

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148819.4 - phase: 0
(1009 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
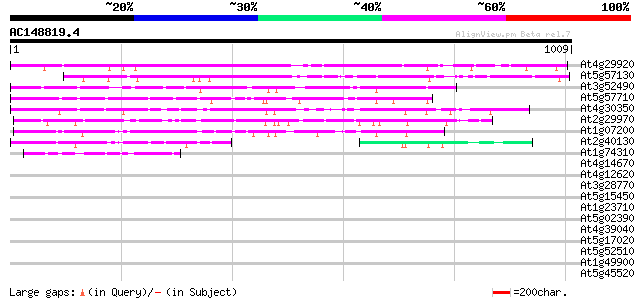
Score E
Sequences producing significant alignments: (bits) Value
At4g29920 putative protein 684 0.0
At5g57130 unknown protein 532 e-151
At3g52490 putative protein 361 1e-99
At5g57710 101 kDa heat shock protein; HSP101-like protein 334 2e-91
At4g30350 unknown protein 328 9e-90
At2g29970 unknown protein 207 2e-53
At1g07200 unknown protein 180 4e-45
At2g40130 unknown protein 161 2e-39
At1g74310 heat shock protein 101 78 2e-14
At4g14670 heat shock protein like 42 0.001
At4g12620 origin recognition complex subunit 1 -like protein 40 0.009
At3g28770 hypothetical protein 39 0.019
At5g15450 clpB heat shock protein-like 38 0.033
At1g23710 unknown protein 37 0.056
At5g02390 putative protein 36 0.13
At4g39040 unknown protein 35 0.21
At5g17020 Exportin1 (XPO1) protein 35 0.28
At5g52510 scarecrow-like 8 (SCL8) 34 0.48
At1g49900 Cys2/His2-type zinc finger protein, putative 34 0.48
At5g45520 putative protein 33 0.62
>At4g29920 putative protein
Length = 1017
Score = 684 bits (1764), Expect = 0.0
Identities = 452/1078 (41%), Positives = 613/1078 (55%), Gaps = 140/1078 (12%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSL-RNACLKS 59
MR+GAYT+ +TLT EAAS+LK SL LARRRGH+Q+TPLHVA+TLL+ S+L R ACLKS
Sbjct: 1 MRTGAYTVHQTLTPEAASVLKQSLTLARRRGHSQVTPLHVASTLLTSSRSNLFRRACLKS 60
Query: 60 Q------QQNNHP-LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQ 112
+Q HP L CRALELCFNV+LNRLPT +PL Q Q PSLSNAL+AALKRAQ
Sbjct: 61 NPFTALGRQMAHPSLHCRALELCFNVSLNRLPTNP-NPLFQTQ--PSLSNALVAALKRAQ 117
Query: 113 AHQRRGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENS 172
AHQRRGC+EQ Q QQ QP L+VKVEL+QL++SILDDPSVSRVMREAG SS SVK+N+E+
Sbjct: 118 AHQRRGCVEQQQSQQNQPFLAVKVELEQLVVSILDDPSVSRVMREAGLSSVSVKSNIEDD 177
Query: 173 STLIN-----SSSVFHSSPSPLSHNHFLSSYGYGSV-----------LFSSQKKEQVVYH 216
S++++ SSS SP S + ++ G G++ L + EQ +
Sbjct: 178 SSVVSPVFYGSSSSVGVFSSPCSPSSSENNQGGGTLSPNPSKIWHAHLTNHHSFEQNPFF 237
Query: 217 PFLKSSESN-------KEDINLVFDVLLRK---KKKNTVIVGDTVSLTEGLVSEIMKRFE 266
F K +ED N V +VLL K KK+NTVIVGD+VSLTEG+V+++M R E
Sbjct: 238 HFPKGKTFTPDQAFPVREDANPVIEVLLGKKNNKKRNTVIVGDSVSLTEGVVAKLMGRIE 297
Query: 267 RGEVPDEMKTTHFVKFHGLSSVSLKYMKKEEVEMNVIRVLKRKVSDYVAL-GVGAIFYVG 325
RGEVPD++K THF+KF S V L +MKKE++E +R LKRK+ + + G G I +G
Sbjct: 298 RGEVPDDLKQTHFIKFQ-FSQVGLNFMKKEDIE-GQVRELKRKIDSFTSWGGKGVIVCLG 355
Query: 326 DLKWIV--DDNDGSLNEKEVVDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMRV 383
DL W V N S + D++VEEIG+L + N K+WL+ TASYQ+YMRCQM+
Sbjct: 356 DLDWAVWGGGNSASSSNYSAADHLVEEIGRLVYDYSNTGAKVWLLGTASYQTYMRCQMKQ 415
Query: 384 PTFENQWCLQAVPVPSGGLGLSLHSSSVHDSKMSISQNPSPMLESKFFSNKEEHE-KLNC 442
