

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148995.8 + phase: 0
(325 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
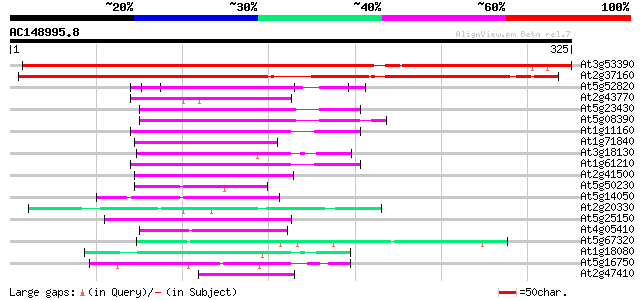
Score E
Sequences producing significant alignments: (bits) Value
At3g53390 putative protein 508 e-144
At2g37160 unknown protein 442 e-124
At5g52820 Notchless protein homolog 54 1e-07
At2g43770 putative splicing factor 50 1e-06
At5g23430 unknown protein 50 2e-06
At5g08390 katanin p80 subunit - like protein 50 2e-06
At1g11160 hypothetical protein 49 5e-06
At1g71840 unknown protein (At1g71840) 48 6e-06
At3g18130 protein kinase C-receptor/G-protein, putative 48 8e-06
At1g61210 47 1e-05
At2g41500 putative U4/U6 small nuclear ribonucleoprotein 45 4e-05
At5g50230 putative protein 45 5e-05
At5g14050 unknown protein 45 5e-05
At2g20330 putative WD-40 repeat protein 45 7e-05
At5g25150 transcription initiation factor IID-associated factor-... 44 9e-05
At4g05410 U3 snoRNP-associated -like protein 44 9e-05
At5g67320 unknown protein 44 2e-04
At1g18080 putative guanine nucleotide-binding protein beta sub... 44 2e-04
At5g16750 WD40-repeat protein 43 2e-04
At2g47410 putative WD-40 repeat protein 43 3e-04
>At3g53390 putative protein
Length = 556
Score = 508 bits (1307), Expect = e-144
Identities = 248/326 (76%), Positives = 279/326 (85%), Gaps = 15/326 (4%)
Query: 8 RCTCISWVPGGDGAFVVAHADGNLYNKDGAGESSFPILKDQTLFSVAHARYSKSNPIARW 67
RCT ISWVPGGDGAFVVAHADGNLYNKDGA ES+FP ++D T FSV A+YSKSNP+ARW
Sbjct: 238 RCTSISWVPGGDGAFVVAHADGNLYNKDGATESTFPAIRDPTQFSVDKAKYSKSNPVARW 297
Query: 68 HICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKY 127
HICQGSINSI+FS DGA+LATVGRDGYLR+FD+ + LVCGGKSYYG LLCC+WSMDGKY
Sbjct: 298 HICQGSINSIAFSNDGAHLATVGRDGYLRIFDFLTQKLVCGGKSYYGALLCCSWSMDGKY 357
Query: 128 ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVG 187
ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDS WSSPNS+ +GE + YRFGSVG
Sbjct: 358 ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSNWSSPNSDGSGEHVMYRFGSVG 417
Query: 188 QDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMR 247
QDTQLLLW+LEMDEIVVPLRR P GSPT+S GS S+HWDN +P+GTLQPAP R
Sbjct: 418 QDTQLLLWDLEMDEIVVPLRRAPG------GSPTYSTGS-SAHWDNVIPMGTLQPAPCKR 470
Query: 248 DVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPGVAESQPSNSE-----T 302
DVPK+SP++AHRVHTEPLS L+FTQESV+TACREGHIKIWTRP +E+Q ++SE
Sbjct: 471 DVPKLSPIIAHRVHTEPLSGLMFTQESVVTACREGHIKIWTRPSESETQTNSSEANPTTA 530
Query: 303 LLATSLKE--KPSLSSK-ISNSIYKQ 325
LL+TS + K SLSSK IS S +KQ
Sbjct: 531 LLSTSFTKDNKASLSSKIISGSTFKQ 556
>At2g37160 unknown protein
Length = 444
Score = 442 bits (1136), Expect = e-124
Identities = 220/313 (70%), Positives = 251/313 (79%), Gaps = 32/313 (10%)
Query: 6 NSRCTCISWVPGGDGAFVVAHADGNLYNKDGAGESSFPILKDQTLFSVAHARYSKSNPIA 65
NSRCT I+WVPGGDG+FV AHADGNLYNK+GA +SSF ++D T FSV A+YSKSNP+A
Sbjct: 150 NSRCTSIAWVPGGDGSFVAAHADGNLYNKEGATDSSFSAIRDPTQFSVDKAKYSKSNPVA 209
Query: 66 RWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDG 125
RWHI QG+INSI+FS DGAYLATVGRDGYLR+FD++ + LVCG KSYYG LLCCAWSMDG
Sbjct: 210 RWHIGQGAINSIAFSNDGAYLATVGRDGYLRIFDFSTQKLVCGVKSYYGALLCCAWSMDG 269
Query: 126 KYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGS 185
KY+LTGGEDDLVQVWSMEDRKVVAW EG +GE + YRFGS
Sbjct: 270 KYLLTGGEDDLVQVWSMEDRKVVAW-EG---------------------SGENVMYRFGS 307
Query: 186 VGQDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPS 245
VGQDTQLLLW+LEMDEIVVPLRR PPG GSPT+S GSQS+HWDN VP+GTLQPAP
Sbjct: 308 VGQDTQLLLWDLEMDEIVVPLRR-PPG-----GSPTYSTGSQSAHWDNIVPMGTLQPAPC 361
Query: 246 MRDVPKISPLVAHRVHTEPLSSLIFTQESVLTACREGHIKIWTRPGVAESQPSNSETLLA 305
MRDVPK+SP+VAHRVHTEPLS LIFTQES++TACREGH+KIWTRP ++QPS+S
Sbjct: 362 MRDVPKLSPVVAHRVHTEPLSGLIFTQESLITACREGHLKIWTRP---DTQPSSSSE-AT 417
Query: 306 TSLKEKPSLSSKI 318
KPSL+SK+
Sbjct: 418 NPTTSKPSLTSKV 430
>At5g52820 Notchless protein homolog
Length = 473
Score = 53.5 bits (127), Expect = 1e-07
Identities = 33/132 (25%), Positives = 67/132 (50%), Gaps = 11/132 (8%)
Query: 77 ISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDL 136
+SFS DG LA+ D +R++D E + K + +L AWS DGK++++G +
Sbjct: 115 VSFSPDGKQLASGSGDTTVRLWDLYTETPLFTCKGHKNWVLTVAWSPDGKHLVSGSKSGE 174
Query: 137 VQVWSMEDRKVVAWG-EGHNSWVSGVAFDS-YWSSPNSNDNGETITYRFGSVGQDTQLLL 194
+ W+ + ++ GH W++G++++ + SSP RF + +D +
Sbjct: 175 ICCWNPKKGELEGSPLTGHKKWITGISWEPVHLSSP---------CRRFVTSSKDGDARI 225
Query: 195 WELEMDEIVVPL 206
W++ + + ++ L
Sbjct: 226 WDITLKKSIICL 237
Score = 49.7 bits (117), Expect = 2e-06
Identities = 33/126 (26%), Positives = 60/126 (47%), Gaps = 13/126 (10%)
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILT 130
