

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148918.4 - phase: 0
(468 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
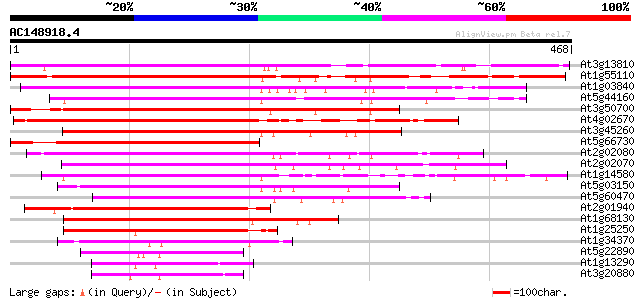
Score E
Sequences producing significant alignments: (bits) Value
At3g13810 zinc finger protein, putative 430 e-121
At1g55110 putative zinc finger protein 396 e-110
At1g03840 putative DNA-binding protein 365 e-101
At5g44160 unknown protein 350 1e-96
At3g50700 zinc finger protein 342 3e-94
At4g02670 putative zinc finger protein 335 2e-92
At3g45260 zinc finger like protein 335 3e-92
At5g66730 zinc finger protein 330 1e-90
At2g02080 C2H2-type zinc finger protein 328 4e-90
At2g02070 putative C2H2-type zinc finger protein 322 2e-88
At1g14580 putative zinc finger protein 322 2e-88
At5g03150 putative protein 312 2e-85
At5g60470 putative zinc finger protein 288 6e-78
At2g01940 putative C2H2-type zinc finger protein 236 3e-62
At1g68130 putative C2H2-type zinc finger protein 233 2e-61
At1g25250 Indeterminate 1 like Zn finger containing protein 228 7e-60
At1g34370 zinc finger protein, putative 106 2e-23
At5g22890 unknown protein 97 1e-20
At1g13290 hypothetical protein 91 2e-18
At3g20880 putative DNA-binding protein 88 1e-17
>At3g13810 zinc finger protein, putative
Length = 513
Score = 430 bits (1106), Expect = e-121
Identities = 252/527 (47%), Positives = 311/527 (58%), Gaps = 93/527 (17%)
Query: 1 MSNLTSASGEASANSSGNRTHEVDAKF-----SQQYFASSQTQTHDETPAKKRRNLPGNP 55
MSNLTSASG+ ++ SSGN T + + QQ Q D KKRRN PGNP
Sbjct: 20 MSNLTSASGDQASVSSGNITEASGSNYFPHHQQQQEQQQQQLVVPDSQTQKKRRNQPGNP 79
Query: 56 DPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNE-IRKK 114
DP++EVIALSPKTLMATNRF+CEICNKGFQRDQNLQLH+RGHNLPWKLKQR++ E IRKK
Sbjct: 80 DPESEVIALSPKTLMATNRFVCEICNKGFQRDQNLQLHRRGHNLPWKLKQRSNKEVIRKK 139
Query: 115 VYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCG 174
VYVCPE +CVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSD KAHSKTCG
Sbjct: 140 VYVCPEASCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDCKAHSKTCG 199
Query: 175 TREYRCDCGTLFSRRDSFITHRAFCDALAEESSRTVI---------PQP-------TQPN 218
T+EYRCDCGTLFSRRDSFITHRAFC+ALAEE++R V+ P P + P+
Sbjct: 200 TKEYRCDCGTLFSRRDSFITHRAFCEALAEETAREVVIPQNQNNNQPNPLLIHQSASHPH 259
Query: 219 SHH---------------------------------NMNNLQTQDIQGFTLKKEHQSFNM 245
HH N +N + F +KKE QS +
Sbjct: 260 HHHQTQPTINVSSSSSSSHNHNIINSLHFDTNNGNTNNSNNSNNHLHTFPMKKEQQSNDH 319
Query: 246 LRPEQEVQIPSWLCQSSIDLSSNYSSLDQDLHLYENPNPRNGPTSTLPSYQPSSAASPHM 305
+ IP WL L+S+ NPNP NG + S ASP M
Sbjct: 320 IMNYHHSIIPPWLAPQPHALTSS------------NPNPSNGGGGGGSLF---SLASPAM 364
Query: 306 SATALLQKAAQMGATSSCSSQSMMSGTHQQGHVSIVDSATNNMINSNGNFSLNLSSCEDQ 365
SATALLQKAAQMG+T + + + ++ + M + +G S S+ +
Sbjct: 365 SATALLQKAAQMGSTKTPPLPPTTAYERSTHNNNLTTTMAAMMTSPSGFIS---SNNNNH 421
Query: 366 MINNSFSSSGF--HG--TSFEDTFAGNILHSNQDHNINHDGDNDIPKTTTNDDDVAAGGN 421
++ +++SGF HG +F+DTF G + N++ ++ +GG
Sbjct: 422 VLFQDYNASGFDNHGREEAFDDTFGGFL------------RTNEVTAAAGSEKSTKSGGG 469
Query: 422 NAFTRDFLGLKPL-SDSDILTIAGMGSCMNPSNSNHQENHSQKPWEG 467
TRDFLGL+PL S ++IL+ AG+GSC+N S S+ KPW+G
Sbjct: 470 EGLTRDFLGLRPLMSHNEILSFAGLGSCINSSASDQLH---PKPWQG 513
>At1g55110 putative zinc finger protein
Length = 455
Score = 396 bits (1017), Expect = e-110
Identities = 241/488 (49%), Positives = 299/488 (60%), Gaps = 79/488 (16%)
Query: 1 MSNLTSASGEASANSSGNRTHEVDAKFSQQYFASSQTQTH-DETPAKKRRNLPGNPDPQA 59
MSNLTSASG+ ++ SSGNRT + +Q + Q Q ++ K++RN PGNPDP+A
