

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148970.12 + phase: 0
(560 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
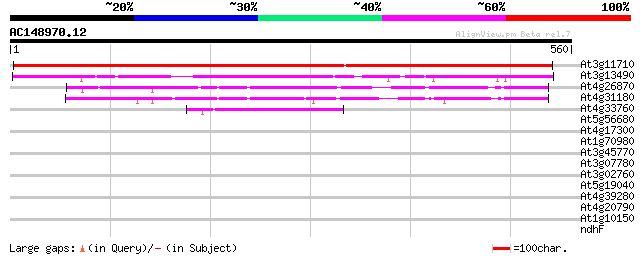
Score E
Sequences producing significant alignments: (bits) Value
At3g11710 lysyl-tRNA synthetase 850 0.0
At3g13490 lysyl-tRNA synthetase like protein 332 3e-91
At4g26870 putative aspartate-tRNA ligase 110 2e-24
At4g31180 aspartate--tRNA ligase - like protein 88 1e-17
At4g33760 aspartate--tRNA ligase like protein 66 6e-11
At5g56680 SYNC1 protein (gb|AAD46681.1) 40 0.003
At4g17300 asparagine--tRNA ligase 38 0.017
At1g70980 37 0.022
At3g45770 nuclear receptor binding factor-like protein 30 2.7
At3g07780 unknown protein 30 2.7
At3g02760 putative histidyl tRNA synthetase 30 3.5
At5g19040 tRNA isopentenyltransferase -like protein 30 4.6
At4g39280 phenylalanyl-trna synthetase - like protein 30 4.6
At4g20790 putative protein 30 4.6
At1g10150 unknown protein 30 4.6
ndhF -chloroplast genome- NADH dehydrogenase ND5 29 6.1
>At3g11710 lysyl-tRNA synthetase
Length = 626
Score = 850 bits (2197), Expect = 0.0
Identities = 411/539 (76%), Positives = 468/539 (86%), Gaps = 1/539 (0%)
Query: 4 AQENRPAAADEEDMDPTQYLENRLKYLSTEKEEGRNPYPHKFSATMTIKQYIKEYEGLSD 63
A + AAD+E+MD TQY ENRLKYL+ EK +G NPYPHKF+ +M+I +YI+ Y L++
Sbjct: 72 ASSQKAVAADDEEMDATQYYENRLKYLAAEKAKGENPYPHKFAVSMSIPKYIETYGSLNN 131
Query: 64 GQHLDDVSVSLTGKIMHKRSSGAKLVFYDLHDDGFKVQVMADASKSDLNEAGFDKVHSNV 123
G H+++ SL G+IM KRSS +KL FYDLH D FKVQVMADASKS L+EA F K+HSN
Sbjct: 132 GDHVENAEESLAGRIMSKRSSSSKLFFYDLHGDDFKVQVMADASKSGLDEAEFLKLHSNA 191
Query: 124 KRGDIVGITGFPGKSRKGELSIFPKTFVVLSHCLHMMPRQKSAAAAAADNAWLPGKTRNP 183
KRGDIVG+ GFPGK+++GELSIFP++F++LSHCLHMMPR+ A W+PG+TRNP
Sbjct: 192 KRGDIVGVIGFPGKTKRGELSIFPRSFILLSHCLHMMPRKADNVNAKKPEIWVPGQTRNP 251
Query: 184 EAYTLKDQETRYRLRHLDLMLNTEVREIFRTRSKIISYVRSFLDKLDFLEVETPMMNMIA 243
EAY LKDQE+RYR RHLD++LN EVR+IFRTR+KIISYVR FLD +FLEVETPMMNMIA
Sbjct: 252 EAYVLKDQESRYRQRHLDMILNVEVRQIFRTRAKIISYVRRFLDNKNFLEVETPMMNMIA 311
Query: 244 GGAAARPFVTHHNDLNMRLFMRIAPELYLKKLVVGGLNRVYEIGKQFRNEGIDLTHNPEF 303
GGAAARPFVTHHNDL+MRL+MRIAPELYLK+L+VGGL RVYEIGKQFRNEGIDLTHNPEF
