

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
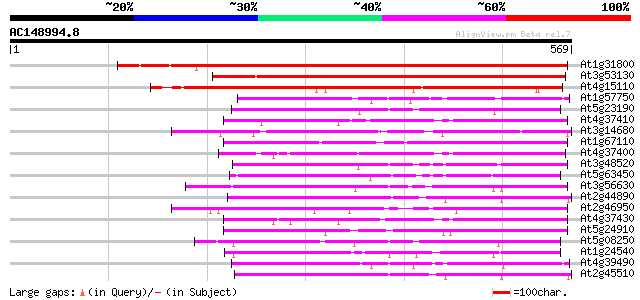
Query= AC148994.8 - phase: 0 /pseudo
(569 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g31800 unknown protein 736 0.0
At3g53130 Cytochrom P450 -like protein 370 e-103
At4g15110 cytochrome P450 (Z97337.39) 352 4e-97
At1g57750 unknown protein 121 9e-28
At5g23190 cytochrome P450-like protein 121 1e-27
At4g37410 cytochrome P450 monooxygenase like protein 118 8e-27
At3g14680 putative cytochrome P450 117 1e-26
At1g67110 unknown protein 117 2e-26
At4g37400 cytochrome P450 monooxygenase -like protein 115 7e-26
At3g48520 cytochrome P450-like protein 115 9e-26
At5g63450 cytochrome P450-like protein 114 2e-25
At3g56630 cytochrome P450-like protein 113 2e-25
At2g44890 putative cytochrome P450 113 3e-25
At2g46950 putative cytochrome P450 112 4e-25
At4g37430 cytochrome P450 monooxygenase (CYP91A2) 112 7e-25
At5g24910 cytochrome P450-like protein 110 2e-24
At5g08250 cytochrome P450-like protein 110 3e-24
At1g24540 putative cytochrome P450 110 3e-24
At4g39490 cytochrome P450 - like protein 109 4e-24
At2g45510 putative cytochrome P450 109 4e-24
>At1g31800 unknown protein
Length = 595
Score = 736 bits (1901), Expect = 0.0
Identities = 370/465 (79%), Positives = 420/465 (89%), Gaps = 11/465 (2%)
Query: 110 SSFIA-CSSSNGRSP--NDSVDDGVVKSADQLLEEKRRAELSAKIASGEFTVKQESGLPS 166
S+F+ SSSNGR P +SV +GV KS ++L EEKRRAELSA+IASG FTV++ S PS
Sbjct: 24 SNFVVFSSSSNGRDPLEENSVPNGV-KSLEKLQEEKRRAELSARIASGAFTVRKSS-FPS 81
Query: 167 ILKKSLSNLGVSNEILEFLFGL------YPKIPEAKGSISAIRSEAFFIPLYELYITYGG 220
+K LS +G+ + +L+F+F YPK+PEAKGSI A+R+EAFFIPLYEL++TYGG
Sbjct: 82 TVKNGLSKIGIPSNVLDFMFDWTGSDQDYPKVPEAKGSIQAVRNEAFFIPLYELFLTYGG 141
Query: 221 IFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIPADGEIWRVRR 280
IFRL FGPKSFLIVSDP+IAKHILKDN+KAYSKGILAEILDFVMGKGLIPADGEIWR RR
Sbjct: 142 IFRLTFGPKSFLIVSDPSIAKHILKDNAKAYSKGILAEILDFVMGKGLIPADGEIWRRRR 201
Query: 281 RTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFN 340
R IVPALH K+VAAMI LFG+A+DRLCQKLD AA GE+VEMESLFSRLTLD+IGKAVFN
Sbjct: 202 RAIVPALHQKYVAAMISLFGEASDRLCQKLDAAALKGEEVEMESLFSRLTLDIIGKAVFN 261
Query: 341 YDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTAALKLVNDTL 400
YDFDSL+NDTG+IEAVYTVLREAEDRS+SPIPVWD+PIWKDISPRQRKV +LKL+NDTL
Sbjct: 262 YDFDSLTNDTGVIEAVYTVLREAEDRSVSPIPVWDIPIWKDISPRQRKVATSLKLINDTL 321
Query: 401 NNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDDLMTMLIAGHET 460
++LIA CKRMV+EEELQFHEEYMNE+DPSILHFLLASGDDV+SKQLRDDLMTMLIAGHET
Sbjct: 322 DDLIATCKRMVEEEELQFHEEYMNERDPSILHFLLASGDDVSSKQLRDDLMTMLIAGHET 381
Query: 461 SAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVINESLRLYPQP 520
SAAVLTWTFYLL+ EPSV++KLQEEVDSV+GDRFPTI+DMKKLKYTTRV+NESLRLYPQP
Sbjct: 382 SAAVLTWTFYLLTTEPSVVAKLQEEVDSVIGDRFPTIQDMKKLKYTTRVMNESLRLYPQP 441
Query: 521 PVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLWNDADKLN 565
PVLIRRSI++D+LGEYPIKRGEDIFISVWNLHRSP W+DA+K N
Sbjct: 442 PVLIRRSIDNDILGEYPIKRGEDIFISVWNLHRSPLHWDDAEKFN 486
>At3g53130 Cytochrom P450 -like protein
Length = 539
Score = 370 bits (951), Expect = e-103
Identities = 188/361 (52%), Positives = 261/361 (72%), Gaps = 4/361 (1%)
Query: 206 AFFIPLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMG 265
A F+PLY+ YG I+RL GP++F+IVSDPAIAKH+L++ K Y+KG++AE+ +F+ G
Sbjct: 96 ALFLPLYKWMNEYGPIYRLAAGPRNFVIVSDPAIAKHVLRNYPK-YAKGLVAEVSEFLFG 154
Query: 266 KGLIPADGEIWRVRRRTIVPALHLKFVAAMIG-LFGQATDRLCQKLDTAASDGEDVEMES 324
G A+G +W RRR +VP+LH ++++ ++ +F + +RL +KL A DG V ME+
Sbjct: 155 SGFAIAEGPLWTARRRAVVPSLHRRYLSVIVERVFCKCAERLVEKLQPYAEDGSAVNMEA 214
Query: 325 LFSRLTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISP 384
FS++TLDVIG ++FNY+FDSL+ D+ +IEAVYT L+EAE RS +P W + I P
Sbjct: 215 KFSQMTLDVIGLSLFNYNFDSLTTDSPVIEAVYTALKEAELRSTDLLPYWKIDALCKIVP 274
Query: 385 RQRKVTAALKLVNDTLNNLIAICKRMVDEEELQFH-EEYMNEQDPSILHFLLASGDDVTS 443
RQ K A+ L+ +T+ +LIA CK +V+ E + + EEY+N+ DPSIL FLLAS ++V+S