Sbjct: 21 MSNLTSASGDQASVSSGNRTETSGSNINQHH----QEQCFVPQSSLKRKRNQPGNPDPEA 76
Query: 60 EVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEI-RKKVYVC 118
EV+ALSPKTLMATNRFICE+CNKGFQRDQNLQLHKRGHNLPWKLKQR++ ++ RKKVYVC
Sbjct: 77 EVMALSPKTLMATNRFICEVCNKGFQRDQNLQLHKRGHNLPWKLKQRSNKDVVRKKVYVC 136
Query: 119 PEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREY 178
PEP CVHH PSRALGDLTGIKKHF RKHGEKKWKC+KCSKKYAVQSDWKAH+KTCGT+EY
Sbjct: 137 PEPGCVHHHPSRALGDLTGIKKHFFRKHGEKKWKCEKCSKKYAVQSDWKAHAKTCGTKEY 196
Query: 179 RCDCGTLFSRRDSFITHRAFCDALAEESSRT-----VIPQPTQPNSHHNMNNLQTQDIQG 233
+CDCGTLFSRRDSFITHRAFCDALAEES+R +I P+ HH+ QTQ G
Sbjct: 197 KCDCGTLFSRRDSFITHRAFCDALAEESARAMPNPIMIQASNSPHHHHH----QTQQNIG 252
Query: 234 FTLKKEH--QSFNMLRPEQEVQ-------IPSWLCQSSIDLSSNYSSLDQDLHLYENPNP 284
F+ ++ + N+ P ++ + IP WL +SSN NPN
Sbjct: 253 FSSSSQNIISNSNLHGPMKQEESQHHYQNIPPWL------ISSN-----------PNPNG 295
Query: 285 RNG----PTSTLPSYQPSS--AASPHMSATALLQKAAQMGATSSCSSQSMMSGTHQQGHV 338
NG P ++ + SS SP MSATALLQKAAQMG+T S + + +
Sbjct: 296 NNGNLFPPVASSVNTGRSSFPHPSPAMSATALLQKAAQMGSTKSTTPEEEERSSR----- 350
Query: 339 SIVDSATNNMINSNGNFSLNLSSCEDQMINNSFSSSGFHGTSFEDTFAGNILHSNQDHNI 398
S+ NN+I + + +S G F+D + N H
Sbjct: 351 ----SSYNNLITTT--------------MAAMMTSPPEPGFGFQDYYMMNHQHHGGGEAF 392
Query: 399 NHDGDNDIPKTTTNDDDVAAGGNNAFTRDFLGLKPL-SDSDILTIA-GMGSCMNPS-NSN 455
N +P ND GG TRDFLGL+ L S ++IL+ A +G+C+N S
Sbjct: 393 N---GGFVPGEEKNDVVDDGGGE---TRDFLGLRSLMSHNEILSFANNLGNCLNTSATEQ 446
Query: 456 HQENHSQK 463
Q+ HS +
Sbjct: 447 QQQQHSHQ 454
>At1g03840 putative DNA-binding protein
Length = 506
Score = 365 bits (936), Expect = e-101
Identities = 221/499 (44%), Positives = 288/499 (57%), Gaps = 89/499 (17%)
Query: 10 EASANSSGNRTHEVDAKFSQQYFASSQTQTHDETPAKKRRNLPGNPDPQAEVIALSPKTL 69
+ + +SSG + Q ++ + KK+RNLPGNPDP+AEVIALSPKTL
Sbjct: 5 DQTISSSGGYVQSSSTTDHVDHHHHDQHESLNPPLVKKKRNLPGNPDPEAEVIALSPKTL 64
Query: 70 MATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPEPTCVHHDPS 129
MATNRF+CEIC KGFQRDQNLQLH+RGHNLPWKLKQRTS E+RK+VYVCPE +CVHH P+
Sbjct: 65 MATNRFLCEICGKGFQRDQNLQLHRRGHNLPWKLKQRTSKEVRKRVYVCPEKSCVHHHPT 124
Query: 130 RALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRR 189
RALGDLTGIKKHFCRKHGEKKWKC+KC+K+YAVQSDWKAHSKTCGTREYRCDCGT+FSRR
Sbjct: 125 RALGDLTGIKKHFCRKHGEKKWKCEKCAKRYAVQSDWKAHSKTCGTREYRCDCGTIFSRR 184
Query: 190 DSFITHRAFCDALAEESSR-------------------------TVIPQPT--------- 215
DSFITHRAFCDALAEE++R T+IP P+
Sbjct: 185 DSFITHRAFCDALAEETARLNAASHLKSFAATAGSNLNYHYLMGTLIPSPSLPQPPSFPF 244
Query: 216 ---QPNSHHN------MNNLQTQDIQ--GFTLK-------KEHQSFNM-----LRPEQEV 252
QP HH+ NN QD+ TL HQ + +P
Sbjct: 245 GPPQPQHHHHHQFPITTNNFDHQDVMKPASTLSLWSGGNINHHQQVTIEDRMAPQPHSPQ 304
Query: 253 QIPSWLCQSS------IDLSSNYSSLDQDLHLYENPNPRNGPTS-TLPSY--------QP 297
+ +W+ ++ I S + + D ++++ ++ NG TS ++PS Q
Sbjct: 305 EDYNWVFGNANNHGELITTSDSLITHDNNINIVQSKENANGATSLSVPSLFSSVDQITQD 364
Query: 298 SSAAS---PHMSATALLQKAAQMGATSSCSSQSMMSGTHQQGHVSIVDSATNNMINSNGN 354
++AAS +MSATALLQKAAQMGATSS S + ++ T Q ++ S +N ++ G+
Sbjct: 365 ANAASVAVANMSATALLQKAAQMGATSSTSPTTTIT-TDQSAYLQSFASKSNQIVEDGGS 423
Query: 355 --FSLNLSSCEDQMINNSFSSSGFHGTSFEDTFAGNILHSNQDHNINHDGDNDIPKTTTN 412
F + S ++++N +++G H GN N ++ G+
Sbjct: 424 DRFFASFGSNSVELMSN--NNNGLHE-------IGN--PRNGVTVVSGMGELQNYPWKRR 472
Query: 413 DDDVAAGGNNAFTRDFLGL 431
D+ G TRDFLG+
Sbjct: 473 RVDIGNAGGGGQTRDFLGV 491
>At5g44160 unknown protein
Length = 466
Score = 350 bits (898), Expect = 1e-96
Identities = 210/458 (45%), Positives = 261/458 (56%), Gaps = 86/458 (18%)
