

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148996.15 + phase: 0
(153 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
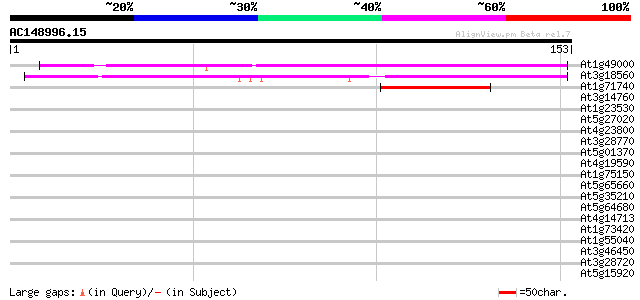
Score E
Sequences producing significant alignments: (bits) Value
At1g49000 hypothetical protein; similar to ESTs gb|R64892.1, gb|... 112 6e-26
At3g18560 unknown protein 97 4e-21
At1g71740 hypothetical protein 47 4e-06
At3g14760 unknown protein 37 0.004
At1g23530 unknown protein 32 0.10
At5g27020 putative protein 31 0.23
At4g23800 98b like protein 31 0.23
At3g28770 hypothetical protein 30 0.67
At5g01370 unknown protein 29 0.87
At4g19590 putative protein 29 0.87
At1g75150 hypothetical protein 28 1.5
At5g65660 unknown protein 28 2.5
At5g35210 putative protein 28 2.5
At5g64680 unknown protein 27 3.3
At4g14713 unknown protein 27 3.3
At1g73420 27 3.3
At1g55040 putative protein 27 3.3
At3g46450 unknown protein 27 4.3
At3g28720 hypothetical protein 27 4.3
At5g15920 putative protein 27 5.7
>At1g49000 hypothetical protein; similar to ESTs gb|R64892.1,
gb|T04316.1
Length = 156
Score = 112 bits (281), Expect = 6e-26
Identities = 64/149 (42%), Positives = 89/149 (58%), Gaps = 9/149 (6%)
Query: 9 SPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRD-----EEWRFDLKTPP 63
SP+ + F V++ + L +R ANRL +K K S + E +R+++ +
Sbjct: 11 SPSHKRLSVSFLVSM---MVLCARHANRLSKKLKLKSKKRTCSGGEGGGERFRWNMISSS 67
Query: 64 KSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGV 123
S +PK+L + +SNKA++ +K EE+ G+WQ+EILMGGKCEPLDFSGV
Sbjct: 68 MSS-PRPKELFTTLSNKAMTMVRRKNPPEEKATAMEEEHGLWQREILMGGKCEPLDFSGV 126
Query: 124 IYYDINGKQTREVPIRSPRASPLPGYLTR 152
IYYD NG+ EVP RSPR +PLP Y TR
Sbjct: 127 IYYDSNGRLLNEVPPRSPRGTPLPSYPTR 155
>At3g18560 unknown protein
Length = 195
Score = 96.7 bits (239), Expect = 4e-21
Identities = 67/183 (36%), Positives = 96/183 (51%), Gaps = 40/183 (21%)
Query: 5 TSPESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKT--- 61
++PE + H +S ++ L +R A+R+ +K K T K E++ K+
Sbjct: 4 STPEHVSSAHKRISVSFLVS-LMVLCARHASRVSKKLKPKKTRKQTHLEDYLESPKSNGN 62
Query: 62 --------------PPK--SPM-AKPKKLLSNISNKALSQFGKKKQR------------- 91
P + SPM +PK+L + +SNKA++ G+K +
Sbjct: 63 GSEDGRGGGRFGWSPARTFSPMRVRPKELYTTLSNKAMTMVGRKNKAYDGGPTKKTAVEM 122
Query: 92 --EEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRASPLPGY 149
EE E+E GVWQ+EILMGGKCEPLD+SGVIYYD +G Q ++VP RSPRAS +P
Sbjct: 123 VMEEDEEEY----GVWQREILMGGKCEPLDYSGVIYYDCSGHQLKQVPPRSPRASLVPER 178
Query: 150 LTR 152
TR
Sbjct: 179 PTR 181
>At1g71740 hypothetical protein
