 LGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVP-TAIPQIFCDNL 1480
LG + ISW +KKQ V RSS EAEYRS+A S+E+ WI SLL EL + T P I+CDN+
Sbjct: 1334 LGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPPVIYCDNV 1393
Query: 1481 STVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLE 1540
L NPV HSR KH+ +D F+R +V S L V H+ Q+AD LTKPLS + F
Sbjct: 1394 GATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRTAFQN 1453
Query: 1541 LRNKLRVS 1548
+K+ V+
Sbjct: 1454 FASKIGVT 1461
>At1g58889 polyprotein, putative
Length = 1466
Score = 928 bits (2399), Expect = 0.0
Identities = 589/1568 (37%), Positives = 825/1568 (52%), Gaps = 181/1568 (11%)
Query: 31 KLDETNYLQWKQQVEGVLRGTKMVRHVV-SPQIPPVFLNDAAREAGTENPAYTEWEEQDS 89
KL TNYL W +QV + G ++ + S +PP + A A NP YT W+ QD
Sbjct: 25 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTTMPPATIGTDA--APRVNPDYTRWKRQDK 82
Query: 90 LLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGTRSI 149
L+ + +L IS S+ R + Q+W+ + QLR++L+ TKGT++I
Sbjct: 83 LIYSAVLGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQLRTQLKQWTKGTKTI 142
Query: 150 AEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELESQ 209
+++ + + + L +G P+ H + +E VLE LPEE+ P++ + ++ +L E+ +
Sbjct: 143 DDYMQGLVTRFDQLALLGKPMDHDEQVERVLENLPEEYKPVIDQIAAKDTPPTLTEIHER 202
Query: 210 LLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNNS 269
LL E++ A V + ++H + N +N + Y ++ N + P S
Sbjct: 203 LLNHESKILAVSSATVIPITANAVSHRNTTTTNNNNNGNRNNRYDNRNNNNNSKPW-QQS 261
Query: 270 SQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVPP 329
S F+PN ++ + + G +CQIC GH A C
Sbjct: 262 STNFHPN-----NNQSKPYLG----------------KCQICGVQGHSAKRC-------- 292
Query: 330 QYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRP 389
+ ++ V + P +P P+ P+
Sbjct: 293 ---------------------------SQLQHFLSSVNSQQPP--SPFTPWQPR------ 317
Query: 390 PAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLA 449
N + S +N W DSGATHH+T D NNL ++G D + + +G +
Sbjct: 318 --------ANLALGSPYSSNNWLLDSGATHHITSDFNNLSLHQPYTGGDDVMVADGSTIP 369
Query: 450 INSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQ 509
I+ GS S S+ P L L+N+L+VP+I KNL+SV + C N V EF VK
Sbjct: 370 ISHTGSTSLSTKSRP---LNLHNILYVPNIHKNLISVYRLCNANGVSVEFFPASFQVKDL 426
Query: 510 DSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCN 569
++ LL+G D+ LY++ VS AS SS AT
Sbjct: 427 NTGVPLLQGKTKDE-LYEWPIASSQPVSLFASPSSKAT---------------------- 463
Query: 570 NGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDF-C 628
H+S WH RLGHP ++ SV+ + + N S C
Sbjct: 464 --------HSS---------------WHARLGHPAPSILNSVISNYSLSVLNPSHKFLSC 500
Query: 629 SACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIF 688
S C + KS+++P S +P E I+ D+W + + SH Y Y++I VD ++RYTW++
Sbjct: 501 SDCLINKSNKVPFSQSTINSTRPLEYIYSDVWS-SPILSHDNYRYYVIFVDHFTRYTWLY 559
Query: 689 PLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTH 748
PLK KS TF FK ++E ++ I + +D GGEF ++ + GI+H + PHT
Sbjct: 560 PLKQKSQVKETFITFKNLLENRFQTRIGTFYSDNGGEFVALWEYFSQHGISHLTSPPHTP 619
Query: 749 HQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLH 808
NG ERKHRHIVETGLTLL++A +P YW +AF A YLINRLP+P+L +SPF L
Sbjct: 620 EHNGLSERKHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQLESPFQKLF 679
Query: 809 LQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFIS 867
P+Y L+ FGC+C+P+ RPYN HKL+ +S++CVFLGYS + Y CL T R++IS
Sbjct: 680 GTSPNYDKLRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHLQTSRLYIS 739
Query: 868 KDVIFNEYKFPYSELFTS-----GQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTP 922
+ V F+E FP+S + Q S HT LP+ L C ++TP
Sbjct: 740 RHVRFDENCFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRTPVLPAPSCSDPHHAATP 799
Query: 923 TTTLHT-------------ASPHSSFPES------NQSNHHHSIQ------DTHAS---S 954
++ +S SSFP S Q+ + Q TH+S S
Sbjct: 800 PSSPSAPFRNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQTQTHSSQNTS 859
Query: 955 HSNHHNISPGPIFN--PTPISTHPPSPSP-----SSHSHNTYHSISVEPVTSQPSTQ-AE 1006
+N N SP + TP + SPSP SS + T SI + P P Q
Sbjct: 860 QNNPTNESPSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHP--PPPLAQIVN 917
Query: 1007 PHRIHPNNTHSMATRAKHGIVQKRKHPTL---LLTHIEPTGYRQAMKQPQWLQAMQLEHE 1063
+ P NTHSM TRAK GI++ +L L EP QA+K +W AM E
Sbjct: 918 NNNQAPLNTHSMGTRAKAGIIKPNPKYSLAVSLAAESEPRTAIQALKDERWRNAMGSEIN 977
Query: 1064 ALMKNNTWTLVPLPADR-QAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKET 1122
A + N+TW LVP P VGC+W+F K N DGS+N+YKAR VAKG++Q PG DY ET
Sbjct: 978 AQIGNHTWDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARFVAKGYNQRPGLDYAET 1037
Query: 1123 FSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PSLVC 1181
FSPV+K ++R VL +AV W I+QLDVNNAFL G L ++VYM+QPPGF D P+ VC
Sbjct: 1038 FSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDKDRPNYVC 1097
Query: 1182 KLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILIT 1241
KL KALYGLKQAPRAW+ L++ LL +GF +S D SLF+L + +MLVYVDDILIT
Sbjct: 1098 KLRKALYGLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVYVDDILIT 1157
Query: 1242 GSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMA 1301
G+ +L+ + L+ FS+KD +L YFLGIE G L LSQ++YI DLL + NM
Sbjct: 1158 GNDPTLLHNTLDNLSQRFSVKDHEELHYFLGIEAKRVPTG-LHLSQRRYILDLLARTNMI 1216
Query: 1302 NANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAP 1361
A + +PMA S KL+ Y ++DPT +R IVG LQY+ TRP+ISY+VN++ QF+ P
Sbjct: 1217 TAKPVTTPMAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRLSQFMHMP 1276
Query: 1362 LEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACIL 1421
E+H +A+KRILRYL GT +HG+ + LSL A+ DADWA D DD ST+G +
Sbjct: 1277 TEEHLQALKRILRYLAGTPNHGIFLKKG---NTLSLHAYSDADWAGDKDDYVSTNGYIVY 1333
Query: 1422 LGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVP-TAIPQIFCDNL 1480
LG + ISW +KKQ V RSS EAEYRS+A S+E+ WI SLL EL + T P I+CDN+
Sbjct: 1334 LGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPPVIYCDNV 1393
Query: 1481 STVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLE 1540
L NPV HSR KH+ +D F+R +V S L V H+ Q+AD LTKPLS + F
Sbjct: 1394 GATYLCANPVFHSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRTAFQN 1453
Query: 1541 LRNKLRVS 1548
+K+ V+
Sbjct: 1454 FASKIGVT 1461
>At1g44510 polyprotein, putative
Length = 1459
Score = 923 bits (2386), Expect = 0.0
Identities = 579/1588 (36%), Positives = 825/1588 (51%), Gaps = 189/1588 (11%)
Query: 7 PVNNSETQRVTAAASKNFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVV-SPQIPPV 65
P E +T N KL TNYL W Q+ +L G + ++ S IPP
Sbjct: 8 PATRDEAIVLTPQTLFNVNTSNVTKLTSTNYLMWSIQIHALLDGYDLAGYLDNSVVIPPE 67
Query: 66 FLNDAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCY 125
+ NP++T W+ QD L+ + ++ IS ++ S R S Q+W ++N
Sbjct: 68 --TTTINSVVSANPSFTLWKRQDKLIFSALIGAISPAVQSLVSRATNSSQIWSTLNNTYA 125
Query: 126 TQMRTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPE 185
+QLR +++ +TKGT++I E++ + + L +G P+ H + +E +L+ LPE
Sbjct: 126 KPSYGHIKQLRQQIQRLTKGTKTIDEYVQSHTTRLDQLAILGKPMEHEEQVEHILKGLPE 185
Query: 186 EFNPIVATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGH 245
E+ +V + + ++ E+ +L+ E+ K V ++S ++ A ++N +
Sbjct: 186 EYKTVVDQIEGKDNTPTITEIHERLINHES---KLLSDEVPPSSSFPMS-ANAVQQRNFN 241
Query: 246 NQPQTGSYPDQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGN 305
N + ++ GN NN++ P+ ++G R F+ G+
Sbjct: 242 NNCNQNQHKNRY---QGNTHNNNTNTNSQPSTYNKSGQR-------TFKPYLGK------ 285
Query: 306 IQCQICYKTGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQG 365
CQIC GH A C PQ +
Sbjct: 286 --CQICSVQGHSARRC-------PQLQA-------------------------------- 304
Query: 366 VGQRNPSYGAPRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDA 425
+ P+ + +PF P P+A N + S N W DSGATHH+T D
Sbjct: 305 --MQLPASSSAHSPFTPW-------QPRA----NLAIGSPYAANPWLLDSGATHHITSDL 351
Query: 426 NNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVS 485
N L ++G + + I +G GL I GS S N L L+ +L+VP I KNL+S
Sbjct: 352 NALSLHQPYNGGEYVMIADGTGLTIKQTGSTFLPSQ---NRDLALHKVLYVPDIRKNLIS 408
Query: 486 VSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSV 545
V + C N V EF VK ++ +LL+G DD LY++ P+
Sbjct: 409 VYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDD-LYEWPVTNPPA---------- 457
Query: 546 ATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHH 605
T + TS S + +S WH+RLGHP
Sbjct: 458 -----------------------------TALFTSPSPKTTLSS------WHSRLGHPSA 482
Query: 606 EVVRSVMKLCNQQLPNKSFTDF-CSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPAS 664
++ +++ + + S CS C + KSH+LP +S + P E IF D+W +
Sbjct: 483 SILNTLLSKFSLPVSVASSNKTSCSDCLINKSHKLPFATSSIHSSSPLEYIFTDVW-TSP 541
Query: 665 VESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGG 724
+ SH Y Y+L+ VD Y+RYTW++PL+ KS TF FK +VE ++ I+++ +D GG
Sbjct: 542 IISHDNYKYYLVLVDHYTRYTWLYPLQQKSQVKATFIAFKALVENRFQAKIRTLYSDNGG 601
Query: 725 EFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFL 784
EF FL + GI+H + PHT NG ERKHRHIVETGLTLL A +P YW +AF
Sbjct: 602 EFIALRDFLVSNGISHLTSPPHTPEHNGLSERKHRHIVETGLTLLTQASVPREYWTYAFA 661
Query: 785 TATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECV 844
TA YLINR+P+P+L +SPF L P+Y+ L+ FGC CFP+ RPY +KLE RSK CV
Sbjct: 662 TAVYLINRMPTPVLCLQSPFQKLFGSSPNYQRLRVFGCLCFPWLRPYTRNKLEERSKRCV 721
Query: 845 FLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQSSS---PPTTSSDHTP 900
FLGYS + Y CLD R++ S+ V+F+E +P++ SS PP +SS +P
Sbjct: 722 FLGYSLTQTAYLCLDVDNNRLYTSRHVMFDESTYPFAASIREQSQSSLVTPPESSSSSSP 781
Query: 901 LPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTHASSHSNHHN 960
S FP C + S P +SP + P Q++ S + T + + S+H
Sbjct: 782 ANSG-FP-----CSVLRLQSPP-----ASSPETPSPPQQQNDSPVSPRQTGSPTPSHHSQ 830
Query: 961 I-----SPGP-IFNPTPISTHPPSPSPSSHSH----------------NTYHSISVEPVT 998
+ SP P + N P + H P P + S+ N SI PV
Sbjct: 831 VRDSTLSPSPSVSNSEPTAPHENGPEPEAQSNPNSPFIGPLPNPNPETNPSSSIEQRPVD 890
Query: 999 SQPSTQAEPHRI-----------HPNNTHSMATRAKHGIVQKRKHPTLLLT----HI-EP 1042
+T P++ P N H M TR+K+ I + + +L + H+ EP
Sbjct: 891 KSTTTALPPNQTTIAATSNSRSQPPKNNHQMKTRSKNNITKPKTKTSLTVALTQPHLSEP 950
Query: 1043 TGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKY 1102
QA+K +W AM E +A +N+TW LVP + VGC+WVF+ K P+G I+KY
Sbjct: 951 NTVTQALKDKKWRFAMSDEFDAQQRNHTWDLVPPNPTQHLVGCRWVFKLKYLPNGLIDKY 1010
Query: 1103 KARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEE 1162
KARLVAKGF+Q G DY ETFSPV+K T+R VL +AV W ++QLDVNNAFL G L E
Sbjct: 1011 KARLVAKGFNQQYGVDYAETFSPVIKATTIRVVLDVAVKKNWPLKQLDVNNAFLQGTLTE 1070
Query: 1163 EVYMTQPPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFI 1221
EVYM QPPGF D PS VC+L KA+YGLKQAPRAW+ LK LL +GF +S D SLFI
Sbjct: 1071 EVYMAQPPGFVDKDRPSHVCRLRKAIYGLKQAPRAWYMELKQHLLNIGFVNSLADTSLFI 1130
Query: 1222 LHANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENG 1281
++LVYVDDI++TGS + ++ L FS+KD L YFLGIE + G
Sbjct: 1131 YSHGTTLLYLLVYVDDIIVTGSDHKSVSAVLSSLAERFSIKDPTDLHYFLGIEATRTNTG 1190
Query: 1282 SLLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVT 1341
L L Q+KY+ DLL K NM +A +A+P+ +S KLT +G ++D + +RS+VG LQY+
Sbjct: 1191 -LHLMQRKYMTDLLAKHNMLDAKPVATPLPTSPKLTLHGGTKLNDASEYRSVVGSLQYLA 1249
Query: 1342 VTRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFC 1401
TRP+I+++VN++ QF+ P DHW+A KR+LRYL GT HG+ +N + P+ L AF
Sbjct: 1250 FTRPDIAFAVNRLSQFMHQPTSDHWQAAKRVLRYLAGTTTHGIFLNSS---SPIHLHAFS 1306
Query: 1402 DADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQS 1461
DADWA D D ST+ I LG N ISW +KKQ V+RSS E+EYR++A A++E+ W+ S
Sbjct: 1307 DADWAGDSADYVSTNAYVIYLGRNPISWSSKKQRGVSRSSTESEYRAVANAASEIRWLCS 1366
Query: 1462 LLKEL--KVPTAIPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHI 1519
LL EL ++P P IFCDN+ + NPV HSR KH+ LD FVR + S+ L VSH+
Sbjct: 1367 LLTELHIRLPHG-PTIFCDNIGATYICANPVFHSRMKHIALDYHFVRGMIQSRALRVSHV 1425
Query: 1520 PAQYQVADILTKPLSASRFLELRNKLRV 1547
Q+AD LTK LS FL R+K+ V
Sbjct: 1426 STNDQLADALTKSLSRPHFLSARSKIGV 1453
>At4g02960 putative polyprotein of LTR transposon
Length = 1456
Score = 918 bits (2372), Expect = 0.0
Identities = 604/1574 (38%), Positives = 817/1574 (51%), Gaps = 212/1574 (13%)