Q +N + FS DG ++A+ D +R+++ V + + G + +WS D + +L+
Sbjct: 360 QQLVNHVYFSPDGKWIASASFDKSVRLWNGITGQFVTVFRGHVGPVYQVSWSADSRLLLS 419
Query: 131 GGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDT 190
G +D +++W + +K+ GH V V WS +GE + S G+D
Sbjct: 420 GSKDSTLKIWEIRTKKLKQDLPGHADEVFAVD----WS-----PDGEKVV----SGGKDR 466
Query: 191 QLLLWE 196
L LW+
Sbjct: 467 VLKLWK 472
Score = 42.7 bits (99), Expect = 3e-04
Identities = 20/78 (25%), Positives = 38/78 (48%), Gaps = 1/78 (1%)
Query: 88 TVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKV 147
T +DG R++D T + + + + C W DG I TG +D +++W K+
Sbjct: 216 TSSKDGDARIWDITLKKSIICLSGHTLAVTCVKWGGDG-IIYTGSQDCTIKMWETTQGKL 274
Query: 148 VAWGEGHNSWVSGVAFDS 165
+ +GH W++ +A +
Sbjct: 275 IRELKGHGHWINSLALST 292
Score = 37.7 bits (86), Expect = 0.009
Identities = 16/48 (33%), Positives = 28/48 (58%)
Query: 116 LLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
+LC ++S DGK + +G D V++W + + +GH +WV VA+
Sbjct: 112 VLCVSFSPDGKQLASGSGDTTVRLWDLYTETPLFTCKGHKNWVLTVAW 159
>At2g43770 putative splicing factor
Length = 343
Score = 50.4 bits (119), Expect = 1e-06
Identities = 31/101 (30%), Positives = 46/101 (44%), Gaps = 8/101 (7%)
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFD---YTKEHLVCG-----GKSYYGGLLCCAWS 122
Q +I +S S DG+YL T G D L V+D Y ++ ++ LL C+WS
Sbjct: 222 QDTITGMSLSPDGSYLLTNGMDNKLCVWDMRPYAPQNRCVKIFEGHQHNFEKNLLKCSWS 281
Query: 123 MDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
DG + G D +V +W R+ + GH V+ F
Sbjct: 282 PDGTKVTAGSSDRMVHIWDTTSRRTIYKLPGHTGSVNECVF 322
Score = 37.4 bits (85), Expect = 0.011
Identities = 25/85 (29%), Positives = 39/85 (45%), Gaps = 4/85 (4%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
I ++SFS + T G D ++V+D K + + + + S DG Y+LT G
Sbjct: 183 ITAVSFSDAADKIFTGGVDNDVKVWDLRKGEATMTLEGHQDTITGMSLSPDGSYLLTNGM 242
Query: 134 DDLVQVWSME----DRKVVAWGEGH 154
D+ + VW M + V EGH
Sbjct: 243 DNKLCVWDMRPYAPQNRCVKIFEGH 267
Score = 33.9 bits (76), Expect = 0.12
Identities = 22/90 (24%), Positives = 42/90 (46%), Gaps = 5/90 (5%)
Query: 73 SINSISFSADGAYLATVGRDGYL---RVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYIL 129
++ ++ F+ G +A+ D + RV K +V K + +L W+ DG I+
Sbjct: 55 AVYTMKFNPAGTLIASGSHDREIFLWRVHGDCKNFMVL--KGHKNAILDLHWTSDGSQIV 112
Query: 130 TGGEDDLVQVWSMEDRKVVAWGEGHNSWVS 159
+ D V+ W +E K + H+S+V+
Sbjct: 113 SASPDKTVRAWDVETGKQIKKMAEHSSFVN 142
Score = 32.0 bits (71), Expect = 0.47
Identities = 19/70 (27%), Positives = 32/70 (45%), Gaps = 13/70 (18%)
Query: 128 ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVG 187
I TGG D+ V+VW + + EGH ++G++ S D +T G
Sbjct: 195 IFTGGVDNDVKVWDLRKGEATMTLEGHQDTITGMSL--------SPDGSYLLTN-----G 241
Query: 188 QDTQLLLWEL 197
D +L +W++
Sbjct: 242 MDNKLCVWDM 251
>At5g23430 unknown protein
Length = 837
Score = 50.1 bits (118), Expect = 2e-06
Identities = 27/128 (21%), Positives = 60/128 (46%), Gaps = 13/128 (10%)
Query: 76 SISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDD 135
S+ F G + A+ D L+++D K+ + K + G+ ++ DG+++++GGED+
Sbjct: 106 SVDFHPFGEFFASGSLDTNLKIWDIRKKGCIHTYKGHTRGVNVLRFTPDGRWVVSGGEDN 165
Query: 136 LVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLW 195
+V+VW + K++ + H + + F + + + D + W
Sbjct: 166 IVKVWDLTAGKLLTEFKSHEGQIQSLDFHPH-------------EFLLATGSADRTVKFW 212
Query: 196 ELEMDEIV 203
+LE E++
Sbjct: 213 DLETFELI 220
Score = 40.8 bits (94), Expect = 0.001
Identities = 22/91 (24%), Positives = 45/91 (49%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
+N + F+ DG ++ + G D ++V+D T L+ KS+ G + + + TG
Sbjct: 146 VNVLRFTPDGRWVVSGGEDNIVKVWDLTAGKLLTEFKSHEGQIQSLDFHPHEFLLATGSA 205
Query: 134 DDLVQVWSMEDRKVVAWGEGHNSWVSGVAFD 164
D V+ W +E +++ G + V ++F+
Sbjct: 206 DRTVKFWDLETFELIGSGGPETAGVRCLSFN 236
Score = 35.8 bits (81), Expect = 0.032
Identities = 35/164 (21%), Positives = 66/164 (39%), Gaps = 25/164 (15%)
Query: 126 KYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGS 185
+ ++TGGED V +W++ + GH+S + V FD+ E + +
Sbjct: 30 RVLVTGGEDHKVNLWAIGKPNAILSLYGHSSGIDSVTFDA----------SEVLVAAGAA 79
Query: 186 VGQDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPS 245
G + LW+LE +IV L G + + G+L
Sbjct: 80 SG---TIKLWDLEEAKIVRTLT----------GHRSNCISVDFHPFGEFFASGSLDTNLK 126
Query: 246 MRDVPKISPLVAHRVHTEPLSSLIFTQES--VLTACREGHIKIW 287
+ D+ K + ++ HT ++ L FT + V++ + +K+W
Sbjct: 127 IWDIRKKGCIHTYKGHTRGVNVLRFTPDGRWVVSGGEDNIVKVW 170
Score = 35.0 bits (79), Expect = 0.055
Identities = 21/73 (28%), Positives = 37/73 (49%), Gaps = 1/73 (1%)
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILT 130
+G I S+ F LAT D ++ +D L+ G G+ C +++ DGK +L
Sbjct: 185 EGQIQSLDFHPHEFLLATGSADRTVKFWDLETFELIGSGGPETAGVRCLSFNPDGKTVLC 244