P + W LQAV +PSGGL L+LH+SS + + P + E + + +EE E KLN
Sbjct: 416 PPLDVHWALQAVSIPSGGLSLTLHASSSEMASQVMEMKPFRVKEEEEGAREEEEEDKLNF 475
Query: 443 CEECVSNYEKEAQLFKPDQKNLLPSWLQSH--STEARQKDELTQLNKKWNRLCQCLHQNK 500
C EC NYEKEA+ F Q +LP WLQ H + QKDEL+ L KKWNR CQ LH K
Sbjct: 476 CGECAFNYEKEAKAFISAQHKILPPWLQPHGDNNNINQKDELSGLRKKWNRFCQALHHKK 535
Query: 501 QPQNHWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKP-AYSSNLIPR 559
W Q SS+LP S DS+ K + +S+ + +
Sbjct: 536 PSMTAWR-------------------AEQSSSVLPG-----SLMDSSLKQNSRASSSVAK 571
Query: 560 FRRGQQSCTIEFNFNDEKAQKNQVTATLELDSLKGM--EGTKEVKTTLALGNSTF---SV 614
FRR Q SCTIEF+F + + + T L LD K EG K K TLALG+S F S
Sbjct: 572 FRR-QNSCTIEFSFGSNRQEGLKKTDELSLDGFKSNNDEGVK-TKITLALGHSPFPSDSE 629
Query: 615 SDQKRMENLTLQRDHIYKVLQENIPWHCETVSSIAEALVDS-KSSKECATWLFLQGNDSV 673
+ ++ ++ + + L ENIPW + + SI EA+ +S K SK W+ + GND
Sbjct: 630 NSEEEEPEKAIKMSKLLEKLHENIPWQKDVLPSIVEAMEESVKRSKRKDAWMLVSGNDVT 689
Query: 674 GKKRLALAIAESVFGSVEMFSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVENADFGD 733
K+RLA+ + S+FGS E +++ + SE E++ LK E+ V+L+E D D
Sbjct: 690 AKRRLAITLTTSLFGSHENMLKINLRTSKASEA--CEELKNALKKKEEVVILIERVDLAD 747
Query: 734 TLIRKMLADEFEIAKF----GTLGQKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKP 789
+L D FE G Q IF+L+ E++ V+ +VL
Sbjct: 748 AQFMNILVDRFEAGDLDGFQGKKSQIIFLLTREDDECVENE--HFVIPMVL--------- 796
Query: 790 TFELSPSSSSSSKSPCLGNKRSAELD---LFSKIKIPRIEENE---------GNKKREFS 837
+ + S S + NKR E D K K PRIEE++ N K+E
Sbjct: 797 -------NCNKSGSGLVNNKRKPEYDAAPTMIKKKNPRIEEDDDESNVACDISNIKKE-- 847
Query: 838 FSRQSSF-NNTLDLNMKADEEDNEDYDEGENSPISSDLTRETLGEHLISNESLDSIENLF 896
FSRQ F +N LDLN++ D +++E+ + + ISS LDSI+N F
Sbjct: 848 FSRQLKFESNALDLNLRVDADEDEEEEAKPATEISSG----------FEERFLDSIQNRF 897
Query: 897 EFNQSPAKNKEMTQMFMSRVKESFEEVLGNVK----FSVQDKVIEEIGVGCGSFTNNMFE 952
+F + ++++T+ F++++K+S EE+LG + F+V ++IE+ GCG F N +FE
Sbjct: 898 DF--TVLSDEDITKFFVTKIKDSCEEILGQREERFGFTVDAELIEKFYKGCGFFANGLFE 955
Query: 953 KWLKGIFQTSLERVNGGDKNGI-VYTLCWGG------KEDRKWDSGFMGSCLPKNIQI 1003
+W+K +FQ L V G K GI V LC GG E + + GFMG+CLP I +
Sbjct: 956 EWVKEVFQRGLVTVKNGGKEGISVINLCLGGIDMIDQGEVYEEEEGFMGTCLPNRIHV 1013
>At5g57130 unknown protein
Length = 920
Score = 532 bits (1370), Expect = e-151
Identities = 393/989 (39%), Positives = 542/989 (54%), Gaps = 155/989 (15%)
Query: 98 PSLSNALIAALKRAQAHQRRGCIEQNQQQQQQP------LLSVKVELDQLILSILDDPSV 151
PSL+NAL+AALKRAQAHQRRGCIEQ QQ Q P LL+VKVEL+QL++SILDDPSV
Sbjct: 6 PSLANALVAALKRAQAHQRRGCIEQQQQTQTHPQTQQTQLLAVKVELEQLVISILDDPSV 65
Query: 152 SRVMREAGFSSPSVKNNLENSSTLI------------------------NSSSVFHSSPS 187
SRVMREAGF+S +VK+ +E+ S NS + H +
Sbjct: 66 SRVMREAGFNSTAVKSCVEDCSVSSVFYGGSAVGVFSSPNSPDQQQQHHNSINRLHHYQN 125
Query: 188 PLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKE---------DINLVFDVLLR 238