Sbjct: 275 RQVKAEKAVTLIRETVEDLIAKCKEIVEREGERINDEEYVNDADPSILRFLLASREEVSS 334
Query: 444 KQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKL 503
QLRDDL++ML+AGHET+ +VLTWT YLLSK S + K QEEVD VL R P ED+K+L
Sbjct: 335 VQLRDDLLSMLVAGHETTGSVLTWTLYLLSKNSSALRKAQEEVDRVLEGRNPAFEDIKEL 394
Query: 504 KYTTRVINESLRLYPQPPVLIRRSIEDDVL-GEYPIKRGEDIFISVWNLHRSPTLWNDAD 562
KY TR INES+RLYP PPVLIRR+ D+L G Y + G+DI ISV+N+HRS +W A+
Sbjct: 395 KYITRCINESMRLYPHPPVLIRRAQVPDILPGNYKVNTGQDIMISVYNIHRSSEVWEKAE 454
Query: 563 K 563
+
Sbjct: 455 E 455
>At4g15110 cytochrome P450 (Z97337.39)
Length = 580
Score = 352 bits (902), Expect = 4e-97
Identities = 198/439 (45%), Positives = 284/439 (64%), Gaps = 33/439 (7%)
Query: 143 RRAELSAKIASGEFTVKQESGLPSILKKSLSNLGVSNEILEFLFG-LYPKIPEAKGSISA 201
RRA +S K S E P L N SN + FL G +P A+GS+S
Sbjct: 45 RRASVSIKCQSTE---------PKTNGNILDN--ASNLLTNFLSGGSLGSMPTAEGSVSD 93
Query: 202 IRSEAFFIPLYELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILD 261
+ + F+ LY+ ++ +GGI++L FGPK+F+++SDP IA+H+L++N+ +Y KG+LAEIL+
Sbjct: 94 LFGKPLFLSLYDWFLEHGGIYKLAFGPKAFVVISDPIIARHVLRENAFSYDKGVLAEILE 153
Query: 262 FVMGKGLIPADGEIWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQK-----LDTAASD 316
+MGKGLIPAD + W++RRR I PA H ++ AM+ +F ++++ K + S
Sbjct: 154 PIMGKGLIPADLDTWKLRRRAITPAFHKLYLEAMVKVFSDCSEKMILKSEKLIREKETSS 213
Query: 317 GED---VEMESLFSRLTLDVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPV 373
GED +++E+ FS L LD+IG +VFNYDF S++ ++ +I+AVY L EAE RS P
Sbjct: 214 GEDTIELDLEAEFSSLALDIIGLSVFNYDFGSVTKESPVIKAVYGTLFEAEHRSTFYFPY 273
Query: 374 WDLPIWKDISPRQRKVTAALKLVNDTLNNLIAICK---RMVDEEELQFHEEYMNEQDPSI 430
W+ P + I PRQRK + LK++ND L+ LI K + D E+LQ +Y N +D S+
Sbjct: 274 WNFPPARWIVPRQRKFQSDLKIINDCLDGLIQNAKETRQETDVEKLQ-ERDYTNLKDASL 332
Query: 431 LHFLL-ASGDDVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSV 489
L FL+ G D+ +QLRDDLMTMLIAGHET+AAVLTW +LLS+ P + K Q E+D+V
Sbjct: 333 LRFLVDMRGVDIDDRQLRDDLMTMLIAGHETTAAVLTWAVFLLSQNPEKIRKAQAEIDAV 392
Query: 490 LGDRFPTIEDMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVL-----GE---YPIKRG 541
LG PT E MKKL+Y ++ E LRL+PQPP+LIRR+++ + L GE + + +G
Sbjct: 393 LGQGPPTYESMKKLEYIRLIVVEVLRLFPQPPLLIRRTLKPETLPGGHKGEKEGHKVPKG 452
Query: 542 EDIFISVWNLHRSPTLWND 560
DIFISV+NLHRSP W++
Sbjct: 453 TDIFISVYNLHRSPYFWDN 471
>At1g57750 unknown protein
Length = 497
Score = 121 bits (304), Expect = 9e-28
Identities = 103/353 (29%), Positives = 165/353 (46%), Gaps = 34/353 (9%)
Query: 232 LIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIPADGEIWRVRRRTIVPALHLK- 290
L +DP HIL N Y KG + + V+G+G++ D E+W R++ H +
Sbjct: 80 LFTADPRNIHHILSSNFGNYPKGPEFKKIFDVLGEGILTVDFELWEEMRKSNHALFHNQD 139
Query: 291 FVAAMIGLF-GQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNYDFDSLSND 349
F+ + + + L LD AA +E++ +F R D + YD SLS
Sbjct: 140 FIELSVSSNKSKLKEGLVPFLDNAAQKNIIIELQDVFQRFMFDTSSILMTGYDPMSLS-- 197
Query: 350 TGIIEAVYTVLREAED--------RSISPIPVWDLPIWKDISPRQRKVTAALKLVNDTLN 401
IE + EA D R P+ +W L W I +RK+ AL VN
Sbjct: 198 ---IEMLEVEFGEAADIGEEAIYYRHFKPVILWRLQNWIGIG-LERKMRTALATVNRMFA 253
Query: 402 NLIA------ICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDDLMTMLI 455
+I+ I + + YMN D S L + D K +RD + ++++
Sbjct: 254 KIISSRRKEEISRAKTEPYSKDALTYYMNV-DTSKYKLLKPNKD----KFIRDVIFSLVL 308
Query: 456 AGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVINESLR 515
AG +T+++VLTW F+LLSK P VM+KL+ E+++ + ED++KL Y ++ES+R
Sbjct: 309 AGRDTTSSVLTWFFWLLSKHPQVMAKLRHEINTKFDN-----EDLEKLVYLHAALSESMR 363
Query: 516 LYPQPPVLIRRSIEDDVL-GEYPIKRGEDIFISVWNLHRSPTLWNDADKLNLK 567
LYP P + + DVL + + I I ++ L R ++W + D L+ K
Sbjct: 364 LYPPLPFNHKSPAKPDVLPSGHKVDANSKIVICIYALGRMRSVWGE-DALDFK 415
>At5g23190 cytochrome P450-like protein
Length = 559
Score = 121 bits (303), Expect = 1e-27
Identities = 95/355 (26%), Positives = 159/355 (44%), Gaps = 28/355 (7%)
Query: 226 FGPKSFLIVSDPAIAKHILKDNSKAYSKG-ILAEILDFVMGKGLIPADGEIWRVRRRTIV 284
F + I DP +H+LK+ + KG + L ++G G+ AD E W+ +R+T