Sbjct: 312 GGAAARPFVTHHNDLDMRLYMRIAPELYLKQLIVGGLERVYEIGKQFRNEGIDLTHNPEF 371
Query: 304 TTCEFYMAYKDYYDLMEITENMLSGMVYELTKGSYKIKYHADGLDKDPIEIDFTPPFRRI 363
TTCEFYMA+ DY DLME+TE MLSGMV ELT G YKIKY+A+G DKDPIEIDFTPPFRRI
Sbjct: 372 TTCEFYMAFADYNDLMEMTEVMLSGMVKELT-GGYKIKYNANGYDKDPIEIDFTPPFRRI 430
Query: 364 DMIEGLEEMAGLSIPKDLTSAEANQYLRDACLKYEIKCPPPETTARLLDKLVGHFLEETC 423
+MI LE++A L+IPKDL S EAN+YL DAC ++++KCPPP+TTARLLDKLVG FLE TC
Sbjct: 431 EMIGELEKVAKLNIPKDLASEEANKYLIDACARFDVKCPPPQTTARLLDKLVGEFLEPTC 490
Query: 424 VNPTFIINHPEIMSPLAKWHRTKPGLTERFELFINKHELANAYTELNDPVVQRQRFADQL 483
VNPTFIIN PEIMSPLAKWHR+K GLTERFELFINKHEL NAYTELNDPVVQRQRFADQL
Sbjct: 491 VNPTFIINQPEIMSPLAKWHRSKSGLTERFELFINKHELCNAYTELNDPVVQRQRFADQL 550
Query: 484 KDRQSGDDEAMALDETFCTALEYGLPPTAGWGLGIDRLTMLLTDSQNIKEVLLFPAMKP 542
KDRQSGDDEAMALDETFC ALEYGL PT GWGLGIDRL+MLLTDS NIKEVL FPAM+P
Sbjct: 551 KDRQSGDDEAMALDETFCNALEYGLAPTGGWGLGIDRLSMLLTDSLNIKEVLFFPAMRP 609
>At3g13490 lysyl-tRNA synthetase like protein
Length = 602
Score = 332 bits (851), Expect = 3e-91
Identities = 210/574 (36%), Positives = 316/574 (54%), Gaps = 70/574 (12%)
Query: 3 SAQENRPAAADEEDMDPTQYLENRLKYLSTEKEEGRNPYPHKFSATMTIKQYIKEYEGLS 62
S + R A++ D RLK + + +G PY +K+ + + Q + Y+ L+
Sbjct: 65 SGRNRRSASSSNSTSDREAIRSIRLKKVEELRGQGLEPYAYKWEKSHSANQLQEIYKHLA 124
Query: 63 DGQHLDDV--SVSLTGKIMHKRSSGAKLVFYDLHDDGFKVQVMADASKSDLNEAGFDKVH 120
+G+ D+ VS+ G+++ +R+ G KL F L DD +Q+ + K L++ F+++
Sbjct: 125 NGEESDNEIDCVSIAGRVVARRAFG-KLAFLTLRDDSGTIQLYCE--KERLSDDQFEQLK 181
Query: 121 SNVKRGDIVGITGFPGKSRKGELSIFPKTFVVLSHCLHMMPRQKSAAAAAADNAWLPGKT 180
+ GDI+G +G ++ KGELSI +F +L+ L +P
Sbjct: 182 QFIDIGDILGASGSMKRTEKGELSICVNSFSILTKSLLPLP------------------- 222
Query: 181 RNPEAYTLKDQETRYRLRHLDLMLNTEVREIFRTRSKIISYVRSFLDKLDFLEVETPMMN 240
+ + L D + RYR R++D++ N EV ++FR R+KI+S +R ++ +LEVETP++
Sbjct: 223 --DKYHGLTDIDKRYRQRYVDMIANPEVADVFRRRAKIVSEIRKTVESFGYLEVETPVLQ 280
Query: 241 MIAGGAAARPFVTHHNDLNMRLFMRIAPELYLKKLVVGGLNRVYEIGKQFRNEGIDLTHN 300
AGGA ARPFVT HN L L++RIA EL+LK+++VGG +VYEIG+ FRNEGI HN
Sbjct: 281 GAAGGAEARPFVTFHNSLGRDLYLRIATELHLKRMLVGGFEKVYEIGRIFRNEGISTRHN 340