P N ++ F +Q +Q +P L SS ++ D+ LV DVL+R
Sbjct: 126 PKDFNFINPNFPLWQTHFLNQSPDQ---NPLLLSSSASHHHQQQRLREIDLKLVVDVLMR 182
Query: 239 K--KKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPD--EMKTTHFVKFHGLSSVSLKYMK 294
K KKKN VIVGD++S TEG VSE+M + ERGE+ E+K THFVKFH S ++ K+M+
Sbjct: 183 KKTKKKNPVIVGDSISFTEGFVSELMAKLERGEIDQTGELKQTHFVKFH-FSPMASKFMR 241
Query: 295 KEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKW----IVDDNDGSLNE----KEVVDY 346
+E+VE+N I+ L++KV G AI + GDLKW I ++N G +NE +D+
Sbjct: 242 REDVELN-IKELRKKVLSLTTSGKNAIIFTGDLKWTVKEITNNNSGGINEISSSYSPLDH 300
Query: 347 VVEEIGKLFGE-------EGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVP- 398
+VEEIGKL E + K K+W++ TAS+Q+YMRCQMR P+ E W L V VP
Sbjct: 301 LVEEIGKLITECNDDGDDDDCKTRKVWVMGTASFQTYMRCQMRQPSLETLWALHPVSVPS 360
Query: 399 SGGLGLSLHSSSVHDSKMSISQNPSPMLESKFFSNKEE--HEKLNCCEECVSNYEKEAQL 456
S LGLSLH++S H+++ + N + L + +EE L+CC ECV+++++EA+
Sbjct: 361 SANLGLSLHATSGHEARNMSTVNATKSLSGYDKAEEEETISHVLSCCPECVTSFDREAKS 420
Query: 457 FKPDQKNLLPSWLQSHSTE-ARQKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAK 515
K +Q LLPSWLQSH + + QKDEL L +KWNR C+ LH + N
Sbjct: 421 LKANQDKLLPSWLQSHDADSSSQKDELMGLKRKWNRFCETLHNQTGQLSMMGN------- 473
Query: 516 IYPYNSSYPYWPNQGSSILPDTSSSISFADS-ATKP-AYSSNLIPRFRRGQQSCTIEFNF 573
YPY Y GSS ++S S S DS KP ++N I +FRR Q SCTIEF+
Sbjct: 474 -YPYGLPY------GSS--HESSKSTSLIDSLGLKPNQRATNSIAKFRR-QNSCTIEFDL 523
Query: 574 NDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTF---SVSDQKRMENLTLQRDHI 630
+ +K + E D KG E TL LG S F SV+D + L+ +
Sbjct: 524 GGNEHEKGESINEAEDD--KGNE-----TVTLDLGRSLFRSDSVTDTR------LKLSAL 570
Query: 631 YKVLQENIPWHCETVSSIAEALVDSKSSKECATWLFLQGNDSVGKKRLALAIAESVFGSV 690
K L+E+IP T+ IAE+L+D S K+ +W+ ++G D+ K+R+A ++ESVFGS
Sbjct: 571 VKALEESIPRQTVTMRLIAESLMDCVSKKK-DSWIIIEGRDTTAKRRVARTVSESVFGSF 629
Query: 691 EMFSHVDMMKREN-SETPFSEKVVGPLKNNEKFVVLVENADFGDTLIRKMLADEFEIAKF 749
E H+D+ K+ N S+ + + LKN EK V L+E+ D D+ K+LAD FE +
Sbjct: 630 ESLVHIDLKKKGNESKASPATLLAYELKNPEKVVFLIEDIDLADSRFLKLLADRFEDKRR 689
Query: 750 GTLG----QKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKPTFELSPSSSSSSKSPC 805
G Q IFIL+ S + +DSV+++ L+I +++SP
Sbjct: 690 IKTGIDHRQAIFILTKEDS--RNVRNRDSVLQIGLEI-----------------TAQSP- 729
Query: 806 LGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQSSFNNT-LDLNMKADEEDNEDYDE 864
G KR E DL IE KK SRQSSFN++ LDLN+KA++E+ E
Sbjct: 730 -GKKRKPESDL-------SIENGFWMKKE--VCSRQSSFNSSYLDLNIKAEDEE----VE 775
Query: 865 GENSPISSDLTRETLGEHLISNESLDSIENLFEFNQS--PAKNKEMTQMFMSRVKESFEE 922
GE SPISSDLT E E S+ L+ I+N F N+S P K M + EE
Sbjct: 776 GEISPISSDLTGEEETEFSSSSNFLNRIQNRFVLNRSCEPGIEKGMITAAFREIFPEREE 835
Query: 923 VLGNVKFSVQDKVIEEIGVGCGSFTNNMFEKWLKGIFQTSLERV-NGGDKNGIVYTLCWG 981
G V+FSV+DK++EE+ N FE+WLK +FQT L V GG K+ V + +G