Query: 131 GGEDDLVQVWSME 143
G ++ L +++S E
Sbjct: 245 GLQESL-KIFSWE 256
Score = 32.0 bits (71), Expect = 0.47
Identities = 24/112 (21%), Positives = 43/112 (37%), Gaps = 13/112 (11%)
Query: 86 LATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDR 145
L T G D + ++ K + + + G+ + + G +++W +E+
Sbjct: 32 LVTGGEDHKVNLWAIGKPNAILSLYGHSSGIDSVTFDASEVLVAAGAASGTIKLWDLEEA 91
Query: 146 KVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWEL 197
K+V GH S V F + GE F S DT L +W++
Sbjct: 92 KIVRTLTGHRSNCISVDFHPF---------GEF----FASGSLDTNLKIWDI 130
>At5g08390 katanin p80 subunit - like protein
Length = 823
Score = 49.7 bits (117), Expect = 2e-06
Identities = 32/143 (22%), Positives = 66/143 (45%), Gaps = 19/143 (13%)
Query: 76 SISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDD 135
S++F G + A+ D L+++D K+ + K + G+ ++ DG++I++GGED+
Sbjct: 199 SVNFHPFGEFFASGSLDTNLKIWDIRKKGCIHTYKGHTRGVNVLRFTPDGRWIVSGGEDN 258
Query: 136 LVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLW 195
+V+VW + K++ + H + + F + + + D + W
Sbjct: 259 VVKVWDLTAGKLLHEFKSHEGKIQSLDFHPH-------------EFLLATGSADKTVKFW 305
Query: 196 ELEMDEIVVPLRRGPPGGSPTFG 218
+LE E++ GG+ T G
Sbjct: 306 DLETFELI------GSGGTETTG 322
Score = 40.4 bits (93), Expect = 0.001
Identities = 22/91 (24%), Positives = 44/91 (48%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
+N + F+ DG ++ + G D ++V+D T L+ KS+ G + + + TG
Sbjct: 239 VNVLRFTPDGRWIVSGGEDNVVKVWDLTAGKLLHEFKSHEGKIQSLDFHPHEFLLATGSA 298
Query: 134 DDLVQVWSMEDRKVVAWGEGHNSWVSGVAFD 164
D V+ W +E +++ G + V + F+
Sbjct: 299 DKTVKFWDLETFELIGSGGTETTGVRCLTFN 329
Score = 34.7 bits (78), Expect = 0.072
Identities = 33/164 (20%), Positives = 66/164 (40%), Gaps = 25/164 (15%)
Query: 126 KYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGS 185
+ ++TGGED V +W++ + GH+S + V FD+ E + +
Sbjct: 123 RVLVTGGEDHKVNLWAIGKPNAILSLYGHSSGIDSVTFDA----------SEGLVAAGAA 172
Query: 186 VGQDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPS 245
G + LW+LE ++V L G + + G+L
Sbjct: 173 SG---TIKLWDLEEAKVVRTLT----------GHRSNCVSVNFHPFGEFFASGSLDTNLK 219
Query: 246 MRDVPKISPLVAHRVHTEPLSSLIFTQES--VLTACREGHIKIW 287
+ D+ K + ++ HT ++ L FT + +++ + +K+W
Sbjct: 220 IWDIRKKGCIHTYKGHTRGVNVLRFTPDGRWIVSGGEDNVVKVW 263
Score = 34.7 bits (78), Expect = 0.072
Identities = 26/103 (25%), Positives = 42/103 (40%), Gaps = 2/103 (1%)
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILT 130
+G I S+ F LAT D ++ +D L+ G + G+ C ++ DGK +L
Sbjct: 278 EGKIQSLDFHPHEFLLATGSADKTVKFWDLETFELIGSGGTETTGVRCLTFNPDGKSVLC 337
Query: 131 GGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSN 173
G ++ L + M H SG D + NS+
Sbjct: 338 GLQESLKRTEPMSGG--ATQSNSHPEKTSGSGRDQAGLNDNSS 378
Score = 32.3 bits (72), Expect = 0.36
Identities = 25/112 (22%), Positives = 43/112 (38%), Gaps = 13/112 (11%)
Query: 86 LATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDR 145
L T G D + ++ K + + + G+ + + G +++W +E+
Sbjct: 125 LVTGGEDHKVNLWAIGKPNAILSLYGHSSGIDSVTFDASEGLVAAGAASGTIKLWDLEEA 184
Query: 146 KVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWEL 197
KVV GH S V F + GE F S DT L +W++
Sbjct: 185 KVVRTLTGHRSNCVSVNFHPF---------GEF----FASGSLDTNLKIWDI 223
>At1g11160 hypothetical protein
Length = 974
Score = 48.5 bits (114), Expect = 5e-06
Identities = 29/133 (21%), Positives = 63/133 (46%), Gaps = 13/133 (9%)
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILT 130
+ + +++ F G +LA+ D LRV+D K+ + K + G+ +S DG+++++
Sbjct: 49 RSNCSAVEFHPFGEFLASGSSDTNLRVWDTRKKGCIQTYKGHTRGISTIEFSPDGRWVVS 108
Query: 131 GGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDT 190
GG D++V+VW + K++ + H + + F + + + D
Sbjct: 109 GGLDNVVKVWDLTAGKLLHEFKCHEGPIRSLDF-------------HPLEFLLATGSADR 155
Query: 191 QLLLWELEMDEIV 203
+ W+LE E++
Sbjct: 156 TVKFWDLETFELI 168
Score = 38.1 bits (87), Expect = 0.007
Identities = 23/90 (25%), Positives = 43/90 (47%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
I++I FS DG ++ + G D ++V+D T L+ K + G + + + TG
Sbjct: 94 ISTIEFSPDGRWVVSGGLDNVVKVWDLTAGKLLHEFKCHEGPIRSLDFHPLEFLLATGSA 153
Query: 134 DDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
D V+ W +E +++ + V +AF
Sbjct: 154 DRTVKFWDLETFELIGTTRPEATGVRAIAF 183
Score = 33.9 bits (76), Expect = 0.12
Identities = 39/170 (22%), Positives = 62/170 (35%), Gaps = 25/170 (14%)
Query: 120 AWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETI 179
A++ + +L G ++++W +E+ K+V GH S S V F + GE +
Sbjct: 14 AFNSEEVLVLAGASSGVIKLWDLEESKMVRAFTGHRSNCSAVEFHPF---------GEFL 64
Query: 180 TYRFGSVGQDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGT 239
S DT L +W+ + + G S SP W V G