Query: 301 PEFTTCEFYMAYKDYYDLMEITENMLSGMVYELTKGSYKIKYHADGLDKDPIEIDFTPPF 360
PEFTT E Y AY DY+ +M++ E ++ G I Y EI P+
Sbjct: 341 PEFTTIEMYEAYSDYHSMMDMAE-LIVTQCSMAVNGKLTIDYQG-------TEICLERPW 392
Query: 361 RRIDMIEGLEEMAGLS---IPKDLTSAEANQYLRDACLKYEIKCPPPETTARLLDKLVGH 417
RR M ++E+ G++ + +DL +A+ L ++ P ++ R L GH
Sbjct: 393 RRETMHNLVKEITGINFSELGEDLGNAKDTVLL----ALQDVLEPKDKSGIRACSSL-GH 447
Query: 418 FLEE--------TCVNPTFIINHPEIMSPLAKWHRTKPGLTERFELFINKHELANAYTEL 469
L E V PTF++++P +SPLAK HR GLTERFELFI E+ANA++EL
Sbjct: 448 LLNEIFEVVVEPKLVQPTFVLDYPIEISPLAKPHRGNAGLTERFELFICGREMANAFSEL 507
Query: 470 NDPVVQRQRFADQLKD------------------RQSGDDEA--MALDETFCTALEYGLP 509
DPV QR R +Q++ + DDE+ + LDE F TALEYG+P
Sbjct: 508 TDPVDQRTRLEEQVRQHNAKRAEAVRESPEPNAKKDDDDDESYEVTLDEDFLTALEYGMP 567
Query: 510 PTAGWGLGIDRLTMLLTDSQNIKEVLLFPAMKPQ 543
P +G GLGIDRL MLLT+S +I++V+ FP +K Q
Sbjct: 568 PASGMGLGIDRLVMLLTNSASIRDVIAFPVLKLQ 601
>At4g26870 putative aspartate-tRNA ligase
Length = 532
Score = 110 bits (275), Expect = 2e-24
Identities = 124/500 (24%), Positives = 209/500 (41%), Gaps = 56/500 (11%)
Query: 57 EYEGLSDGQHLDDVS----------VSLTGKIMHKRSSGAKLVFYDLHDDGFKVQVMADA 106
E + +G+ L DVS VS+ G++ R G KL F L + GF VQ + +
Sbjct: 63 ELQSAVEGKELTDVSNLVEEIVGSEVSIRGRLHKNRLVGTKL-FVILRESGFTVQCVVEE 121
Query: 107 SKSDLNEAGFDKVHSNVKRGDIVGITGFPGKSRKG---ELSIFPKTFVVLSHCLHMMPRQ 163
++ N F K S +++G+ P K G ++ I + LS L +P
Sbjct: 122 TRVGANMIKFVKQLSRESVVELIGVVSHPKKPLTGTTQQVEIHVRKMYCLSRSLPNLPLV 181
Query: 164 KSAAAAAADNAWLPGKTRNPEAYTLKDQETRYRLRHLDLMLNTEVREIFRTRSKIISYVR 223
AA + + GK A L Q+TR R LD+ + IFR + ++ R
Sbjct: 182 VEDAARSESDIEKSGKDGKQAARVL--QDTRLNNRVLDIRTPAN-QAIFRIQCQVQIAFR 238
Query: 224 SFLDKLDFLEVETPMMNMIAGGAAARPFVTHHNDLNMRLFMRIAPELYLKKLVVGGLNRV 283
+L FLE+ TP +IAG + V + + +P+L+ + + G + RV
Sbjct: 239 EYLQSKGFLEIHTP--KLIAGSSEGGSAVFRLDYKGQPACLAQSPQLHKQMAICGDMRRV 296
Query: 284 YEIGKQFRNE-GIDLTHNPEFTTCEFYMAYKDYY-DLMEITENMLSGMVYELTKGSYKIK 341
+E+G FR E H EF + M + +Y ++M++ + + TK +
Sbjct: 297 FEVGPVFRAEDSFTHRHLCEFVGLDVEMEIRMHYSEIMDLVGELFP---FIFTKIEERCP 353
Query: 342 YHADGLDKD-PIE-IDFTPPFRRIDMIEGLEEMAGLSIPKDLTSAEANQYLRDACLKYEI 399
+ + K P + + F P R LT AE Q L++A + +