Sbjct: 104 FSSLNSTITCDPRNVEHLLKNRFSVFPKGSYFRDNLRDLLGDGIFNADDETWQRQRKTAS 163
Query: 285 PALHLKFVAAMI--GLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNYD 342
H + LF RL L+T+ ++++ + RLT D + F D
Sbjct: 164 IEFHSAKFRQLTTQSLFELVHKRLLPVLETSVKSSSPIDLQDVLLRLTFDNVCMIAFGVD 223
Query: 343 FDSLSNDTGII---EAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTAALKLVNDT 399
L D +I +A A R + P VW + DI ++K+ ++K V+D
Sbjct: 224 PGCLGPDQPVIPFAKAFEDATEAAVVRFVMPTCVWKFMRYLDIGT-EKKLKESIKGVDDF 282
Query: 400 LNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLAS--GDDVTSKQLRDDLMTMLIAG 457
+ +I K+ EL E D + L G+ + K LRD + ++AG
Sbjct: 283 ADEVIRTRKK-----ELSLEGETTKRSDLLTVFMGLRDEKGESFSDKFLRDICVNFILAG 337
Query: 458 HETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVL-----------GDRFPTI--EDMKKLK 504
+TS+ L+W F+LL K P V K+ E+ +L D P E++KK+
Sbjct: 338 RDTSSVALSWFFWLLEKNPEVEEKIMVEMCKILRQRDDHGNAEKSDYEPVFGPEEIKKMD 397
Query: 505 YTTRVINESLRLYPQPPVLIRRSIEDDVLGE-YPIKRGEDIFISVWNLHRSPTLW 558
Y ++E+LRLYP PV + EDDV + +K+G+ + +++ + R +W
Sbjct: 398 YLQAALSEALRLYPSVPVDHKEVQEDDVFPDGTMLKKGDKVIYAIYAMGRMEAIW 452
>At4g37410 cytochrome P450 monooxygenase like protein
Length = 501
Score = 118 bits (296), Expect = 8e-27
Identities = 89/361 (24%), Positives = 168/361 (45%), Gaps = 27/361 (7%)
Query: 218 YGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKG----ILAEILDFVMGKGLIPADG 273
+G IF L G + +++S ++A+ + + ++ + + + G
Sbjct: 62 HGPIFYLRLGSRRAVVISSSSLARECFTGQNDVIVSNRPRFLTSKYIAYNYTTIATTSYG 121
Query: 274 EIWR-VRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLD 332
+ WR +RR + + K +A + + + R+ +L A G++VE+ES+ LT +
Sbjct: 122 DHWRNLRRICSLEIVSSKRLANFLHIRKEEIQRMLTRLSRDARVGKEVELESILYDLTFN 181
Query: 333 -----VIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPR-Q 386
V GK + D + + + ++T + S + P LP K +
Sbjct: 182 NIVRMVTGKIYYGDDVSD-KEEAELFKKLFTFITT---NSGARHPGEYLPFMKIFGGSFE 237
Query: 387 RKVTAALKLVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQL 446
++V AA K++++ L L+ CK D + H + + DP D+ K L
Sbjct: 238 KEVKAAAKVIDEMLQRLLDECKSDKDGNTMVNHLLSLQQDDPEYY-------TDIIIKGL 290
Query: 447 RDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLG-DRFPTIEDMKKLKY 505
++ +++A ETSA + W L P V+ K++ E+D ++G DR D+ L Y
Sbjct: 291 ---MLGIMVASSETSALTIEWAMASLLNHPKVLDKVKLEIDEIIGQDRLIEESDIANLPY 347
Query: 506 TTRVINESLRLYPQPPVLIRRSIEDDV-LGEYPIKRGEDIFISVWNLHRSPTLWNDADKL 564
V++E+LRL+P PVL+ RS +D+ +G Y + R + ++ W +HR P LW + ++
Sbjct: 348 LQNVVSETLRLHPAAPVLVPRSTAEDIKIGGYDVPRDTMVMVNAWAIHRDPDLWTEPERF 407
Query: 565 N 565
N
Sbjct: 408 N 408
>At3g14680 putative cytochrome P450
Length = 512
Score = 117 bits (294), Expect = 1e-26
Identities = 111/429 (25%), Positives = 193/429 (44%), Gaps = 45/429 (10%)
Query: 165 PSILKKSLSNLGVSNEILEFLFGLYPKIPEA--KGSISAIRSEAFFIPLY-----ELYIT 217
P +L++SL G+S L G + K+ + + I+ P ++ T
Sbjct: 32 PKMLERSLRRQGLSGTSYTPLIGDFKKMISMFIEATSKPIKPTDDITPRVMPHPLQMLKT 91
Query: 218 YGGIFRLNFGPKSFLIVSDPAIAKHILK---DNSKAYSKGILAEILDFVMGKGLIPADGE 274
+G FGP + + DP K + D KA++ L ++G GL+ DG+
Sbjct: 92 HGRTNLTWFGPIPTITIMDPEQIKEVFNKVYDFQKAHTFP-----LSKILGTGLVSYDGD 146
Query: 275 IWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASD-GEDVEMESL--FSRLTL 331
W RR I PA HL+ + M+ +F ++ L + D SD G E++ + +T
Sbjct: 147 KWAQHRRIINPAFHLEKIKNMVHVFHESCSELVGEWDKLVSDKGSSCEVDVWPGLTSMTA 206
Query: 332 DVIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTA 391
DVI + F + + + ++ +A + P ++ LP + R++
Sbjct: 207 DVISRTAFGSSYREGHRIFELQAELAQLVMQAFQKFFIPGYIY-LP-----TKGNRRMKT 260
Query: 392 ALKLVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLAS------GDDVTSKQ 445
A + + D L +I +R + E + +L LL S G+ ++++
Sbjct: 261 AAREIQDILRGIINKRERARESGEAPSED---------LLGILLESNLGQTEGNGMSTED 311
Query: 446 LRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKY 505
+ ++ +AG ET++ +L WT LLS+ ++ +EEV V GD+ P E + +LK
Sbjct: 312 MMEECKLFYLAGQETTSVLLVWTMVLLSQHQDWQARAREEVKQVFGDKQPDTEGLNQLKV 371