Sbjct: 65 ----ASGSSDTNLRVWDTRKKGCIQTYKGHTRGISTIEFSP-------DGRW---VVSGG 110
Query: 240 LQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQESVL--TACREGHIKIW 287
L + D+ L + H P+ SL F L T + +K W
Sbjct: 111 LDNVVKVWDLTAGKLLHEFKCHEGPIRSLDFHPLEFLLATGSADRTVKFW 160
>At1g71840 unknown protein (At1g71840)
Length = 407
Score = 48.1 bits (113), Expect = 6e-06
Identities = 25/83 (30%), Positives = 41/83 (49%)
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
S++ ++FS DG LA+ G DG +++FD + L C G+ W G +L G
Sbjct: 115 SVSCLAFSYDGQLLASGGLDGVVQIFDASSGTLKCVLDGPGAGIEWVRWHPRGHIVLAGS 174
Query: 133 EDDLVQVWSMEDRKVVAWGEGHN 155
ED + +W+ + + GHN
Sbjct: 175 EDCSLWMWNADKEAYLNMFSGHN 197
Score = 35.8 bits (81), Expect = 0.032
Identities = 33/123 (26%), Positives = 55/123 (43%), Gaps = 13/123 (10%)
Query: 79 FSADGAYLATVGRDGYLRVFD---YTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDD 135
F+ DG + T D L V++ H+V G + GL C + + ++G +D
Sbjct: 205 FTPDGKLICTGSDDASLIVWNPKTCESIHIVKGHPYHTEGLTCLDINSNSSLAISGSKDG 264
Query: 136 LVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLW 195
V + ++ KVV+ H V V F SP+S TI + G D +L++W
Sbjct: 265 SVHIVNIVTGKVVSSLNSHTDSVECVKF-----SPSS----ATIPLA-ATGGMDKKLIIW 314
Query: 196 ELE 198
+L+
Sbjct: 315 DLQ 317
Score = 30.4 bits (67), Expect = 1.4
Identities = 23/95 (24%), Positives = 38/95 (39%), Gaps = 8/95 (8%)
Query: 73 SINSISFSADGAYL---ATVGRDGYLRVFD--YTKEHLVCGGKSYYGGLLCCAWSMDGKY 127
S+ + FS A + AT G D L ++D ++ +C + G+ W KY
Sbjct: 286 SVECVKFSPSSATIPLAATGGMDKKLIIWDLQHSTPRFIC---EHEEGVTSLTWIGTSKY 342
Query: 128 ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVA 162
+ TG + V +W V GH V ++
Sbjct: 343 LATGCANGTVSIWDSLLGNCVHTYHGHQDAVQAIS 377
Score = 28.1 bits (61), Expect = 6.7
Identities = 19/60 (31%), Positives = 24/60 (39%), Gaps = 2/60 (3%)
Query: 104 HLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
H G K L C D + TGG DD +W + + A GH VS +AF
Sbjct: 64 HTFTGHKGELYALACSP--TDATLVATGGGDDKAFLWKIGNGDWAAELPGHKDSVSCLAF 121
>At3g18130 protein kinase C-receptor/G-protein, putative
Length = 326
Score = 47.8 bits (112), Expect = 8e-06
Identities = 32/127 (25%), Positives = 57/127 (44%), Gaps = 13/127 (10%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
+ + S+DG + + DG LR++D + +L A+S D + I++
Sbjct: 66 VEDVVLSSDGQFALSGSWDGELRLWDLATGETTRRFVGHTKDVLSVAFSTDNRQIVSASR 125
Query: 134 DDLVQVWSM--EDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
D +++W+ E + ++ G+GH WVS V F SPN T+ S D
Sbjct: 126 DRTIKLWNTLGECKYTISEGDGHKEWVSCVRF-----SPN------TLVPTIVSASWDKT 174
Query: 192 LLLWELE 198
+ +W L+
Sbjct: 175 VKVWNLQ 181
Score = 38.1 bits (87), Expect = 0.007
Identities = 20/81 (24%), Positives = 41/81 (49%), Gaps = 10/81 (12%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYY----GGLLCCAWSMDGKY 127
G +N+++ S DG+ A+ G+DG + ++D + GK Y G ++ +Y
Sbjct: 194 GYLNTVAVSPDGSLCASGGKDGVILLWDLAE------GKKLYSLEAGSIIHSLCFSPNRY 247
Query: 128 ILTGGEDDLVQVWSMEDRKVV 148
L ++ +++W +E + VV
Sbjct: 248 WLCAATENSIRIWDLESKSVV 268
Score = 35.0 bits (79), Expect = 0.055
Identities = 31/138 (22%), Positives = 65/138 (46%), Gaps = 17/138 (12%)
Query: 76 SISFSADGAYLATVGRDGYLRVFDYTKE--HLVCGGKSYYGGLLCCAWSMDGKY--ILTG 131
S++FS D + + RD +++++ E + + G + + C +S + I++
Sbjct: 110 SVAFSTDNRQIVSASRDRTIKLWNTLGECKYTISEGDGHKEWVSCVRFSPNTLVPTIVSA 169
Query: 132 GEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQ 191
D V+VW++++ K+ GH+ +++ VA SP+ + S G+D
Sbjct: 170 SWDKTVKVWNLQNCKLRNSLVGHSGYLNTVAV-----SPDGS--------LCASGGKDGV 216
Query: 192 LLLWELEMDEIVVPLRRG 209
+LLW+L + + L G
Sbjct: 217 ILLWDLAEGKKLYSLEAG 234
Score = 30.4 bits (67), Expect = 1.4
Identities = 30/124 (24%), Positives = 46/124 (36%), Gaps = 18/124 (14%)
Query: 122 SMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITY 181
S DG++ L+G D +++W + + GH V VAF S DN + +
Sbjct: 72 SSDGQFALSGSWDGELRLWDLATGETTRRFVGHTKDVLSVAF--------STDNRQIV-- 121
Query: 182 RFGSVGQDTQLLLWELEMDEIVVPLRRGPPG----GSPTFGSPTFSAGSQSSHWDNAVPL 237
S +D + LW + E + G F T S+ WD V +
Sbjct: 122 ---SASRDRTIKLWN-TLGECKYTISEGDGHKEWVSCVRFSPNTLVPTIVSASWDKTVKV 177
Query: 238 GTLQ 241
LQ
Sbjct: 178 WNLQ 181
>At1g61210
Length = 282
Score = 47.0 bits (110), Expect = 1e-05
Identities = 26/133 (19%), Positives = 63/133 (46%), Gaps = 13/133 (9%)
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILT 130
+ + +++ F G +LA+ D L+++D K+ + K + G+ ++ DG+++++
Sbjct: 130 RSNCSAVEFHPFGEFLASGSSDANLKIWDIRKKGCIQTYKGHSRGISTIRFTPDGRWVVS 189
Query: 131 GGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDT 190
GG D++V+VW + K++ + H + + F + + + D
Sbjct: 190 GGLDNVVKVWDLTAGKLLHEFKFHEGPIRSLDF-------------HPLEFLLATGSADR 236
Query: 191 QLLLWELEMDEIV 203
+ W+LE E++