Sbjct: 836 G-GGVRFSVEDKLVEEL----YGIQNGAFERWLKEVFQTGLLTVKKGGKKDTGVIRMVFG 890
Query: 982 GKEDRK----WDSGFMGSCLPKNIQIVNY 1006
G D K G+M + LP +Q+ +
Sbjct: 891 GIVDNKGYGGGVGGYMATFLPNKVQVSKF 919
>At3g52490 putative protein
Length = 815
Score = 361 bits (926), Expect = 1e-99
Identities = 266/833 (31%), Positives = 417/833 (49%), Gaps = 102/833 (12%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+G T+++ LTA+AA+++K ++GLARRRGHAQ+TPLHVA+T+LS P+ LR ACL
Sbjct: 1 MRAGGCTVEQALTADAANVVKQAMGLARRRGHAQVTPLHVASTMLSAPTGLLRTACL--- 57
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTTTSPLL--QPQHVPSLSNALIAALKRAQAHQRRG 118
Q + HPLQCRALELCFNVALNRLPT+T SP+L PS+SNAL AA KRAQAHQRRG
Sbjct: 58 QSHTHPLQCRALELCFNVALNRLPTSTGSPMLGVPTSPFPSISNALGAAFKRAQAHQRRG 117
Query: 119 CIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINS 178
IE QQQP+L+VK+E++QLI+SILDDPSVSRVMREAGFSSP VK +E + +L
Sbjct: 118 SIE----SQQQPILAVKIEVEQLIISILDDPSVSRVMREAGFSSPQVKTKVEQAVSLEIC 173
Query: 179 SSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLR 238
S SS+ KE + P ED+ V + L+
Sbjct: 174 S----------------------KTTSSSKPKEGKLLTPV------RNEDVMNVINNLVD 205
Query: 239 KKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMKKEEV 298
KK++N VIVG+ ++ +G+V +M++ ++ +VP+ +K VKF LS S + +V
Sbjct: 206 KKRRNFVIVGECLATIDGVVKTVMEKVDKKDVPEVLKD---VKFITLSFSSFGQPSRADV 262
Query: 299 EMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDD--NDGSLNEKE----VVDYVVEEIG 352
E + L+ V V G G I +GDL W V+ SL VV++++ EIG
Sbjct: 263 ERK-LEELETLVKSCV--GKGVILNLGDLNWFVESRTRGSSLYNNNDSYCVVEHMIMEIG 319
Query: 353 KL-FGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLSLHSSSV 411
KL G +G+ WL+ A+ Q+Y+RC+ P+ E+ WCL + +P+ L L S
Sbjct: 320 KLACGLVMGDHGRFWLMGLATSQTYVRCKSGQPSLESLWCLTTLTIPATSNSLRLSLVSE 379
Query: 412 HDSKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEAQLFKPDQKNL----LPS 467
+ ++ S+N S L+ + ++L+ CEEC +E EA+ K N+ LP+
Sbjct: 380 SELEVKKSENVSLQLQ-------QSSDQLSFCEECSVKFESEARFLKSSNSNVTTVALPA 432
Query: 468 WLQSHSTEAR----QKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKIYPYNSSY 523
WLQ + E + D + +L KWN +C +H+ + ++ +S+
Sbjct: 433 WLQQYKKENQNSHTDSDSIKELVVKWNSICDSIHKRPSLKTLTLSSPTSSF--------- 483
Query: 524 PYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRGQQSCTIEFNFNDEKAQKNQV 583
S P S+ + P +N ++ + + +++
Sbjct: 484 ------SGSTQPSISTLHHLQTNGDWPVIETNTHRHHSVVHETSHLRLFIPEHDSEQKTE 537
Query: 584 TATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLTLQRDHIYKVLQENIPWHCE 643
+S E + L +S F + ENL + L+ +PW +
Sbjct: 538 LVCSNPNSTMNSEASSSDAMELEHASSRFK---EMNAENLAT----LCAALESKVPWQKD 590
Query: 644 TVSSIAEALVDSKS-----------SKECATWLFLQGNDSVGKKRLALAIAESVFGSVEM 692
V +A+ ++ +S K+ TW+F QG D K+++A +A+ VFGS +
Sbjct: 591 LVPELAKTVLKCRSGSSTRKINGNEDKKEDTWMFFQGLDVDAKEKIARELAKLVFGSQDS 650