Query: 506 TTRVINESLRLYPQPPVLIRRSIEDDV-LGEYPIKRGEDIFISVWNLHRSPTLW-NDADK 563
T ++ E LRLYP P V + R+I ++ LG+ + G I + V +HR LW NDA +
Sbjct: 372 MTMILYEVLRLYP-PVVQLTRAIHKEMKLGDLTLPGGVQISLPVLLVHRDTELWGNDAGE 430
Query: 564 L---NLKDG 569
KDG
Sbjct: 431 FKPERFKDG 439
>At1g67110 unknown protein
Length = 512
Score = 117 bits (292), Expect = 2e-26
Identities = 92/351 (26%), Positives = 162/351 (45%), Gaps = 14/351 (3%)
Query: 218 YGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEI-LDFVMGKGLIPADGEIW 276
YG F + G + L +++ + K +L ++ K L + +G+GL+ A+GE W
Sbjct: 94 YGKRFIMWNGTEPRLCLTETEMIKELLTKHNPVTGKSWLQQQGTKGFIGRGLLMANGEAW 153
Query: 277 RVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGK 336
+R PA + + T + ++L GE+VE+ RLT D+I +
Sbjct: 154 HHQRHMAAPAFTRDRLKGYAKHMVECTKMMAERLRKEV--GEEVEIGEEMRRLTADIISR 211
Query: 337 AVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTAALKLV 396
F D ++ + + +A P + + + + +LK
Sbjct: 212 TEFGSSCDKGKELFSLLTVLQRLCAQATRHLCFPGS-------RFLPSKYNREIKSLKTE 264
Query: 397 NDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDDLMTMLIA 456
+ L L+ I D E+ Y ++ +L+ + ++ +++ + + D+ T
Sbjct: 265 VERL--LMEIIDSRKDSVEIGRSSSYGDDLLGLLLNQMDSNKNNLNVQMIMDECKTFFFT 322
Query: 457 GHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLG-DRFPTIEDMKKLKYTTRVINESLR 515
GHET++ +LTWT LL+ P+ +++EV V G D P++E + L +VINESLR
Sbjct: 323 GHETTSLLLTWTLMLLAHNPTWQDNVRDEVRQVCGQDGVPSVEQLSSLTSLNKVINESLR 382
Query: 516 LYPQPPVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLW-NDADKLN 565
LYP +L R + ED LG+ I +G I+I V +H S LW DA++ N
Sbjct: 383 LYPPATLLPRMAFEDIKLGDLIIPKGLSIWIPVLAIHHSNELWGEDANEFN 433
>At4g37400 cytochrome P450 monooxygenase -like protein
Length = 501
Score = 115 bits (288), Expect = 7e-26
Identities = 99/367 (26%), Positives = 167/367 (44%), Gaps = 33/367 (8%)
Query: 212 YELYITYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGK----- 266
+ L T+G IF L FG + +++S ++A + IL+ F+ K
Sbjct: 56 HRLSKTHGPIFSLQFGSRRAVVISSSSLATQCFTGQNDI----ILSNRPCFLTAKYVAYN 111
Query: 267 ----GLIPADGEIWRVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAASD-GEDVE 321
G P G+ WR RR + +L + + D + + L + D +++E
Sbjct: 112 YTTVGTAPY-GDHWRNLRR--ICSLEILSSNRLTNFLHIRKDEIHRMLTRLSRDVNKEIE 168
Query: 322 MESLFSRLTLDVIGKAVFN--YDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIW 379
+E L S LT + I + V Y D + N+ ++ + D S + P LP
Sbjct: 169 LEPLLSDLTFNNIVRMVTGKRYYGDEVHNEEEA-NVFKKLVADINDCSGARHPGDYLPFM 227
Query: 380 KDISPR-QRKVTAALKLVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASG 438
K ++KV A + +++ L L+ CKR D + H + + +P
Sbjct: 228 KMFGGSFEKKVKALAEAMDEILQRLLEECKRDKDGNTMVNHLLSLQQNEPEYY------- 280
Query: 439 DDVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLG-DRFPTI 497
DVT K L ++ M+IAG +TSA L W L P + K + E+D +G +R
Sbjct: 281 TDVTIKGL---MLGMMIAGTDTSAVTLEWAMSSLLNHPEALEKAKLEIDEKIGQERLIDE 337
Query: 498 EDMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDV-LGEYPIKRGEDIFISVWNLHRSPT 556
D+ L Y +++E+ RLYP P+L+ RS +D+ +G Y + RG + ++ W +HR P
Sbjct: 338 PDIANLPYLQNIVSETFRLYPAAPLLVPRSPTEDIKVGGYDVPRGTMVMVNAWAIHRDPE 397
Query: 557 LWNDADK 563
LWN+ +K
Sbjct: 398 LWNEPEK 404
>At3g48520 cytochrome P450-like protein
Length = 506
Score = 115 bits (287), Expect = 9e-26
Identities = 96/350 (27%), Positives = 174/350 (49%), Gaps = 24/350 (6%)
Query: 227 GPKSFLIVSDPAIAKHILKDNSKAYSKGI-LAEILDFVMGKGLIPADGEIWRVRRRTIVP 285
G + +I ++P ++ILK N + KG ++L ++G G+ DG W +R+
Sbjct: 77 GNRRTIITTNPLNVEYILKTNFFNFPKGKPFTDLLGDLLGGGIFNVDGHSWSSQRKLASH 136
Query: 286 ALHLKFVA--AMIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNYDF 343
+ + A L + +RL L TAA G V+++ + R DV+ K +D
Sbjct: 137 EFSTRSLRSFAFEVLKDEVENRLVPVLSTAADVGTTVDLQDVLKRFAFDVVCKVSLGWDP 196
Query: 344 DSLSNDTGI---IEAVYTVLREAEDRSISPI-PVWDLPIWKDISPRQRKVTAALKLVNDT 399
D L + +EA T + R+ PI VW ++ +RK+ A++ V+
Sbjct: 197 DCLDLTRPVNPLVEAFDTAAEISARRATEPIYAVWKTKRVLNVGS-ERKLREAIRTVHVL 255
Query: 400 LNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDDLMTMLIAGHE 459