Sbjct: 237 TVKFWDLETFELI 249
Score = 31.2 bits (69), Expect = 0.80
Identities = 36/162 (22%), Positives = 58/162 (35%), Gaps = 25/162 (15%)
Query: 128 ILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVG 187
+L G ++++W +E+ K+V GH S S V F + GE + S
Sbjct: 103 VLAGASSGVIKLWDVEEAKMVRAFTGHRSNCSAVEFHPF---------GEFL----ASGS 149
Query: 188 QDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMR 247
D L +W++ + + G S +P W V G L +
Sbjct: 150 SDANLKIWDIRKKGCIQTYKGHSRGISTIRFTP-------DGRW---VVSGGLDNVVKVW 199
Query: 248 DVPKISPLVAHRVHTEPLSSLIFTQESVL--TACREGHIKIW 287
D+ L + H P+ SL F L T + +K W
Sbjct: 200 DLTAGKLLHEFKFHEGPIRSLDFHPLEFLLATGSADRTVKFW 241
Score = 28.5 bits (62), Expect = 5.2
Identities = 35/174 (20%), Positives = 68/174 (38%), Gaps = 35/174 (20%)
Query: 126 KYILTGGEDDLVQVWSM--------EDRKVVAWGE--GHNSWVSGVAFDSYWSSPNSNDN 175
+ +TGG+D V +W++ D W GH S V VAFDS
Sbjct: 49 RLFITGGDDYKVNLWAIGKPTSLMKNDAIAFYWQSLCGHTSAVDSVAFDS---------- 98
Query: 176 GETITYRFGSVGQDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAV 235
E + S G + LW++E ++V G + + + + +
Sbjct: 99 AEVLVLAGASSG---VIKLWDVEEAKMVRAFT----------GHRSNCSAVEFHPFGEFL 145
Query: 236 PLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQES--VLTACREGHIKIW 287
G+ + D+ K + ++ H+ +S++ FT + V++ + +K+W
Sbjct: 146 ASGSSDANLKIWDIRKKGCIQTYKGHSRGISTIRFTPDGRWVVSGGLDNVVKVW 199
>At2g41500 putative U4/U6 small nuclear ribonucleoprotein
Length = 554
Score = 45.4 bits (106), Expect = 4e-05
Identities = 25/92 (27%), Positives = 46/92 (49%)
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
S+ I+F DGA A+ G D RV+D + + + + +S +G ++ +GG
Sbjct: 383 SVYGIAFQQDGALAASCGLDSLARVWDLRTGRSILVFQGHIKPVFSVNFSPNGYHLASGG 442
Query: 133 EDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFD 164
ED+ ++W + RK + H + VS V ++
Sbjct: 443 EDNQCRIWDLRMRKSLYIIPAHANLVSQVKYE 474
Score = 35.4 bits (80), Expect = 0.042
Identities = 39/174 (22%), Positives = 77/174 (43%), Gaps = 8/174 (4%)
Query: 29 GNLYNKDGAGESSFPILKDQTLFSVAHARYSKSNPIARWHICQGSINSISFSADGAYLAT 88
G + +DGA +S + +L V R +S + + HI + S++FS +G +LA+
Sbjct: 386 GIAFQQDGALAASCGL---DSLARVWDLRTGRSILVFQGHI--KPVFSVNFSPNGYHLAS 440
Query: 89 VGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWS-MDGKYILTGGEDDLVQVWSMEDRKV 147
G D R++D + ++ + + +G ++ T D V +WS D +
Sbjct: 441 GGEDNQCRIWDLRMRKSLYIIPAHANLVSQVKYEPQEGYFLATASYDMKVNIWSGRDFSL 500
Query: 148 VAWGEGHNSWVSG--VAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWELEM 199
V GH S V+ + DS + S+D + G+ +D + ++++
Sbjct: 501 VKSLAGHESKVASLDITADSSCIATVSHDRTIKLWTSSGNDDEDEEKETMDIDL 554
Score = 35.0 bits (79), Expect = 0.055
Identities = 57/262 (21%), Positives = 93/262 (34%), Gaps = 50/262 (19%)
Query: 77 ISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDL 136
+ FS LAT D +++ T L+ + + L A+ GKY+ T D
Sbjct: 304 VVFSPVDDCLATASADRTAKLWK-TDGTLLQTFEGHLDRLARVAFHPSGKYLGTTSYDKT 362
Query: 137 VQVWSMEDRKVVAWGEGHNSWVSGVAFDS---------------YWS------------- 168
++W + + EGH+ V G+AF W
Sbjct: 363 WRLWDINTGAELLLQEGHSRSVYGIAFQQDGALAASCGLDSLARVWDLRTGRSILVFQGH 422
Query: 169 -----SPNSNDNGETITYRFGSVGQDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFS 223
S N + NG Y S G+D Q +W+L M + + + S P
Sbjct: 423 IKPVFSVNFSPNG----YHLASGGEDNQCRIWDLRMRKSLYIIPAHANLVSQVKYEPQEG 478
Query: 224 AGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQES--VLTACRE 281
++ +D V + S RD + L H ++SL T +S + T +
Sbjct: 479 YFLATASYDMKVNIW------SGRDFSLVKSLAGHE---SKVASLDITADSSCIATVSHD 529
Query: 282 GHIKIWTRPGVAESQPSNSETL 303
IK+WT G + + ET+
Sbjct: 530 RTIKLWTSSG-NDDEDEEKETM 550
>At5g50230 putative protein
Length = 515
Score = 45.1 bits (105), Expect = 5e-05
Identities = 27/82 (32%), Positives = 38/82 (45%), Gaps = 6/82 (7%)
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSM-----DGKY 127
++ S+S S +G + T GRD VFD T+ +CG G L WS D Y
Sbjct: 401 AVTSVSLSRNGNRILTSGRDNVHNVFD-TRTLEICGTLRASGNRLASNWSRSCISPDDDY 459
Query: 128 ILTGGEDDLVQVWSMEDRKVVA 149
+ G D V VWS+ +V+
Sbjct: 460 VAAGSADGSVHVWSLSKGNIVS 481
Score = 38.1 bits (87), Expect = 0.007
Identities = 19/62 (30%), Positives = 32/62 (50%)
Query: 80 SADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQV 139
S D Y+A DG + V+ +K ++V K +LCC+WS GK + + ++ V
Sbjct: 454 SPDDDYVAAGSADGSVHVWSLSKGNIVSILKEQTSPILCCSWSGIGKPLASADKNGYVCT 513
Query: 140 WS 141
W+
Sbjct: 514 WT 515
Score = 35.0 bits (79), Expect = 0.055
Identities = 29/106 (27%), Positives = 45/106 (42%), Gaps = 6/106 (5%)
Query: 66 RWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGL---LCCAWS 122
R H +G SI F + L T G+D ++++D L+ KS YG L L A +
Sbjct: 226 RIHAHEGGCGSIVFEYNSGTLFTGGQDRAVKMWDTNSGTLI---KSLYGSLGNILDMAVT 282