Query: 31 KLDETNYLQWKQQVEGVLRGTKMVRHV--VSPQIPPVFLNDAAREAGTENPAYTEWEEQD 88
KL TNYL W +QV + G ++ + +P P DA NP YT W QD
Sbjct: 25 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTPMPPATIGTDAVPRV---NPDYTRWRRQD 81
Query: 89 SLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGTRS 148
L+ + IL IS S+ R + Q+W+ + QLR
Sbjct: 82 KLIYSAILGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQLR------------ 129
Query: 149 IAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISLDELES 208
FI R + L +G P+ H + +E VLE LP+++ P++ + ++ SL E+
Sbjct: 130 ---FITRF----DQLALLGKPMDHDEQVERVLENLPDDYKPVIDQIAAKDTPPSLTEIHE 182
Query: 209 QLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNN 268
+L+ +E++ A V + +TH + +N +N+ +Y + NN
Sbjct: 183 RLINRESKLLALNSAEVVPITANVVTHRNTNTNRNQNNRGDNRNYNNN----------NN 232
Query: 269 SSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVP 328
S + P+ +GSR + + GR CQIC GH A C
Sbjct: 233 RSNSWQPS---SSGSRSDNRQPKPYLGR-----------CQICSVQGHSAKRC------- 271
Query: 329 PQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSR 388
PQ + Q + +PF P
Sbjct: 272 PQLHQF---------------------------------QSTTNQQQSTSPFTPW----- 293
Query: 389 PPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGL 448
P+A + N ++N+ W DSGATHH+T D NNL ++G D + I +G +
Sbjct: 294 --QPRANLAVNSPYNANN----WLLDSGATHHITSDFNNLSFHQPYTGGDDVMIADGSTI 347
Query: 449 AINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKS 508
I GS S + + +L LN +L+VP+I KNL+SV + C N V EF VK
Sbjct: 348 PITHTGSASLPTS---SRSLDLNKVLYVPNIHKNLISVYRLCNTNRVSVEFFPASFQVKD 404
Query: 509 QDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQC 568
++ LL+G D+ LY++ +VS AS S AT
Sbjct: 405 LNTGVPLLQGKTKDE-LYEWPIASSQAVSMFASPCSKAT--------------------- 442
Query: 569 NNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLP--NKSFTD 626
H+S WH+RLGHP ++ SV+ N LP N S
Sbjct: 443 ---------HSS---------------WHSRLGHPSLAILNSVIS--NHSLPVLNPSHKL 476
Query: 627 F-CSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYT 685
CS C + KSH++P +S +KP E I+ D+W + + S Y Y++I VD ++RYT
Sbjct: 477 LSCSDCFINKSHKVPFSNSTITSSKPLEYIYSDVWS-SPILSIDNYRYYVIFVDHFTRYT 535
Query: 686 WIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCP 745
W++PLK KS TF FK++VE ++ I ++ +D GGEF +L+ GI+H + P
Sbjct: 536 WLYPLKQKSQVKDTFIIFKSLVENRFQTRIGTLYSDNGGEFVVLRDYLSQHGISHFTSPP 595
Query: 746 HTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFF 805
HT NG ERKHRHIVE GLTLL++A +P YW +AF A YLINRLP+P+L +SPF
Sbjct: 596 HTPEHNGLSERKHRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQLQSPFQ 655
Query: 806 LLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRM 864
L Q P+Y+ LK FGC+C+P+ RPYN HKLE +SK+C F+GYS + Y CL PTGR+
Sbjct: 656 KLFGQPPNYEKLKVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRL 715
Query: 865 FISKDVIFNEYKFPYSE----LFTSGQ--SSSPPTTSSDHTPLPSFLFPLNNKQC----- 913
+ S+ V F+E FP+S + TS + S S P S HT LP+ L C
Sbjct: 716 YTSRHVQFDERCFPFSTTNFGVSTSQEQRSDSAPNWPS-HTTLPTTPLVLPAPPCLGPHL 774
Query: 914 ---PTTQSSSTPTTTLHTAS---PHSSFPESNQSN----HHHSIQDTHASSHSNHHNISP 963
P SS +P T +S P SS + S H+ Q T A H ++ S
Sbjct: 775 DTSPRPPSSPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPT-AQPHQTQNSNSN 833
Query: 964 GPIF-NPTPISTHPPSP--------SPSSHSHNTYHSISV----EPVTSQPSTQAEP--- 1007
PI NP P S P SP SP S H S S+ P +S ST P
Sbjct: 834 SPILNNPNPNSPSPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSSSSTSTPPLPPVL 893
Query: 1008 --------HRIHPNNTHSMATRAKHGI---VQKRKHPTLLLTHIEPTGYRQAMKQPQWLQ 1056
+ P NTHSMATRAK GI QK + T L + EP QAMK +W Q
Sbjct: 894 PAPPIIQVNAQAPVNTHSMATRAKDGIRKPNQKYSYATSLAANSEPRTAIQAMKDDRWRQ 953
Query: 1057 AMQLEHEALMKNNTWTLVPLPADR-QAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMP 1115
AM E A + N+TW LVP P VGC+W+F K N DGS+N+YKARLVAKG++Q P
Sbjct: 954 AMGSEINAQIGNHTWDLVPPPPPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGYNQRP 1013
Query: 1116 GFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAV 1175
G DY ETFSPV+K ++R VL +AV W I+QLDVNNAFL G L +EVYM+QPPGF
Sbjct: 1014 GLDYAETFSPVIKSTSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDEVYMSQPPGFVDK 1073
Query: 1176 D-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVY 1234
D P VC+L KA+YGLKQAPRAW+ L++ LL +GF +S D SLF+L + +MLVY
Sbjct: 1074 DRPDYVCRLRKAIYGLKQAPRAWYVELRTYLLTVGFVNSISDTSLFVLQRGRSIIYMLVY 1133
Query: 1235 VDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDL 1294
VDDILITG+ L++ + L+ FS+K+ L YFLGIE G L LSQ++Y DL
Sbjct: 1134 VDDILITGNDTVLLKHTLDALSQRFSVKEHEDLHYFLGIEAKRVPQG-LHLSQRRYTLDL 1192
Query: 1295 LVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKV 1354
L + NM A +A+PMA+S KLT + + DPT +R IVG LQY+ TRP++SY+VN++
Sbjct: 1193 LARTNMLTAKPVATPMATSPKLTLHSGTKLPDPTEYRGIVGSLQYLAFTRPDLSYAVNRL 1252
Query: 1355 CQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRS 1414
Q++ P +DHW A+KR+LRYL GT HG+ + LSL A+ DADWA D DD S
Sbjct: 1253 SQYMHMPTDDHWNALKRVLRYLAGTPDHGIFLKKG---NTLSLHAYSDADWAGDTDDYVS 1309
Query: 1415 TSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIPQ 1474
T+G + LG + ISW +KKQ V RSS EAEYRS+A S+E+ WI SLL EL + + P
Sbjct: 1310 TNGYIVYLGHHPISWSSKKQKGVVRSSTEAEYRSVANTSSELQWICSLLTELGIQLSHPP 1369
Query: 1475 -IFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPL 1533
I+CDN+ L NPV HSR KH+ LD F+R +V S L V H+ Q+AD LTKPL
Sbjct: 1370 VIYCDNVGATYLCANPVFHSRMKHIALDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPL 1429
Query: 1534 SASRFLELRNKLRV 1547
S F K+ V
Sbjct: 1430 SRVAFQNFSRKIGV 1443
>At4g16870 retrotransposon like protein
Length = 1474
Score = 909 bits (2350), Expect = 0.0
Identities = 594/1607 (36%), Positives = 826/1607 (50%), Gaps = 203/1607 (12%)
Query: 2 SSPSSPVNNSETQRVTAAASKNFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHV---V 58
S+ P E T N KL NYL W Q+ +L G ++ H+ +
Sbjct: 4 SANGLPATTDEAIVFTPQTIFNINTSNVTKLTSNNYLMWSLQIHALLDGYELAGHLDGSI 63
Query: 59 SPQIPPVFLNDAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWD 118
P + N+ + NP YT W+ QD L+ + ++ IS + R + Q+W
Sbjct: 64 ETPAPTLTTNNVV----SANPQYTLWKRQDRLIFSALIGAISPPVQPLVSRATKASQIWK 119
Query: 119 EIHNYCYTQMRTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIET 178
+ N +QLR++++ + KGT++I E++ ++ + L +G P+ H + +E
Sbjct: 120 TLTNTYAKSSYDHIKQLRTQIKQLKKGTKTIDEYVLSHTTLLDQLAILGKPMEHEEQVER 179
Query: 179 VLEALPEEFNPIVATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAEN 238
+LE LPE++ +V + + S+ E+ +L+ EA K +S +L + N
Sbjct: 180 ILEGLPEDYKTVVDQIEGKDNTPSITEIHERLINHEA-----KLLSTAALSSSSLPMSAN 234
Query: 239 SGEKNGHNQPQTGSYPDQQFNISGNPT-GNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRG 297
++ HN + N + N T GN + + P+ ++G R F+
Sbjct: 235 VAQQRHHNNNRNN-------NQNKNRTQGNTYTNNWQPSANNKSGQRP-------FKPYL 280
Query: 298 GRNFGRGNIQCQICYKTGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGT 357
G+ CQIC GH A C ++ P + S + P
Sbjct: 281 GK--------CQICNVQGHSARRCPQLQAMQPS-----------SSSSASTFTP------ 315
Query: 358 HSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNGWYPDSGA 417
W + N + GAP N W DSGA
Sbjct: 316 ----WQP---RANLAMGAPYTA-----------------------------NNWLLDSGA 339
Query: 418 THHVTPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTT--LTLNNLLH 475
THH+T D N L ++G D M I +G L I GS F P+ LTLN +L+
Sbjct: 340 THHITSDLNALALHQPYNGDDVM-IADGTSLKITKTGST-----FLPSNARDLTLNKVLY 393
Query: 476 VPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPS 535
VP I KNLVSV + C N V EF VK ++ +LL+G D+ LY++ P
Sbjct: 394 VPDIQKNLVSVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDE-LYEW-----PV 447
Query: 536 VSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKV 595
+ A T + T+ S + +S
Sbjct: 448 TNPKA----------------------------------TALFTTPSPKTTLSS------ 467
Query: 596 WHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDF-CSACCLGKSHRLPSVSSKTVYNKPFEL 654
WH+RLGHP ++ +++ + + + CS C + KSH+LP S P E
Sbjct: 468 WHSRLGHPSSSILNTLISKFSLPVSVSASNKLACSDCFINKSHKLPFSISSIKSTSPLEY 527
Query: 655 IFCDLWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLP 714
IF D+W + + S Y Y+L+ VD ++RYTW++PL+ KS TF FK +VE ++
Sbjct: 528 IFSDVW-MSPILSPDNYKYYLVLVDHHTRYTWLYPLQQKSQVKSTFIAFKALVENRFQAK 586
Query: 715 IKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKL 774
I+++ +D GGEF +FL + GI+H + PHT NG ERKHRHIVETGLTLL A +
Sbjct: 587 IRTLYSDNGGEFIALREFLVSNGISHLTSPPHTPEHNGLSERKHRHIVETGLTLLTQASV 646
Query: 775 PLHYWDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNH 834
P YW +AF A YLINR+P+P+L+ +SPF L P+Y+ L+ FGC CFP+ RPY ++
Sbjct: 647 PREYWPYAFAAAVYLINRMPTPVLSMESPFQKLFGSKPNYERLRVFGCLCFPWLRPYTHN 706
Query: 835 KLELRSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSEL--------FTS 885
KLE RS+ CVFLGYS + Y C D R++ S+ V+F+E FP+S L T
Sbjct: 707 KLEERSRRCVFLGYSLTQTAYLCFDVEHKRLYTSRHVVFDEASFPFSNLTSQNSLPTVTF 766
Query: 886 GQSSSPPTTS--SDHTPLPSFLFP----LNNKQCPTT----QSSSTPTTTLHTASPHSSF 935
QSSSP T S + LPS L L+ +Q P T SS PTT+ SPH S
Sbjct: 767 EQSSSPLVTPILSSSSVLPSCLSSPCTVLHQQQPPVTTPNSPHSSQPTTSPAPLSPHRST 826
Query: 936 PESNQSNHHHSIQDTHASSHS--------NHHNISP-------GPIFNPTPISTHPPSPS 980
Q S +SS S N + P GP+ NPT + P P+
Sbjct: 827 TMDFQVPQVRSSSPLLSSSSSLNSEPTAPNENGPEPEAQSPPIGPLSNPTHEAFIGPLPN 886
Query: 981 PSSHSHNTY------HSISVEP--VTSQPSTQAEPHRIH-----PNNTHSMATRAKHGIV 1027
P+ + N H V+P T+ P+ H N H+M TRAK+ I
Sbjct: 887 PNRNPTNEIEPTPAPHPKPVKPTTTTTTPNRTTVSDASHQPTAPQQNQHNMKTRAKNNIK 946
Query: 1028 QKRKHPTLLLT-----HIEPTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQA 1082
+ +L T EPT QA+K +W AM E +A +N+TW LVP +
Sbjct: 947 KPNTKFSLTATLPNRSPSEPTNVTQALKDKKWRFAMSDEFDAQQRNHTWDLVP-HESQLL 1005
Query: 1083 VGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTN 1142
VGCKWVF+ K P+G+I+KYKARLVAKGF+Q G DY ETFSPV+K T+R VL +AV
Sbjct: 1006 VGCKWVFKLKYLPNGAIDKYKARLVAKGFNQQYGVDYAETFSPVIKSTTIRLVLDVAVKK 1065
Query: 1143 KWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERL 1201
W I+QLDVNNAFL G L EEVYM QPPGF D P+ VC+L KA+YGLKQAPRAW+ L
Sbjct: 1066 DWEIKQLDVNNAFLQGTLTEEVYMAQPPGFIDKDRPTHVCRLRKAIYGLKQAPRAWYMEL 1125
Query: 1202 KSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSL 1261
K L +GF +S D SLFI ++LVYVDDI++TGS S I ++ L FS+
Sbjct: 1126 KQHLFNIGFVNSLSDASLFIYCHGTTFVYVLVYVDDIIVTGSDKSSIDAVLTSLAERFSI 1185
Query: 1262 KDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGS 1321
KD L YFLGIE ++ G L L Q+KYI+DLL K NMA+A + +P+ +S KLT +G
Sbjct: 1186 KDPTDLHYFLGIEATRTKQG-LHLMQRKYIKDLLAKHNMADAKPVLTPLPTSPKLTLHGG 1244
Query: 1322 NHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIH 1381
++D + +RS+VG LQY+ TRP+I+Y+VN++ Q + P EDHW+A KR+LRYL GT
Sbjct: 1245 TKLNDASEYRSVVGSLQYLAFTRPDIAYAVNRLSQLMPQPTEDHWQAAKRVLRYLAGTST 1304
Query: 1382 HGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSS 1441
HG+ ++ PL+L AF DADWA D DD ST+ I LG N ISW +KKQ VARSS
Sbjct: 1305 HGIFLDTT---SPLNLHAFSDADWAGDSDDYVSTNAYVIYLGKNPISWSSKKQRGVARSS 1361
Query: 1442 AEAEYRSLAQASAEVLWIQSLLKELKVPTAI-PQIFCDNLSTVSLAHNPVLHSRTKHMEL 1500
E+EYR++A A++EV W+ SLL +L + I P IFCDN+ L NPV HSR KH+ +
Sbjct: 1362 TESEYRAVANAASEVKWLCSLLSKLHIRLPIRPSIFCDNIGATYLCANPVFHSRMKHIAI 1421
Query: 1501 DIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLELRNKLRV 1547
D FVR + S L VSH+ + Q+AD LTKPLS + F R K+ V
Sbjct: 1422 DYHFVRNMIQSGALRVSHVSTRDQLADALTKPLSRAHFQSARFKIGV 1468
>At1g53810
Length = 1522
Score = 867 bits (2241), Expect = 0.0
Identities = 549/1576 (34%), Positives = 808/1576 (50%), Gaps = 190/1576 (12%)
Query: 23 NFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSPQIPPVFLNDAAREAGTE---NP 79
N ++V L++ NY+ WK Q E L G ++ V P T NP
Sbjct: 10 NISNCVTVTLNQQNYILWKSQFESFLSGQGLLGFVTGSISAPAQTRSVTHNNVTSEEPNP 69
Query: 80 AYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSEL 139
+ W + D ++ +W+L + + +LS V SHQVW + N+ +R +L+ L
Sbjct: 70 EFYTWHQTDQVVKSWLLGSFAEDILSVVVNCFTSHQVWLTLANHFNRVSSSRLFELQRRL 129
Query: 140 RTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTE 199
+T+ K ++ F+ ++ I + L S+G PV + I + L L E+ PI T+ + +
Sbjct: 130 QTLEKKDNTMEVFLKDLKHICDQLASVGSPVPEKMKIFSALNGLGREYEPIKTTIENSVD 189