++ ++ K+ L+ +QD +L LA+G + + +RD +++ ++AG +
Sbjct: 256 VSEIVRAKKK-----SLEIGTGAEAKQD--LLSRFLAAGHN--GEAVRDMVISFIMAGRD 306
Query: 460 TSAAVLTWTFYLLSKEPSVMSKLQEEVDSV--LGDRFPTIEDMKKLKYTTRVINESLRLY 517
T++A +TW F+LL++ V K+ EEVD + LG F ED+K++ YT + E++RLY
Sbjct: 307 TTSAAMTWLFWLLTENDDVERKILEEVDPLVSLGLGF---EDLKEMAYTKACLCEAMRLY 363
Query: 518 PQPPVLIRRSIEDDVLGE-YPIKRGEDIFISVWNLHRSPTLW-NDADKLN 565
P + + DDVL + +KRG+ + + + R TLW D+++ N
Sbjct: 364 PPVSWDSKHAANDDVLPDGTRVKRGDKVTYFPYGMGRMETLWGTDSEEFN 413
>At5g63450 cytochrome P450-like protein
Length = 510
Score = 114 bits (284), Expect = 2e-25
Identities = 89/343 (25%), Positives = 174/343 (49%), Gaps = 20/343 (5%)
Query: 224 LNFGPKSFLIVSDPAIAKHILKDNSKAYSKGI-LAEILDFVMGKGLIPADGEIWRVRRRT 282
L FG ++ +I ++P +HILK N + KG ++L ++G G+ +DGE+W +R+
Sbjct: 77 LLFGRRT-IITANPENVEHILKTNFYNFPKGKPFTDLLGDLLGGGIFNSDGELWSSQRKL 135
Query: 283 IVPALHLKFVAAMIG--LFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFN 340
++ + L + +RL L +A GE V+ + + R DV+ K
Sbjct: 136 ASHEFTMRSLREFTFEILREEVQNRLIPVLSSAVDCGETVDFQEVLKRFAFDVVCKVSLG 195
Query: 341 YDFDSLSNDTGIIEAVYTVLREAE---DRSISPI-PVWDLPIWKDISPRQRKVTAALKLV 396
+D D L + E V AE R+ P+ VW + + ++ +R + A+K V
Sbjct: 196 WDPDCLDLTRPVPELVKAFDVAAEISARRATEPVYAVWKVKRFLNVGSEKR-LREAIKTV 254
Query: 397 NDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDDLMTMLIA 456
+ +++ +I K+ +D + ++QD +L LA+G + +RD +++ ++A
Sbjct: 255 HLSVSEIIRAKKKSLD-----IGGDVSDKQD--LLSRFLAAGHG--EEAVRDSVISFIMA 305
Query: 457 GHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMKKLKYTTRVINESLRL 516
G +T++A +TW F+LLS+ V +K+ +E+ + G ED++++ YT + E++RL
Sbjct: 306 GRDTTSAAMTWLFWLLSQNDDVETKILDELRN-KGSLGLGFEDLREMSYTKACLCEAMRL 364
Query: 517 YPQPPVLIRRSIEDDVLGE-YPIKRGEDIFISVWNLHRSPTLW 558
YP + + DD+L + P+K+G+ + + + R +W
Sbjct: 365 YPPVAWDSKHAANDDILPDGTPLKKGDKVTYFPYGMGRMEKVW 407
>At3g56630 cytochrome P450-like protein
Length = 499
Score = 113 bits (283), Expect = 2e-25
Identities = 102/404 (25%), Positives = 192/404 (47%), Gaps = 29/404 (7%)
Query: 179 NEILEFLFGLYPKIPEAKGSIS-AIRSEAFFIPLYELYITYGGIFRLNFGPKSFLIVSDP 237
N EF F YP + G ++ R + + T IFR G F++ ++P
Sbjct: 24 NSSSEFGFKSYPIVGSLPGLVNNRHRFLDWTVETLSRCPTQTAIFRRP-GKLQFVMTANP 82
Query: 238 AIAKHILKDNSKAYSKGI-LAEILDFVMGKGLIPADGEIWRVRRRTIVPALHLKFVA--A 294
A +++LK +++ KG IL+ +G+G+ +DGE+W +R+T K +
Sbjct: 83 ANVEYMLKTKFESFPKGERFISILEDFLGRGIFNSDGEMWWKQRKTASYEFSTKSLRDFV 142
Query: 295 MIGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNYDFDSLSND----T 350
M + + RL L AA++G+ ++++ + R D I K FN D L +D
Sbjct: 143 MSNVTVEINTRLVPVLAEAATNGKLIDLQDILERFAFDNICKLAFNVDSACLGDDGAAGV 202
Query: 351 GIIEAVYTVLREAEDRSISPIPV-WDLPIWKDISPRQRKVTAALKLVNDTLNNLIAICKR 409
++A T R S I W + +I +R + ++ +V+ + ++ +
Sbjct: 203 NFMQAFETAATIISQRFQSVISYSWKIKKKLNIGS-ERVLRESIMIVHKFADEIV---RN 258
Query: 410 MVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDDLMTMLIAGHETSAAVLTWTF 469
+++ ++ H+E +L ++ + + + LRD +++ ++AG +T+++ L+W F
Sbjct: 259 RIEQGKVSDHKE-------DLLSRFISKEEMNSPEILRDIVISFILAGRDTTSSALSWFF 311
Query: 470 YLLSKEPSVMSKLQEEVDSV---LGDRFPTI---EDMKKLKYTTRVINESLRLYPQPPVL 523
+LLS P V K+ +E++S+ G R + ED+K + Y I ESLRLYP PV
Sbjct: 312 WLLSMHPEVKDKILQELNSIRERTGKRIGEVYGFEDLKLMNYLHAAITESLRLYPPVPVD 371
Query: 524 IRRSIEDDVLGEYP-IKRGEDIFISVWNLHRSPTLW-NDADKLN 565
ED+VL + I + I + + + R ++W D D+ +
Sbjct: 372 TMSCAEDNVLPDGTFIGKDWGISYNAYAMGRMESIWGKDCDRFD 415
>At2g44890 putative cytochrome P450
Length = 490
Score = 113 bits (282), Expect = 3e-25
Identities = 91/372 (24%), Positives = 173/372 (46%), Gaps = 31/372 (8%)
Query: 222 FRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEI-LDFVMGKGLIPADGEIWRVRR 280
FR +S + +DP +HILK YSKG + + L ++G G+ DGE W+ +R
Sbjct: 50 FRFLSPGQSEIFTADPRNVEHILKTRFHNYSKGPVGTVNLADLLGHGIFAVDGEKWKQQR 109
Query: 281 RTIVPALHLKFVAAM-IGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVF 339
+ + + + +F + +L + A G+ + + + + TLD I K F