Query: 34 SSQTQTHDETP-----AKKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQRDQ 88
SS T HDE+ KK+RNLPGNPDP+AEVIALSP TLMATNRF+CE+C KGFQRDQ
Sbjct: 20 SSSTLDHDESLINPPLVKKKRNLPGNPDPEAEVIALSPTTLMATNRFLCEVCGKGFQRDQ 79
Query: 89 NLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGE 148
NLQLH+RGHNLPWKLKQRTS E+RK+VYVCPE TCVHH SRALGDLTGIKKHFCRKHGE
Sbjct: 80 NLQLHRRGHNLPWKLKQRTSKEVRKRVYVCPEKTCVHHHSSRALGDLTGIKKHFCRKHGE 139
Query: 149 KKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALAEESSR 208
KKW C+KC+K+YAVQSDWKAHSKTCGTREYRCDCGT+FSRRDSFITHRAFCDALAEE+++
Sbjct: 140 KKWTCEKCAKRYAVQSDWKAHSKTCGTREYRCDCGTIFSRRDSFITHRAFCDALAEETAK 199
Query: 209 -----------------------------------TVIPQP-TQPNSHHNMNNLQTQDIQ 232
+PQP T PN HH T
Sbjct: 200 INAVSHLNGLAAAGAPGSVNLNYQYLMGTFIPPLQPFVPQPQTNPNHHHQHFQPPTSSSL 259
Query: 233 GFTLKKEHQSFNMLRPEQEVQIPSWLCQSSIDLSSNYSSLD-QDLHLYENPNPRNGPTST 291
+ ++ + P Q W+ ++ S+ + + D + +N N T+T
Sbjct: 260 SLWMGQD------IAPPQPQPDYDWVFGNAKAASACIDNNNTHDEQITQNANASLTTTTT 313
Query: 292 L------PSYQPSSA---ASPHMSATALLQKAAQMGATSSCSSQSMMSGTHQQGH-VSIV 341
L S QP +A ++ +MSATALLQKAA++GATS+ ++ + T Q +
Sbjct: 314 LSAPSLFSSDQPQNANANSNVNMSATALLQKAAEIGATSTTTAATNDPSTFLQSFPLKST 373
Query: 342 DSATN--------NMINSNGNFSLNLSSCEDQMINNSFSSSGFHGTSFEDTFAGNILHSN 393
D T+ + SN N L S + Q I N+ + D + L
Sbjct: 374 DQTTSYDSGEKFFALFGSNNNIGLMSRSHDHQEIENARN----------DVTVASALDEL 423
Query: 394 QDHNINHDGDNDIPKTTTNDDDVAAGGNNAFTRDFLGL 431
Q++ + +V GG TRDFLG+
Sbjct: 424 QNYPWKR-------RRVDGGGEVGGGGQ---TRDFLGV 451
>At3g50700 zinc finger protein
Length = 452
Score = 342 bits (877), Expect = 3e-94
Identities = 189/359 (52%), Positives = 231/359 (63%), Gaps = 50/359 (13%)
Query: 1 MSNLTSASGEASANSSGNRTHEVDAKFSQQYFASSQTQTHDETPAKKRRNLPGNPDPQAE 60
+ N ++ SGEAS + S +Q + T KK+RNLPG PDP++E
Sbjct: 5 LDNSSTVSGEASVSISST---------------GNQNPLPNST-GKKKRNLPGMPDPESE 48
Query: 61 VIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPE 120
VIALSPKTL+ATNRF+CEICNKGFQRDQNLQLH+RGHNLPWKL+Q+++ E++KKVYVCPE
Sbjct: 49 VIALSPKTLLATNRFVCEICNKGFQRDQNLQLHRRGHNLPWKLRQKSNKEVKKKVYVCPE 108
Query: 121 PTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRC 180
+CVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSK CGT+EY+C
Sbjct: 109 VSCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKICGTKEYKC 168
Query: 181 DCGTLFSRRDSFITHRAFCDALAEESSRTVIPQPTQ------------PNSHHNMNNLQT 228
DCGTLFSRRDSFITHRAFCDALAEE++R+ Q + PN + ++
Sbjct: 169 DCGTLFSRRDSFITHRAFCDALAEENARSHHSQSKKQNPEILTRKNPVPNPVPAPVDTES 228
Query: 229 QDIQGFTLKKEHQSFNMLRPEQEVQ---------------IPSWLCQSSIDLSSNYSSLD 273
I+ + QS + P + VQ + + L +SS S Y++
Sbjct: 229 AKIKSSSTLTIKQSESPKTPPEIVQEAPKPTSLNVVTSNGVFAGLFESSSASPSIYTTSS 288
Query: 274 QDLHLYENPNPRNGPTSTLPSYQPSS-------AASPHMSATALLQKAAQMGATSSCSS 325
L+ + + + L + SS A P MSATALLQKAAQMGA SS S
Sbjct: 289 SSKSLFASSSSIEPISLGLSTSHGSSFLGSNRFHAQPAMSATALLQKAAQMGAASSGGS 347
>At4g02670 putative zinc finger protein
Length = 402
Score = 335 bits (860), Expect = 2e-92
Identities = 198/375 (52%), Positives = 238/375 (62%), Gaps = 53/375 (14%)
Query: 4 LTSASGEASANSSGNRTHEVDAKFSQQYFASSQTQTHDET-PAKKRRNLPGNPDPQAEVI 62
L+S S EASA SSGN T +FS + S TH ET KK+R LPGNPDP AEVI
Sbjct: 11 LSSLSTEASA-SSGNNTLSTIQEFSGFHNVISSVCTHTETHKPKKKRGLPGNPDPDAEVI 69
Query: 63 ALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQR-TSNEIRKKVYVCPEP 121
ALSPKTL+ATNRF+CEICNKGFQRDQNLQLH+RGHNLPWKLKQ+ T + +KKVYVCPE
Sbjct: 70 ALSPKTLLATNRFVCEICNKGFQRDQNLQLHRRGHNLPWKLKQKNTKEQQKKKVYVCPET 129