Query: 200 V---ISLDELESQLLTQEARNEKF-KKALVGETASVNLTHAENSGEKNGHNQPQTGSYPD 255
+SLDE+ S+L + R + + + + + N+TH++
Sbjct: 190 SNPSLSLDEVASKLRGYDDRLQSYVTEPTISPHVAFNVTHSD------------------ 231
Query: 256 QQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTG 315
S Y N N G + G G RGRG
Sbjct: 232 -------------SGYYHNNNRGKGRSNSGSGKSSFSTRGRG------------------ 260
Query: 316 HDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHS-NVWMQGVGQRNPSYG 374
+H+ P G+ GN G G H+ W +
Sbjct: 261 ------FHQQISPTS--------GSQAGNSGLVCQICGKAGHHALKCWHR---------- 296
Query: 375 APRAPFPPQFGNS--RPPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAA 432
F NS P A T + ++ + W PDS A+ HVT + + L +
Sbjct: 297 ---------FDNSYQHEDLPMALATMRITDVTDHHGHEWIPDSAASAHVTNNRHVLQQSQ 347
Query: 433 SFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKD 492
+ GSD + + +G L I GS S +S + + L +L P I K+L+SVS+ D
Sbjct: 348 PYHGSDSIMVADGNFLPITHTGSGSIASS---SGKIPLKEVLVCPDIVKSLLSVSKLTSD 404
Query: 493 NNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSL 552
EF ++ + + + K+L+ G DGLY ++P + + T +S+ +
Sbjct: 405 YPCSVEFDADSVRINDKATKKLLVMGR-NRDGLYSLEEPKLQVLYSTRQNSASS------ 457
Query: 553 NNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVM 612
+VWH RLGH + EV+ +
Sbjct: 458 -----------------------------------------EVWHRRLGHANAEVLHQLA 476
Query: 613 KLCNQQLPNKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYS 672
+ + NK C AC LGKS RLP + S ++P E I CDLWGP+ S G+
Sbjct: 477 SSKSIIIINKVVKTVCEACHLGKSTRLPFMLSTFNASRPLERIHCDLWGPSPTSSVQGFR 536
Query: 673 YFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEF--RPFT 730
Y+++ +D YSR+TW +PLKLKS TF F+ +VE Q IK Q DGGGEF F
Sbjct: 537 YYVVFIDHYSRFTWFYPLKLKSDFFSTFVMFQKLVENQLGHKIKIFQCDGGGEFISSQFL 596
Query: 731 QFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLI 790
+ L GI ++CP+T QNG ERKHRHIVE GL+++ +KLPL YW +F TA ++I
Sbjct: 597 KHLQDHGIQQNMSCPYTPQQNGMAERKHRHIVELGLSMIFQSKLPLKYWLESFFTANFVI 656
Query: 791 NRLPSPIL-NNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYS 849
N LP+ L NN+SP+ L+ + P+Y L+ FGC+C+P R Y + K + RS +CVFLGY+
Sbjct: 657 NLLPTSSLDNNESPYQKLYGKAPEYSALRVFGCACYPTLRDYASTKFDPRSLKCVFLGYN 716
Query: 850 PSHKGYKCL-DPTGRMFISKDVIFNEYKFPYSELFTS--GQSSSP------------PTT 894
+KGY+CL PTGR++IS+ V+F+E P+ +++ Q +P T
Sbjct: 717 EKYKGYRCLYPPTGRIYISRHVVFDENTHPFESIYSHLHPQDKTPLLEAWFKSFHHVTPT 776
Query: 895 SSDHTPLPSFLFPLNNKQCPTTQSSSTPTTTL-HTASPHSSFPESNQSNHHHSIQDTHAS 953
D + P P Q TT S+ P + TA P++S +++Q N S+
Sbjct: 777 QPDQSRYPVSSIP----QPETTDLSAAPASVAAETAGPNAS-DDTSQDNETISVVSGSPE 831
Query: 954 SHSNHHNISPGPIFN-PTPISTHPPSPSPSSHSHNTYHSISVEPVTSQPSTQAEPHRIHP 1012
+ + S G ++ PT S+HP SP+ SS + + S P+ P+ Q +
Sbjct: 832 RTTGLDSASIGDSYHSPTADSSHP-SPARSSPASSPQGS----PIQMAPAQQVQAP---V 883
Query: 1013 NNTHSMATRAKHGIVQKRKHPTLLLTHI---EPTGYRQAMKQPQWLQAMQLEHEALMKNN 1069
N H+M TR K GI + K LL + EP +A+K P W AMQ E +
Sbjct: 884 TNEHAMVTRGKEGISKPNKRYVLLTHKVSIPEPKTVTEALKHPGWNNAMQEEMGNCKETE 943
Query: 1070 TWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKP 1129
TWTLVP + +G WVFRTK + DGS++K KARLVAKGF Q G DY ET+SPVV+
Sbjct: 944 TWTLVPYSPNMNVLGSMWVFRTKLHADGSLDKLKARLVAKGFKQEEGIDYLETYSPVVRT 1003
Query: 1130 VTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGF-EAVDPSLVCKLNKALY 1188
TVR +L +A KW ++Q+DV NAFL+G L E VYM QP GF + P VC L+K+LY
Sbjct: 1004 PTVRLILHVATVLKWELKQMDVKNAFLHGDLTETVYMRQPAGFVDKSKPDHVCLLHKSLY 1063
Query: 1189 GLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSASLI 1248
GLKQ+PRAWF+R + LL+ GF S DPSLF+ +N +L+YVDD++ITG+++ +
Sbjct: 1064 GLKQSPRAWFDRFSNFLLEFGFICSLFDPSLFVYSSNNDVILLLLYVDDMVITGNNSQSL 1123
Query: 1249 QQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIAS 1308
L+ LN EF +KD+G++ YFLGI++ + +G L +SQ+KY +DLL+ A+MAN + + +
Sbjct: 1124 THLLAALNKEFRMKDMGQVHYFLGIQIQ-TYDGGLFMSQQKYAEDLLITASMANCSPMPT 1182
Query: 1309 PMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKA 1368
P+ SDPT+FRS+ G LQY+T+TRP+I ++VN VCQ + P +
Sbjct: 1183 PLPLQLDRVSNQDEVFSDPTYFRSLAGKLQYLTLTRPDIQFAVNFVCQKMHQPSVSDFNL 1242
Query: 1369 VKRILRYLKGTIHHGLLINP------APMHQPLSLTAFCDADWASDPDDRRSTSGACILL 1422
+KRILRY+KGT+ G+ N + L+A+ D+D+A+ + RRS G C +
Sbjct: 1243 LKRILRYIKGTVSMGIQYNSNSSSVVSAYESDYDLSAYSDSDYANCKETRRSVGGYCTFM 1302
Query: 1423 GPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTA-IPQIFCDNLS 1481
G N+ISW +KKQ V+RSS EAEYRSL++ ++E+ W+ S+L+E+ V P++FCDNLS
Sbjct: 1303 GQNIISWSSKKQPTVSRSSTEAEYRSLSETASEIKWMSSILREIGVSLPDTPELFCDNLS 1362
Query: 1482 TVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLEL 1541
V L NP H+RTKH ++D ++RE+V K L+V HIP Q+ADI TK L F L
Sbjct: 1363 AVYLTANPAFHARTKHFDVDHHYIRERVALKTLVVKHIPGHLQLADIFTKSLPFEAFTRL 1422
Query: 1542 RNKLRVSDP--MSLRG 1555
R KL V P SLRG
Sbjct: 1423 RFKLGVDFPPTPSLRG 1438
>At4g10690 retrotransposon like protein
Length = 1515
Score = 860 bits (2221), Expect = 0.0
Identities = 556/1524 (36%), Positives = 801/1524 (52%), Gaps = 204/1524 (13%)
Query: 78 NPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRS 137
N + +W D L+ WI ++S L + L + +VW + TR L+
Sbjct: 66 NQEFLKWTRIDQLVKAWIFGSLSEEALKVVIGLNSAQEVWLGLARRFNRFSTTRKYDLQK 125
Query: 138 ELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQ 197
L T +K +++ +++ +++I + L SIG PV ++ I VL L +E+ I +
Sbjct: 126 RLGTCSKAGKTMDAYLSEVKNICDQLDSIGFPVTEQEKIFGVLNGLGKEYESIATVIEHS 185
Query: 198 TEVIS---LDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYP 254
+V D++ +L T F L TA+ +T P Y
Sbjct: 186 LDVYPGPCFDDVVYKLTT-------FDDKLSTYTANSEVT-------------PHLAFYT 225
Query: 255 DQQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKT 314
D+ ++ GN NNS GGR G+ FRGRG
Sbjct: 226 DKSYSSRGN---NNSR-------GGRYGN---------FRGRGS---------------- 250
Query: 315 GHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYG 374
Y S G F GSG +G G+ Q YG
Sbjct: 251 -------------------YSSRGRGFHQQFGSGSNNGSGNGSKPTC------QICRKYG 285
Query: 375 APRAPFPPQFGNSRPPA--PQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAA 432
+F + P P A+ S + + ++ W PDS AT H+T + L ++
Sbjct: 286 HSAFKCYTRFEENYLPEDLPNAFAAMRVSDQNQASSHEWLPDSAATAHITNTTDGLQNSQ 345
Query: 433 SFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKD 492
++SG D + +GNG L I IG++ + TL L ++L P ITK+L+SVS+ D
Sbjct: 346 TYSGDDSVIVGNGDFLPITHIGTIPLNIS---QGTLPLEDVLVCPGITKSLLSVSKLTDD 402
Query: 493 NNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQF-DQPYVPSVSRTASSSSVATSSLS 551
F F S+ +K + + ++L +G+ GLY D P+ S
Sbjct: 403 YPCSFTFDSDSVVIKDKRTQQLLTQGNK-HKGLYVLKDVPFQTYYS-------------- 447
Query: 552 LNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSV 611
+R Q SS+D +VWH RLGHP+ EV++ +
Sbjct: 448 ------------TRQQ--------------SSDD--------EVWHQRLGHPNKEVLQHL 473
Query: 612 MKLCNQQLPNKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGY 671
+K + NK+ ++ C AC +GK RLP V+S+ V ++P E I CDLWGPA V S G+
Sbjct: 474 IKT-KAIVVNKTSSNMCEACQMGKVCRLPFVASEFVSSRPLERIHCDLWGPAPVTSAQGF 532
Query: 672 SYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEF--RPF 729
Y++I +D YSR+TW +PLKLKS F F+ +VE QY I Q DGGGEF F
Sbjct: 533 QYYVIFIDNYSRFTWFYPLKLKSDFFSVFVLFQQLVENQYQHKIAMFQCDGGGEFVSYKF 592
Query: 730 TQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYL 789
L + GI ++CPHT QNG ER+HR++ E GL+L+ ++K+P W AF T+ +L
Sbjct: 593 VAHLASCGIKQLISCPHTPQQNGIAERRHRYLTELGLSLMFHSKVPHKLWVEAFFTSNFL 652
Query: 790 INRLPSPILN-NKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGY 848
N LPS L+ NKSP+ +LH P Y L+ FG +C+P+ RPY +K + +S CVFLGY
Sbjct: 653 SNLLPSSTLSDNKSPYEMLHGTPPVYTALRVFGSACYPYLRPYAKNKFDPKSLLCVFLGY 712
Query: 849 SPSHKGYKCLDP-TGRMFISKDVIFNEYKFPYSE------------LFTSGQSSSPPTTS 895
+ +KGY+CL P TG+++I + V+F+E KFPYS+ LFT+ Q T
Sbjct: 713 NNKYKGYRCLHPPTGKVYICRHVLFDERKFPYSDIYSQFQTISGSPLFTAWQKGFSSTAL 772
Query: 896 SDHTP---LPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSI----- 947
S TP + +FP T SSS PT + ++ P+ + + H +
Sbjct: 773 SRETPSTNVEDIIFP------SATVSSSVPTGCAPNIAETATAPDVDVAAAHDMVVPPSP 826
Query: 948 --------QDTHASSHSNHHNISPGPIFNPTPISTHPPSPSPSSHSHNTYHSISVEPVTS 999
Q ++S NH++ + T IS+ + +P S + + + P+ S
Sbjct: 827 ITSTSLPTQPEESTSDQNHYSTD-----SETAISS---AMTPQSINVSLFEDSDFPPLQS 878
Query: 1000 Q-PSTQAEPHRIHPNNTHSMATRAKHGIVQKRKHPTLLLT---HIEPTGYRQAMKQPQWL 1055
ST A P HP M TRAK GI + L + EP ++A+K W
Sbjct: 879 VISSTTAAPETSHP-----MITRAKSGITKPNPKYALFSVKSNYPEPKSVKEALKDEGWT 933
Query: 1056 QAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMP 1115
AM E + + +TW LVP + +GCKWVF+TK N DGS+++ KARLVA+G+ Q
Sbjct: 934 NAMGEEMGTMHETDTWDLVPPEMVDRLLGCKWVFKTKLNSDGSLDRLKARLVARGYEQEE 993
Query: 1116 GFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAV 1175
G DY ET+SPVV+ TVRS+L +A NKW ++QLDV NAFL+ L+E V+MTQPPGFE
Sbjct: 994 GVDYVETYSPVVRSATVRSILHVATINKWSLKQLDVKNAFLHDELKETVFMTQPPGFEDP 1053
Query: 1176 D-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVY 1234
P VCKL KA+Y LKQAPRAWF++ S LLK GF S DPSLF+ + F+L+Y
Sbjct: 1054 SRPDYVCKLKKAIYDLKQAPRAWFDKFSSYLLKYGFICSFSDPSLFVYLKGRDVMFLLLY 1113
Query: 1235 VDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDL 1294
VDD+++TG++ L+QQL+ L+ EF +KD+G L YFLGI+ HY +G L LSQ+KY DL
Sbjct: 1114 VDDMILTGNNDVLLQQLLNILSTEFRMKDMGALHYFLGIQAHYHNDG-LFLSQEKYTSDL 1172
Query: 1295 LVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKV 1354
LV A M++ + + +P+ L + + +PT+FR + G LQY+T+TRP+I ++VN V
Sbjct: 1173 LVNAGMSDCSSMPTPL--QLDLLQGNNKPFPEPTYFRRLAGKLQYLTLTRPDIQFAVNFV 1230
Query: 1355 CQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRS 1414
CQ + AP + +KRIL YLKGT+ G+ ++ + L + D+DWA D RRS
Sbjct: 1231 CQKMHAPTMSDFHLLKRILHYLKGTMTMGINLS---SNTDSVLRCYSDSDWAGCKDTRRS 1287
Query: 1415 TSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVP-TAIP 1473
T G C LG N+ISW AK+ V++SS EAEYR+L+ A++EV WI LL+E+ +P IP
Sbjct: 1288 TGGFCTFLGYNIISWSAKRHPTVSKSSTEAEYRTLSFAASEVSWIGFLLQEIGLPQQQIP 1347
Query: 1474 QIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPL 1533
+++CDNLS V L+ NP LHSR+KH ++D ++VRE+V L V HIPA Q+ADI TK L
Sbjct: 1348 EMYCDNLSAVYLSANPALHSRSKHFQVDYYYVRERVALGALTVKHIPASQQLADIFTKSL 1407
Query: 1534 SASRFLELRNKLRVSDP--MSLRG 1555
+ F +LR KL V P SLRG
Sbjct: 1408 PQAPFCDLRFKLGVVLPPDTSLRG 1431
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 834 bits (2154), Expect = 0.0
Identities = 543/1546 (35%), Positives = 774/1546 (49%), Gaps = 209/1546 (13%)
Query: 28 ISVKLDETNYLQWKQQVEGVLRGTKMVRHVV----SPQIPPVFLNDAAREAGTENPAYTE 83
+++KL ++NYL WK Q E +L K++ V +P + +N NP Y
Sbjct: 17 VTLKLTDSNYLLWKTQFESLLSSQKLIGFVNGAVNAPSQSRLVVNGEVTSE-EPNPLYES 75
Query: 84 WEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTIT 143
W D L+ +W+ T+S +L L S Q+W + R LR L+ ++
Sbjct: 76 WFCTDQLVRSWLFGTLSEEVLGHVHNLSTSRQIWVSLAENFNKSSVAREFSLRQNLQLLS 135
Query: 144 KGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISL 203
K + + + ++I ++L SIG PV I L L +++PI + S +
Sbjct: 136 KKEKPFSVYCREFKTICDALSSIGKPVDESMKIFGFLNGLGRDYDPITTVIQSSLSKLPT 195
Query: 204 DELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGN 263
+ E + K E ASV P FNI +
Sbjct: 196 PTFND--VVSEVQGFDSKLQSYEEAASVT---------------------PHLAFNIERS 232
Query: 264 PTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYH 323
+G S +NPN GR GR G+N GRG
Sbjct: 233 ESG---SPQYNPNQKGR--------------GRSGQNKGRG------------------- 256
Query: 324 RLSVPPQYEGYGSLGGNFGGNLGSGY--GPATGFGTHSNVWMQGVGQRNPSYGAPRAPFP 381
GY + G F + S GP + V Q G
Sbjct: 257 ---------GYSTRGRGFSQHQSSPQVSGP------------RPVCQICGRTGHTALKCY 295
Query: 382 PQFGNSRPPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFSGSDQMY 441
+F N+ QA+ T S + W+PDS AT HVT N L A + G D +
Sbjct: 296 NRFDNNYQAEIQAFSTLRVSDDTGK---EWHPDSAATAHVTSSTNGLQSATEYEGDDAVL 352
Query: 442 IGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHS 501
+G+G L I GS + S N + LN +L VP+I K+L+SVS+ C D F +
Sbjct: 353 VGDGTYLPITHTGSTTIKSS---NGKIPLNEVLVVPNIQKSLLSVSKLCDDYPCGVYFDA 409
Query: 502 NICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSL 561
N + + K++ G + GLY + ++
Sbjct: 410 NKVCIIDLQTQKVVTTGPRRN-GLYVLENQEFVAL------------------------- 443