Sbjct: 110 KLVSFEFSTRVLRNFSYSVFRTSASKLVGFIAEFALSGKSFDFQDMLMKCTLDSIFKVGF 169
Query: 340 NYDFDSLSNDTGIIEAVYTVLREAEDRSISPI--PVWDLPIWKDISPRQRKVTAALKLVN 397
+ L + E E + S + P W L + +I R + ++ +++
Sbjct: 170 GVELGCLDGFSKEGEEFMKAFDEGNGATSSRVTDPFWKLKCFLNIGSESR-LKKSIAIID 228
Query: 398 DTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGDD---VTSKQLRDDLMTML 454
+ +LI ++ + +E+ + ++ + FLL S D + K LRD ++ ++
Sbjct: 229 KFVYSLITTKRKELSKEQ------NTSVREDILSKFLLESEKDPENMNDKYLRDIILNVM 282
Query: 455 IAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTI-----------EDMKKL 503
+AG +T+AA L+W Y+L K P V K+ +E+ V T E + ++
Sbjct: 283 VAGKDTTAASLSWFLYMLCKNPLVQEKIVQEIRDVTSSHEKTTDVNGFIESVTEEALAQM 342
Query: 504 KYTTRVINESLRLYPQPPVLIRRSIEDDVLGE-YPIKRGEDIFISVWNLHRSPTLW-NDA 561
+Y ++E++RLYP P +R + DDVL + + + +G++I+ + + R +W DA
Sbjct: 343 QYLHAALSETMRLYPPVPEHMRCAENDDVLPDGHRVSKGDNIYYISYAMGRMTYIWGQDA 402
Query: 562 DKLN----LKDG 569
++ LKDG
Sbjct: 403 EEFKPERWLKDG 414
>At2g46950 putative cytochrome P450
Length = 517
Score = 112 bits (281), Expect = 4e-25
Identities = 116/425 (27%), Positives = 178/425 (41%), Gaps = 46/425 (10%)
Query: 165 PSILKKSLSNLGVSNEILEFLFGLYPKIPEAKGSISAI---RSEAFFIP-----LYELYI 216
P +L + G+S L+G +I + K + + +P L +
Sbjct: 33 PWMLSRRFKKQGISGPKYRILYGNLREIRKMKNEAKLMVLDPNSNDIVPRVLPHLQQWKS 92
Query: 217 TYGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIPADGEIW 276
YG F G L +SD +AK IL + +SK + + G GLI +G W
Sbjct: 93 QYGETFLYWQGTDPRLCISDHELAKQILSNKFVFFSKSKTKPEILKLSGNGLIFVNGLDW 152
Query: 277 RVRRRTIVPALHLKFVAAMIGLFGQATDRLC---QKLDTAASDGEDVEMESLFSRLTLDV 333
RR + PA + + M L T R+ +K + V + F RLT D+
Sbjct: 153 VRHRRILNPAFSMDKLKLMTQLMVDCTFRMFLEWKKQRNGVETEQFVLISREFKRLTADI 212
Query: 334 IGKAVFNYDF----DSLSNDTGIIEAVYTVLREAEDRSISPIPV-WDLPIWKDISPRQRK 388
I A F + + + + + L + I +P +L IWK K
Sbjct: 213 IATAAFGSSYAEGIEVFKSQLELQKCCAAALTDLYFPGIQYLPTPSNLQIWK----LDMK 268
Query: 389 VTAALKLVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLL--ASGDDVTSKQL 446
V +++K R++D ++Y N+ +L +L AS ++ K
Sbjct: 269 VNSSIK--------------RIIDARLTSESKDYGND----LLGIMLTAASSNESEKKMS 310
Query: 447 RDDLM----TMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLG-DRFPTIEDMK 501
D+++ T AGHET+A +LTW+ LLS KL+EEV + G D+ P E
Sbjct: 311 IDEIIEECKTFFFAGHETTANLLTWSTMLLSLHQDWQEKLREEVFNECGKDKIPDAETCS 370
Query: 502 KLKYTTRVINESLRLYPQPPVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLW-ND 560
KLK V ESLRLY L+R + ED LG I +G I + + +HR +W +D
Sbjct: 371 KLKLMNTVFMESLRLYGPVLNLLRLASEDMKLGNLEIPKGTTIILPIAKMHRDKAVWGSD 430
Query: 561 ADKLN 565
ADK N
Sbjct: 431 ADKFN 435
>At4g37430 cytochrome P450 monooxygenase (CYP91A2)
Length = 500
Score = 112 bits (279), Expect = 7e-25
Identities = 100/368 (27%), Positives = 166/368 (44%), Gaps = 35/368 (9%)
Query: 218 YGGIFRLNFGPKSFLIVSDPAIAKH-ILKDNSKAYSKGILAEILDFVMGK----GLIPAD 272
YG IF L FG + ++++ P++A+ N S L +V G P
Sbjct: 59 YGPIFSLRFGSRRVVVITSPSLAQESFTGQNDIVLSSRPLQLTAKYVAYNHTTVGTAPY- 117
Query: 273 GEIWRVRRRTI---VPALH--LKFVAAMIGLFGQATDRLCQKLDTA--ASDGEDVEMESL 325
G+ WR RR + + H + F + RL + T+ ++D +E+E L
Sbjct: 118 GDHWRNLRRICSQEILSSHRLINFQHIRKDEILRMLTRLSRYTQTSNESNDFTHIELEPL 177
Query: 326 FSRLTLD-----VIGKAVFNYDFDSLSNDTGIIEAVYTV-LREAEDRSISPIPVWDLPIW 379
S LT + V GK + D ++ + VY + + + S +P+ L
Sbjct: 178 LSDLTFNNIVRMVTGKRYYGDDVNNKEEAELFKKLVYDIAMYSGANHSADYLPILKL--- 234
Query: 380 KDISPRQRKVTAALKLVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASGD 439
+ +++V A K ++D L L+ C+R + + H + +Q P
Sbjct: 235 -FGNKFEKEVKAIGKSMDDILQRLLDECRRDKEGNTMVNHLISLQQQQPEYY-------T 286
Query: 440 DVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLG-DRFPTIE 498
DV K L +M+M++AG ETSA L W L + P V+ K + E+D +G DR
Sbjct: 287 DVIIKGL---MMSMMLAGTETSAVTLEWAMANLLRNPEVLEKARSEIDEKIGKDRLIDES 343
Query: 499 DMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDV-LGEYPIKRGEDIFISVWNLHRSPTL 557
D+ L Y V++E+ RL+P P LI RS DD+ +G Y + R + ++ W +HR P +
Sbjct: 344 DIAVLPYLQNVVSETFRLFPVAPFLIPRSPTDDMKIGGYDVPRDTIVMVNAWAIHRDPEI 403