Query: 123 MDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWS 168
D K ++ + + VW + +V GH V V + S
Sbjct: 283 HDNKSVIAATSSNNLFVWDVSSGRVRHTLTGHTDKVCAVDVSKFSS 328
Score = 32.0 bits (71), Expect = 0.47
Identities = 30/133 (22%), Positives = 53/133 (39%), Gaps = 9/133 (6%)
Query: 75 NSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGED 134
N+I S DG + + DG LR++D L+ + + + S +G ILT G D
Sbjct: 361 NAICLSIDGLTVFSGHMDGNLRLWDIQTGKLLSEVAGHSSAVTSVSLSRNGNRILTSGRD 420
Query: 135 DLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLL 194
++ V+ ++ SG S WS + + + + + D + +
Sbjct: 421 NVHNVFDTRTLEICG-----TLRASGNRLASNWSRSCISPDDDYV----AAGSADGSVHV 471
Query: 195 WELEMDEIVVPLR 207
W L IV L+
Sbjct: 472 WSLSKGNIVSILK 484
>At5g14050 unknown protein
Length = 546
Score = 45.1 bits (105), Expect = 5e-05
Identities = 30/106 (28%), Positives = 48/106 (44%), Gaps = 3/106 (2%)
Query: 51 FSVAHARYSKSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGK 110
F + A++ K P+ + S+ S D +A VG +GY+ + TK + G
Sbjct: 315 FDLEKAKFDKIGPLVGRE--EKSLEYFEVSQDSNTIAFVGNEGYILLVS-TKTKELIGTL 371
Query: 111 SYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNS 156
G + A+S DGK++L+ G D V VW + K + G S
Sbjct: 372 KMNGSVRSLAFSEDGKHLLSSGGDGQVYVWDLRTMKCLYKGVDEGS 417
>At2g20330 putative WD-40 repeat protein
Length = 648
Score = 44.7 bits (104), Expect = 7e-05
Identities = 50/211 (23%), Positives = 78/211 (36%), Gaps = 27/211 (12%)
Query: 12 ISWVP-GGDGAFVVAHADGNLYNKDGAGESSFPILKDQTLFSVAHARYSKSNPIARWHIC 70
+SW P G V A ++++DG F + Y + + HIC
Sbjct: 230 VSWSPTSGQFLCVTGSAQAKIFDRDGLTLGEF----------MKGDMYIRDLKNTKGHIC 279
Query: 71 QGSINSISFSADGAYLATVGRDGYLRVFD---YTKEHLVCGGKSYYGG---LLCCAWSMD 124
+ L T DG LR++D + + V K G + CAW D
Sbjct: 280 GLTCGEWHPRTKETVL-TSSEDGSLRIWDVNNFLSQTQVIKPKLARPGRVPVTTCAWDRD 338
Query: 125 GKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFG 184
GK I G D +Q+WS++ WG + +V D S S+D ++ F
Sbjct: 339 GKRIAGGVGDGSIQIWSLKP----GWGSRPDIYVGKAHTDDITSVKFSSDGRILLSRSF- 393
Query: 185 SVGQDTQLLLWELEMDEIVVPLRRGPPGGSP 215
D L +W+L + + + G P P
Sbjct: 394 ----DGSLKVWDLRQMKEALKVFEGLPNYYP 420
>At5g25150 transcription initiation factor IID-associated
factor-like protein
Length = 700
Score = 44.3 bits (103), Expect = 9e-05
Identities = 27/108 (25%), Positives = 50/108 (46%)
Query: 56 ARYSKSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGG 115
AR + I I G ++ + + + Y+AT D +R++D V +
Sbjct: 484 ARIWSMDRIQPLRIMAGHLSDVDWHPNCNYIATGSSDKTVRLWDVQTGECVRIFIGHRSM 543
Query: 116 LLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
+L A S DG+Y+ +G ED + +W + + + GHNS V +++
Sbjct: 544 VLSLAMSPDGRYMASGDEDGTIMMWDLSTARCITPLMGHNSCVWSLSY 591
Score = 38.5 bits (88), Expect = 0.005
Identities = 31/128 (24%), Positives = 51/128 (39%), Gaps = 16/128 (12%)
Query: 79 FSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQ 138
FS G Y A+ D R++ + + + G L W + YI TG D V+
Sbjct: 468 FSPFGHYFASCSHDRTARIWSMDRIQPL---RIMAGHLSDVDWHPNCNYIATGSSDKTVR 524
Query: 139 VWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWELE 198
+W ++ + V GH S V +A SP+ S +D +++W+L
Sbjct: 525 LWDVQTGECVRIFIGHRSMVLSLAM-----SPDGR--------YMASGDEDGTIMMWDLS 571
Query: 199 MDEIVVPL 206
+ PL
Sbjct: 572 TARCITPL 579
Score = 28.5 bits (62), Expect = 5.2
Identities = 31/158 (19%), Positives = 57/158 (35%), Gaps = 40/158 (25%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTK-----------------EHLVCGGKSYY--- 113
+N S S DG+ +A D ++V+D K + + G+ Y
Sbjct: 355 LNCSSISHDGSLVAGGFSDSSIKVWDMAKIGQAGSGALQAENDSSDQSIGPNGRRSYTLL 414
Query: 114 ----GGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSS 169
G + +S G ++L+ D +++WS + + +GHN V F +
Sbjct: 415 LGHSGPVYSATFSPPGDFVLSSSADTTIRLWSTKLNANLVCYKGHNYPVWDAQFSPF--- 471
Query: 170 PNSNDNGETITYRFGSVGQDTQLLLWELEMDEIVVPLR 207
+ F S D +W ++ + PLR
Sbjct: 472 ----------GHYFASCSHDRTARIWSMDR---IQPLR 496
>At4g05410 U3 snoRNP-associated -like protein
Length = 504
Score = 44.3 bits (103), Expect = 9e-05
Identities = 24/87 (27%), Positives = 45/87 (51%), Gaps = 2/87 (2%)
Query: 76 SISFSADGAYLATVGRDGYLRVFDY-TKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGED 134
+++ S+DG YLAT G D ++ ++D T+EH V + + C + + +G D
Sbjct: 227 ALAVSSDGRYLATGGVDRHVHIWDVRTREH-VQAFPGHRNTVSCLCFRYGTSELYSGSFD 285
Query: 135 DLVQVWSMEDRKVVAWGEGHNSWVSGV 161
V+VW++ED+ + GH + +
Sbjct: 286 RTVKVWNVEDKAFITENHGHQGEILAI 312
Score = 38.1 bits (87), Expect = 0.007
Identities = 19/47 (40%), Positives = 26/47 (54%)
Query: 117 LCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAF 163
L A S DG+Y+ TGG D V +W + R+ V GH + VS + F
Sbjct: 226 LALAVSSDGRYLATGGVDRHVHIWDVRTREHVQAFPGHRNTVSCLCF 272
>At5g67320 unknown protein