Query: 562 SLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPN 621
S QC + +VWH+RLGH + + ++ + Q+
Sbjct: 444 -YSNRQC---------------------AATEEVWHHRLGHANSKALQHLQNSKAIQINK 481
Query: 622 KSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAY 681
+ C C +GKS RLP + S + P + I CDLWGP+ V S+ G Y+ I VD Y
Sbjct: 482 SRTSPVCEPCQMGKSSRLPFLISDSRVLHPLDRIHCDLWGPSPVVSNQGLKYYAIFVDDY 541
Query: 682 SRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEF--RPFTQFLTTLGIT 739
SRY+W +PL KS L F +F+ +VE Q N IK Q+DGGGEF L+ GI
Sbjct: 542 SRYSWFYPLHNKSEFLSVFISFQKLVENQLNTKIKVFQSDGGGEFVSNKLKTHLSEHGIH 601
Query: 740 HRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILN 799
HR++CP+T QNG ERKHRH+VE GL++L ++ P +W +F TA Y+INRLPS +L
Sbjct: 602 HRISCPYTPQQNGLAERKHRHLVELGLSMLFHSHTPQKFWVESFFTANYIINRLPSSVLK 661
Query: 800 NKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCL- 858
N SP+ L + PDY L+ FG +C+P RP +K + RS +CVFLGY+ +KGY+C
Sbjct: 662 NLSPYEALFGEKPDYSSLRVFGSACYPCLRPLAQNKFDPRSLQCVFLGYNSQYKGYRCFY 721
Query: 859 DPTGRMFISKDVIFNEYKFPYSELFTS--GQSSSPPTTSSDHTPLPSFLFPLNNKQCPTT 916
PTG+++IS++VIFNE + P+ E + S Q S+P + H + P
Sbjct: 722 PPTGKVYISRNVIFNESELPFKEKYQSLVPQYSTPLLQAWQHNKISEISVP--------- 772
Query: 917 QSSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTHASSHSNHHNISPGPIFNPTPISTHP 976
A+P F + N T+A S P P
Sbjct: 773 ------------AAPVQLFSKPIDLN-------TYAGSQVTEQLTDPEPT---------- 803
Query: 977 PSPSPSSHSHNTYHSISVEPVTSQPSTQAEPHRIHPNNTHSMATRAKHGIVQKRKHPTLL 1036
S+N V PV + + E N+H+M TR+K GI + L+
Sbjct: 804 --------SNNEGSDEEVNPVAEEIAANQE----QVINSHAMTTRSKAGIQKPNTRYALI 851
Query: 1037 LTHI---EPTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQ 1093
+ + EP AMK P W +A+ E + +TW+LVP D + KWVF+TK
Sbjct: 852 TSRMNTAEPKTLASAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDMNILSSKWVFKTKL 911
Query: 1094 NPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNN 1153
+PDGSI+K KARLVAKGF Q G DY ETFSPVV+ T+R VL ++ + W I+QLDV+N
Sbjct: 912 HPDGSIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLDVSTSKGWPIKQLDVSN 971
Query: 1154 AFLNGYLEEEVYMTQPPGF-EAVDPSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCS 1212
AFL+G L+E V+M QP GF + P+ VC+L KA+YGLKQAPRAWF+ + LL GF
Sbjct: 972 AFLHGELQEPVFMYQPSGFIDPQKPTHVCRLTKAIYGLKQAPRAWFDTFSNFLLDYGFVC 1031
Query: 1213 SKCDPSLFILHANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLG 1272
SK DPSLF+ H + ++L+YVDDIL+TGS SL++ L++ L FS+KDLG YFLG
Sbjct: 1032 SKSDPSLFVCHQDGKILYLLLYVDDILLTGSDQSLLEDLLQALKNRFSMKDLGPPRYFLG 1091
Query: 1273 IEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRS 1332
I++ NG L L Q Y D+L +A M++ N + +P+ +L S ++PT+FRS
Sbjct: 1092 IQIEDYANG-LFLHQTAYATDILQQAGMSDCNPMPTPLPQ--QLDNLNSELFAEPTYFRS 1148
Query: 1333 IVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMH 1392
+ G LQY+T+TRP+I ++VN +CQ + +P + +KRILRY+KGTI GL P +
Sbjct: 1149 LAGKLQYLTITRPDIQFAVNFICQRMHSPTTSDFGLLKRILRYIKGTIGMGL---PIKRN 1205
Query: 1393 QPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQA 1452
L+L+A+ D+D A + RRST+G CILLG NLISW AK+Q V+ SS EAEYR+L A
Sbjct: 1206 STLTLSAYSDSDHAGCKNTRRSTTGFCILLGSNLISWSAKRQPTVSNSSTEAEYRALTYA 1265
Query: 1453 SAEVLWIQSLLKELKVPTAIP-QIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVIS 1511
+ E+ WI LL++L +P +P Q++CDNLS V L+ NP LH+R+KH + D ++RE+V
Sbjct: 1266 AREITWISFLLRDLGIPQYLPTQVYCDNLSAVYLSANPALHNRSKHFDTDYHYIREQVAL 1325
Query: 1512 KDLIVSHIPAQYQVADILTKPLSASRFLELRNKLRV--SDPMSLRG 1555
+ HI A +Q+AD+ TK L F++LR+KL V S SLRG
Sbjct: 1326 GLIETQHISATFQLADVFTKSLPRRAFVDLRSKLGVSGSPTPSLRG 1371
>At4g27210 putative protein
Length = 1318
Score = 768 bits (1984), Expect = 0.0
Identities = 463/1190 (38%), Positives = 651/1190 (53%), Gaps = 144/1190 (12%)
Query: 396 ITGNESTSSNSFNNG----------WYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNG 445
IT ++ + NS+N G W PDS AT HVT +L + + G+D + + +G
Sbjct: 152 ITTSDDSYRNSYNRGKDVTDQHGNEWLPDSAATAHVTNSPRSLQQSQPYHGTDAIMVDDG 211
Query: 446 QGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICY 505
L I GS + +S + T+ L ++L PSITK+L+S+S+ +D EF +
Sbjct: 212 NYLPITHTGSTNLASS---SGTVPLTDVLVCPSITKSLLSMSKLTQDFPCTVEFEYDGVR 268
Query: 506 VKSQDSTKILLKGHLGDDGLY------QFDQPYVPSVSRTASSSSVATSSLSLNNCFSPS 559
V + + K+LL G DGLY QF Q + + R+AS
Sbjct: 269 VNDKATKKLLLMGS-NRDGLYCLKDDKQF-QAFFSTRQRSASD----------------- 309
Query: 560 SLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQL 619
+VWH RLGHPH ++++
Sbjct: 310 ----------------------------------EVWHRRLGHPHPQILQ---------- 325
Query: 620 PNKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVD 679
P E + CDLWGP ++ S G+ Y+ + +D
Sbjct: 326 -------------------------------PLERVHCDLWGPTTITSVQGFRYYAVFID 354
Query: 680 AYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEF--RPFTQFLTTLG 737
YSR++WI+PLKLKS F F +VE Q + I Q DGGGEF F Q L + G
Sbjct: 355 HYSRFSWIYPLKLKSDFYNIFLAFHKLVENQLSQKISVFQCDGGGEFVSHKFLQHLQSHG 414
Query: 738 ITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPI 797
I +L+CPHT QNG ERKHRH+VE GL++L + +P +W AF TA +LIN LP+
Sbjct: 415 IQQQLSCPHTPQQNGLAERKHRHLVELGLSMLFQSHVPHKFWVEAFFTANFLINLLPTSA 474
Query: 798 LNNK-SPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYK 856
L SP+ L+ + PDY L+SFG +CFP R Y +K S +CVFLGY+ +KGY+
Sbjct: 475 LKESISPYEKLYDKKPDYTSLRSFGSACFPTLRDYAENKFNPCSLKCVFLGYNEKYKGYR 534
Query: 857 CL-DPTGRMFISKDVIFNEYKFPYSELFTSGQSSSPPTTSSDHTPLPSFLFPLNNKQCPT 915
CL PTGR++IS+ VIF+E +P+S + TPL + ++ P+
Sbjct: 535 CLYPPTGRLYISRHVIFDESVYPFSHTYKH-------LHPQPRTPLLAAWLRSSDSPAPS 587
Query: 916 TQSSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTHASSHSNHHNISPGPIFNPTPISTH 975
T +S + + L T++ P+ ++ ++ + SH+++ P F+ + +T
Sbjct: 588 TSTSPSSRSPLFTSADFPPLPQ-RKTPLLPTLVPISSVSHASNITTQQSPDFD-SERTTD 645
Query: 976 PPSPSPSSHSHNTYHSISVEPVTSQPSTQAEPHRIHPN-NTHSMATRAKHGIVQKRKHPT 1034
S S SH++ E Q S H+ H + N H M TRAK GI +
Sbjct: 646 FDSASIGDSSHSSQAGSDSEETIQQASVNV--HQTHASTNVHPMVTRAKVGISKPNPRYV 703
Query: 1035 LL---LTHIEPTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRT 1091
L +++ EP A+K P W AM E + TW+LVP +D +G KWVFRT
Sbjct: 704 FLSHKVSYPEPKTVTAALKHPGWTGAMTEEIGNCSETQTWSLVPYKSDMHVLGSKWVFRT 763
Query: 1092 KQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDV 1151
K + DG++NK KAR+VAKGF Q G DY ET+SPVV+ TVR VL LA W I+Q+DV
Sbjct: 764 KLHADGTLNKLKARIVAKGFLQEEGIDYLETYSPVVRTPTVRLVLHLATALNWDIKQMDV 823
Query: 1152 NNAFLNGYLEEEVYMTQPPGFEAVDPSL---VCKLNKALYGLKQAPRAWFERLKSTLLKL 1208
NAFL+G L+E VYMTQP GF VDPS VC L+K++YGLKQ+PRAWF++ + LL+
Sbjct: 824 KNAFLHGDLKETVYMTQPAGF--VDPSKPDHVCLLHKSIYGLKQSPRAWFDKFSTFLLEF 881
Query: 1209 GFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLD 1268
GF SK DPSLFI N + +L+YVDD++ITG+S+ + L+ LN EF + D+G+L
Sbjct: 882 GFFCSKSDPSLFIYAHNNNLILLLLYVDDMVITGNSSQTLTSLLAALNKEFRMTDMGQLH 941
Query: 1269 YFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKYGSNHVSDPT 1328
YFLGI+V +NG L +SQ+KY +DLL+ A+M + + +P+ + SDPT
Sbjct: 942 YFLGIQVQRQQNG-LFMSQQKYAEDLLIAASMEHCTPLPTPLPVQLDRVPHQEELFSDPT 1000
Query: 1329 FFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINP 1388
+FRSI G LQY+T+TRP+I ++VN VCQ + P + +KRILRY+KGTI G+ +
Sbjct: 1001 YFRSIAGKLQYLTLTRPDIQFAVNFVCQKMHQPTISDFHLLKRILRYIKGTITMGISYS- 1059
Query: 1389 APMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRS 1448
P L A+ D+DW + RRS G C +G NL+SW +KK V+RSS EAEY+S
Sbjct: 1060 --RDSPTLLQAYSDSDWGNCKQTRRSVGGLCTFMGTNLVSWSSKKHPTVSRSSTEAEYKS 1117
Query: 1449 LAQASAEVLWIQSLLKELKVPTA-IPQIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVRE 1507
L+ A++E+LW+ +LL+EL++P P++FCDNLS V L NP H+RTKH ++D FVRE
Sbjct: 1118 LSDAASEILWLSTLLRELRIPLPDTPELFCDNLSAVYLTANPAFHARTKHFDIDFHFVRE 1177
Query: 1508 KVISKDLIVSHIPAQYQVADILTKPLSASRFLELRNKLRV--SDPMSLRG 1555
+V K L+V HIP Q+ADI TK L F+ LR KL V S SLRG
Sbjct: 1178 RVALKALVVKHIPGSEQIADIFTKSLPYEAFIHLRGKLGVTLSPTPSLRG 1227
Score = 41.2 bits (95), Expect = 0.005
Identities = 26/124 (20%), Positives = 64/124 (50%), Gaps = 5/124 (4%)
Query: 128 MRTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEF 187
+ R +L+ L+ ++K +++ ++ +++I + L S+G PV + I L L E+
Sbjct: 43 LHMRLFELQRRLQNVSKRDKTMDAYLNDLKNICDQLASVGSPVTEKMKIFAALNGLGREY 102
Query: 188 NPIVATVNSQTEV---ISLDELESQLLTQEARNEKF-KKALVGETASVNLTHAENSGEKN 243
PI T+ + + SL+ + +L + R + + ++ + + N+T +++S +N
Sbjct: 103 EPIKTTIENSMDTQPGPSLENVIPKLTGYDDRLQGYLEETTISPYVAFNITTSDDS-YRN 161
Query: 244 GHNQ 247
+N+
Sbjct: 162 SYNR 165
>At4g28900 putative protein
Length = 1415
Score = 714 bits (1842), Expect = 0.0
Identities = 501/1564 (32%), Positives = 747/1564 (47%), Gaps = 275/1564 (17%)
Query: 24 FKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSPQIPP-----VFLNDAAREAGTEN 78
F +++KL NYL WK Q E L +++ V P + D EA N
Sbjct: 13 FSHYVTLKLSTANYLLWKIQFETWLNNQRLLGFVTGANPCPNATRSIRNGDQVTEA--TN 70
Query: 79 PAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSE 138
P + W + D + W+L ++S L L S +VW + +R L+
Sbjct: 71 PDFLTWVQNDQKIMGWLLGSLSEDALRSVYGLHTSREVWFSLAKKYNRVSASRKSDLQRR 130
Query: 139 LRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPEEFNPIVATV---N 195
L ++K +S+ E++ ++ I + L SIG PV + I VL L +E+ +V+T+ +
Sbjct: 131 LNPVSKNEKSMLEYLNCVKQICDQLDSIGCPVPENEKIFGVLNGLGQEY-MLVSTMIKGS 189
Query: 196 SQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPD 255
T +S +++ +L+ + + L + ++ G + +N G
Sbjct: 190 MDTYPMSFEDVVFKLINFDDK----------------LQNGQSGGNRGRNNYTTKGRGFP 233
Query: 256 QQFNISGNPTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTG 315
QQ + SG+P+ + + CQIC K G
Sbjct: 234 QQIS-SGSPSDSGTRP-----------------------------------TCQICNKYG 257
Query: 316 HDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGA 375
H A C+ R Q E + + SN W+ G
Sbjct: 258 HSAYKCWKRFDHAFQSEDF-----------SKAFAAMRVSDQKSNPWVTDSG-------- 298
Query: 376 PRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLMDAASFS 435
+ T+S S P SG
Sbjct: 299 ---------------------ATSHITNSTSQLQSAQPYSG------------------- 318
Query: 436 GSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNV 495
D + +GN L I IGS + S L L ++L P+ITK+L+SVS+ D
Sbjct: 319 -EDSVIVGNSDFLPITHIGSAVLT---SNQGNLPLRDVLVCPNITKSLLSVSKLTSDYPC 374
Query: 496 FFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNC 555
EF S+ VK + + ++L KG +D LY + P C
Sbjct: 375 VIEFDSDGVIVKDKLTKQLLTKGTRHND-LYLLENP-------------------KFMAC 414
Query: 556 FSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLC 615
+S SR Q + +VWH RLGHP+ +V++ +++
Sbjct: 415 YS------SRQQATSD----------------------EVWHMRLGHPNQDVLQQLLR-- 444
Query: 616 NQQLP-NKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYF 674
N+ + +K+ C AC +GK +LP SS V ++ E + CDLWGPA V S G+ Y+
Sbjct: 445 NKAIVISKTSHSLCDACQMGKICKLPFASSDFVSSRLLERVHCDLWGPAPVVSSQGFRYY 504
Query: 675 LICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEF--RPFTQF 732
+I +D YSR+TW +PL+LKS F F+ MVE Q I S Q DGGGEF F
Sbjct: 505 VIFIDNYSRFTWFYPLRLKSDFFSVFLTFQKMVENQCQQKIASFQCDGGGEFISNQFVSH 564
Query: 733 LTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINR 792
L GI ++CP+T QNG ERKHRHI E G +++ K+P W AF T+ +L N
Sbjct: 565 LAECGIRQLISCPYTPQQNGIAERKHRHITELGSSMMFQGKVPQFLWVEAFYTSNFLCNL 624
Query: 793 LPSPIL-NNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPS 851
LPS +L + KSP+ +L + P Y L+ FGC+C+P RPY ++K + +S CVF GY+
Sbjct: 625 LPSSVLKDQKSPYEVLMGKAPVYTSLRVFGCACYPNLRPYASNKFDPKSLLCVFTGYNEK 684
Query: 852 HKGYKCL-DPTGRMFISKDVIFNEYKFPYSELFTSGQSSSPPTTSSDHTPLPSFLFPLNN 910
+KGYKC PTG+++I++ V+F+E KF +S++++ S T S+ + S P
Sbjct: 685 YKGYKCFHPPTGKIYINRHVLFDESKFLFSDIYSDKVSG---TNSTLVSAWQSNFLP--- 738
Query: 911 KQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTHASSHSNHHNISPGPIFNPT 970
K P T L ++ +SF + Q ++ ++ ++ G +
Sbjct: 739 KSIPATPE------VLDISNTAASFSD-EQGEFSGAVGGGGCGCTADLDSVPIGNSLPSS 791
Query: 971 PI----STHPPSPSPSSHSHNTYHSI--------SVEPVTSQPSTQAEPHRIHPNNTHSM 1018
P+ S P +P S+ S N S V S+ +T+ E + +H M
Sbjct: 792 PVTQQNSPQPETPISSAGSGNDAEDSELSENSENSESSVFSEATTETEAADNTNDQSHPM 851