Query: 693 FSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVENADFGDTL--IRKMLADEFEIAKFG 750
F + + ++ + +E + +E+ + +E +L R +L ++ E A +
Sbjct: 651 FVSICLSSFSSTRSDSAEDLRNKRLRDEQSLSYIERFSEAVSLDPNRVILVEDIEQADY- 709
Query: 751 TLGQKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKPTFELSPSSSSSSKS 803
L Q F + V +++ +K + I E+ + + S S+ KS
Sbjct: 710 -LSQVGFKRAVERGRVCNSSGEEASLKDAIVILSCERFRSRSRACSPPSNQKS 761
>At5g57710 101 kDa heat shock protein; HSP101-like protein
Length = 990
Score = 334 bits (856), Expect = 2e-91
Identities = 262/810 (32%), Positives = 413/810 (50%), Gaps = 97/810 (11%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+G T+Q+TLT EAA++L S+ A RR H Q TPLHVAATLL+ P+ LR AC++S
Sbjct: 1 MRAGLSTIQQTLTPEAATVLNQSIAEAARRNHGQTTPLHVAATLLASPAGFLRRACIRSH 60
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCI 120
++HPLQCRALELCF+VAL RLPT TT+ P + P +SNAL+AALKRAQAHQRRGC
Sbjct: 61 PNSSHPLQCRALELCFSVALERLPTATTT----PGNDPPISNALMAALKRAQAHQRRGCP 116
Query: 121 EQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSS 180
E QQQQPLL+VKVEL+QLI+SILDDPSVSRVMREA FSSP+VK +E S +N+S
Sbjct: 117 E----QQQQPLLAVKVELEQLIISILDDPSVSRVMREASFSSPAVKATIEQS---LNNSV 169
Query: 181 VFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYH-PFLKSSESNKEDINLVFDVLLRK 239
PS S G G + +S ++ + ++S S +D+ V D+L R
Sbjct: 170 TPTPIPSVSSVGLNFRPGGGGPMTRNSYLNPRLQQNASSVQSGVSKNDDVERVMDILGRA 229
Query: 240 KKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPD-EMKTTHFVKFHGLSSVSLKYMKKEEV 298
KKKN V+VGD S ++ EI+K+ E GEV + +K + V +SS +K
Sbjct: 230 KKKNPVLVGD--SEPGRVIREILKKIEVGEVGNLAVKNSKVVSLEEISSDKALRIK---- 283
Query: 299 EMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKEVVDYVVEEIGKLFGEE 358
E++ + + K SD + G G I +GDLKW+V+ + + V EIG+ E
Sbjct: 284 ELDGLLQTRLKNSDPIG-GGGVILDLGDLKWLVEQPSST----QPPATVAVEIGRTAVVE 338
Query: 359 GNK-----NGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLSLHSSSVHD 413
+ G++W + TA+ ++Y+RCQ+ P+ E W LQAV V + +S V
Sbjct: 339 LRRLLEKFEGRLWFIGTATCETYLRCQVYHPSVETDWDLQAVSVAA-----KAPASGVFP 393
Query: 414 SKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEA----QLFKPD------QKN 463
+ ++ +P L+S +N+ L CC +C+ +YE+E + P+ Q
Sbjct: 394 RLANNLESFTP-LKSFVPANR----TLKCCPQCLQSYERELAEIDSVSSPEVKSEVAQPK 448
Query: 464 LLPSW-LQSHSTEARQKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKIYPYN-- 520
LP W L++ + + ++ ++ KKWN C LH + H+ N +I P
Sbjct: 449 QLPQWLLKAKPVDRLPQAKIEEVQKKWNDACVRLH---------PSFHNKNERIVPIPVP 499
Query: 521 ---SSYPYWPNQ--GSSILPDTSSSISFADSA-TKPAYSSNLIPRFRRGQQSCTIEFNFN 574
++ PY PN + P + + KP ++ ++ +
Sbjct: 500 ITLTTSPYSPNMLLRQPLQPKLQPNRELRERVHLKPMSPLVAEQAKKKSPPGSPVQTDLV 559
Query: 575 DEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLTLQRDHIYKVL 634
+A+ ++ +++ G ++ V+ N+ SV ++ + N +L D K+L