Length = 613
Score = 43.5 bits (101), Expect = 2e-04
Identities = 50/245 (20%), Positives = 88/245 (35%), Gaps = 32/245 (13%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
+ ++ ++ +G LAT DG R++ E L+ + G + W+ G Y+LTG
Sbjct: 327 VTTLDWNGEGTLLATGSCDGQARIWTLNGE-LISTLSKHKGPIFSLKWNKKGDYLLTGSV 385
Query: 134 DDLVQVWSMEDRKVVAWGEGHN------SWVSGVAFDS-------YWSSPNSNDNGETIT 180
D VW ++ + E H+ W + V+F + Y +T T
Sbjct: 386 DRTAVVWDVKAEEWKQQFEFHSGPTLDVDWRNNVSFATSSTDSMIYLCKIGETRPAKTFT 445
Query: 181 YRFGSV---------------GQDTQLLLWELEMDEIVVPLRRGPPGGSPTFGSPTFSAG 225
G V D+ +W ++ V LR SPT G
Sbjct: 446 GHQGEVNCVKWDPTGSLLASCSDDSTAKIWNIKQSTFVHDLREHTKEIYTIRWSPT-GPG 504
Query: 226 SQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHRVHTEPLSSLIFTQ--ESVLTACREGH 283
+ + + + + + D L + H EP+ SL F+ E + + +
Sbjct: 505 TNNPNKQLTLASASFDSTVKLWDAELGKMLCSFNGHREPVYSLAFSPNGEYIASGSLDKS 564
Query: 284 IKIWT 288
I IW+
Sbjct: 565 IHIWS 569
Score = 37.4 bits (85), Expect = 0.011
Identities = 20/83 (24%), Positives = 42/83 (50%), Gaps = 5/83 (6%)
Query: 86 LATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDR 145
LA+ D ++++D ++C + + A+S +G+YI +G D + +WS+++
Sbjct: 514 LASASFDSTVKLWDAELGKMLCSFNGHREPVYSLAFSPNGEYIASGSLDKSIHIWSIKEG 573
Query: 146 KVVAWGEGHNSWVSGVAFDSYWS 168
K+V G +G F+ W+
Sbjct: 574 KIVKTYTG-----NGGIFEVCWN 591
Score = 29.3 bits (64), Expect = 3.0
Identities = 26/99 (26%), Positives = 38/99 (38%), Gaps = 9/99 (9%)
Query: 119 CAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSND---- 174
CAWS + +G D ++WS+ + A G N + S+ S D
Sbjct: 271 CAWSPSASLLASGSGDATARIWSIPEGSFKAVHTGRNINALILKHAKGKSNEKSKDVTTL 330
Query: 175 --NGETITYRFGSVGQDTQLLLWELEMDEI-VVPLRRGP 210
NGE GS D Q +W L + I + +GP
Sbjct: 331 DWNGEGTLLATGSC--DGQARIWTLNGELISTLSKHKGP 367
>At1g18080 putative guanine nucleotide-binding protein beta
subunit
Length = 327
Score = 43.5 bits (101), Expect = 2e-04
Identities = 37/157 (23%), Positives = 63/157 (39%), Gaps = 23/157 (14%)
Query: 44 ILKDQTLFSVAHARYSKSNPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDYTKE 103
+ KD + VA R + + + + S+DG + + DG LR++D
Sbjct: 45 LTKDDKAYGVAQRRLTGHSHF---------VEDVVLSSDGQFALSGSWDGELRLWDLAAG 95
Query: 104 HLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDR---KVVAWGEGHNSWVSG 160
+ +L A+S+D + I++ D +++W+ + GEGH WVS
Sbjct: 96 VSTRRFVGHTKDVLSVAFSLDNRQIVSASRDRTIKLWNTLGECKYTISEGGEGHRDWVSC 155
Query: 161 VAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWEL 197
V F SPN T+ S D + +W L
Sbjct: 156 VRF-----SPN------TLQPTIVSASWDKTVKVWNL 181
Score = 37.0 bits (84), Expect = 0.015
Identities = 32/127 (25%), Positives = 61/127 (47%), Gaps = 18/127 (14%)
Query: 76 SISFSADGAYLATVGRDGYLRVFDYTKE---HLVCGGKSYYGGLLCCAWSMDGKY--ILT 130
S++FS D + + RD +++++ E + GG+ + + C +S + I++
Sbjct: 110 SVAFSLDNRQIVSASRDRTIKLWNTLGECKYTISEGGEGHRDWVSCVRFSPNTLQPTIVS 169
Query: 131 GGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDT 190
D V+VW++ + K+ + GH +VS VA SP+ + S G+D
Sbjct: 170 ASWDKTVKVWNLSNCKLRSTLAGHTGYVSTVAV-----SPDGS--------LCASGGKDG 216
Query: 191 QLLLWEL 197
+LLW+L
Sbjct: 217 VVLLWDL 223
Score = 32.0 bits (71), Expect = 0.47
Identities = 17/81 (20%), Positives = 39/81 (47%), Gaps = 10/81 (12%)
Query: 72 GSINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYY----GGLLCCAWSMDGKY 127
G +++++ S DG+ A+ G+DG + ++D + GK Y ++ +Y
Sbjct: 195 GYVSTVAVSPDGSLCASGGKDGVVLLWDLAE------GKKLYSLEANSVIHALCFSPNRY 248
Query: 128 ILTGGEDDLVQVWSMEDRKVV 148
L + +++W +E + +V
Sbjct: 249 WLCAATEHGIKIWDLESKSIV 269
>At5g16750 WD40-repeat protein
Length = 876
Score = 43.1 bits (100), Expect = 2e-04
Identities = 43/164 (26%), Positives = 72/164 (43%), Gaps = 27/164 (16%)
Query: 47 DQTLFSVAHARYSKS------NPIARWHICQGSINSISFSADGAYLATVGRDGYLRVFDY 100
D+ LFS H+R + I W +G + ++ A G LAT G D + V+D
Sbjct: 72 DKLLFSAGHSRQIRVWDLETLKCIRSWKGHEGPVMGMACHASGGLLATAGADRKVLVWDV 131
Query: 101 TK---EHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSME----DRKVVAWGEG 153
H G K +L S + +++G +D V+VW + ++K +A E
Sbjct: 132 DGGFCTHYFRGHKGVVSSILFHPDS-NKNILISGSDDATVRVWDLNAKNTEKKCLAIMEK 190
Query: 154 HNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWEL 197
H S V+ +A +++G T+ S G+D + LW+L
Sbjct: 191 HFSAVTSIAL---------SEDGLTLF----SAGRDKVVNLWDL 221
Score = 40.8 bits (94), Expect = 0.001
Identities = 46/210 (21%), Positives = 84/210 (39%), Gaps = 21/210 (10%)
Query: 85 YLATVGRDGYLRVFDYTK---EHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQVWS 141
+LA +RV+D +++ G K L C S I+TG +D V++W+
Sbjct: 373 FLAVATNLEEVRVYDVATMSCSYVLAGHKEVVLSLDTCVSSSGNVLIVTGSKDKTVRLWN 432
Query: 142 MEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLLLWELE--M 199