Sbjct: 354 KELESVRKQYPFQSLKFLPQTLR------------------LTFAEGIQMLKEAGEEVDP 395
Query: 400 KCPPPETTARLLDKLVGHFLEETCVNPTFIINHPEIMSPLAKW-HRTKPGLTERFELFIN 458
+ R L +LV LE+ + +P + P + + F++FI
Sbjct: 396 LGDLNTESERKLGQLV---LEKYKTEFYMLHRYPSAVRPFYTMPYENDSNYSNSFDVFIR 452
Query: 459 KHELANAYTELNDPVVQRQRFADQLKDRQSGDDEAMALDETFCTALEYGLPPTAGWGLGI 518
E+ + ++DP + +R R+ G D + T+ A YG PP G+G+G+
Sbjct: 453 GEEIMSGAQRIHDPELLEKR------ARECGID--VKTISTYIDAFRYGAPPHGGFGVGL 504
Query: 519 DRLTMLLTDSQNIKEVLLFP 538
+R+ MLL NI++ LFP
Sbjct: 505 ERVVMLLCALNNIRKTSLFP 524
>At4g31180 aspartate--tRNA ligase - like protein
Length = 558
Score = 88.2 bits (217), Expect = 1e-17
Identities = 117/498 (23%), Positives = 207/498 (41%), Gaps = 55/498 (11%)
Query: 56 KEYEGLSD-GQHLDDVSVSLTGKIMHKRSSGAKLVFYDLHDDGFKVQVMADASKSDLNEA 114
KE+ +SD + + + V + G++ R + KL F L + G VQ + S+ A
Sbjct: 93 KEWTDVSDLVEEMLESEVLIRGRVHTNRPTSNKLGFVVLRESGSTVQCVVSQSEKTKVGA 152
Query: 115 GFDKVHSNVKRG---DIVGITGFPGKSRKG---ELSIFPKTFVVLSHCLHMMPRQKSAAA 168
K + R D++G+ P + G ++ I + ++ L +P S
Sbjct: 153 NMVKYLKQLSRESFVDVIGVVTLPKEPLTGTTQQVEIQVRKVYCINKSLAKLPL--SVED 210
Query: 169 AAADNAWLPGKTRNPEAYTLKDQETRYRLRHLDLMLNTEVREIFRTRSKIISYVRSFLDK 228
AA A + + P +Q+TR R LDL + IF+ + ++ R L
Sbjct: 211 AARSEADIEASLQTPSPAARVNQDTRLNYRVLDLRTPAN-QAIFQLQYEVEYAFREKLRF 269
Query: 229 LDFLEVETPMMNMIAGGAAARPFVTHHNDLNMRLFMRIAPELYLKKLVVGGLNRVYEIGK 288
+F+ + TP ++AG + V + +P+L+ + + G L RV+E+G
Sbjct: 270 KNFVGIHTP--KLMAGSSEGGSAVFRLEYKGQPACLAQSPQLHKQMAICGDLRRVFEVGP 327
Query: 289 QFRNEGIDLTHNP--EFTTCEFYMAYKDYY-DLMEITENMLSGMVYELTKGSYKIKYHAD 345
FR E TH EF + M + +Y ++M++ + + V+ T + + K
Sbjct: 328 VFRAED-SFTHRHLCEFVGLDVEMEIRKHYSEIMDLVDELF---VFIFTSLNERCKKELQ 383
Query: 346 GLDKD-PIE-IDFTPPFRRIDMIEGLEEMAGLSIPKDLTSAEANQYLRDACLKYEIKCPP 403
+ K P E + F P R+ EG+ Q L++A ++ +
Sbjct: 384 AVGKQYPFEPLKFLPKTLRLTFEEGV------------------QMLKEAGVEVDPLGDL 425
Query: 404 PETTARLLDKLVGHFLEETCVNPTFIINH--PEIMSPLAKWH-RTKPGLTERFELFINKH 460
+ R L +LV LE+ N F I H P+ + P P + F++FI
Sbjct: 426 NTESERKLGQLV---LEK--YNTEFYILHRYPKAVRPFYTMTCADNPLYSNSFDVFIRGE 480
Query: 461 ELANAYTELNDPVVQRQRFADQLKDRQSGDDEAMALDETFCTALEYGLPPTAGWGLGIDR 520