Query: 1019 ATRAKHGIVQ---KRKHPTLLLTHIEPTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVP 1075
TR+K GI + K T+ + P + A+K P W AM E+++ + +TW LVP
Sbjct: 852 ITRSKSGIFKPNPKYAMFTVKSNYPVPKTVKTALKDPGWTDAMGEEYDSFEETHTWDLVP 911
Query: 1076 LPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSV 1135
+ +GC+WVF+TK DG++++ KARLVAKG+ Q G DY ET+SPVV+ TVR++
Sbjct: 912 PDSFITPLGCRWVFKTKLKADGTLDRLKARLVAKGYEQEEGVDYMETYSPVVRTATVRTI 971
Query: 1136 LTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD-PSLVCKLNKALYGLKQAP 1194
L +A NKW I+QLDV NAFL+G L+E VYM QPPGFE D P VCKLNKA+YGLKQAP
Sbjct: 972 LHVATINKWEIKQLDVKNAFLHGDLKETVYMYQPPGFENQDRPDYVCKLNKAIYGLKQAP 1031
Query: 1195 RAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSASLIQQLVKK 1254
RAWF++ + LL+ GF + DPSLF+ + F+L+Y+DD+L+TG++
Sbjct: 1032 RAWFDKFSTFLLEFGFICTYSDPSLFVFLKGRDLMFLLLYMDDMLLTGNN---------- 1081
Query: 1255 LNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPMASST 1314
KKY DLLV A MA+ + +P+
Sbjct: 1082 ---------------------------------KKYAMDLLVAAGMADCAPMPTPLPLQL 1108
Query: 1315 KLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVKRILR 1374
+DPT+FRS+ +VN VCQ + +P + +KR+LR
Sbjct: 1109 DKVPGQQESFADPTYFRSL----------------AVNLVCQKMHSPTVADFNLLKRVLR 1152
Query: 1375 YLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWWAKKQ 1434
YLKG + GL ++ + ++L A+ D+DWA+ + RRS G C LG N+ISW AK+
Sbjct: 1153 YLKGKVQMGLNLH---NNTDITLRAYSDSDWANCKETRRSVGGFCTFLGTNIISWSAKRH 1209
Query: 1435 TLVARSSAEAEYRSLAQASAEVLWIQSLLKELKV-PTAIPQIFCDNLSTVSLAHNPVLHS 1493
V+RSS EAEYR+L+ A+ EV WI SLL+E+ + A P+++CDNLS V L NP +H+
Sbjct: 1210 PTVSRSSTEAEYRTLSIAATEVKWISSLLREIGIYQPAPPELYCDNLSAVYLTANPAMHN 1269
Query: 1494 RTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLELRNKLRVSDP--M 1551
R+K ++D +VRE+V L+V H+PA +Q+ADI TK L F +LR KL V P
Sbjct: 1270 RSKAFDVDFHYVRERVALGALVVKHVPASHQLADIFTKSLPQRPFFDLRYKLGVVLPPTP 1329
Query: 1552 SLRG 1555
SLRG
Sbjct: 1330 SLRG 1333
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 704 bits (1816), Expect = 0.0
Identities = 493/1560 (31%), Positives = 738/1560 (46%), Gaps = 219/1560 (14%)
Query: 27 IISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSPQIPPVFLNDAAREAGTENPAYTEWEE 86
IIS +LDET Y W + + K V +P +D P + W
Sbjct: 78 IISHRLDETTYGDWSVAMR-ISLDAKNKLGFVDGSLPRPLESD---------PNFRLWSR 127
Query: 87 QDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGT 146
+S++ +W+L+++S + +RL + +W ++ + R+ L E++ + +GT
Sbjct: 128 CNSMVKSWLLNSVSPQIYRSILRLNDATDIWRDLFDRFNLTNLPRTYNLTQEIQDLRQGT 187
Query: 147 RSIAEFIARIRSISESLMS---IGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISL 203
S++E+ ++++ + L S + DP + +A + +A +N ++
Sbjct: 188 MSLSEYYTLLKTLWDQLDSTEALDDPCTCGKAVRLYQKAEKAKIMKFLAGLNESYAIV-- 245
Query: 204 DELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGN 263
Q++ KKAL ++ +NS + G Q +S +
Sbjct: 246 ---RRQIIA--------KKALPSLAEVYHILDQDNS--QKGFFNVVAPPAAFQVSEVSHS 292
Query: 264 PTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYH 323
P + Y + G N GR C C + GH A CY
Sbjct: 293 PITSPEIMYV----------------------QSGPNKGRPT--CSFCNRVGHIAERCYK 328
Query: 324 RLSVPPQYEGYGSLGGN------FGGNLGSGYGPATG-----FGTHSNVWMQGV-----G 367
+ PP + G + TG G S +Q +
Sbjct: 329 KHGFPPGFTPKGKSSDKPPKPQAVAAQVTLSPDKMTGQLETLAGNFSPDQIQNLIALFSS 388
Query: 368 QRNPSYGAPR-APFPPQFGNSRPPAPQAYI--------TGNESTSSNSFNNG-WYPDSGA 417
Q P +P+ A + +S+ AP + G + S NS ++ W DSGA
Sbjct: 389 QLQPQIVSPQTASSQHEASSSQSVAPSGILFSPSTYCFIGILAVSHNSLSSDTWVIDSGA 448
Query: 418 THHVTPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVP 477
THHV+ D L S + + G + I+ +G++ N + L N+L +P
Sbjct: 449 THHVSHD-RKLFQTLDTSIVSFVNLPTGPNVRISGVGTVLI------NKDIILQNVLFIP 501
Query: 478 SITKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDG-LYQFDQPYVPSV 536
NL+S+S D F + C + QD TK L G G LY D
Sbjct: 502 EFRLNLISISSLTTDLGTRVIFDPSCCQI--QDLTKGLTLGEGKRIGNLYVLD------- 552
Query: 537 SRTASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVW 596
+ ++S+N S VW
Sbjct: 553 --------TQSPAISVNAVVDVS-----------------------------------VW 569
Query: 597 HNRLGHPHHEVVRSVMKLCNQQLPNKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIF 656
H RLGHP + S+ ++ + +C C L K +L S+ + N FEL+
Sbjct: 570 HKRLGHPSFSRLDSLSEVLGTTRHKNKKSAYCHVCHLAKQKKLSFPSANNICNSTFELLH 629
Query: 657 CDLWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIK 716
D+WGP SVE+ GY YFL VD +SR TWI+ LK KS L F F +VE QY+ +K
Sbjct: 630 IDVWGPFSVETVEGYKYFLTIVDDHSRATWIYLLKSKSDVLTVFPAFIDLVENQYDTRVK 689
Query: 717 SVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPL 776
SV++D E FT+F GI +CP T QN VERKH+HI+ L+ + + L
Sbjct: 690 SVRSDNAKEL-AFTEFYKAKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSNMSL 748
Query: 777 HYWDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKL 836
YW LTA +LINR PS +L+NK+PF +L ++PDY LK+FGC C+ T HK
Sbjct: 749 PYWGDCVLTAVFLINRTPSALLSNKTPFEVLTGKLPDYSQLKTFGCLCYSSTSSKQRHKF 808
Query: 837 ELRSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQSSSPPTTS 895
RS+ CVFLGY KGYK LD + + IS++V F+E ELF S TT+
Sbjct: 809 LPRSRACVFLGYPFGFKGYKLLDLESNVVHISRNVEFHE------ELFPLASSQQSATTA 862
Query: 896 SD-HTPLPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTHASS 954
SD TP+ PL SS + T H SP S P + S + H
Sbjct: 863 SDVFTPMD----PL----------SSGNSITSHLPSPQIS-PSTQISKRRITKFPAHLQD 907
Query: 955 HSNHHNISPGPIFNPTPISTHPPSPSPSSHSHNTYHSISVEPVTSQPSTQAEPHRIHPNN 1014
+ + +HP S S S + H + + ++ P Q+
Sbjct: 908 YH---------CYFVNKDDSHPISSSLSYSQISPSHMLYINNISKIPIPQS--------- 949
Query: 1015 THSMATRAKHGIVQKRKHPTLLLTHIEPTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLV 1074
Y +A +W A+ E A+ + +TW +
Sbjct: 950 ------------------------------YHEAKDSKEWCGAIDQEIGAMERTDTWEIT 979
Query: 1075 PLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRS 1134
LP ++AVGCKWVF K + DGS+ ++KAR+VAKG+ Q G DY ETFSPV K TV+
Sbjct: 980 SLPPGKKAVGCKWVFTVKFHADGSLERFKARIVAKGYTQKEGLDYTETFSPVAKMATVKL 1039
Query: 1135 VLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGF-----EAVDPSLVCKLNKALYG 1189
+L ++ + KW + QLD++NAFLNG LEE +YM P G+ ++ P++VC+L K++YG
Sbjct: 1040 LLKVSASKKWYLNQLDISNAFLNGDLEETIYMKLPDGYADIKGTSLPPNVVCRLKKSIYG 1099
Query: 1190 LKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSASLIQ 1249
LKQA R WF + ++LL LGF D +LF+ +LVYVDDI+I ++ Q
Sbjct: 1100 LKQASRQWFLKFSNSLLALGFEKQHGDHTLFVRCIGSEFIVLLVYVDDIVIASTTEQAAQ 1159
Query: 1250 QLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASP 1309
L + L A F L++LG L YFLG+EV + G + LSQ+KY +LL A+M + + P
Sbjct: 1160 SLTEALKASFKLRELGPLKYFLGLEVARTSEG-ISLSQRKYALELLTSADMLDCKPSSIP 1218
Query: 1310 MASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAV 1369
M + +L+K + D +R +VG L Y+T+TRP+I+++VNK+CQF SAP H AV
Sbjct: 1219 MTPNIRLSKNDGLLLEDKEMYRRLVGKLMYLTITRPDITFAVNKLCQFSSAPRTAHLAAV 1278
Query: 1370 KRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISW 1429
++L+Y+KGT+ GL + L+L + DADW + PD RRST+G + +G +LISW
Sbjct: 1279 YKVLQYIKGTVGQGLFYS---AEDDLTLKGYTDADWGTCPDSRRSTTGFTMFVGSSLISW 1335
Query: 1430 WAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIPQIFCDNLSTVSLAHNP 1489
+KKQ V+RSSAEAEYR+LA AS E+ W+ +LL L+V + +P ++ D+ + V +A NP
Sbjct: 1336 RSKKQPTVSRSSAEAEYRALALASCEMAWLSTLLLALRVHSGVPILYSDSTAAVYIATNP 1395
Query: 1490 VLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLELRNKLRVSD 1549
V H RTKH+E+D VREK+ + L + H+ + QVADILTKPL +F L +K+ + +
Sbjct: 1396 VFHERTKHIEIDCHTVREKLDNGQLKLLHVKTKDQVADILTKPLFPYQFAHLLSKMSIQN 1455
>At2g13940 putative retroelement pol polyprotein
Length = 1501
Score = 697 bits (1799), Expect = 0.0
Identities = 494/1611 (30%), Positives = 749/1611 (45%), Gaps = 195/1611 (12%)
Query: 2 SSPSSPVNNSETQRVTAAASKNFKQIIS-VKLDETNYLQWKQQVEGVLRGTKMVRHVVSP 60
+S S V++ T A+S N +IS V+L+ NY QW ++ L+ +
Sbjct: 18 ASGGSRVDSLMVSPYTLASSDNPGAVISSVELNGDNYNQWATEMLNALQAKRKTG----- 72
Query: 61 QIPPVFLNDAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEI 120
F+N +P Y W +S++ WI ++I + + + +H +W ++
Sbjct: 73 -----FINGTIPRPPPNDPNYENWTAVNSMIVGWIRTSIEPKVKATVTFISDAHLLWKDL 127
Query: 121 HNYCYTQMRTRSRQLRSELRTITKGTRSIAEFIARIRSISESLM---------------- 164
+ R Q+R++L + + +++ E+ R+ ++ E
Sbjct: 128 KQRFSVGNKVRIHQIRAQLSSCRQDGQAVIEYYGRLSNLWEEYNIYKPVTVCTCGLCRCG 187
Query: 165 SIGDPVAHRD---LIETVLEALPEEFNPIVATVNSQTEVISLDELESQLLTQEARNEKFK 221
+ +P R+ + + VL F + AT+ + + SL E+ S+++ +E R
Sbjct: 188 ATSEPTKEREEEKIHQFVLGLDESRFGGLCATLINMDPLPSLGEIYSRVIREEQRLASVH 247
Query: 222 KALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPTGNNSSQYFNPNFGGRN 281
E A L E + + H++ S
Sbjct: 248 VREQKEEAVGFLARRE---QLDHHSRVDASS----------------------------- 275
Query: 282 GSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRLSVPPQY-EGYGSLGGN 340
SR G+R + +G + C C +TGH+ C+ + P + E G G N
Sbjct: 276 -SRSEHTGGSR-----SNSIIKGRVTCSNCGRTGHEKKECWQIVGFPDWWSERNGGRGSN 329
Query: 341 FGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQFGNSRPPAPQAYITGNE 400
G G G G G V N S P+F +
Sbjct: 330 GRGRGGRGSNGGRGQG---QVMAAHATSSNSSVF-------PEFTEEHMRVLSQLVKEKS 379
Query: 401 STSSNSFNNG-----------WYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLA 449
++ S S NN DSGA+HH+T ++L + + + A
Sbjct: 380 NSGSTSNNNSDRLSGKTKLGDIILDSGASHHMTGTLSSLTNVVPVPPCPVGFADGSKAFA 439
Query: 450 INSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQ 509
+ S+G ++ S+ T++L N+L VPS+ L+SVS+ K F +C+++ +
Sbjct: 440 L-SVGVLTLSN------TVSLTNVLFVPSLNCTLISVSKLLKQTQCLATFTDTLCFLQDR 492
Query: 510 DSTKILLKGHLGDDGLYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCN 569
S+K L+ G+Y Y+ V+
Sbjct: 493 -SSKTLIGSGEERGGVY-----YLTDVTPAK----------------------------- 517
Query: 570 NGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDFCS 629
IHT+ +D + +WH RLGHP V+ S+ S + C
Sbjct: 518 -------IHTANVDSDQA-------LWHQRLGHPSFSVLSSLPLFSKTSSTVTSHS--CD 561
Query: 630 ACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIFP 689
C K R S + F LI CD+WGP V + G YFL VD YSR W +
Sbjct: 562 VCFRAKQTREVFPESINKTEECFSLIHCDVWGPYRVPASCGAVYFLTIVDDYSRAVWTYL 621
Query: 690 LKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHH 749
L KS NF E Q+ +K V++D G EF + + GI H+ +C T
Sbjct: 622 LLEKSEVRQVLTNFLKYAEKQFGKTVKMVRSDNGTEFMCLSSYFRENGIIHQTSCVGTPQ 681
Query: 750 QNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLHL 809
QNG VERKHRHI+ LL A LP+ +W + LTA YLINR PS IL+ ++P+ +LH
Sbjct: 682 QNGRVERKHRHILNVARALLFQASLPIKFWGESILTAAYLINRTPSSILSGRTPYEVLHG 741
Query: 810 QIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLDPTGRMF-ISK 868
P Y L+ FG +C+ + K RS+ C+F+GY KG+K D F +S+
Sbjct: 742 SKPVYSQLRVFGSACYVHRVTRDKDKFGQRSRSCIFVGYPFGKKGWKVYDIERNEFLVSR 801
Query: 869 DVIFNEYKFPYSELFTS--GQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTPTTTL 926
DVIF E FPY+ + +S +S P + D +P PL + + + T
Sbjct: 802 DVIFREEVFPYAGVNSSTLASTSLPTVSEDDDWAIP----PLEVRGSIDSVETERVVCTT 857
Query: 927 HTASPHSSFPESNQSNHHHSIQDTHASSHSNHHNISPG------------PIFNPTPIS- 973
+S +S N DT SS + +SP P+ +P P+S
Sbjct: 858 DEVVLDTSVSDSEIPNQEFVPDDTPPSSPLS---VSPSGSPNTPTTPIVVPVASPIPVSP 914
Query: 974 ---------THPPSPSPSSHSHNTYH---SISVEPVTSQPSTQAEPHRIHPNNTH--SMA 1019
THPP +N + SI P S+ + P + A
Sbjct: 915 PKQRKSKRATHPPPKLNDYVLYNAMYTPSSIHALPADPSQSSTVPGKSLFPLTDYVSDAA 974
Query: 1020 TRAKHGIVQKRKHPTLLLTHIEPTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVPLPAD 1079
+ H R + + ++EP +++A++ W AM E +AL N TW +V LP
Sbjct: 975 FSSSH-----RAYLAAITDNVEPKHFKEAVQIKVWNDAMFTEVDALEINKTWDIVDLPPG 1029
Query: 1080 RQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSVLTLA 1139
+ A+G +WVF+TK N DG++ +YKARLV +G Q+ G DYKETF+PVV+ TVR++L
Sbjct: 1030 KVAIGSQWVFKTKYNSDGTVERYKARLVVQGNKQVEGEDYKETFAPVVRMTTVRTLLRNV 1089
Query: 1140 VTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVDPSLVCKLNKALYGLKQAPRAWFE 1199
N+W + Q+DV+NAFL+G LEEEVYM PPGF P VC+L K+LYGLKQAPR WF+
Sbjct: 1090 AANQWEVYQMDVHNAFLHGDLEEEVYMKLPPGFRHSHPDKVCRLRKSLYGLKQAPRCWFK 1149
Query: 1200 RLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSASLIQQLVKKLNAEF 1259
+L +LL+ GF S D SLF N +L+YVDD+LI G+ ++Q+ L+ F