+ + G GHN + VAF ++ ++ F S D L +W L+
Sbjct: 433 ATSKSCIGVGTGHNGDILAVAFAK-----------KSFSF-FVSGSGDRTLKVWSLDGIS 480
Query: 200 DEIVVPLRRGPPGGSPTFGSPTFSAGSQSSHWDNAVPLGTLQPAPSMRDVPKISPLVAHR 259
++ P+ S + D+ V G+ S+ +P + +V +
Sbjct: 481 EDSEEPINLKTRSVVAAHDKDINSVAVARN--DSLVCTGSEDRTASIWRLPDLVHVVTLK 538
Query: 260 VHTEPLSSLIFT--QESVLTACREGHIKIW 287
H + S+ F+ + V+TA + +KIW
Sbjct: 539 GHKRRIFSVEFSTVDQCVMTASGDKTVKIW 568
Score = 40.8 bits (94), Expect = 0.001
Identities = 26/103 (25%), Positives = 44/103 (42%), Gaps = 5/103 (4%)
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
++ +++ S D L + G +RV+D + K + G ++ A G + T G
Sbjct: 62 TLTALALSPDDKLLFSAGHSRQIRVWDLETLKCIRSWKGHEGPVMGMACHASGGLLATAG 121
Query: 133 EDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDN 175
D V VW ++ + GH VS + F P+SN N
Sbjct: 122 ADRKVLVWDVDGGFCTHYFRGHKGVVSSILF-----HPDSNKN 159
Score = 32.7 bits (73), Expect = 0.27
Identities = 28/130 (21%), Positives = 51/130 (38%), Gaps = 13/130 (10%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
INS++ + + + + T D ++ V K + + +S + ++T
Sbjct: 502 INSVAVARNDSLVCTGSEDRTASIWRLPDLVHVVTLKGHKRRIFSVEFSTVDQCVMTASG 561
Query: 134 DDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSSPNSNDNGETITYRFGSVGQDTQLL 193
D V++W++ D + EGH S V +F T +F S G D L
Sbjct: 562 DKTVKIWAISDGSCLKTFEGHTSSVLRASF-------------ITDGTQFVSCGADGLLK 608
Query: 194 LWELEMDEIV 203
LW + E +
Sbjct: 609 LWNVNTSECI 618
Score = 32.3 bits (72), Expect = 0.36
Identities = 34/145 (23%), Positives = 62/145 (42%), Gaps = 18/145 (12%)
Query: 71 QGSINSISFSADGA--YLATVGRDGYLRVFDY----TKEHLVCGGKSYYGGLLCCAWSMD 124
+G ++SI F D L + D +RV+D T++ + + ++ + A S D
Sbjct: 144 KGVVSSILFHPDSNKNILISGSDDATVRVWDLNAKNTEKKCLAIMEKHFSAVTSIALSED 203
Query: 125 GKYILTGGEDDLVQVWSMEDRKVVAWG------EGHNSWVSGVAFDSYWSS-----PNSN 173
G + + G D +V +W + D A E + SG F S+ +S
Sbjct: 204 GLTLFSAGRDKVVNLWDLHDYSCKATVATYEVLEAVTTVSSGTPFASFVASLDQKKSKKK 263
Query: 174 DNGETITYRFGSVGQDTQLLLWELE 198
++ TY F +VG+ + +W+ E
Sbjct: 264 ESDSQATY-FITVGERGVVRIWKSE 287
Score = 31.6 bits (70), Expect = 0.61
Identities = 16/68 (23%), Positives = 29/68 (42%)
Query: 73 SINSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGG 132
S+ SF DG + G DG L++++ + + + A + I TGG
Sbjct: 585 SVLRASFITDGTQFVSCGADGLLKLWNVNTSECIATYDQHEDKVWALAVGKKTEMIATGG 644
Query: 133 EDDLVQVW 140
D ++ +W
Sbjct: 645 GDAVINLW 652
Score = 28.1 bits (61), Expect = 6.7
Identities = 17/89 (19%), Positives = 41/89 (45%)
Query: 74 INSISFSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGE 133
I S+ FS + T D ++++ + + + + +L ++ DG ++ G
Sbjct: 544 IFSVEFSTVDQCVMTASGDKTVKIWAISDGSCLKTFEGHTSSVLRASFITDGTQFVSCGA 603
Query: 134 DDLVQVWSMEDRKVVAWGEGHNSWVSGVA 162
D L+++W++ + +A + H V +A
Sbjct: 604 DGLLKLWNVNTSECIATYDQHEDKVWALA 632
>At2g47410 putative WD-40 repeat protein
Length = 1389
Score = 42.7 bits (99), Expect = 3e-04
Identities = 17/56 (30%), Positives = 31/56 (55%)
Query: 110 KSYYGGLLCCAWSMDGKYILTGGEDDLVQVWSMEDRKVVAWGEGHNSWVSGVAFDS 165
+ + + C + G+Y++TG +D LV++WSME +A GH ++ +A S
Sbjct: 233 RGHRNAVYCAIFDRSGRYVITGSDDRLVKIWSMETALCLASCRGHEGDITDLAVSS 288
Score = 34.3 bits (77), Expect = 0.094
Identities = 17/85 (20%), Positives = 37/85 (43%)
Query: 79 FSADGAYLATVGRDGYLRVFDYTKEHLVCGGKSYYGGLLCCAWSMDGKYILTGGEDDLVQ 138
F G Y+ T D ++++ + + + G + A S + + + D +++
Sbjct: 244 FDRSGRYVITGSDDRLVKIWSMETALCLASCRGHEGDITDLAVSSNNALVASASNDFVIR 303
Query: 139 VWSMEDRKVVAWGEGHNSWVSGVAF 163
VW + D ++ GH V+ +AF
Sbjct: 304 VWRLPDGMPISVLRGHTGAVTAIAF 328
Score = 29.6 bits (65), Expect = 2.3
Identities = 21/88 (23%), Positives = 37/88 (41%), Gaps = 18/88 (20%)
Query: 116 LLCCAWSMDGKYILTGGEDD------LVQVWSMEDRKVVAWGEGHNSWVSGVAFDSYWSS 169
+LCCA++ +G +TG D + VW+ D +V GH+ S D + +
Sbjct: 394 ILCCAYNANGTIFVTGSSDSNARVYCRICVWNAADGSLVHCLTGHSE--SSYVLDVHPFN 451
Query: 170 PNSNDNGETITYRFGSVGQDTQLLLWEL 197
P S G D + ++W++
Sbjct: 452 PRI----------AMSAGYDGKTIIWDI 469
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.133 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,660,610
Number of Sequences: 26719
Number of extensions: 416705
Number of successful extensions: 1568
Number of sequences better than 10.0: 170
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 62
Number of HSP's that attempted gapping in prelim test: 1068
Number of HSP's gapped (non-prelim): 476
length of query: 325
length of database: 11,318,596
effective HSP length: 100
effective length of query: 225
effective length of database: 8,646,696
effective search space: 1945506600
effective search space used: 1945506600
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC148995.8