E+ + ++ P V QR + G D + T+ + YG P G+G+G++R
Sbjct: 481 EIISGAQRVHIPEVLEQRAGE------CGID--VKTISTYIDSFRYGAPLHGGFGVGLER 532
Query: 521 LTMLLTDSQNIKEVLLFP 538
+ ML NI++ LFP
Sbjct: 533 VVMLFCALNNIRKTSLFP 550
>At4g33760 aspartate--tRNA ligase like protein
Length = 664
Score = 65.9 bits (159), Expect = 6e-11
Identities = 42/165 (25%), Positives = 81/165 (48%), Gaps = 9/165 (5%)
Query: 177 PGKTRNPEAYTLKDQ-------ETRYRLRHLDLMLNTEVREIFRTRSKIISYVRSFL-DK 228
P +T+ P T D+ E R R R LDL +++ R ++ +R +L D+
Sbjct: 178 PVRTKLPFLVTTADENKDLIKEEIRLRFRCLDLR-RQQMKNNIVLRHNVVKLIRRYLEDR 236
Query: 229 LDFLEVETPMMNMIAGGAAARPFVTHHNDLNMRLFMRIAPELYLKKLVVGGLNRVYEIGK 288
F+E+ETP+++ A V + +P+L+ + L+V G ++ Y+I +
Sbjct: 237 HGFIEIETPILSRSTPEGARDYLVPSRIQSGTFYALPQSPQLFKQMLMVSGFDKYYQIAR 296
Query: 289 QFRNEGIDLTHNPEFTTCEFYMAYKDYYDLMEITENMLSGMVYEL 333
FR+E + PEFT + MA+ D++++ E+++ + E+
Sbjct: 297 CFRDEDLRADRQPEFTQLDMEMAFMPMEDMLKLNEDLIRKVFSEI 341
Score = 37.4 bits (85), Expect = 0.022
Identities = 15/36 (41%), Positives = 24/36 (66%)
Query: 503 ALEYGLPPTAGWGLGIDRLTMLLTDSQNIKEVLLFP 538
AL+ G PP G G+DR+ M+L + +I++V+ FP
Sbjct: 600 ALDMGAPPHGGIAYGLDRMVMMLGGASSIRDVIAFP 635
>At5g56680 SYNC1 protein (gb|AAD46681.1)
Length = 572
Score = 40.4 bits (93), Expect = 0.003
Identities = 64/288 (22%), Positives = 108/288 (37%), Gaps = 45/288 (15%)
Query: 263 FMRIAPELYLKKLVVGGLNRVYEIGKQFRNEGIDLT-HNPEFTTCEFYMAYKDYYDLMEI 321
F+ ++ +L ++ L+ VY G FR E + H EF E +A+ D D M
Sbjct: 310 FLTVSGQLQVETYACA-LSNVYTFGPTFRAENSHTSRHLAEFWMVEPEIAFADLEDDMNC 368
Query: 322 TENMLSGMVYELTKGSYK-IKYHADGLDK---DPIEIDFTPPFRRIDMIEGLEEMAGLSI 377
E + M L + Y ++ A D D +++ + PF RI
Sbjct: 369 AEAYVKYMCNWLLEKCYADMELMAKNFDSGCIDRLKLVASTPFGRI-------------- 414
Query: 378 PKDLTSAEANQYLRDACLKYEIKCPPPETTARLLDKLVGHFLEETCVNPTFIINHPEIMS 437
T +A + L +A K + E L + + E P + N+P+ +
Sbjct: 415 ----TYTKAIELLEEAVAKGKEFDNNVEWGIDLASEHERYLTEVLFQKPLIVYNYPKGI- 469
Query: 438 PLAKWHRTKPGLTERFELFINKHELANAYTELNDPVV-------QRQRFADQLKDRQSGD 490
+ F + +N E A ++ P V QR+ D +K R
Sbjct: 470 -------------KAFYMRLNDDEKTVAAMDVLVPKVGELIGGSQREERYDVIKKRIEEM 516
Query: 491 DEAMALDETFCTALEYGLPPTAGWGLGIDRLTMLLTDSQNIKEVLLFP 538