Sbjct: 560 LGRAEDSEKAGDVQVRDFLGCISSESVQ-----NNNNISVLQKENLGN-SLDIDLFKKLL 613
Query: 635 Q---ENIPWHCETVSSIAEALVDSKSS--------KECATWLFLQGNDSVGKKRLALAIA 683
+ E + W + +++A + K + WL G D VGK+++ A++
Sbjct: 614 KGMTEKVWWQNDAAAAVAATVSQCKLGNGKRRGVLSKGDVWLLFSGPDRVGKRKMVSALS 673
Query: 684 ESVFGS----VEMFSHVDMMKRENS--ETPFSEKVVGPLKNNEKFVVLVENADFGDTLIR 737
V+G+ +++ S D +S +K+ +K + V+L+E+ D D L+R
Sbjct: 674 SLVYGTNPIMIQLGSRQDAGDGNSSFRGKTALDKIAETVKRSPFSVILLEDIDEADMLVR 733
Query: 738 KMLADEFEIAKFG-------TLGQKIFILS 760
+ + + +LG IF+++
Sbjct: 734 GSIKQAMDRGRIRDSHGREISLGNVIFVMT 763
>At4g30350 unknown protein
Length = 924
Score = 328 bits (841), Expect = 9e-90
Identities = 308/1004 (30%), Positives = 470/1004 (46%), Gaps = 183/1004 (18%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+ T+Q+TLT EAA++L S+ A RR H TPLHVAATLLS S LR AC+KS
Sbjct: 1 MRADLITIQQTLTPEAATVLNQSIAEATRRNHGHTTPLHVAATLLSSSSGYLRQACIKSH 60
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTTT-------SPLLQPQHV--PSLSNALIAALKRA 111
++HPLQCRALELCF+VAL RLPTT+T S P P LSNAL AALKRA
Sbjct: 61 PNSSHPLQCRALELCFSVALERLPTTSTTTTTTSSSSSSSPSQTQEPLLSNALTAALKRA 120
Query: 112 QAHQRRGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLEN 171
QAHQRRGC E QQQQPLL+VKVEL+QLI+SILDDPSVSRVMREA FSSP+VK+ +E
Sbjct: 121 QAHQRRGCPE----QQQQPLLAVKVELEQLIISILDDPSVSRVMREASFSSPAVKSAIE- 175
Query: 172 SSTLINSSSVFHSSPSPLSHNHFLSSYGYGSV-------LFSSQKKEQVVYHPFLKSSES 224
S + NS S + SP N +GY SV L+ + + +Q
Sbjct: 176 QSLIGNSVSNSRQTGSPGIINPSAIGFGYRSVPAPVNRNLYLNPRLQQPGVGMQSGMMIQ 235
Query: 225 NKEDINLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHG 284
++ V ++++R +K+N V+VGD S LV EI+++ E GE D
Sbjct: 236 RTDEAKRVIEIMIRTRKRNPVLVGD--SEPHILVKEILEKIENGEFSD----------GA 283
Query: 285 LSSVSLKYMKKEEVEMNVIRVLKRKVSDYVAL---GVGAIFYVGDLKWIVDD---NDGSL 338
L + + ++KE V R+ ++S V G G + +GDLKW+V+ N G+
Sbjct: 284 LRNFQVIRLEKELVSQLATRL--GEISGLVETRIGGGGVVLDLGDLKWLVEHPAANGGA- 340
Query: 339 NEKEVVDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVP 398
V E+ KL G++ + TA+ ++Y+RCQ+ P+ EN W LQA+P+
Sbjct: 341 ---------VVEMRKLL---ERYKGRLCFIGTATCETYLRCQVYYPSMENDWDLQAIPIA 388
Query: 399 SGGLGLSLH---SSSVHDSKMSISQNPSPMLE-SKFFSNKEEHEKLNCCEECVSNYEKEA 454
+ ++ S+ +++ M +S N + S S + K++CC C+ +YE +
Sbjct: 389 AKSSLPAIFPRLGSNNNNNAMLLSNNIISIESISPTRSFQIPMSKMSCCSRCLQSYENDV 448
Query: 455 QLFKP----DQKNLLPSWLQSHST------EARQKDELTQLNKKWNRLCQCLHQNKQPQN 504
+ D +++LP WLQ+ + + ++ +L KKWN LC LH N+
Sbjct: 449 AKVEKDLTGDNRSVLPQWLQNAKANDDGDKKLTKDQQIVELQKKWNDLCLRLHPNQSVSE 508
Query: 505 HWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRGQ 564
+ + S KI N S I P S P + ++ R RG
Sbjct: 509 RIAPSTLSMMKI-----------NTRSDITPPGS-----------PVGTDLVLGRPNRGL 546
Query: 565 QSCTIEFNFNDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLT 624
S EK T+E + LG+S F + K+