Query: 122 TCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCD 181
C HH PSRALGDLTGIKKHFCRKHGEKKWKC+KCSK YAVQSDWKAH+K CGTR+YRCD
Sbjct: 130 NCAHHHPSRALGDLTGIKKHFCRKHGEKKWKCEKCSKFYAVQSDWKAHTKICGTRDYRCD 189
Query: 182 CGTLFSRRDSFITHRAFCDALAEESSRTVIPQPTQPNSHHNMNNLQTQDIQGFTLKKEHQ 241
CGTLFSR+D+FITHRAFCDALAEES+R S N+ N + QG H
Sbjct: 190 CGTLFSRKDTFITHRAFCDALAEESAR------LHSTSSSNLTN-PNPNFQG-----HHF 237
Query: 242 SFNMLRPEQEVQIPSWLCQSSIDLSSNYSSLDQDLHLYENPNPRNGPTSTLPSYQPSSAA 301
FN SS+ +S+ ++ L ST P +AA
Sbjct: 238 MFNK--------------SSSLLFTSSPLFIEPSL-------------STAALSTPPTAA 270
Query: 302 SPHMSATALLQKAAQMGATS-SCSSQSMMSGTHQQGHVSIVDSATNNMINSNGNFSLNLS 360
+SATALLQKA + +T+ Q+ G H+ H++ V N + + + S
Sbjct: 271 ---LSATALLQKATSLSSTTFGGGGQTRSIGHHR--HLTNV----NEFLGVDRVMMTSAS 321
Query: 361 SCE-DQMINNSFSSS 374
S E DQ++ + F+S+
Sbjct: 322 SSEYDQLVVDGFTST 336
>At3g45260 zinc finger like protein
Length = 446
Score = 335 bits (859), Expect = 3e-92
Identities = 185/337 (54%), Positives = 213/337 (62%), Gaps = 54/337 (16%)
Query: 45 AKKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLK 104
AK++RNLPGNPDP AEVIALSP +LM TNRFICE+CNKGF+RDQNLQLH+RGHNLPWKLK
Sbjct: 38 AKRKRNLPGNPDPDAEVIALSPNSLMTTNRFICEVCNKGFKRDQNLQLHRRGHNLPWKLK 97
Query: 105 QRTSNE-IRKKVYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQ 163
QRT+ E ++KKVY+CPE TCVHHDP+RALGDLTGIKKHF RKHGEKKWKCDKCSKKYAV
Sbjct: 98 QRTNKEQVKKKVYICPEKTCVHHDPARALGDLTGIKKHFSRKHGEKKWKCDKCSKKYAVM 157
Query: 164 SDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALAEESSR--TVIPQPTQPN--- 218
SDWKAHSK CGT+EYRCDCGTLFSR+DSFITHRAFCDALAEES+R +V P P N
Sbjct: 158 SDWKAHSKICGTKEYRCDCGTLFSRKDSFITHRAFCDALAEESARFVSVPPAPAYLNNAL 217
Query: 219 ----SHHNMN-NLQTQDIQGFTLKKEHQSFNMLRPE-----QEVQIPSWLCQSSIDLSSN 268
+H N+N N Q + + + + + FN R Q + + SS S
Sbjct: 218 DVEVNHGNINQNHQQRQLNTTSSQLDQPGFNTNRNNIAFLGQTLPTNVFASSSSPSPRSA 277
Query: 269 YSSLDQDLHLY----------ENPNPRNG----------------------------PTS 290
SL HL EN N N +
Sbjct: 278 SDSLQNLWHLQGQSSHQWLLNENNNNNNNILQRGISKNQEEHEMKNVISNGSLFSSEARN 337
Query: 291 TLPSYQPSSAASPHMSATALLQKAAQMGATSSCSSQS 327
+Y + MSATALLQKAAQMG+ S SS S
Sbjct: 338 NTNNYNQNGGQIASMSATALLQKAAQMGSKRSSSSSS 374
>At5g66730 zinc finger protein
Length = 500
Score = 330 bits (845), Expect = 1e-90
Identities = 152/208 (73%), Positives = 173/208 (83%), Gaps = 18/208 (8%)
Query: 1 MSNLTSASGEASANSSGNRTHEVDAKFSQQYFASSQTQTHDETPAKKRRNLPGNPDPQAE 60
+ N ++ SG+AS +S+GN+ ++ KK+RNLPG PDP AE
Sbjct: 5 LDNSSTVSGDASVSSTGNQN------------------LTPKSVGKKKRNLPGMPDPDAE 46
Query: 61 VIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPE 120
VIALSPKTLMATNRF+CEICNKGFQRDQNLQLH+RGHNLPWKL+QR++ E+RKKVYVCP
Sbjct: 47 VIALSPKTLMATNRFVCEICNKGFQRDQNLQLHRRGHNLPWKLRQRSTKEVRKKVYVCPV 106
Query: 121 PTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRC 180
CVHHDPSRALGDLTGIKKHFCRKHGEKKWKC+KCSKKYAVQSDWKAHSK CGT+EY+C
Sbjct: 107 SGCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCEKCSKKYAVQSDWKAHSKICGTKEYKC 166
Query: 181 DCGTLFSRRDSFITHRAFCDALAEESSR 208
DCGTLFSRRDSFITHRAFCDALAEES++
Sbjct: 167 DCGTLFSRRDSFITHRAFCDALAEESAK 194
>At2g02080 C2H2-type zinc finger protein
Length = 516
Score = 328 bits (841), Expect = 4e-90
Identities = 196/450 (43%), Positives = 255/450 (56%), Gaps = 77/450 (17%)
Query: 15 SSGNRTHEVDAKFSQQYFASSQTQTHDETPAKKRRNLPGNPDPQAEVIALSPKTLMATNR 74
SSG ++ + K + + Q P KKRRN PGNP+P AEV+ALSPKTLMATNR
Sbjct: 25 SSGTGDNDFNRK--DTFMSMIQQPNSSAPPPKKRRNQPGNPNPDAEVVALSPKTLMATNR 82