+ E + YG G+GLG +R+ + T NI++V+ FP
Sbjct: 517 GLPIEPYEWYLDLRRYGTVKHCGFGLGFERMILFATGLDNIRDVIPFP 564
>At4g17300 asparagine--tRNA ligase
Length = 567
Score = 37.7 bits (86), Expect = 0.017
Identities = 60/264 (22%), Positives = 106/264 (39%), Gaps = 32/264 (12%)
Query: 280 LNRVYEIGKQFRNEGIDLT-HNPEFTTCEFYMAYKDYYDLMEITENMLSGMVYELTKGSY 338
L+ VY G FR E + + H EF E +A+ D D M L Y
Sbjct: 323 LSDVYTFGPTFRAENSNTSRHLAEFWMIEPELAFADLDDDMACATAYLQ----------Y 372
Query: 339 KIKYHADGLDKDPIEIDFTPPFRRIDMIEGLEEMAGLSIPKDLTSAEANQYLRDACLKYE 398
+KY D +D ++F + +I L ++A + L +A + L A K++
Sbjct: 373 VVKYVLDNCKED---MEFFDTWIEKGIIRRLSDVAEKEFLQ-LGYTDAIEILLKANKKFD 428
Query: 399 IKCPPPETTARLLDKLVGHFLEETCVN-PTFIINHPEIMSPLAKWHRTKPGLTERFELFI 457
P + L + + EE P I ++P+ + ++ +
Sbjct: 429 F---PVKWGLDLQSEHERYITEEAFGGRPVIIRDYPKEIKAFYMRENDDGKTVAAMDMLV 485
Query: 458 NKHELANAYTELNDPVVQRQRFADQLKDRQSGDDEAMALDETFCTALE---YGLPPTAGW 514
+ + + + QR ++L+ ++ DE E++ L+ YG P AG+
Sbjct: 486 PR---------IGELIGGSQR-EERLEVLEARLDELKLNKESYWWYLDLRRYGSVPHAGF 535
Query: 515 GLGIDRLTMLLTDSQNIKEVLLFP 538
GLG +RL +T NI++V+ FP
Sbjct: 536 GLGFERLVQFVTGIDNIRDVIPFP 559
>At1g70980
Length = 571
Score = 37.4 bits (85), Expect = 0.022
Identities = 65/299 (21%), Positives = 114/299 (37%), Gaps = 42/299 (14%)
Query: 252 VTHHNDL-NMRLFMRIAPELYLKKLVVGGLNRVYEIGKQFRNEGIDLT-HNPEFTTCEFY 309
+ + ND + F+ ++ +L ++ L+ VY G FR E + H EF E
Sbjct: 295 IDYSNDFFGRQAFLTVSGQLQVETYACA-LSSVYTFGPTFRAENSHTSRHLAEFWMVEPE 353
Query: 310 MAYKDYYDLMEITENMLSGMVYELTKGSYKIKYHADGLDKDPIEIDFTPPFRRIDMIEGL 369
+A+ D +D M E + Y K+ D D +D + +D EG
Sbjct: 354 IAFADIHDDMNCAEAYVK----------YMCKWLMDKCGDDMELMD-----KNVD--EGC 396
Query: 370 EE---MAGLSIPKDLTSAEANQYLRDACLKYEIKCPPPETTARLLDKLVGHFLEETCVNP 426
+ M + K +T EA + L A + ++ + D V ++ +
Sbjct: 397 TKRLNMVAKASFKRVTYTEAIERLEKAVAQGKV----------VFDNKVEWGIDLASEHE 446
Query: 427 TFIINHPEIMSPLAKWHRTKPGLTERFELFINKHELANAYTELNDPVV-------QRQRF 479
++ P+ ++ K + F + +N E A ++ P V QR+
Sbjct: 447 RYLTEVEFDQKPIIVYNYPKG--IKAFYMRLNDDEKTVAAMDVLVPKVGELIGGSQREER 504
Query: 480 ADQLKDRQSGDDEAMALDETFCTALEYGLPPTAGWGLGIDRLTMLLTDSQNIKEVLLFP 538
D +K R M E + YG G+GLG +R+ T NI++V+ FP