Length = 130
Score = 47.0 bits (110), Expect = 4e-06
Identities = 20/30 (66%), Positives = 24/30 (79%)
Query: 102 GGVWQKEILMGGKCEPLDFSGVIYYDINGK 131
G +WQK ILMGGKC+ DFSGVI YD +G+
Sbjct: 86 GPLWQKNILMGGKCQLPDFSGVILYDADGQ 115
>At3g14760 unknown protein
Length = 168
Score = 37.0 bits (84), Expect = 0.004
Identities = 31/98 (31%), Positives = 41/98 (41%), Gaps = 14/98 (14%)
Query: 44 SSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQF-----GKKKQREERE--- 95
+S K + D EW + ++ K K S + + F K + E R
Sbjct: 42 NSRGKQVADAEWASETAFGSRTMEEKRKCKWSKVKRTLMGSFCWSSAAKWMEMETRRQPP 101
Query: 96 ----KEGWGNG--GVWQKEILMGGKCEPLDFSGVIYYD 127
KE N VWQ+ ILMG KCE FSG+I YD
Sbjct: 102 LLAVKERSLNAVDPVWQRPILMGEKCELPRFSGLILYD 139
>At1g23530 unknown protein
Length = 189
Score = 32.3 bits (72), Expect = 0.10
Identities = 36/128 (28%), Positives = 50/128 (38%), Gaps = 35/128 (27%)
Query: 5 TSPESPNRHHSNGGFFVTISAILALLS-------RKANRLKEKAK------------SSS 45
T P S + N G + A++ +LS R NR AK SS
Sbjct: 29 TKPSSSSGSSGNFGTVFIVLAVIFVLSALACVFGRLCNRESRAAKQQHSKHPKHEKASSK 88
Query: 46 TTKPIR----------DEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQFGKKKQREERE 95
++ IR D ++ FD+K P P+ KP S +FG +R ER
Sbjct: 89 KSREIRPVEREPRERGDMDFGFDMKRP--EPIEKP----SGRDGGEDIEFGFDNKRGERG 142
Query: 96 KEGWGNGG 103
+EG G GG
Sbjct: 143 REGGGGGG 150
>At5g27020 putative protein
Length = 164
Score = 31.2 bits (69), Expect = 0.23
Identities = 15/44 (34%), Positives = 26/44 (59%), Gaps = 2/44 (4%)
Query: 87 KKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDING 130
+++ R EG + +K ILMG +C+P+ +GV+ YD +G
Sbjct: 117 RRRNRRRSLSEGMEEEELCKKRILMGERCKPM--NGVLQYDGDG 158
>At4g23800 98b like protein
Length = 456
Score = 31.2 bits (69), Expect = 0.23
Identities = 19/70 (27%), Positives = 33/70 (47%), Gaps = 1/70 (1%)
Query: 23 ISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKAL 82
+SA L + + L+E+ KS I EEW+ +L K+P K K +A+
Sbjct: 259 VSAFLVYANERRAALREENKSVVEVAKITGEEWK-NLSDKKKAPYEKVAKKNKETYLQAM 317
Query: 83 SQFGKKKQRE 92
++ + K+ E
Sbjct: 318 EEYKRTKEEE 327
>At3g28770 hypothetical protein
Length = 2081
Score = 29.6 bits (65), Expect = 0.67
Identities = 19/64 (29%), Positives = 29/64 (44%)
Query: 33 KANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQFGKKKQRE 92
K KEK KS + +D E R K +S K KK K S+ K K++E
Sbjct: 1023 KEEAKKEKKKSQDKKREEKDSEERKSKKEKEESRDLKAKKKEEETKEKKESENHKSKKKE 1082
Query: 93 EREK 96
++++
Sbjct: 1083 DKKE 1086
Score = 27.7 bits (60), Expect = 2.5
Identities = 19/64 (29%), Positives = 29/64 (44%), Gaps = 6/64 (9%)