Query: 75 FICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPEPTCVHHDPSRALGD 134
FIC++CNKGFQR+QNLQLH+RGHNLPWKLKQ+++ E+++KVY+CPEPTCVHHDPSRALGD
Sbjct: 83 FICDVCNKGFQREQNLQLHRRGHNLPWKLKQKSTKEVKRKVYLCPEPTCVHHDPSRALGD 142
Query: 135 LTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFIT 194
LTGIKKH+ RKHGEKKWKC+KCSK+YAVQSDWKAHSKTCGT+EYRCDCGT+FSRRDS+IT
Sbjct: 143 LTGIKKHYYRKHGEKKWKCEKCSKRYAVQSDWKAHSKTCGTKEYRCDCGTIFSRRDSYIT 202
Query: 195 HRAFCDALAEESSRTVIPQPTQPN---------------------SHHNMN---NLQTQD 230
HRAFCDAL +E++R T SHH+++ N
Sbjct: 203 HRAFCDALIQETARNPTVSFTSMTAASSGVGSGGIYGRLGGGSALSHHHLSDHPNFGFNP 262
Query: 231 IQGFTLKKEHQSFNMLRPEQEVQIPSWLCQSSIDL--------SSNYSSLDQDLHLYENP 282
+ G+ L S N + P++L QS+ ++N S ++Q + +P
Sbjct: 263 LVGYNLNIA-SSDNRRDFIPQSSNPNFLIQSASSQGMLNTTPNNNNQSFMNQHGLIQFDP 321
Query: 283 -----------------------NPRNGPTSTLPSYQPSSA----------ASPHMSATA 309
N +N TS LPS + A ++SATA
Sbjct: 322 VDNINLKSSGTNNSFFNLGFFQENTKNSETS-LPSLYSTDVLVHHREENLNAGSNVSATA 380
Query: 310 LLQKAAQMGATSSCSSQSMMSGTHQQGHVSIVDSATNNMINSNGNFSLNLSSCEDQMINN 369
LLQKA QMG+ +S ++ G + S V N+ G N ++ Q + N
Sbjct: 381 LLQKATQMGSVTSNDPSALFRGLASSSNSSSV--IANHF--GGGRIMENDNNGNLQGLMN 436
Query: 370 SFSS----SGFHGTSFEDTFAGNILHSNQD 395
S ++ G G+ F+ F N S D
Sbjct: 437 SLAAVNGGGGSGGSIFDVQFGDNGNMSGSD 466
>At2g02070 putative C2H2-type zinc finger protein
Length = 602
Score = 322 bits (826), Expect = 2e-88
Identities = 193/420 (45%), Positives = 238/420 (55%), Gaps = 53/420 (12%)
Query: 44 PAKKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKL 103
P KK+RN P P+ AEVIALSPKTLMATNRFICE+CNKGFQR+QNLQLH+RGHNLPWKL
Sbjct: 50 PQKKKRNQPRTPNSDAEVIALSPKTLMATNRFICEVCNKGFQREQNLQLHRRGHNLPWKL 109
Query: 104 KQRTSNEIRKKVYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQ 163
KQ+++ E+++KVY+CPEP+CVHHDPSRALGDLTGIKKH+ RKHGEKKWKCDKCSK+YAVQ
Sbjct: 110 KQKSTKEVKRKVYLCPEPSCVHHDPSRALGDLTGIKKHYYRKHGEKKWKCDKCSKRYAVQ 169
Query: 164 SDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALAEESSRTVIPQPTQPNSH--- 220
SDWKAHSKTCGT+EYRCDCGTLFSRRDSFITHRAFCDALA+ES+R + P+ H
Sbjct: 170 SDWKAHSKTCGTKEYRCDCGTLFSRRDSFITHRAFCDALAQESARHPTSLTSLPSHHFPY 229
Query: 221 ----HNMNNLQTQDIQGFT-----LKKEHQSFNMLRPEQEVQIPSWLCQSSIDL------ 265
+N NN + I G + +HQ ++LR +SS DL
Sbjct: 230 GQNTNNSNNNASSMILGLSHMGAPQNLDHQPGDVLRLGSGGGGGGAASRSSSDLIAANAS 289
Query: 266 -----SSNYSSLDQDLH--------LYENPNPRNGPTS---TLPSYQPSSAASPHMSATA 309
N S DQ H L N N + P S L + + S +
Sbjct: 290 GYFMQEQNPSFHDQQDHHHHHQQGFLAGNNNIKQSPMSFQQNLMQFSHDNHNSAPSNVFN 349
Query: 310 LLQKAAQMGATS-----------SCSSQSMMSGTHQQGHVSIVDSATNNMINSNGNFSLN 358
L + G TS + SS ++M H G ++ S G F N
Sbjct: 350 LSFLSGNNGVTSATSNPNAAAAAAVSSGNLMISNHYDGENAVGGGGE----GSTGLFPNN 405
Query: 359 LSSCEDQMINNS----FSSSGFHGTSFEDTFAGNILHSNQDHNINHDGDNDIPKTTTNDD 414
L S D++ + S FSSS S A +L +N+ T N++
Sbjct: 406 LMSSADRISSGSVPSLFSSSMQSPNSAPHMSATALLQKAAQMGSTSSNNNNGSNTNNNNN 465
>At1g14580 putative zinc finger protein
Length = 467
Score = 322 bits (826), Expect = 2e-88
Identities = 198/478 (41%), Positives = 268/478 (55%), Gaps = 82/478 (17%)
Query: 27 FSQQYFASSQTQTHDET----PAKKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNK 82
F++Q A + Q + P KKRRN PGNP+P AEVIALSPKT+MATNRF+CE+CNK
Sbjct: 30 FNRQEAAMTMVQQQPTSSVAPPPKKRRNQPGNPNPDAEVIALSPKTIMATNRFLCEVCNK 89
Query: 83 GFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPEPTCVHHDPSRALGDLTGIKKHF 142
GFQR+QNLQLH+RGHNLPWKLKQ+++ E+R+KVY+CPEP+CVHHDP+RALGDLTGIKKH+
Sbjct: 90 GFQREQNLQLHRRGHNLPWKLKQKSNKEVRRKVYLCPEPSCVHHDPARALGDLTGIKKHY 149