Sbjct: 505 YDVIKQRIEEMGLPMEPYEWYLDLRRYGTVKHCGFGLGFERMIQFATGIDNIRDVIPFP 563
>At3g45770 nuclear receptor binding factor-like protein
Length = 375
Score = 30.4 bits (67), Expect = 2.7
Identities = 29/97 (29%), Positives = 41/97 (41%), Gaps = 6/97 (6%)
Query: 324 NMLSGMVYELTKGSYKIKYHADGLDKDPIEIDFTPPFRRIDMIEGLEEMAGLSIPKDLTS 383
N S ++ L +G + Y G+ K PI + T + + G + LS+ K
Sbjct: 269 NAASLVLKYLREGGTMVTY--GGMSKKPITVSTTSFIFKDLALRGFWLQSWLSMGKVKEC 326
Query: 384 AEANQYL----RDACLKYEIKCPPPETTARLLDKLVG 416
E YL RD LKYE + P E LDK +G
Sbjct: 327 REMIDYLLGLARDGKLKYETELVPFEEFPVALDKALG 363
>At3g07780 unknown protein
Length = 566
Score = 30.4 bits (67), Expect = 2.7
Identities = 31/129 (24%), Positives = 56/129 (43%), Gaps = 15/129 (11%)
Query: 162 RQKSAAAAAADNAWLPGKT----RNPEAYTLKDQETRYRLRHLDLMLNTEVREIFRTRSK 217
R+ A +A++ W K+ + T D ++ +RH+ + +R+I R R
Sbjct: 42 RESPAESASSQETWPTSKSIMGRKTDSGKTGPDSHDQHVIRHVSIADKVSLRDIARERLD 101
Query: 218 IISYVRSFLDKLDFLEVETPMMNMIAGGAAARP---------FVTHHNDLNMRLFMRIAP 268
I++ L + ++LE + I G A+P FV +DL + +R A
Sbjct: 102 IVAERMHRLPE-EYLEELKNGLKAILEGNGAQPIDEFMFLQKFVQTRSDLTSKTLVR-AH 159
Query: 269 ELYLKKLVV 277
+ L+ LVV
Sbjct: 160 RVQLEVLVV 168
>At3g02760 putative histidyl tRNA synthetase
Length = 479
Score = 30.0 bits (66), Expect = 3.5
Identities = 28/103 (27%), Positives = 51/103 (49%), Gaps = 6/103 (5%)
Query: 284 YEIGKQFRNEGIDLTHNPEFTTCEFYMA--YKDYYDLMEITENMLSGMVYELTKGSYKIK 341
Y+I K +R + EF C+F +A ++ EI + +L+ ++ EL G Y++K
Sbjct: 136 YQIAKVYRRDNPSKGRYREFYQCDFDIAGLFEPMGPDFEIVK-ILTELLDELEIGDYEVK 194
Query: 342 YHADGLDKDPIEIDFTPP--FRRI-DMIEGLEEMAGLSIPKDL 381
+ L +EI PP FR I I+ L++ + + K++
Sbjct: 195 LNHRKLLDGMLEICGVPPEKFRTICSSIDKLDKQSFEQVKKEM 237
>At5g19040 tRNA isopentenyltransferase -like protein
Length = 330
Score = 29.6 bits (65), Expect = 4.6
Identities = 18/62 (29%), Positives = 29/62 (46%)
Query: 392 DACLKYEIKCPPPETTARLLDKLVGHFLEETCVNPTFIINHPEIMSPLAKWHRTKPGLTE 451
D L+ E++ P ETT RLL+ + E TC+ + + + KW+ + TE
Sbjct: 209 DEFLRSEMRNYPAETTERLLETAIEKIKENTCLLACRQLQKIQRLYKQWKWNMHRVDATE 268
Query: 452 RF 453
F
Sbjct: 269 VF 270
>At4g39280 phenylalanyl-trna synthetase - like protein