Query: 32 RKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQFGKKKQR 91
+K+ K + K S K +++E DLK K K KK N +K KK+ +
Sbjct: 1031 KKSQDKKREEKDSEERKSKKEKEESRDLKAKKKEEETKEKKESENHKSK------KKEDK 1084
Query: 92 EERE 95
+E E
Sbjct: 1085 KEHE 1088
>At5g01370 unknown protein
Length = 427
Score = 29.3 bits (64), Expect = 0.87
Identities = 23/71 (32%), Positives = 32/71 (44%), Gaps = 3/71 (4%)
Query: 27 LALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQ-- 84
L L K + +E SSS ++ R E+ D K PP P A KK ++ K +
Sbjct: 170 LLLYLDKLDEKRELVGSSSFSRSKRFEKVVEDSKKPPLPPSAPKKKENEKVAKKFKDEPR 229
Query: 85 -FGKKKQREER 94
KKK+ E R
Sbjct: 230 KVAKKKKSENR 240
>At4g19590 putative protein
Length = 345
Score = 29.3 bits (64), Expect = 0.87
Identities = 18/66 (27%), Positives = 32/66 (48%), Gaps = 3/66 (4%)
Query: 10 PNRHHSNG--GFFVTISAILALLSRKANRLK-EKAKSSSTTKPIRDEEWRFDLKTPPKSP 66
P++++ G G F + A LLS K R+ ++ + + KP R + + K PP P
Sbjct: 85 PDKNNCEGAEGAFKLVLAAWCLLSDKVKRIAYDQKRKLNEVKPKRSRKQKQPPKKPPNQP 144
Query: 67 MAKPKK 72
+P +
Sbjct: 145 KQQPNQ 150
>At1g75150 hypothetical protein
Length = 753
Score = 28.5 bits (62), Expect = 1.5
Identities = 20/59 (33%), Positives = 28/59 (46%), Gaps = 4/59 (6%)
Query: 29 LLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQFGK 87
L R +++ KAKSSS+T EE +K P AKP S+ +AL + K
Sbjct: 576 LQQRLYKKMELKAKSSSSTADENSEEILRHIKKPEIKKKAKP----SSFKERALMEINK 630
>At5g65660 unknown protein
Length = 136
Score = 27.7 bits (60), Expect = 2.5
Identities = 11/24 (45%), Positives = 16/24 (65%)
Query: 49 PIRDEEWRFDLKTPPKSPMAKPKK 72
P R E+ D++TPP+SP KP +
Sbjct: 107 PPRPEKLTVDVQTPPQSPPVKPAR 130
>At5g35210 putative protein
Length = 1606
Score = 27.7 bits (60), Expect = 2.5
Identities = 17/47 (36%), Positives = 25/47 (53%), Gaps = 9/47 (19%)
Query: 7 PESPNRHHSNGGFFVTISAILALLS---------RKANRLKEKAKSS 44
P + N H++NG V+ +A LA+LS RK N K+ A S+
Sbjct: 648 PNTYNNHYTNGELAVSAAASLAVLSSEETHEPDLRKYNSAKKAASSN 694
>At5g64680 unknown protein
Length = 203
Score = 27.3 bits (59), Expect = 3.3
Identities = 15/67 (22%), Positives = 29/67 (42%)
Query: 31 SRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQFGKKKQ 90
S+K++ L + ++ S T+ R++ PKSP K + S + S +
Sbjct: 131 SKKSSNLTQSSRHKSGTRSSREKSMFSGFTETPKSPKRKNSESSSGKHRSSTSTVSGSSE 190
Query: 91 REEREKE 97
R + K+
Sbjct: 191 RSAKSKK 197
>At4g14713 unknown protein
Length = 313
Score = 27.3 bits (59), Expect = 3.3
Identities = 23/86 (26%), Positives = 38/86 (43%), Gaps = 5/86 (5%)
Query: 61 TPPKSPMAKPKKLLSNISNKALSQFGKKKQREER--EKEGWGNGGVWQKEILMGGKCEPL 118