Sbjct: 1150 KLSDSLLRFGFVQSYEDYSLFSYTRNNIELRVLIYVDDLLICGNDGYMLQKFKDYLSRCF 1209
Query: 1260 SLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPMASSTKLTKY 1319
S+KDLGKL YFLGIEV G + LSQ+KY D++ + + +P+ + L
Sbjct: 1210 SMKDLGKLKYFLGIEVSRGPEG-IFLSQRKYALDVIADSGNLGSRPAHTPLEQNHHLASD 1268
Query: 1320 GSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVKRILRYLKGT 1379
+SDP +R +VG L Y+ TRPE+SYSV+ + QF+ P E H+ A R++RYLKG+
Sbjct: 1269 DGPLLSDPKPYRRLVGRLLYLLHTRPELSYSVHVLAQFMQNPREAHFDAALRVVRYLKGS 1328
Query: 1380 IHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWWAKKQTLVAR 1439
G+L+N P L+L +CD+DW S P RRS S +LLG + ISW KKQ V+
Sbjct: 1329 PGQGILLNADP---DLTLEVYCDSDWQSCPLTRRSISAYVVLLGGSPISWKTKKQDTVSH 1385
Query: 1440 SSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIP-QIFCDNLSTVSLAHNPVLHSRTKHM 1498
SSAEAEYR+++ A E+ W++ LLKEL + + P +++CD+ + + +A NPV H RTKH+
Sbjct: 1386 SSAEAEYRAMSYALKEIKWLRKLLKELGIEQSTPARLYCDSKAAIHIAANPVFHERTKHI 1445
Query: 1499 ELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLELRNKLRVSD 1549
E D VR+ V + H+ Q+AD+ TK L ++FL L +KL V +
Sbjct: 1446 ESDCHSVRDAVRDGIITTQHVRTTEQLADVFTKALGRNQFLYLMSKLGVQN 1496
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 681 bits (1757), Expect = 0.0
Identities = 474/1557 (30%), Positives = 736/1557 (46%), Gaps = 220/1557 (14%)
Query: 27 IISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSPQIPPVFLNDAAREAGTENPAYTEWEE 86
IIS +LDETNY W + L F++ + + W
Sbjct: 74 IISHRLDETNYGDWSVAMLISLDAKNKTG----------FIDGTLSRPLESDLNFRLWSR 123
Query: 87 QDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGT 146
+S++ +W+L+++S + +R+ + +W ++++ R+ L E++ +GT
Sbjct: 124 CNSMVKSWLLNSVSPQIYRSILRMNDASDIWRDLNSRFNVTNLPRTYNLTQEIQDFRQGT 183
Query: 147 RSIAEFIARIRSISESLMS---IGDPVAHRDLIETVLEALPEEFNPIVATVNSQTEVISL 203
S++E+ R++++ + L S + +P + +A + +A +N ++
Sbjct: 184 LSLSEYYTRLKTLWDQLDSTEALDEPCTCGKAMRLQQKAEQAKIVKFLAGLNESYAIV-- 241
Query: 204 DELESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGN 263
Q++ KKAL ++ +NS + + ++ Q I+ +
Sbjct: 242 ---RRQIIA--------KKALPSLGEVYHILDQDNSQQSFSNVVAPPAAF--QVSEITQS 288
Query: 264 PTGNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYH 323
P+ + + Y + G N GR C + GH A CY
Sbjct: 289 PSMDPTVCYV----------------------QNGPNKGRPI--CSFYNRVGHIAERCYK 324
Query: 324 RLSVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYGAPRAPFPPQ 383
+ PP + G G A ++++ A F Q
Sbjct: 325 KHGFPPGFTPKGKAGEKLQKPKPLAANVAESSEVNTSLESMVGNLSKEQLQQFIAMFSSQ 384
Query: 384 FGNSRPPAPQAYITGNESTSSN----------SF------------NNGWYPDSGATHHV 421
N+ P Y T + S S N SF + W DSGATHHV
Sbjct: 385 LQNT---PPSTYATASTSQSDNLGICFSPSTYSFIGILTVARHTLSSATWVIDSGATHHV 441
Query: 422 TPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITK 481
+ D +L + S + + G + I+ +G++ N + L N+L +P
Sbjct: 442 SHD-RSLFSSLDTSVLSAVNLPTGPTVKISGVGTLKL------NDDILLKNVLFIPEFRL 494
Query: 482 NLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDG-----LYQFDQPYVPSV 536
NL+S+S D F N C ++ L+KG + G LY D
Sbjct: 495 NLISISSLTDDIGSRVIFDKNSCEIQD------LIKGRMLGQGRRVANLYLLD------- 541
Query: 537 SRTASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVW 596
V S+S+N S +W
Sbjct: 542 --------VGDQSISVNAVVDIS-----------------------------------MW 558
Query: 597 HNRLGHPHHEVVRSVMKLCNQQLPNKSFTDFCSACCLGKSHRLPSVSSKTVYNKPFELIF 656
H RLGH + + ++ +DFC C L K +L +S V + F+L+
Sbjct: 559 HRRLGHASLQRLDAISDSLGTTRHKNKGSDFCHVCHLAKQRKLSFPTSNKVCKEIFDLLH 618
Query: 657 CDLWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIK 716
D+WGP SVE+ GY YFL VD +SR TW++ LK KS L F F VE QY + +K
Sbjct: 619 IDVWGPFSVETVEGYKYFLTIVDDHSRATWMYLLKTKSEVLTVFPAFIQQVENQYKVKVK 678
Query: 717 SVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPL 776
+V++D E + FT F GI +CP T QN VERKH+HI+ L+ +++PL
Sbjct: 679 AVRSDNAPELK-FTSFYAEKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSQVPL 737
Query: 777 HYWDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKL 836
W LTA +LINR PS +L NK+P+ +L P Y+ L++FGC C+ T P HK
Sbjct: 738 SLWGDCVLTAVFLINRTPSQLLMNKTPYEILTGTAPVYEQLRTFGCLCYSSTSPKQRHKF 797
Query: 837 ELRSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQSSSPPTTS 895
+ RS+ C+FLGY +KGYK +D + +FIS++V F+E FP L + S S
Sbjct: 798 QPRSRACLFLGYPSGYKGYKLMDLESNTVFISRNVQFHEEVFP---LAKNPGSESSLKLF 854
Query: 896 SDHTPLPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTHASSH 955
+ P+ S + TT S S+ + + P S + H + D H ++
Sbjct: 855 TPMVPVSSGII------SDTTHSPSSLPSQISDLPPQISSQRVRKPPAH--LNDYHCNTM 906
Query: 956 SNHHNISPGPIFNPTPISTHPPSPSPSSHSHNTYHSISVEPVTSQPSTQAEPHRIHPNNT 1015
+ H PIS+ S S S SH Y + +T P
Sbjct: 907 QSDHKY---------PISS-TISYSKISPSHMCY----INNITKIPI------------- 939
Query: 1016 HSMATRAKHGIVQKRKHPTLLLTHIEPTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVP 1075
PT Y +A +W +A+ E A+ K NTW +
Sbjct: 940 --------------------------PTNYAEAQDTKEWCEAVDAEIGAMEKTNTWEITT 973
Query: 1076 LPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSV 1135
LP ++AVGCKWVF K DG++ +YKARLVAKG+ Q G DY +TFSPV K T++ +
Sbjct: 974 LPKGKKAVGCKWVFTLKFLADGNLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTIKLL 1033
Query: 1136 LTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFE-----AVDPSLVCKLNKALYGL 1190
L ++ + KW ++QLDV+NAFLNG LEEE++M P G+ + ++V +L +++YGL
Sbjct: 1034 LKVSASKKWFLKQLDVSNAFLNGELEEEIFMKIPEGYAERKGIVLPSNVVLRLKRSIYGL 1093
Query: 1191 KQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSASLIQQ 1250
KQA R WF++ S+LL LGF + D +LF+ + +LVYVDDI+I +S + Q
Sbjct: 1094 KQASRQWFKKFSSSLLSLGFKKTHGDHTLFLKMYDGEFVIVLVYVDDIVIASTSEAAAAQ 1153
Query: 1251 LVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPM 1310
L ++L+ F L+DLG L YFLG+EV + G + + Q+KY +LL M ++ PM
Sbjct: 1154 LTEELDQRFKLRDLGDLKYFLGLEVARTTAG-ISICQRKYALELLQSTGMLACKPVSVPM 1212
Query: 1311 ASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVK 1370
+ K+ K + + D +R IVG L Y+T+TRP+I+++VNK+CQF SAP H A
Sbjct: 1213 IPNLKMRKDDGDLIEDIEQYRRIVGKLMYLTITRPDITFAVNKLCQFSSAPRTTHLTAAY 1272
Query: 1371 RILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWW 1430
R+L+Y+KGT+ GL + + L+L F D+DWAS D RRST+ + +G +LISW
Sbjct: 1273 RVLQYIKGTVGQGLFYSAS---SDLTLKGFADSDWASCQDSRRSTTSFTMFVGDSLISWR 1329
Query: 1431 AKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIPQIFCDNLSTVSLAHNPV 1490
+KKQ V+RSSAEAEYR+LA A+ E++W+ +LL L+ +P ++ D+ + + +A NPV
Sbjct: 1330 SKKQHTVSRSSAEAEYRALALATCEMVWLFTLLVSLQASPPVPILYSDSTAAIYIATNPV 1389
Query: 1491 LHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLELRNKLRV 1547
H RTKH++LD VRE++ + +L + H+ + QVADILTKPL +F L++K+ +
Sbjct: 1390 FHERTKHIKLDCHTVRERLDNGELKLLHVRTEDQVADILTKPLFPYQFEHLKSKMSI 1446
>At4g17450 retrotransposon like protein
Length = 1433
Score = 628 bits (1620), Expect = e-180
Identities = 473/1576 (30%), Positives = 701/1576 (44%), Gaps = 286/1576 (18%)
Query: 27 IISVKLDETNYLQWKQQVEGVLRGTKMVRHVVSPQIPPVFLNDAAREAGTENPAYTEWEE 86
I+S LD TNY W + L + F++ + + + W
Sbjct: 80 IVSHILDGTNYNNWSIAMRMSLDAKNKLS----------FVDGSLPRPDVSDRMFKIWSR 129
Query: 87 QDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHNYCYTQMRTRSRQLRSELRTITKGT 146
+S++ TW+L+ ++ ++W+++ + R QL + T+ +G
Sbjct: 130 CNSMVKTWLLNVVT--------------EMWNDLFSRFRVSNLPRKYQLEQSIHTLKQGN 175
Query: 147 RSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEALPE-EFNPIVATVNSQTEVISLDE 205
++ + + +++ E L + + E V E L E E + I+ + + +
Sbjct: 176 LDLSTYYTKKKTLWEQLANTRVLTVRKCNCEHVKELLEEAETSRIIQFLMGLND--NFAH 233
Query: 206 LESQLLTQEARNEKFKKALVGETASVNLTHAENSGEKNGHNQPQTGSYPDQQFNISGNPT 265
+ Q+L + R LT N ++ D+ + GNPT
Sbjct: 234 IRGQILNMKPRP--------------GLTEIYNMLDQ------------DESQRLVGNPT 267
Query: 266 GNNSSQYFNPNFGGRNGSRGRGFRGNRFRGRGGRNFGRGNIQCQICYKTGHDASICYHRL 325
+N + F S+ +G+ + + C C K GH CY +
Sbjct: 268 LSNPTAAFQVQASPIIDSQVNMAQGSYKKPK-----------CSYCNKLGHLVDKCYKKH 316
Query: 326 SVPPQYEGYGSLGGNFGGNLGSGYGPATGFGTHSNVWMQGVGQRNPSYG----------- 374
PP G + G G T + ++ SY
Sbjct: 317 GYPP--------GSKW--TKGQTIGSTNLASTQLQPVNETPNEKTDSYEEFSTDQIQTMI 366
Query: 375 ---------APRAPFPPQFGNSRPPAPQA-YITGNESTSSNSFNN--------------- 409
A +P P S +P I+ T + F+N
Sbjct: 367 SYLSTKLHIASASPMPTTSSASISASPSVPMISQISGTFLSLFSNAYYDMLISSVSQEPA 426
Query: 410 ----GWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPN 465
GW DSGATHHVT + + ++ S + + + N + I IG + S S
Sbjct: 427 VSPRGWVIDSGATHHVTHNRDLYLNFRSLENTF-VRLPNDCTVKIAGIGFIQLSDAIS-- 483
Query: 466 TTLTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDG- 524
L+N+L++P NL+S + TK L+ G G
Sbjct: 484 ----LHNVLYIPEFKFNLIS------------------------ELTKELMIGRGSQVGN 515
Query: 525 LYQFDQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSN 584
LY D N SL + S C SV + + +
Sbjct: 516 LYVLD----------------------FNENNHTVSLKGTTSMCPEFSVCSSVVVDSVT- 552
Query: 585 DSSNSLSLYKVWHNRLGHPHHEVVRSVMKLCNQQLP--NKSFTDFCSAC--C-LGKSHRL 639
WH RLGHP + + + + N ++ NK + C C C L K L
Sbjct: 553 -----------WHKRLGHPAYSKIDLLSDVLNLKVKKINKEHSPVCHVCHVCHLSKQKHL 601
Query: 640 PSVSSKTVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLIT 699
S + + + F+L+ D WGP SV ++ TWI+ LK KS L
Sbjct: 602 SFQSRQNMCSAAFDLVHIDTWGPFSVPTNDA--------------TWIYLLKNKSDVLHV 647
Query: 700 FQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHR 759
F F MV QY +KSV++D E + FT GI +CP T QN VERKH+
Sbjct: 648 FPAFINMVHTQYQTKLKSVRSDNAHELK-FTDLFAAHGIVAYHSCPETPEQNSVVERKHQ 706
Query: 760 HIVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKS 819
HI+ LL + +PL +W LTA +LINRLP+P+LNNKSP+ L P Y+ LK+
Sbjct: 707 HILNVARALLFQSNIPLEFWGDCVLTAVFLINRLPTPVLNNKSPYEKLKNIPPAYESLKT 766
Query: 820 FGCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFP 878
FGC C+ T P HK E R++ CVFLGY +KGYK LD T + IS+ VIF+E FP
Sbjct: 767 FGCLCYSSTSPKQRHKFEPRARACVFLGYPLGYKGYKLLDIETHAVSISRHVIFHEDIFP 826
Query: 879 YSELFTSGQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPES 938
+ SS+ D PL FP P Q+S
Sbjct: 827 FI-------SSTIKDDIKDFFPL--LQFPARTDDLPLEQTS------------------- 858
Query: 939 NQSNHHHSIQDTHASSHSNHHNISPGPIFNP-TPISTHPPSPSPSSHSHNTYHSISVEPV 997
I DTH H ++S P P+S P + Y++ +
Sbjct: 859 --------IIDTHP-----HQDVSSSKALVPFDPLSKRQKKPPKHLQDFHCYNNTT---- 901
Query: 998 TSQPSTQAEPHRIHPNNTHSMATRAKHGIVQKRKHPTLLLTHIEPTGYRQAMKQPQWLQA 1057
EP NN + + P Y +A W A
Sbjct: 902 --------EPFHAFINN---------------------ITNAVIPQRYSEAKDFKAWCDA 932
Query: 1058 MQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGF 1117
M+ E A+++ NTW++V LP +++A+GCKWVF K N DGSI +YKARLVAKG+ Q G
Sbjct: 933 MKEEIGAMVRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKARLVAKGYTQEEGL 992
Query: 1118 DYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGF----- 1172
DY+ETFSPV K +VR +L LA KW + QLD++NAFLNG L+EE+YM PPG+
Sbjct: 993 DYEETFSPVAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEIYMKIPPGYADLVG 1052
Query: 1173 EAVDPSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFML 1232
EA+ P +C+L+K++YGLKQA R W+ +L +TL +GF S D +LFI +AN +L
Sbjct: 1053 EALPPHAICRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTLFIKYANGVLMGVL 1112
Query: 1233 VYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQ 1292
VYVDDI+I +S + Q +L + F L+DLG YFLGIE+ SE G + + Q+KYI
Sbjct: 1113 VYVDDIMIVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSEKG-ISICQRKYIL 1171
Query: 1293 DLLVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVN 1352
+LL + + P+ S KL K ++D T +R +VG L Y+ +TRP+I+Y+VN
Sbjct: 1172 ELLSTTGFLGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYLQITRPDIAYAVN 1231
Query: 1353 KVCQFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDR 1412
+CQF AP H AV ++LRYLKGT+ GL + L + D+D+ S D R
Sbjct: 1232 TLCQFSHAPTSVHLSAVHKVLRYLKGTVGQGLFYS---ADDKFDLRGYTDSDFGSCTDSR 1288
Query: 1413 RSTSGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAI 1472
R + C+ +G L+SW +KKQ V+ S+AEAE+R+++Q + E++W+ L + KVP
Sbjct: 1289 RCVAAYCMFIGDYLVSWKSKKQDTVSMSTAEAEFRAMSQGTKEMIWLSRLFDDFKVPFIP 1348
Query: 1473 P-QIFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTK 1531
P ++CDN + + + +N V H RTK +ELD + RE V S L + QVAD LTK
Sbjct: 1349 PAYLYCDNTAALHIVNNSVFHERTKFVELDCYKTREAVESGFLKTMFVETGEQVADPLTK 1408