Length = 485
Score = 29.6 bits (65), Expect = 4.6
Identities = 16/55 (29%), Positives = 27/55 (49%), Gaps = 7/55 (12%)
Query: 253 THHNDLNMRLFMRIAPELYLKKLVVGGLNRVYEIGKQFRNEGIDLTHNPEFTTCE 307
TH ++ R+ +A + ++ K + + I + FRNE +D TH EF E
Sbjct: 315 THTTAVSSRMLYALAQKPFVPK-------KYFSIDRVFRNEAVDRTHLAEFHQIE 362
>At4g20790 putative protein
Length = 518
Score = 29.6 bits (65), Expect = 4.6
Identities = 19/69 (27%), Positives = 29/69 (41%), Gaps = 2/69 (2%)
Query: 369 LEEMAGLSIPKDLTSAEANQYLRDACLKY--EIKCPPPETTARLLDKLVGHFLEETCVNP 426
+E++A LS P D + +N LRD L+Y P + +V L NP
Sbjct: 29 VEDLANLSPPPDFNTTISNNCLRDPMLRYCNNTSSSPMDIVEIFRSTIVASHLCNESKNP 88
Query: 427 TFIINHPEI 435
+ + P I
Sbjct: 89 NCVESFPRI 97
>At1g10150 unknown protein
Length = 414
Score = 29.6 bits (65), Expect = 4.6
Identities = 28/87 (32%), Positives = 41/87 (46%), Gaps = 15/87 (17%)
Query: 48 TMTIKQYIKEYEGLSDGQHLD---DVSVSLTGKIMHK---------RSSGAKLVFYDLHD 95
TM I Y + +EGL++ +H D DV VS+ + SS +K V ++ D
Sbjct: 229 TMDINTYDQIFEGLTNPKHQDSVKDVLVSVCNGALETIVRTSHDVFTSSRSKNVIEEIED 288
Query: 96 DGFKVQVMADASKSDLNEAGFDKVHSN 122
D FK A ++E+G D V SN
Sbjct: 289 DDFKSN--GSARSKMVSESG-DGVKSN 312
>ndhF -chloroplast genome- NADH dehydrogenase ND5
Length = 746
Score = 29.3 bits (64), Expect = 6.1
Identities = 20/78 (25%), Positives = 36/78 (45%), Gaps = 3/78 (3%)
Query: 281 NRVYEIGKQFRNEGIDLTHNPEFTTCEFYMAYKDYYDLMEITENMLSGMVYELTKGSYKI 340
N+ Y+I RN+ N T FY ++ ++ ML +++ L G+ I
Sbjct: 510 NKTYKISNNVRNQIFITVENFGLNTRTFYYPHESDNTILF---PMLILVLFTLFIGAIGI 566
Query: 341 KYHADGLDKDPIEIDFTP 358
++ +G+D D + FTP
Sbjct: 567 PFNQEGIDFDILSKFFTP 584
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,861,971
Number of Sequences: 26719
Number of extensions: 560400
Number of successful extensions: 1422
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1400
Number of HSP's gapped (non-prelim): 20
length of query: 560
length of database: 11,318,596
effective HSP length: 104
effective length of query: 456
effective length of database: 8,539,820
effective search space: 3894157920
effective search space used: 3894157920
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148970.12