Query: 558 WNDADKLN 565
W + +K N
Sbjct: 404 WEEPEKFN 411
>At5g24910 cytochrome P450-like protein
Length = 532
Score = 110 bits (276), Expect = 2e-24
Identities = 89/365 (24%), Positives = 165/365 (44%), Gaps = 30/365 (8%)
Query: 218 YGGIFRLNFGPKSFLIVSDPAIAKHILKDNSKAYSK-GILAEILDFVMGKGLIPADGEIW 276
YG ++ + G K L ++ P + K + + N+ K + + L ++G+G+I ++G W
Sbjct: 101 YGRVYTYSTGVKQHLYMNHPELVKELNQANTLNLGKVSYVTKRLKSILGRGVITSNGPHW 160
Query: 277 RVRRRTIVPALHLKFVAAMIGLFGQATDRLCQKLDTAAS-DGE---DVEMESLFSRLTLD 332
+RR I P L V M+GL ++ + K + +GE D+ ++ + D
Sbjct: 161 AHQRRIIAPEFFLDKVKGMVGLVVESAMPMLSKWEEMMKREGEMVCDIIVDEDLRAASAD 220
Query: 333 VIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLPIWKDISPRQRKVTAA 392
VI +A F F + +++ LR + ++ L + D+ V
Sbjct: 221 VISRACFGSSFSK-------GKEIFSKLRCLQKAITHNNILFSLNGFTDV------VFGT 267
Query: 393 LKLVNDTLNNLIAICKRMVDEEELQFHEEYMNEQDPSILHFLLASG--------DDVTSK 444
K N ++ L + ++ E + E + + ++ +L +D T
Sbjct: 268 KKHGNGKIDELERHIESLIWETVKERERECVGDHKKDLMQLILEGARSSCDGNLEDKTQS 327
Query: 445 Q---LRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIEDMK 501
+ D+ ++ AGHETSA ++W LL+ PS +++++EV + P + +
Sbjct: 328 YKSFVVDNCKSIYFAGHETSAVAVSWCLMLLALNPSWQTRIRDEVFLHCKNGIPDADSIS 387
Query: 502 KLKYTTRVINESLRLYPQPPVLIRRSIEDDVLGEYPIKRGEDIFISVWNLHRSPTLWN-D 560
LK T VI E+LRLYP + R ++ED LG + +G I+ + LHR P +W D
Sbjct: 388 NLKTVTMVIQETLRLYPPAAFVSREALEDTKLGNLVVPKGVCIWTLIPTLHRDPEIWGAD 447
Query: 561 ADKLN 565
A++ N
Sbjct: 448 ANEFN 452
>At5g08250 cytochrome P450-like protein
Length = 550
Score = 110 bits (274), Expect = 3e-24
Identities = 97/397 (24%), Positives = 172/397 (42%), Gaps = 37/397 (9%)
Query: 188 LYPKIPEAKGSISAIRSEAFFIPLYELYITYGGIFRLN---FGPKSFLIVSDPAIAKHIL 244
++P + ISA+RS + L ++ I+ G FR F + ++ DP +H+L
Sbjct: 65 VWPLVGMLPSLISAVRSNIYEW-LSDVLISQNGTFRFRGPWFSTLNCVVTCDPRNVEHLL 123
Query: 245 KDNSKAYSKG-ILAEILDFVMGKGLIPADGEIWRVRRRTIVPALHLKFVAAMIG--LFGQ 301
K Y KG E + ++G G+ D W+ +R+ H + L
Sbjct: 124 KTRFSIYPKGSYFRETMQDLLGDGIFNTDDGTWQRQRKAASVEFHSAKFRQLTSQSLHEL 183
Query: 302 ATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNYDFDSLSN---DTGIIEAVYT 358
+RL L+T+ ++++ + RLT D + F D LS + +A
Sbjct: 184 VHNRLLPVLETSGK----IDLQDILLRLTFDNVCMIAFGVDPGCLSPKLPEIPFAKAFED 239
Query: 359 VLREAEDRSISPIPVWDLPIWKDISPRQRKVTAALKLVNDTLNNLIAICKRMVDEEELQF 418
R + P VW L ++ ++K+ ++ V+D +I K+ E+
Sbjct: 240 ATEATVVRFVMPKFVWKLMRSLNLGT-EKKLKESINGVDDFAEEVIRTRKK-----EMSL 293
Query: 419 HEEYMNEQDPSILHFLLA--SGDDVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEP 476
E D + L +G + K LRD + ++AG +TS+ L+W F+L+ K P
Sbjct: 294 ETEIAKRPDLLTIFMGLRDENGQKFSDKFLRDICVNFILAGRDTSSVALSWFFWLIEKNP 353
Query: 477 SVMSKLQEEVDSVLGDRF------------PTI--EDMKKLKYTTRVINESLRLYPQPPV 522
V K+ + +L R P E++KK+ Y ++E+LRLYP PV
Sbjct: 354 EVEEKIMMGICKILEQRVDHGDTKKNMEYEPVFRPEEIKKMDYLQAALSETLRLYPSVPV 413
Query: 523 LIRRSIEDDVLGE-YPIKRGEDIFISVWNLHRSPTLW 558
+ +EDDV + +K+GE + +++ + R T+W
Sbjct: 414 DHKEVLEDDVFPDGTKLKKGEKVIYAIYAMGRMETIW 450
>At1g24540 putative cytochrome P450
Length = 522
Score = 110 bits (274), Expect = 3e-24
Identities = 95/371 (25%), Positives = 170/371 (45%), Gaps = 43/371 (11%)
Query: 219 GGIFRLN---FGPKSFLIVSDPAIAKHILKDNSKAYSKGIL-AEILDFVMGKGLIPADGE 274
GG F FG ++ +DPA +HILK N K Y KG E ++ G+ AD E
Sbjct: 74 GGTFHYRGIWFGGAYGIMTADPANVEHILKTNFKNYPKGAFYRERFRDLLEDGIFNADDE 133
Query: 275 IWRVRRRTIVPALHL-KFVAAMIGLFGQATD-RLCQKLDTAASDGEDVEMESLFSRLTLD 332
+W+ RR +H +F+ D +L ++ ++ +++ L R T D
Sbjct: 134 LWKEERRVAKTEMHSSRFLEHTFTTMRDLVDQKLVPLMENLSTSKRVFDLQDLLLRFTFD 193
Query: 333 VIGKAVFNYDFDSLSNDTGIIEAVYTVLREAEDRSISPIPVWDLP--IWKDIS----PRQ 386
I + F SL +TG+ E + + ED + + + +P +WK + +
Sbjct: 194 NICISAFGVYPGSL--ETGLPEIPFA--KAFEDATEYTLARFLIPPFVWKPMRFLGIGYE 249
Query: 387 RKVTAALKLVNDTLNNLIAICKRMV----------DEEELQFHEEYMNEQDPSILHFLLA 436
RK+ A+++V+ N + + + D EY E+D +
Sbjct: 250 RKLNNAVRIVHAFANKTVRERRNKMRKLGNLNDYADLLSRLMQREYEKEEDTT------- 302