Query: 143 CRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDAL 202
RKHGEKKWKCDKCSK+YAVQSDWKAHSKTCGT+EYRCDCGT+FSRRDS+ITHRAFCDAL
Sbjct: 150 YRKHGEKKWKCDKCSKRYAVQSDWKAHSKTCGTKEYRCDCGTIFSRRDSYITHRAFCDAL 209
Query: 203 AEESSR------TVIPQPTQPNSHHNMNNLQTQDIQGFTLKKEHQSFNMLRPEQEVQIPS 256
+ES+R T + + H G + H F P +
Sbjct: 210 IQESARNPTVSFTAMAAGGGGGARHGFYG-------GASSALSHNHFGN-NPNSGF---T 258
Query: 257 WLCQSSIDLSSNYSSLDQDLHLYENPNPRNGPTSTLPSYQPSSAASPHMSATALLQKAAQ 316
L + +L+ + S +D + + NP GPT+ L P+ LL +
Sbjct: 259 PLAAAGYNLNRSSSDKFEDF-VPQATNPNPGPTNFLMQCSPNQ---------GLLAQ--- 305
Query: 317 MGATSSCSSQSMMSGTHQQGHVSIVDSATNNMINSNGNFSLNLSSCEDQMINN-----SF 371
++QS+M+ H ++ NN N+N NF NL+ +D ++ S
Sbjct: 306 -------NNQSLMN------HHGLISLGDNN--NNNHNF-FNLAYFQDTKNSDQTGVPSL 349
Query: 372 SSSGFHGTSFEDTFAGNILHSNQDHNINHDGD----------NDIPKTTTND-------- 413
++G G S+ +N GD N + TT
Sbjct: 350 FTNGADNNGPSALLRGLTSSSSSSVVVNDFGDCDHGNLQGLMNSLAATTDQQGRSPSLFD 409
Query: 414 ----DDVAAGGNNAFTRDFLGLKPLSDSDILTIAGMG--SCMNPSNSNHQENHSQKPW 465
++++ GG++ T DFLG ++ + T+ G G S P ++ + +H P+
Sbjct: 410 LHFANNLSMGGSDRLTLDFLG---VNGGIVSTVNGRGGRSGGPPLDAEMKFSHPNHPY 464
>At5g03150 putative protein
Length = 501
Score = 312 bits (800), Expect = 2e-85
Identities = 176/342 (51%), Positives = 205/342 (59%), Gaps = 59/342 (17%)
Query: 41 DETPAKKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLP 100
+ + AKK+RN PG PD A+VIALSP TLMATNRF+CEICNKGFQRDQNLQLH+RGHNLP
Sbjct: 48 NSSSAKKKRNQPGTPD--ADVIALSPTTLMATNRFVCEICNKGFQRDQNLQLHRRGHNLP 105
Query: 101 WKLKQRTSNE-IRKKVYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKK 159
WKLKQR+ E I+KKVY+CP TCVHHD SRALGDLTGIKKH+ RKHGEKKWKC+KCSKK
Sbjct: 106 WKLKQRSKQEVIKKKVYICPIKTCVHHDASRALGDLTGIKKHYSRKHGEKKWKCEKCSKK 165
Query: 160 YAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALAEESSR----------- 208
YAVQSDWKAH+KTCGTREY+CDCGTLFSR+DSFITHRAFCDAL EE +R
Sbjct: 166 YAVQSDWKAHAKTCGTREYKCDCGTLFSRKDSFITHRAFCDALTEEGARMSSLSNNNPVI 225
Query: 209 -----------TVIPQPTQPNS-------HHNMN------------NLQTQDIQGFT--- 235
V+ P P+ H ++N +L QG +
Sbjct: 226 STTNLNFGNESNVMNNPNLPHGFVHRGVHHPDINAAISQFGLGFGHDLSAMHAQGLSEMV 285
Query: 236 ----------LKKEHQSFNMLRPEQEVQIPSWLCQSSIDLSSNYSSLDQDLHLYEN--PN 283
S + QIP S+ LSS+ +S L +
Sbjct: 286 QMASTGNHHLFPSSSSSLPDFSGHHQFQIPMTSTNPSLTLSSSSTSQQTSASLQHQTLKD 345
Query: 284 PRNGPTSTLPSYQPSSAASPHMSATALLQKAAQMGATSSCSS 325
P + S + MSATALLQKAAQMG+T S SS
Sbjct: 346 SSFSPLFSSSSENKQNKPLSPMSATALLQKAAQMGSTRSNSS 387
>At5g60470 putative zinc finger protein
Length = 392
Score = 288 bits (736), Expect = 6e-78
Identities = 162/340 (47%), Positives = 202/340 (58%), Gaps = 64/340 (18%)
Query: 70 MATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTS-NEIRKKVYVCPEPTCVHHDP 128
MATNRF CEICNKGFQR+QNLQLHKRGHNLPWKLKQ+T+ N+++KKVY+CPE +CVHHDP
Sbjct: 1 MATNRFFCEICNKGFQREQNLQLHKRGHNLPWKLKQKTNKNQVKKKVYICPEKSCVHHDP 60
Query: 129 SRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSR 188
+RALGDLTGIKKHF RKHGEKKWKCDKCSKKYAV SDWKAH+K CG+RE+RCDCGTLFSR
Sbjct: 61 ARALGDLTGIKKHFSRKHGEKKWKCDKCSKKYAVISDWKAHNKICGSREFRCDCGTLFSR 120
Query: 189 RDSFITHRAFCDALAEESSRTV-IPQPTQPNS--------------------------HH 221
+DSFI+HR+FCD LAEESS+ +P P NS
Sbjct: 121 KDSFISHRSFCDVLAEESSKFFSVPSPLAANSTIATVTDTNNPILIQSQLDQSSTGTADL 180
Query: 222 NMNNLQTQDI-QGFTLKKEHQS--------------------FNMLRPEQEVQIPSWLCQ 260
N+NN T Q FT Q N+ + ++E WL
Sbjct: 181 NVNNNHTTLFGQKFTNSNPTQQQPNALALSSPPSPRSTSDSVHNLWKLQEEECAHQWLLN 240