Sbjct: 547 SS--------PEKK-------------------TREARFG-KLGDS-FDIDLFKK----- 572
Query: 625 LQRDHIYKVLQENIPWHCETVSSIAEALVDSK---SSKECATWLFLQGNDSVGKKRLALA 681
+ K L +++ W + SS+A A+ + K + WL G D GK ++A A
Sbjct: 573 -----LLKGLAKSVWWQHDAASSVAAAITECKHGNGKSKGDIWLMFTGPDRAGKSKMASA 627
Query: 682 IAESVFGSVEMFSHVDMMKRENSETPFS-----EKVVGPLKNNEKFVVLVENADFGDTLI 736
+++ V GS + + R + ++ ++ N V+++E+ D D L+
Sbjct: 628 LSDLVSGSQPITISLGSSSRMDDGLNIRGKTALDRFAEAVRRNPFAVIVLEDIDEADILL 687
Query: 737 RKMLADEFEIAK----FG---TLGQKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKP 789
R + E + +G +LG I IL+ S+ S K V I ET +
Sbjct: 688 RNNVKIAIERGRICDSYGREVSLGNVIIILTANSSLGS--------AKNVASIDETRLES 739
Query: 790 T----FELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKRE---FSFSRQS 842
+EL S +SSK+ KR L+S +N+ K+R+ F + +
Sbjct: 740 LVNKGWELRLSVCNSSKT----RKRKPNW-LYS--------DNDQTKQRKEICFDLNEAA 786
Query: 843 SFNNTLDLNMKADEEDNEDY---------DEGENSPISSDLTRETLGEHLISNESLDSIE 893
F+++ D+ ++ D+EDN + D P+ D + E L S +
Sbjct: 787 EFDSSSDVTVEHDQEDNGNLVHKLVGLVDDAILFRPVDFDSIKSKTAESLKKRFSNGLAD 846
Query: 894 NLFEFNQSPAKNKEMTQMFMSRV--KESFEEVLGNVKFSVQDKV 935
L + A + +++S++ +E EE +G+ SV+ +V
Sbjct: 847 GLTVEIEDDALERIAGAIWLSKISLEEWLEEAMGSSLNSVKSRV 890
>At2g29970 unknown protein
Length = 1002
Score = 207 bits (527), Expect = 2e-53
Identities = 259/937 (27%), Positives = 405/937 (42%), Gaps = 141/937 (15%)
Query: 7 TLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHP 66
T ++ LT E A L ++ +ARRR HAQ T LH + LL++PSS LR C+ S+ +N P
Sbjct: 7 TARQCLTEETARALDDAVSVARRRSHAQTTSLHAVSGLLTMPSSILREVCI-SRAAHNTP 65
Query: 67 ----LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRR----- 117
LQ RALELC V+L+RLP++ ++P + P +SN+L+AA+KR+QA QRR
Sbjct: 66 YSSRLQFRALELCVGVSLDRLPSSKSTPTTTVEEDPPVSNSLMAAIKRSQATQRRHPETY 125
Query: 118 --GCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTL 175
I N + +L KVEL ILSILDDP VSRV EAGF S +K ++ +
Sbjct: 126 HLHQIHGNNNTETTSVL--KVELKYFILSILDDPIVSRVFGEAGFRSTDIKLDVLHPPVT 183
Query: 176 INSSSVFHS-SPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFD 234
SS F S S P L G V F PF E+ + + +
Sbjct: 184 SQFSSRFTSRSRIPPLFLCNLPESDSGRVRFG---------FPFGDLDENCRR----IGE 230
Query: 235 VLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMK 294
VL RK KKN ++VG ++ + R + G +P E+ GLS VS+K
Sbjct: 231 VLARKDKKNPLLVGVCGVEALKTFTDSINRGKFGFLPLEIS--------GLSVVSIKI-- 280
Query: 295 KEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKEVVDYVVEEIGKL 354
EV ++ R+ K D L G + +G+LK + D VD + + + KL
Sbjct: 281 -SEVLVDGSRI-DIKFDDLGRLKSGMVLNLGELKVLASDVFS-------VDVIEKFVLKL 331
Query: 355 FGEEGNKNGKIWLV-ATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLSLHSSSVHD 413
K+W + + +S ++Y++ R PT + W L +P+ S GL SS+
Sbjct: 332 ADLLKLHREKLWFIGSVSSNETYLKLIERFPTIDKDWNLHLLPITSSSQGL-YPKSSLMG 390