Query: 437 SGDDVTSKQLRDDLMTMLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSV------- 489
G+ + K R+ + +IAG +T++ L W F+L+ K P V ++ E+ +
Sbjct: 303 RGNYFSDKYFREFCTSFIIAGRDTTSVALVWFFWLVQKHPEVEKRILREIREIKRKLTTQ 362
Query: 490 -LGDRFPTIEDMKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVLGE-YPIKRGEDIFIS 547
D+F ED +++ Y + ESLRLYP P+ +++++EDDVL + +K+G I S
Sbjct: 363 ETEDQFEA-EDFREMVYLQAALTESLRLYPSVPMEMKQALEDDVLPDGTRVKKGARIHYS 421
Query: 548 VWNLHRSPTLW 558
V+++ R ++W
Sbjct: 422 VYSMGRIESIW 432
>At4g39490 cytochrome P450 - like protein
Length = 479
Score = 109 bits (273), Expect = 4e-24
Identities = 99/370 (26%), Positives = 165/370 (43%), Gaps = 36/370 (9%)
Query: 226 FGPKSFLIVSDPAIAKHILKDNSKAYSKGILAEILDFVMGKGLIPADGEIWR-VRRRTIV 284
F L+ DPA HI+ N Y KG + L ++G G+ AD E+W+ +R+
Sbjct: 34 FANLDMLVTVDPANIHHIMSSNFANYPKGPEFKKLFDILGDGIFNADSELWKDLRKSAQS 93
Query: 285 PALHLKFVAAMIGLFGQATDR-LCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNYDF 343
++ +F + + ++ L LD A + V+++ +F R T D YD
Sbjct: 94 MMMNPEFQKFSLATSLKKLEKGLVPLLDHVAKEKLAVDLQDMFQRFTFDTTFVLATGYDP 153
Query: 344 DSLSNDTGIIEAVYTVLREAED----RSISPIPVWDLPIWKDISPRQRKVTAALKLVNDT 399
LS + +E L +AE+ R + P W L + ++K+T A ++
Sbjct: 154 GCLSVEMPEVEFA-RALDDAEEAIFFRHVKPEIFWRLQGLLGLGD-EKKMTKARSTLDRV 211
Query: 400 LNNLIAICK-------RMVDEEELQFHEEYMNEQDPSILHFLLASGDDVTSKQLRDDLMT 452
+ IAI + VD YMN + LL D+ + LRD ++T
Sbjct: 212 CSKYIAIKRDEVSRGTNNVDSHSKDLLTSYMNLDTTK--YKLLNPSDE---RFLRDTILT 266
Query: 453 MLIAGHETSAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTIED------------- 499
++AG +T+ + LTW F+LL K P V++K+++E+++ L R +D
Sbjct: 267 FMLAGRDTTGSGLTWFFWLLIKNPEVIAKIRQEINTALFQRSKVDDDASNNNDSDSFSPQ 326
Query: 500 -MKKLKYTTRVINESLRLYPQPPVLIRRSIEDDVL-GEYPIKRGEDIFISVWNLHRSPTL 557
+KKL Y I ESLRLYP P + + DVL + + I +++L R ++
Sbjct: 327 ELKKLVYLHGAICESLRLYPPVPFQHKSPTKPDVLPSGHKVDANSKILFCLYSLGRMKSV 386
Query: 558 WNDADKLNLK 567
W + D L K
Sbjct: 387 WGE-DALEFK 395
>At2g45510 putative cytochrome P450
Length = 511
Score = 109 bits (273), Expect = 4e-24
Identities = 96/366 (26%), Positives = 169/366 (45%), Gaps = 33/366 (9%)
Query: 229 KSFLIVSDPAIAKHILKDNSKAYSKGILA-EILDFVMGKGLIPADGEIWRVRRRTIVPAL 287
+S ++ +DP +HILK YSKG + E + ++G G+ DGE WR +R+
Sbjct: 78 QSEILTADPRNVEHILKTRFDNYSKGHSSRENMADLLGHGIFAVDGEKWRQQRKLSSFEF 137
Query: 288 HLKFVAAM-IGLFGQATDRLCQKLDTAASDGEDVEMESLFSRLTLDVIGKAVFNYDFDSL 346
+ + +F + +L + A G+ + + L R TLD I K F + L
Sbjct: 138 STRVLRDFSCSVFRRNASKLVGFVSEFALSGKAFDAQDLLMRCTLDSIFKVGFGVELKCL 197
Query: 347 SNDTGIIEAVYTVLREAEDRSISPI--PVWDLPIWKDISPRQRKVTAALKLVNDTLNNLI 404
+ + E + S P+W L + +I Q K+ ++ ++ + +LI
Sbjct: 198 DGFSKEGQEFMEAFDEGNVATSSRFIDPLWKLKWFFNIGS-QSKLKKSIATIDKFVYSLI 256
Query: 405 AIC-KRMVDEEELQFHEEYMNEQDPSILHFLLASGDD---VTSKQLRDDLMTMLIAGHET 460
K + E+ E+ ++ FL+ S D + K LRD ++ +IAG +T
Sbjct: 257 TTKRKELAKEQNTVVREDILSR-------FLVESEKDPENMNDKYLRDIILNFMIAGKDT 309
Query: 461 SAAVLTWTFYLLSKEPSVMSKLQEEVDSVLGDRFPTI-----------EDMKKLKYTTRV 509
+AA+L+W Y+L K P V K+ +E+ V T E + ++ Y
Sbjct: 310 TAALLSWFLYMLCKNPLVQEKIVQEIRDVTFSHEKTTDVNGFVESINEEALDEMHYLHAA 369
Query: 510 INESLRLYPQPPVLIRRSIEDDVLGE-YPIKRGEDIFISVWNLHRSPTLW-NDADKLN-- 565
++E+LRLYP PV +R + DDVL + + + +G++I+ + + R +W DA++
Sbjct: 370 LSETLRLYPPVPVDMRCAENDDVLPDGHRVSKGDNIYYIAYAMGRMTYIWGQDAEEFKPE 429
Query: 566 --LKDG 569
LKDG
Sbjct: 430 RWLKDG 435
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,053,586
Number of Sequences: 26719
Number of extensions: 572229
Number of successful extensions: 2360
Number of sequences better than 10.0: 252
Number of HSP's better than 10.0 without gapping: 232
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 1602
Number of HSP's gapped (non-prelim): 336
length of query: 569
length of database: 11,318,596
effective HSP length: 105
effective length of query: 464
effective length of database: 8,513,101
effective search space: 3950078864
effective search space used: 3950078864
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148994.8