Query: 1532 PLSASRFLELRNKLRV 1547
+ ++F +L K+ V
Sbjct: 1409 AIHPAQFHKLIGKMGV 1424
>At1g60020 hypothetical protein
Length = 1194
Score = 626 bits (1614), Expect = e-179
Identities = 390/1010 (38%), Positives = 532/1010 (52%), Gaps = 115/1010 (11%)
Query: 370 NPSYGAPRAPFPPQFGNSRPPAPQAYITGNESTSSNSFNNGWYPDSGATHHVTPDANNLM 429
N P +PF P P+A N + +S N W DSGATHH+T D NNL
Sbjct: 258 NSQQSIPASPFTPW-------QPRA----NAAIASPYNANNWLLDSGATHHITSDLNNLS 306
Query: 430 DAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLHVPSITKNLVSVSQF 489
++G + + I +G GL+I+ GS S+P + +L L ++L+VP+I KNL+SV +
Sbjct: 307 LHQPYTGGEDVTIADGSGLSISHTGSALISTP---SRSLALTDVLYVPNIHKNLISVYRM 363
Query: 490 CKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFD-QPYVPSVSRTASSSSVATS 548
C N V EF VK + LL+G D+ LY++ P PS T ++ +
Sbjct: 364 CNANKVSVEFFPAHFQVKDLKTGVQLLQGRTKDE-LYEWPVNPPKPSSHFTTTTPKTDLT 422
Query: 549 SLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKVWHNRLGHPHHEVV 608
S WH+RLGHP +
Sbjct: 423 S----------------------------------------------WHSRLGHPSLSTL 436
Query: 609 RSVMKLCNQQLPNKSFTDF-CSACCLGKSHRLPSVSSKTVYNKPFELIFCDLWGPASVES 667
+ V+ + + N F CS C L K+H+LP ++ +P E ++ DLW + + S
Sbjct: 437 KVVVSQFSLPVSNSLQKQFNCSDCLLNKTHKLPFHTNTITSTQPLEYLYIDLW-TSPIVS 495
Query: 668 HGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFR 727
+ Y+L+ VD Y+RY+W +P+K KSH F FK +V ++ I + +D GGEF
Sbjct: 496 IDNFKYYLVIVDHYTRYSWFYPIKQKSHVKDVFMTFKALVANKFQRKIIHLYSDNGGEFI 555
Query: 728 PFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLANAKLPLHYWDHAFLTAT 787
FL++ GITH T PHT NG ERKHRHIVETGLTLL A +P YW +AF A
Sbjct: 556 ALRSFLSSNGITHLTTPPHTPEHNGISERKHRHIVETGLTLLGQASMPKSYWSYAFTIAI 615
Query: 788 YLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRPYNNHKLELRSKECVFLG 847
YLINR+ S ++ SP+ L Q P+Y L+ FGC CFP+ RPY HKL+ R CVFLG
Sbjct: 616 YLINRMSSDVIGGISPYKRLFGQAPNYLKLRVFGCLCFPWLRPYTTHKLDDRPAPCVFLG 675
Query: 848 YSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYS----ELFTSGQSSS------------ 890
YS + Y CL+ TGR++ S+ V F E +P++ + FT+ + S+
Sbjct: 676 YSQTQSAYLCLNRTTGRVYTSRHVQFVENTYPFTKPTLDPFTNLEESNNHSITTTVPSPP 735
Query: 891 -------PPTTSSDHTPLPSFLFPLNNKQCPTTQSS----STP-------TTTLHTASPH 932
PP T H P PS P + P + SS S+P +TTLH H
Sbjct: 736 FVQLPSVPPPTRDPHQPPPSQPAPSPSPLSPPSMSSPVMTSSPQFSSNRDSTTLHGDYSH 795
Query: 933 SSFPESNQSNHHHSIQDTHASSHSNHHNISPGPIFNPTPISTHPPSPSPSSHSHNTYHSI 992
+ S+ SN I S + SP + P T P SPS SS S S
Sbjct: 796 VDYGLSSPSNPPGPITSPTTSKSPSEPTSSPS--HSNQPNKTPPNSPSSSSSSPTPIPSP 853
Query: 993 SVEPVTSQPSTQAEPHRIHPNNTHSMATRAKHGIVQKRKHPTLLLT----HIEPTGYRQA 1048
S + S P P N HSM TRAK+ I + K TL T PT +A
Sbjct: 854 SPQSSNSPPPP--------PQNQHSMRTRAKNNITKPIKKLTLAATPKGKSKIPTTVAEA 905
Query: 1049 MKQPQWLQAMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVA 1108
++ P W AM E A ++N+T+ LVP + VG +W+F K NPDGSIN+YKAR +A
Sbjct: 906 LRDPNWRNAMSEEFNAGLRNSTYDLVPPKPHQNFVGTRWIFTIKYNPDGSINRYKARFLA 965
Query: 1109 KGFHQMPGFDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQ 1168
KGFHQ G DY TFSPV+K TV++VL +AV+ W I+QLD+NNAFL G L E+VY+ Q
Sbjct: 966 KGFHQQHGLDYSNTFSPVIKSTTVQTVLDIAVSRSWDIRQLDINNAFLQGRLTEDVYVAQ 1025
Query: 1169 PPGFEAVD-PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQH 1227
PPGF D P+ VC L KAL+GLKQAPRAW++ L+ LL GF +S + SLFI N+
Sbjct: 1026 PPGFINPDRPNYVCHLKKALHGLKQAPRAWYQELRGFLLTCGFTNSVANTSLFIRQHNKD 1085
Query: 1228 STFMLVYVDDILITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQ 1287
++LVYVDD LITGS+++LI Q + L FSLKDLG+L YFL IE ++ G L L Q
Sbjct: 1086 YIYILVYVDDFLITGSNSNLIAQFITCLANRFSLKDLGQLSYFLEIEATRTKAG-LHLMQ 1144
Query: 1288 KKYIQDLLVKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGL 1337
++Y+ DLL K M +A +++PM+ + KLT + +P +R I+G L
Sbjct: 1145 RRYVLDLLTKTKMLDAKTVSTPMSPTPKLTLTSGTPIDNPGGYRQILGSL 1194
Score = 93.6 bits (231), Expect = 8e-19
Identities = 70/274 (25%), Positives = 127/274 (45%), Gaps = 9/274 (3%)
Query: 5 SSPVNNSETQRVTAAASK-NFKQIISVKLDETNYLQWKQQVEGVLRGTKMVRHVV-SPQI 62
S+P ++ET + + N KL TNY+ W +QV +L G + ++ S
Sbjct: 3 STPSGSTETIAFSETPTLLNVNMANITKLTPTNYIMWNRQVHALLDGYDLAGYIDGSVTA 62
Query: 63 PPVFLNDAAREAGTENPAYTEWEEQDSLLCTWILSTISSSLLSRFVRLRFSHQVWDEIHN 122
P + A A NPAY W+ QD L+ + IL TI++++ R + ++W+++ +
Sbjct: 63 PSEMITTAGVSAA--NPAYKFWKRQDKLIYSAILGTITTTIQPLLSRSNTAAEIWEKLKS 120
Query: 123 YCYTQMRTRSRQLRSELRTITKGTRSIAEFIARIRSISESLMSIGDPVAHRDLIETVLEA 182
T +Q+R ++ +KGT++I E+ + + L +G P+ H + IE +L
Sbjct: 121 IYATPSWGHIQQMRQHIKQWSKGTKTITEYFQGHTTRFDELALLGKPLEHAEQIEFLLGG 180
Query: 183 LPEEFNPIVATVNSQTEVISLDELESQLLTQEARNEKFKKALVGETASVNLT-HAENSGE 241
L E++ +V + + +L EL +LL +EA+ T S+ T HA N
Sbjct: 181 LSEDYKSVVDQTEIRDKPPTLTELLEKLLNREAK----LMCAAATTPSLPATAHAANYKG 236
Query: 242 KNGHNQPQTGSYPDQQFNISGNPTGNNSSQYFNP 275
+ +NQ + ++ S N + + F P
Sbjct: 237 NSNNNQYNNNNRNNKSHGSSYNSQQSIPASPFTP 270
>At2g16670 putative retroelement pol polyprotein
Length = 1333
Score = 616 bits (1588), Expect = e-176
Identities = 366/967 (37%), Positives = 529/967 (53%), Gaps = 41/967 (4%)
Query: 593 YKVWHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDFCSACCLGKSHR--LPSVSSKTVYNK 650
Y++WH+R+GHP VV S++ + + + C C K R P +KT+ +
Sbjct: 389 YELWHSRMGHPAARVV-SLIPESSVSVSSTHLNKACDVCHRAKQTRNSFPLSINKTL--R 445
Query: 651 PFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQ 710
FELI+CDLWGP SH G YFL +D YSR W++ L KS +NF M + Q
Sbjct: 446 IFELIYCDLWGPYRTPSHTGARYFLTIIDDYSRGVWLYLLNDKSEAPCHLKNFFAMTDRQ 505
Query: 711 YNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETGLTLLA 770
+N+ IK+V++D G EF T+F G+ H +C T +N VERKHRH++ L
Sbjct: 506 FNVKIKTVRSDNGTEFLCLTKFFQEQGVIHERSCVATPERNDRVERKHRHLLNVARALRF 565
Query: 771 NAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCFPFTRP 830
A LP+ +W LTA YLINR PS +LN+ +P+ LH + P + L+ FG C+ R
Sbjct: 566 QANLPIQFWGECVLTAAYLINRTPSSVLNDSTPYERLHKKQPRFDHLRVFGSLCYAHNRN 625
Query: 831 YNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQSS 889
K RS+ CVF+GY KG++ D F+S+DV+F+E +FP+ + Q+
Sbjct: 626 RGGDKFAERSRRCVFVGYPHGQKGWRLFDLEQNEFFVSRDVVFSELEFPFR--ISHEQNV 683
Query: 890 SPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSIQD 949
+ P+ L ++ Q++ PT + +SP I
Sbjct: 684 IEEEEEALWAPIVDGLI---EEEVHLGQNAG-PTPPICVSSP---------------ISP 724
Query: 950 THASSHSNHHNISP-GPIFNPTPISTHPPSPSPSSHSHNTYHSIS-VEPVTSQ---PSTQ 1004
+ SS S H SP PTP ++ + SPSS ++ + +S +P T+Q P
Sbjct: 725 SATSSRSEHSTSSPLDTEVVPTPATSTTSASSPSSPTNLQFLPLSRAKPTTAQAVAPPAV 784
Query: 1005 AEPHRIHPNNTHSMATRAKHGIVQK---RKHPTLLLTHIEPTGYRQAMKQPQWLQAMQLE 1061
P R N T K +V ++ P+ L + + R ++ +
Sbjct: 785 PPPRRQSTRNKAPPVT-LKDFVVNTTVCQESPSKLNSILYQLQKRDDTRRFSASHTTYVA 843
Query: 1062 HEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKE 1121
+A +N+TWT+ LP ++A+G +WV++ K N DGS+ +YKARLVA G Q G DY E
Sbjct: 844 IDAQEENHTWTIEDLPPGKRAIGSQWVYKVKHNSDGSVERYKARLVALGNKQKEGEDYGE 903
Query: 1122 TFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVDPSLVC 1181
TF+PV K TVR L +AV W I Q+DV+NAFL+G L EEVYM PPGFEA P+ VC
Sbjct: 904 TFAPVAKMATVRLFLDVAVKRNWEIHQMDVHNAFLHGDLREEVYMKLPPGFEASHPNKVC 963
Query: 1182 KLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILIT 1241
+L KALYGLKQAPR WFE+L + L + GF S D SLF L +L+YVDD++IT
Sbjct: 964 RLRKALYGLKQAPRCWFEKLTTALKRYGFQQSLADYSLFTLVKGSVRIKILIYVDDLIIT 1023
Query: 1242 GSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMA 1301
G+S QQ + L + F +KDLG L YFLGIEV S G + + Q+KY D++ + +
Sbjct: 1024 GNSQRATQQFKEYLASCFHMKDLGPLKYFLGIEVARSTTG-IYICQRKYALDIISETGLL 1082
Query: 1302 NANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAP 1361
P+ + KL S ++DP +R +VG L Y+ VTR ++++SV+ + +F+ P
Sbjct: 1083 GVKPANFPLEQNHKLGLSTSPLLTDPQRYRRLVGRLIYLAVTRLDLAFSVHILARFMQEP 1142
Query: 1362 LEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACIL 1421
EDHW A R++RYLK G+ + + Q +T +CD+DWA DP RRS +G +
Sbjct: 1143 REDHWAAALRVVRYLKADPGQGVFLRRSGDFQ---ITGWCDSDWAGDPMSRRSVTGYFVQ 1199
Query: 1422 LGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIPQIF-CDNL 1480
G + ISW KKQ V++SSAEAEYR+++ ++E+LW++ LL L V P I CD+
Sbjct: 1200 FGDSPISWKTKKQDTVSKSSAEAEYRAMSFLASELLWLKQLLFSLGVSHVQPMIMCCDSK 1259
Query: 1481 STVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLE 1540
S + +A NPV H RTKH+E+D FVR++ + + H+ Q+ADI TKPL F
Sbjct: 1260 SAIYIATNPVFHERTKHIEIDYHFVRDEFVKGVITPRHVGTTSQLADIFTKPLGRDCFSA 1319
Query: 1541 LRNKLRV 1547
R KL +
Sbjct: 1320 FRIKLGI 1326
>At1g26990 polyprotein, putative
Length = 1436
Score = 584 bits (1506), Expect = e-166
Identities = 393/1144 (34%), Positives = 577/1144 (50%), Gaps = 147/1144 (12%)
Query: 411 WYPDSGATHHVTPDANNLMDAASFSGSDQMYIG--NGQGLAINSIGSMSFSSPFSPNTTL 468
W DSGA+HHVT + N ++ D+ ++ NG + I G + + L
Sbjct: 421 WVIDSGASHHVTHERNLYH---TYKALDRTFVRLPNGHTVKIEGTGFIQLTD------AL 471
Query: 469 TLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQF 528
+L+N+L +P NL+SVS K F S+ C +++ +L KG Q
Sbjct: 472 SLHNVLFIPEFKFNLLSVSVLTKTLQSKVSFTSDECMIQALTKELMLGKGS-------QV 524
Query: 529 DQPYVPSVSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSN 588
Y+ ++ + S V SS +S C S N+S
Sbjct: 525 GNLYILNLDK----SLVDVSSFP------------GKSVC-----------SSVKNES-- 555
Query: 589 SLSLYKVWHNRLGHPHH---EVVRSVMKLCNQQLPNKSFTDFCSACCLGKSHRLPSVSSK 645
++WH RLGHP + + V+ L Q++ S C C L K LP S
Sbjct: 556 -----EMWHKRLGHPSFAKIDTLSDVLMLPKQKINKDS--SHCHVCHLSKQKHLPFKSVN 608
Query: 646 TVYNKPFELIFCDLWGPASVESHGGYSYFLICVDAYSRYTWIFPLKLKSHTLITFQNFKT 705
+ K FEL+ D WGP SV + Y YFL VD +SR TWI+ LK KS L F +F
Sbjct: 609 HIREKAFELVHIDTWGPFSVPTVDSYRYFLTIVDDFSRATWIYLLKQKSDVLTVFPSFLK 668
Query: 706 MVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRHIVETG 765
MVE QY+ + SV++D E + F + GI CP T QN VERKH+H++
Sbjct: 669 MVETQYHTKVCSVRSDNAHELK-FNELFAKEGIKADHPCPETPEQNFVVERKHQHLLNVA 727
Query: 766 LTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSFGCSCF 825
L+ + +PL YW LTA +LINRL SP++NN++P+ L PDY LK+FGC C+
Sbjct: 728 RALMFQSGIPLEYWGDCVLTAVFLINRLLSPVINNETPYERLTKGKPDYSSLKAFGCLCY 787
Query: 826 PFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFT 884
T P + K + R+K C+FLGY +KGYK LD T + IS+ VIF E FP++
Sbjct: 788 CSTSPKSRTKFDPRAKACIFLGYPMGYKGYKLLDIETYSVSISRHVIFYEDIFPFA---- 843
Query: 885 SGQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHH 944
SS+ + D P P N++ P QSSS +PH+
Sbjct: 844 ---SSNITDAAKDFFPHIYLPAPNNDEHLPLVQSSSD--------APHN----------- 881
Query: 945 HSIQDTHASSHSNHHNISPGPIFNPTPISTHPPSPSPSSHSHNTYHSISVEPVTSQPSTQ 1004
H+ S IF P+ + PS H + +H + P T T+
Sbjct: 882 --------------HDESSSMIFVPSEPKSTRQRKLPS-HLQD-FHCYNNTPTT----TK 921
Query: 1005 AEPHRIHPNNTHSMATRAKHGIVQKRKHPTLLLTHIEPTGYRQAMKQPQWLQAMQLEHEA 1064
P+ + ++S + + ++ P Y +A W AM E A
Sbjct: 922 TSPYPLTNYISYSYLSEPFGAFIN------IITATKLPQKYSEARLDKVWNDAMGKEISA 975
Query: 1065 LMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFS 1124
++ TW++ LPA + AVGCKW+ K DGSI ++KARLVAKG+ Q G D+ TFS
Sbjct: 976 FVRTGTWSICDLPAGKVAVGCKWIITIKFLADGSIERHKARLVAKGYTQQEGIDFFNTFS 1035
Query: 1125 PVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVDPSLVCKLN 1184
PV K VTV+ +L+LA KW + QLD++NA LNG LEEE+YM PPG+ +
Sbjct: 1036 PVAKMVTVKVLLSLAPKMKWYLHQLDISNALLNGDLEEEIYMKLPPGYSEIQ-------- 1087
Query: 1185 KALYGLKQAPRAWFERLKSTLLKLGFCSSKC--DPSLFILHANQHSTFMLVYVDDILITG 1242
G + +P A KC D +LF+ + +LVYVDDILI
Sbjct: 1088 ----GQEVSPNA-----------------KCHGDHTLFVKAQDGFFLVVLVYVDDILIAS 1126
Query: 1243 SSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMAN 1302
++ + +L +L++ F L+DLG+ +FLGIE+ + +G + L Q+KY+ DLL ++ ++
Sbjct: 1127 TTEAASAELTSQLSSFFQLRDLGEPKFFLGIEIARNADG-ISLCQRKYVLDLLASSDFSD 1185
Query: 1303 ANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPL 1362
+ PM + KL+K + D +R I+G LQY+ +TRP+I+++V+K+ Q+ SAP
Sbjct: 1186 CKPSSIPMEPNQKLSKDTGTLLEDGKQYRRILGKLQYLCLTRPDINFAVSKLAQYSSAPT 1245
Query: 1363 EDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILL 1422
+ H +A+ +ILRYLKGTI GL L F D+DW + PD RR +G I +
Sbjct: 1246 DIHLQALHKILRYLKGTIGQGLFYG---ADTNFDLRGFSDSDWQTCPDTRRCVTGFAIFV 1302