+P KS +AKP KLL+ L++ +K +++ + W Q+ + + EP
Sbjct: 6 SPAKSILAKPLKLLTEEDISQLTREDCRKFLKDKGMRRPSWNKSQAIQQVLSLKALYEPG 65
Query: 119 DFSGVIYYDINGKQTREVPIRSPRAS 144
D SG I K P+ PR +
Sbjct: 66 DDSGA---GIFRKILVSQPVNPPRVT 88
>At1g73420
Length = 89
Score = 27.3 bits (59), Expect = 3.3
Identities = 17/68 (25%), Positives = 31/68 (45%)
Query: 30 LSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQFGKKK 89
LSRK R + K + +++ R +++T PK+ + + K Q G +
Sbjct: 13 LSRKKPREHMQRKKFQIPGEMDEKQRRIEIRTIPKTMRNSTEIKREQVKEKRELQQGMHE 72
Query: 90 QREEREKE 97
+ EE E+E
Sbjct: 73 EEEEEEEE 80
>At1g55040 putative protein
Length = 849
Score = 27.3 bits (59), Expect = 3.3
Identities = 21/85 (24%), Positives = 35/85 (40%), Gaps = 7/85 (8%)
Query: 5 TSPESPNR-----HHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDL 59
TSP+S +H +S + + R EK ++ + R + L
Sbjct: 36 TSPQSNTESTHEPNHKPSSLSSRLSFVFDQIDAIEKRHSEKDETLERIRAWRQSKQTHQL 95
Query: 60 KTPPKSPMAKP--KKLLSNISNKAL 82
KTPP +P P +++ S +SN L
Sbjct: 96 KTPPSAPQQDPDLREVESVVSNDKL 120
>At3g46450 unknown protein
Length = 486
Score = 26.9 bits (58), Expect = 4.3
Identities = 11/33 (33%), Positives = 19/33 (57%)
Query: 38 KEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKP 70
+ K +SSS T+ + D W ++ +TP S +P
Sbjct: 417 RRKRRSSSETEKLDDSHWSYNAQTPDLSYEDEP 449
>At3g28720 hypothetical protein
Length = 687
Score = 26.9 bits (58), Expect = 4.3
Identities = 12/31 (38%), Positives = 20/31 (63%)
Query: 91 REEREKEGWGNGGVWQKEILMGGKCEPLDFS 121
++E EKE G+GGV+ I +G + +P +S
Sbjct: 188 KQEYEKEKHGDGGVYIYLISLGSQAKPYAYS 218
>At5g15920 putative protein
Length = 1053
Score = 26.6 bits (57), Expect = 5.7
Identities = 13/31 (41%), Positives = 20/31 (63%)
Query: 66 PMAKPKKLLSNISNKALSQFGKKKQREEREK 96
P+AK ++L S ++ S GKK Q+E+ EK
Sbjct: 367 PVAKLEELSSQVTELHHSINGKKNQKEDNEK 397
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.133 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,904,564
Number of Sequences: 26719
Number of extensions: 169671
Number of successful extensions: 484
Number of sequences better than 10.0: 37
Number of HSP's better than 10.0 without gapping: 21
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 458
Number of HSP's gapped (non-prelim): 42
length of query: 153
length of database: 11,318,596
effective HSP length: 90
effective length of query: 63
effective length of database: 8,913,886
effective search space: 561574818
effective search space used: 561574818
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148996.15