Query: 261 SSIDLSSN------YSSLDQDL---HLYENPNPRNGPTSTLPSYQPSSAASPHMSATALL 311
++ + N + + + ++ ++Y NP +G ++L SY + SAT LL
Sbjct: 241 EYMNNNKNIFHKGIFKNQEDEIKKGNIYSGSNPTDGNIASLFSYNQEAVNMASFSATTLL 300
Query: 312 QKAAQMGATSSCSSQSMMSGTHQQGHVSIVDSATNNMINS 351
QK AQ G SS + + M G Q SI + N M+NS
Sbjct: 301 QKVAQTGTPSSSETSTTMFG---QMTSSIFN---NTMLNS 334
>At2g01940 putative C2H2-type zinc finger protein
Length = 439
Score = 236 bits (601), Expect = 3e-62
Identities = 115/210 (54%), Positives = 152/210 (71%), Gaps = 12/210 (5%)
Query: 13 ANSSGNRTHEVDAKFSQQYFASSQ---TQTHDETPAKKRRNLPGNPDPQAEVIALSPKTL 69
+N + N V + S + +SS+ T T+ T +KRR G PDP AEV++LSP+TL
Sbjct: 3 SNKNTNTCCVVSSSSSDPFLSSSENGVTTTNTSTQKRKRRPA-GTPDPDAEVVSLSPRTL 61
Query: 70 MATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSN-EIRKKVYVCPEPTCVHHDP 128
+ ++R+ICEICN+GFQRDQNLQ+H+R H +PWKL +R +N E++K+VYVCPEPTC+HH+P
Sbjct: 62 LESDRYICEICNQGFQRDQNLQMHRRRHKVPWKLLKRDNNIEVKKRVYVCPEPTCLHHNP 121
Query: 129 SRALGDLTGIKKHFCRKH-GEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFS 187
ALGDL GIKKHF RKH K+W C++CSK YAVQSD+KAH KTCGTR + CDCG +FS
Sbjct: 122 CHALGDLVGIKKHFRRKHSNHKQWVCERCSKGYAVQSDYKAHLKTCGTRGHSCDCGRVFS 181
Query: 188 RRDSFITHRAFCDALAEESSRTVIPQPTQP 217
R +SFI H+ C S+R V +P +P
Sbjct: 182 RVESFIEHQDNC------SARRVHREPPRP 205
>At1g68130 putative C2H2-type zinc finger protein
Length = 419
Score = 233 bits (593), Expect = 2e-61
Identities = 123/254 (48%), Positives = 164/254 (64%), Gaps = 25/254 (9%)
Query: 46 KKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWK-LK 104
K++R G PDP+AEV++LSP+TL+ ++R++CEICN+GFQRDQNLQ+H+R H +PWK LK
Sbjct: 41 KRKRRPAGTPDPEAEVVSLSPRTLLESDRYVCEICNQGFQRDQNLQMHRRRHKVPWKLLK 100
Query: 105 QRTSNEIRKKVYVCPEPTCVHHDPSRALGDLTGIKKHFCRKH-GEKKWKCDKCSKKYAVQ 163
+ T+ E+RK+VYVCPEPTC+HH+P ALGDL GIKKHF RKH K+W C++CSK YAVQ
Sbjct: 101 RETNEEVRKRVYVCPEPTCLHHNPCHALGDLVGIKKHFRRKHSNHKQWICERCSKGYAVQ 160
Query: 164 SDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDA---------LAEESSRTVIPQP 214
SD+KAH KTCGTR + CDCG +FSR +SFI H+ C L E+ T
Sbjct: 161 SDYKAHLKTCGTRGHSCDCGRVFSRVESFIEHQDTCTVRRSQPSNHRLHEQQQHTTNATQ 220
Query: 215 TQPNSHHNMN-NLQTQDI-QGFTLKK------EHQSFNMLRP-----EQEVQ-IPSWLCQ 260
T + +N N +L I G L++ E Q +L P E+Q +PS C
Sbjct: 221 TASTAENNENGDLSIGPILPGHPLQRRQSPPSEQQPSTLLYPFVTNGSIELQLLPSRNCA 280
Query: 261 SSIDLSSNYSSLDQ 274
LS + ++DQ
Sbjct: 281 DETSLSLSIGTMDQ 294
>At1g25250 Indeterminate 1 like Zn finger containing protein
Length = 385
Score = 228 bits (580), Expect = 7e-60
Identities = 104/181 (57%), Positives = 135/181 (74%), Gaps = 14/181 (7%)
Query: 46 KKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKL-- 103
K++R G PDP AEV++LSP+TL+ ++R++CEICN+GFQRDQNLQ+H+R H +PWKL
Sbjct: 33 KRKRRPAGTPDPDAEVVSLSPRTLLESDRYVCEICNQGFQRDQNLQMHRRRHKVPWKLLK 92
Query: 104 KQRTSNEIRKKVYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHG-EKKWKCDKCSKKYAV 162
+ + E+RK+VYVCPEPTC+HHDP ALGDL GIKKHF RKH K+W C++CSK YAV
Sbjct: 93 RDKKDEEVRKRVYVCPEPTCLHHDPCHALGDLVGIKKHFRRKHSVHKQWVCERCSKGYAV 152
Query: 163 QSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALAEESSRTVIPQPTQPNSHHN 222
QSD+KAH KTCG+R + CDCG +FSR +SFI H+ C I QP QP +H +
Sbjct: 153 QSDYKAHLKTCGSRGHSCDCGRVFSRVESFIEHQDTC----------TIRQP-QPTNHRH 201
Query: 223 M 223
+
Sbjct: 202 L 202
>At1g34370 zinc finger protein, putative
Length = 499
Score = 106 bits (265), Expect = 2e-23
Identities = 67/214 (31%), Positives = 100/214 (46%), Gaps = 23/214 (10%)
Query: 41 DETPAKKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLP 100