Query: 1423 GPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIP-QIFCDNLS 1481
G +L+SW +KKQ +V+ SSAEAEYR+++ A+ E++W+ +L K+P P ++CDN +
Sbjct: 1303 GNSLVSWRSKKQDVVSMSSAEAEYRAMSVATKELIWLGYILTAFKIPFTHPAYLYCDNEA 1362
Query: 1482 TVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRFLEL 1541
+ +A+N V H RTKH+E D VRE + + L + Q+AD LTKPL F E
Sbjct: 1363 ALHIANNSVFHERTKHIENDCHKVRECIEAGILKTIFVRTDNQLADTLTKPLYPKPFREN 1422
Query: 1542 RNKL 1545
+KL
Sbjct: 1423 NSKL 1426
>At4g27200 putative protein
Length = 819
Score = 518 bits (1333), Expect = e-146
Identities = 302/743 (40%), Positives = 426/743 (56%), Gaps = 37/743 (4%)
Query: 821 GCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCL-DPTGRMFISKDVIFNEYKFPY 879
G +CFP R Y +K S +CVFLGY+ +KGY+CL PTGR++IS+ VIF+E +P+
Sbjct: 15 GSACFPTLRDYAENKFNPCSLKCVFLGYNEKYKGYRCLYPPTGRLYISRHVIFDESVYPF 74
Query: 880 SELFTSGQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESN 939
S + TPL + ++ P+T +S + + L T++ P+
Sbjct: 75 SHTYKH-------LHPQPRTPLLAAWLRSSDSPAPSTSTSPSSRSPLFTSADFPPLPQ-R 126
Query: 940 QSNHHHSIQDTHASSHSNHHNISPGPIFNPTPISTHPPSPSPSSHSHNTYHSISVEPVTS 999
++ ++ + SH+++ P F+ + +T S S SH++ E
Sbjct: 127 KTPLLPTLVPISSVSHASNITTQQSPDFD-SERTTDFDSASIGDSSHSSQAGSDSEETIQ 185
Query: 1000 QPSTQAEPHRIHPNNTHSMATRAKHGIVQKRKHPTLL---LTHIEPTGYRQAMKQPQWLQ 1056
Q S N H M TRAK GI + L +++ EP A+K P W
Sbjct: 186 QASVNVHQTPAS-TNVHPMVTRAKVGISKPNPRYVFLSHKVSYPEPKTVTAALKHPGWTG 244
Query: 1057 AMQLEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPG 1116
AM E + TW+LVP +D +G KWVFRTK + DG++NK KAR+VAKGF Q G
Sbjct: 245 AMTEEIGNCSETQTWSLVPYKSDMHVLGSKWVFRTKLHADGTLNKLKARIVAKGFLQEEG 304
Query: 1117 FDYKETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVD 1176
DY ET+SPVV+ TVR VL LA W I+Q+DV NAFL+G L+E VYMTQP A +
Sbjct: 305 IDYLETYSPVVRTPTVRLVLHLATALNWDIKQMDVKNAFLHGDLKETVYMTQP----AAN 360
Query: 1177 PSLVCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVD 1236
VC L+K++YGLKQ+PRAWF++ + LL+ GF K DPSLFI N + +L+
Sbjct: 361 RDHVCLLHKSIYGLKQSPRAWFDKFSTFLLEFGFFCRKSDPSLFIYAHNNNLILLLL--- 417
Query: 1237 DILITGSSASLIQQLVKKLNAEFSLKDLGKLDY-FLGIEVHYSENGSLLLSQKKYIQDLL 1295
+ + L+ LN EF + D+G+ FLGI+V +NG L +SQ+KY +DLL
Sbjct: 418 --------SQTLTSLLAALNKEFRMTDMGQHSLTFLGIQVQRQQNG-LFMSQQKYAEDLL 468
Query: 1296 VKANMANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVC 1355
+ A+M + + +P+ + SDPT+FRSI G LQY+T+TRP+I ++VN VC
Sbjct: 469 IAASMEHCTPLPTPLPVQLDRVPHQEELFSDPTYFRSIAGKLQYLTLTRPDIQFAVNFVC 528
Query: 1356 QFLSAPLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRST 1415
Q + P + +KRILRY+KGTI G+ + P L A+ D+DW + RRS
Sbjct: 529 QKMHQPTISDFHLLKRILRYIKGTITMGISYS---RDSPTLLQAYSDSDWGNCKQTRRSV 585
Query: 1416 SGACILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTA-IPQ 1474
G C +G NL+SW +KK V+RSS EAEY+SL+ A++E+LW+ +LL+EL++P P+
Sbjct: 586 GGLCTFMGTNLVSWSSKKHPTVSRSSTEAEYKSLSDAASEILWLSTLLRELRIPLPDTPE 645
Query: 1475 IFCDNLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLS 1534
+FCDNLS V L NP H+RTKH ++D FVRE+V K L+V HIP Q+ADI TK L
Sbjct: 646 LFCDNLSAVYLTANPAFHARTKHFDIDFHFVRERVALKALVVKHIPGSEQIADIFTKSLP 705
Query: 1535 ASRFLELRNKLRVSDP--MSLRG 1555
F+ LR KL V+ P SLRG
Sbjct: 706 YEAFIHLRGKLGVTLPPTPSLRG 728
>At2g06840 putative retroelement pol polyprotein
Length = 1102
Score = 460 bits (1183), Expect = e-129
Identities = 296/847 (34%), Positives = 415/847 (48%), Gaps = 111/847 (13%)
Query: 701 QNFKTMVELQYNLPIKSVQTDGGGEFRPFTQFLTTLGITHRLTCPHTHHQNGSVERKHRH 760
+NF M + Q+ P++S+++D G EF T + GI H +C T QN VERKHRH
Sbjct: 356 RNFCAMADRQFRKPVRSIRSDNGTEFMCHTSYFQEHGILHETSCVDTPQQNARVERKHRH 415
Query: 761 IVETGLTLLANAKLPLHYWDHAFLTATYLINRLPSPILNNKSPFFLLHLQIPDYKFLKSF 820
I+ T L P PSP+L K+P+ +L + P Y L++F
Sbjct: 416 ILNVARTCLFQGNFPT-----------------PSPVLKGKTPYEVLFGKQPSYDMLRTF 458
Query: 821 GCSCFPFTRPYNNHKLELRSKECVFLGYSPSHKGYKCLDPTGRMFISKDVIFNEYKFPYS 880
GC C+ RP + K RS++C+F+GY +E P +
Sbjct: 459 GCLCYAHIRPRDKDKFASRSRKCIFIGYP-----------------------HETATPNT 495
Query: 881 ELFTSGQSSSPPTTSSDHTPLPSFLFPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQ 940
S P +TSSD N P T + P + +SPH +
Sbjct: 496 H-----DSIDPTSTSSDEN---------NTPPEPVTPQAEQPHSPSSISSPH--IVHNKG 539
Query: 941 SNHHHSIQDTHASSHSNHHNISPGPIFNPTPIST-HPPSPSPSSHSHNTYHSISVEPVTS 999
S H + + H SS SPG P + H P P H + P TS
Sbjct: 540 SVHSRHLNEDHDSS-------SPG---LPELLGKGHRPKHPPVYLKDYVAHKVHSSPHTS 589
Query: 1000 QPSTQAEPHRIHPNNTHSMATRAKHGIVQKRKHPTLLLTHIEPTGYRQAMKQPQWLQAMQ 1059
P ++ S A H +L E ++ + +W AMQ
Sbjct: 590 SPGLS--------DSNVSPTVSANHIAFM-----AAILDSNEQNHFKDDVLIKEWCDAMQ 636
Query: 1060 LEHEALMKNNTWTLVPLPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDY 1119
E EAL N+TW + LP ++A+ KWV++ K N DG++ ++KARLV G HQ G D+
Sbjct: 637 KEIEALEANHTWDVTDLPHGKKAISSKWVYKLKFNSDGTLERHKARLVVMGNHQKEGIDF 696
Query: 1120 KETFSPVVKPVTVRSVLTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFEAVDPSL 1179
KETF+PV K TVR +L +A W + Q+DV+NAFL+G LE+E
Sbjct: 697 KETFAPVAKMTTVRLLLAVAAAKDWDVFQMDVHNAFLHGDLEQE---------------- 740
Query: 1180 VCKLNKALYGLKQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDIL 1239
+LYGLKQAPR WF +L + L KLGF S D SLF L+ + LVYVDD +
Sbjct: 741 ------SLYGLKQAPRCWFAKLSTALRKLGFTQSYEDYSLFSLNRDGTVIHFLVYVDDFI 794
Query: 1240 ITGSSASLIQQLVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKAN 1299
I G++ I + L+ F +KDLGKL YFLG+EV +G LSQ+KY D++ +A
Sbjct: 795 IVGNNLKAIDHFKEHLHKCFHMKDLGKLKYFLGLEVSRGADG-FCLSQQKYALDIINEAG 853
Query: 1300 MANANGIASPMASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLS 1359
+ A PM KL S +P +R +V Y+T+TRP++SY+V+ + QF+
Sbjct: 854 LLGYKPSAVPMELHHKLGSISSPVFDNPAQYRRLVDRFIYLTITRPDLSYAVHILSQFMQ 913
Query: 1360 APLEDHWKAVKRILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGAC 1419
PLE HW A R++RYLKG+ G+L+ + LSLTA+CD+D+ P RRS S
Sbjct: 914 TPLEAHWHATLRLVRYLKGSPDQGILLR---SDRALSLTAYCDSDYNPCPRTRRSLSAYV 970
Query: 1420 ILLGPNLISWWAKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIP-QIFCD 1478
+ LG ISW KKQ V+ SSAEAEYR++A E+ W+++L+ L V P +FCD
Sbjct: 971 LYLGDTPISWKTKKQDTVSSSSAEAEYRAMAYTLKEIKWLKALMTTLGVDHTQPILLFCD 1030
Query: 1479 NLSTVSLAHNPVLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPLSASRF 1538
+ + + +A NPV H RTKH+E D VR+ V K + HI D+LTK L F
Sbjct: 1031 SQAAIHIAANPVFHERTKHIEKDCHQVRDAVTDKVISTPHI----STTDLLTKALPRPTF 1086
Query: 1539 LELRNKL 1545
L + L
Sbjct: 1087 ERLLSTL 1093
Score = 45.4 bits (106), Expect = 3e-04
Identities = 76/347 (21%), Positives = 121/347 (33%), Gaps = 88/347 (25%)
Query: 308 CQICYKTGHDASICYHRLSVPPQYEGYGSLGGNFGGNLGSGY------GPATGFGTHSNV 361
C C++ GHD + C+ P Y Y GG+ + PA G S
Sbjct: 84 CTHCHRQGHDVTDCFLVHGFPEWY--YEQKGGSRVSSDNREVVSRLENKPAKREGRSS-- 139
Query: 362 WMQGVGQRNPSYGAPRAPFPPQFGNSRPPA------PQAYITGNESTSSNSFNNGWYPDS 415
+G G+ + RAP G+ + Q + +E S N+ DS
Sbjct: 140 --KGNGRGRGRVNSARAPLSSSNGSDQITQLISLLQAQRPKSTSERLSGNTCLTDVIIDS 197
Query: 416 GATHHVTPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTTLTLNNLLH 475
GA+HH+T D + L+D S + +G+ ++ SS + L ++L
Sbjct: 198 GASHHMTGDCSILVDVFDIIPS-AVTKPDGKASCATKCVTLLLSSSYK------LQDVLF 250
Query: 476 VPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQFDQPYVPS 535
VP L+SVS+ K F ++ L+ + +Y F V S
Sbjct: 251 VPDFDCTLISVSKLLKQTGPF---------------SRTLIGAGEVRERVYYFTGVLVAS 295
Query: 536 VSRTASSSSVATSSLSLNNCFSPSSLSLSRSQCNNGSVYTPIHTSGSSNDSSNSLSLYKV 595
+T+S S+ +SG+ +
Sbjct: 296 AHKTSSDST----------------------------------SSGA------------L 309
Query: 596 WHNRLGHPHHEVVRSVMKLCNQQLPNKSFTDFCSACCLGKSHRLPSV 642
WH RLGHP + S+ + C+Q + D C C K LPS+
Sbjct: 310 WHRRLGHPSTSFLLSLPE-CHQSSKDLGKIDSCDTCSRAK-QTLPSL 354
>At4g10990 putative retrotransposon polyprotein
Length = 1203
Score = 453 bits (1165), Expect = e-127
Identities = 269/704 (38%), Positives = 401/704 (56%), Gaps = 38/704 (5%)
Query: 848 YSPSHKGYKCLD-PTGRMFISKDVIFNEYKFPYSELFTSGQSSSPPTTSSDHTPLPSFL- 905
Y +KGYK LD + + I+++V+F+E KFP+ + S + PLP+ L
Sbjct: 335 YPSGYKGYKVLDLESHSISITRNVVFHETKFPFKT--SKFLKESVDMFPNSILPLPAPLH 392
Query: 906 ----FPLNNKQCPTTQSSSTPTTTLHTASPHSSFPESNQSNHHHSIQDTHASSHSNHHNI 961
PL++ ++ +T ++AS SS P + + + ++S
Sbjct: 393 FVESMPLDDD----LRADDNNASTSNSASSASSIPPLPSTVNTQNTDALDIDTNSV---- 444
Query: 962 SPGPIFNPTPISTHPPSPSPSSHSHNTYHSISVEPVTSQ----PSTQAEPHRIHPNNTHS 1017
PI P + P+ H ++ S+ P TS PS+ P +I +
Sbjct: 445 ---PIARPKR-NAKAPAYLSEYHCNSVPFLSSLSPTTSTSIETPSSSIPPKKI--TTPYP 498
Query: 1018 MATRAKHGIVQKRKHPTLLLTHIE--PTGYRQAMKQPQWLQAMQLEHEALMKNNTWTLVP 1075
M+T + + H + ++E P + QAMK +W +A E AL +N TW +
Sbjct: 499 MSTAISYDKLTPLFHSYICAYNVETEPKAFTQAMKSEKWTRAANEELHALEQNKTWIVES 558
Query: 1076 LPADRQAVGCKWVFRTKQNPDGSINKYKARLVAKGFHQMPGFDYKETFSPVVKPVTVRSV 1135
L + VGCKWVF K NPDGSI +YKARLVA+GF Q G DY ETFSPV K +V+ +
Sbjct: 559 LTEGKNVVGCKWVFTIKYNPDGSIERYKARLVAQGFTQQEGIDYMETFSPVAKFGSVKLL 618
Query: 1136 LTLAVTNKWCIQQLDVNNAFLNGYLEEEVYMTQPPGFE-----AVDPSLVCKLNKALYGL 1190
L LA W + Q+DV+NAFL+G L+EE+YM+ P G+ ++ VC+L K+LYGL
Sbjct: 619 LGLAAATGWSLTQMDVSNAFLHGELDEEIYMSLPQGYTPPTGISLPSKPVCRLLKSLYGL 678
Query: 1191 KQAPRAWFERLKSTLLKLGFCSSKCDPSLFILHANQHSTFMLVYVDDILITGSSASLIQQ 1250
KQA R W++RL S L F S D ++F+ + +LVYVDD++I + +S ++
Sbjct: 679 KQASRQWYKRLSSVFLGANFIQSPADNTMFVKVSCTSIIVVLVYVDDLMIASNDSSAVEN 738
Query: 1251 LVKKLNAEFSLKDLGKLDYFLGIEVHYSENGSLLLSQKKYIQDLLVKANMANANGIASPM 1310
L + L +EF +KDLG +FLG+E+ S G + + Q+KY Q+LL ++ + PM
Sbjct: 739 LKELLRSEFKIKDLGPARFFLGLEIARSSEG-ISVCQRKYAQNLLEDVGLSGCKPSSIPM 797
Query: 1311 ASSTKLTKYGSNHVSDPTFFRSIVGGLQYVTVTRPEISYSVNKVCQFLSAPLEDHWKAVK 1370
+ LTK + + T +R +VG L Y+ +TRP+I+++V+ + QFLSAP + H +A
Sbjct: 798 DPNLHLTKEMGTLLPNATSYRELVGRLLYLCITRPDITFAVHTLSQFLSAPTDIHMQAAH 857
Query: 1371 RILRYLKGTIHHGLLINPAPMHQPLSLTAFCDADWASDPDDRRSTSGACILLGPNLISWW 1430
++LRYLKG GL+ + + L L F DADW + D RRS +G CI LG +LI+W
Sbjct: 858 KVLRYLKGNPGQGLMYSAS---SELCLNGFSDADWGTCKDSRRSVTGFCIYLGTSLITWK 914
Query: 1431 AKKQTLVARSSAEAEYRSLAQASAEVLWIQSLLKELKVPTAIP-QIFCDNLSTVSLAHNP 1489
+KKQ++V+RSS E+EYRSLAQA+ E++W+Q LLK+L V P ++FCDN S + LA NP
Sbjct: 915 SKKQSVVSRSSTESEYRSLAQATCEIIWLQQLLKDLHVTMTCPAKLFCDNKSALHLATNP 974
Query: 1490 VLHSRTKHMELDIFFVREKVISKDLIVSHIPAQYQVADILTKPL 1533
V H RTKH+E+D VR+++ + L H+P Q+ADILTKPL
Sbjct: 975 VFHERTKHIEIDCHTVRDQIKAGKLKTLHVPTGNQLADILTKPL 1018
Score = 62.0 bits (149), Expect = 3e-09
Identities = 32/84 (38%), Positives = 48/84 (57%), Gaps = 1/84 (1%)
Query: 673 YFLICVDAYSRYTWIFPLKLKSHTLITFQNFKTMVELQYNLPIKSVQTDGGGEFRPFTQF 732
YFL VD +R TW++ +K KS F F ++ QYN IK++++D E FT+F
Sbjct: 252 YFLTLVDDCTRTTWVYMMKNKSEVSNIFPVFVKLIFTQYNAKIKAIRSDNVKEL-AFTKF 310
Query: 733 LTTLGITHRLTCPHTHHQNGSVER 756
+ G+ H+ +C +T QN VER
Sbjct: 311 VKEQGMIHQFSCAYTPQQNSVVER 334
Score = 50.8 bits (120), Expect = 6e-06
Identities = 41/142 (28%), Positives = 67/142 (46%), Gaps = 14/142 (9%)
Query: 408 NNGWYPDSGATHHVTPDANNLMDAASFSGSDQMYIGNGQGLAINSIGSMSFSSPFSPNTT 467
++ W DSGA+ HV D + SG + + NG +AI G++ +S T
Sbjct: 96 SDAWIIDSGASSHVCSDLTMFRELIHVSGVT-VTLPNGTRVAITHTGTICITS------T 148
Query: 468 LTLNNLLHVPSITKNLVSVSQFCKDNNVFFEFHSNICYVKSQDSTKILLKGHLGDDGLYQ 527
L L+N+L VP NL+SV K + F ++ CY+ Q+ T+ L+ G
Sbjct: 149 LILHNVLLVPDFKFNLISVCCLVKTLSYSAHFFADCCYI--QELTRGLMIGR-----GKT 201
Query: 528 FDQPYVPSVSRTASSSSVATSS 549
++ Y+ RT+ S S+ +S
Sbjct: 202 YNNLYILETQRTSFSPSLPAAS 223
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.132 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,623,493
Number of Sequences: 26719
Number of extensions: 1779900
Number of successful extensions: 9611
Number of sequences better than 10.0: 321
Number of HSP's better than 10.0 without gapping: 139
Number of HSP's successfully gapped in prelim test: 196
Number of HSP's that attempted gapping in prelim test: 7146
Number of HSP's gapped (non-prelim): 1290
length of query: 1557
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1445
effective length of database: 8,326,068
effective search space: 12031168260
effective search space used: 12031168260
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC149038.3