DE ++ NLP E++ L + ++A + C IC KGF+RD NL++H RGH
Sbjct: 213 DEDDVEEGENLPPG---SYEILQLEKEEILAPHTHFCTICGKGFKRDANLRMHMRGHGDE 269
Query: 101 WKLKQRTSNEIRKKV----------YVCPEPTCVH---HDPSRALGDLTGIKKHFCRKHG 147
+K + ++ V Y CP C H + L + +K H+ R H
Sbjct: 270 YKTAAALAKPNKESVPGSEPMLIKRYSCPFLGCKRNKEHKKFQPLKTILCVKNHYKRTHC 329
Query: 148 EKKWKCDKC-SKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAF----CDAL 202
+K + C +C +KK++V +D K H K CG ++ C CGT FSR+D H A A+
Sbjct: 330 DKSFTCSRCHTKKFSVIADLKTHEKHCGKNKWLCSCGTTFSRKDKLFGHIALFQGHTPAI 389
Query: 203 AEESSRTVIPQPTQPNSHHNMNNLQTQDIQGFTL 236
E ++ TQ S NN Q + GF L
Sbjct: 390 PLEETKPSASTSTQRGSSEGGNN--NQGMVGFNL 421
>At5g22890 unknown protein
Length = 235
Score = 97.4 bits (241), Expect = 1e-20
Identities = 51/150 (34%), Positives = 80/150 (53%), Gaps = 14/150 (9%)
Query: 60 EVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQR----TSNE----- 110
+++ L L+A C+IC KGF+RD NL++H R H +K ++ TS +
Sbjct: 64 DILELDVADLLAKYTHYCQICGKGFKRDANLRMHMRAHGDEYKTREALISPTSQDKKGGY 123
Query: 111 -IRKKVYVCPEPTC---VHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCS-KKYAVQSD 165
++K Y CP+ C H+ + L + K H+ R H K + C +CS K ++V SD
Sbjct: 124 SLKKHYYSCPQHGCRWNQRHEKFQPLKSVICAKNHYKRSHCPKMYMCRRCSVKHFSVLSD 183
Query: 166 WKAHSKTCGTREYRCDCGTLFSRRDSFITH 195
+ H K CG ++ C CGT FSR+D ++H
Sbjct: 184 LRTHEKHCGDIKWVCSCGTKFSRKDKLMSH 213
>At1g13290 hypothetical protein
Length = 302
Score = 90.5 bits (223), Expect = 2e-18
Identities = 51/146 (34%), Positives = 76/146 (51%), Gaps = 12/146 (8%)
Query: 69 LMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKL-------KQRTSNEIRKKVYVCPE- 120
L+ +F C +CNK F R N+Q+H GH ++ + +S+ +R Y C E
Sbjct: 95 LVGPTQFSCSVCNKTFNRFNNMQMHMWGHGSQYRKGPESLRGTKSSSSILRLPCYCCAEG 154
Query: 121 -PTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKC-DKCSKKYAVQSDWKAHSKTCGTREY 178
+ H S+ L D ++ H+ RKHG K ++C KC K +AV+ DW+ H K CG + +
Sbjct: 155 CKNNIDHPRSKPLKDFRTLQTHYKRKHGAKPFRCRKKCEKTFAVRGDWRTHEKNCG-KLW 213
Query: 179 RCDCGTLFSRRDSFITH-RAFCDALA 203
C CG+ F + S H RAF D A
Sbjct: 214 FCVCGSDFKHKRSLKDHVRAFGDGHA 239
>At3g20880 putative DNA-binding protein
Length = 348
Score = 87.8 bits (216), Expect = 1e-17
Identities = 46/135 (34%), Positives = 65/135 (48%), Gaps = 9/135 (6%)
Query: 69 LMATNRFICEICNKGFQRDQNLQLHKRGHNL-----PWKLKQRTSNEIRKKVYVCPEPTC 123
LM +F C +C K F R N+Q+H GH P L+ + K C P C
Sbjct: 186 LMGPTQFSCPLCFKTFNRYNNMQMHMWGHGSQYRKGPESLRGTQPTAMLKLPCYCCAPGC 245
Query: 124 ---VHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRC 180
+ H +R L D ++ H+ RKHG + + C +C K +AV+ DW+ H K CG Y C
Sbjct: 246 KNNIDHPRARPLKDFRTLQTHYKRKHGVRPFACRRCGKAFAVKGDWRTHEKNCGKLWY-C 304
Query: 181 DCGTLFSRRDSFITH 195
CG+ F + S H
Sbjct: 305 SCGSDFKHKRSLKDH 319
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.126 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,727,451
Number of Sequences: 26719
Number of extensions: 542070
Number of successful extensions: 1654
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 34
Number of HSP's successfully gapped in prelim test: 32
Number of HSP's that attempted gapping in prelim test: 1548
Number of HSP's gapped (non-prelim): 96
length of query: 468
length of database: 11,318,596
effective HSP length: 103
effective length of query: 365
effective length of database: 8,566,539
effective search space: 3126786735
effective search space used: 3126786735
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148918.4