Query: 414 SKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEAQLFK------PDQ-KNLLP 466
S + S + + S+ ++ L C C YE+E F DQ LP
Sbjct: 391 SFVPFGGFFSSTSDFRIPSSSSMNQTLPRCHLCNEKYEQEVTAFAKSGSMIDDQCSEKLP 450
Query: 467 SWLQSHSTE------ARQKDE-------LTQLNKKWNRLCQCLHQNK----------QPQ 503
SWL++ E + KD+ + L KKW+ +CQ +HQ +PQ
Sbjct: 451 SWLRNVEHEHEKGNLGKVKDDPNVLASRIPALQKKWDDICQRIHQTPAFPKLSFQPVRPQ 510
Query: 504 NHWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRG 563
SS K+ + P + + S P + L + +
Sbjct: 511 FPLQLGSSSQTKMSLGS------PTEKIVCTRTSESFQGMVALPQNPPHQPGLSVKISKP 564
Query: 564 QQSCTIEFNFNDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENL 623
+ T + + + + + VT L L ++ + +E T +++ F V +K++ L
Sbjct: 565 KH--TEDLSSSTTNSPLSFVTTDLGLGTIYASK-NQEPSTPVSVERRDFEVIKEKQL--L 619
Query: 624 TLQR-----DHIYKVLQENIPWHCETVSSIAEALV-----DSKSSKECAT----WLFLQG 669
+ R + ++L + + E V++I+E + + + AT WL L G
Sbjct: 620 SASRYCKDFKSLRELLSRKVGFQNEAVNAISEIVCGYRDESRRRNNHVATTSNVWLALLG 679
Query: 670 NDSVGKKRLALAIAESVFGSVEMFSHVDMMKRENSETPFSEKVV-----GPLKNNEKFVV 724
D GKK++ALA+AE G + F VD +++ + F K V G + VV
Sbjct: 680 PDKAGKKKVALALAEVFCGGQDNFICVDFKSQDSLDDRFRGKTVVDYIAGEVARRADSVV 739
Query: 725 LVENADFGDTLIRKMLADEFEIAKFGTLGQKIFILSN---GGSMVSEDQKKD-SVMKLVL 780
+EN + + + L++ K + + N ++ D+ D V++ +
Sbjct: 740 FIENVEKAEFPDQIRLSEAMRTGKLRDSHGREISMKNVIVVATISGSDKASDCHVLEEPV 799
Query: 781 KISE----TEKKPTFE--LSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKR 834
K SE K T + L+ +S+ + P NKR R EE E
Sbjct: 800 KYSEERVLNAKNWTLQIKLADTSNVNKNGP---NKR-------------RQEEAETEVTE 843
Query: 835 EFSFSRQSSFNNTLDLNMKADE-EDNED--YDEGENS 868
+ Q SF LDLN+ DE E NED Y EN+
Sbjct: 844 LRALKSQRSF---LDLNLPVDEIEANEDEAYTMSENT 877
>At1g07200 unknown protein
Length = 979
Score = 180 bits (456), Expect = 4e-45
Identities = 217/840 (25%), Positives = 354/840 (41%), Gaps = 128/840 (15%)
Query: 7 TLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQN--- 63
T +E LT EAA L ++ +ARRR HAQ T LH + LL++PSS LR C+ ++
Sbjct: 7 TARECLTEEAARALDDAVVVARRRSHAQTTSLHAVSALLAMPSSILREVCVSRAARSVPY 66
Query: 64 NHPLQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQN 123
+ LQ RALELC V+L+RLP ++ SP + P +SN+L+AA+KR+QA+QRR +
Sbjct: 67 SSRLQFRALELCVGVSLDRLP-SSKSPATEED--PPVSNSLMAAIKRSQANQRRHPESYH 123
Query: 124 QQQQQQ--------PLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTL 175
QQ +KVEL ILSILDDP V+RV EAGF S +K ++ +
Sbjct: 124 LQQIHASNNGGGGCQTTVLKVELKYFILSILDDPIVNRVFGEAGFRSSEIKLDVLHPPVT 183
Query: 176 INSSSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDV 235
SS PL FL + L +S + PF SS + E+ + +V
Sbjct: 184 QLSSRFSRGRCPPL----FLCN------LPNSDPNRE---FPFSGSSGFD-ENSRRIGEV 229
Query: 236 LLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMKK 295
L RK KKN +++G+ + ++ + + G + ++ + S L K
Sbjct: 230 LGRKDKKNPLLIGNCANEALKTFTDSINSGKLGFLQMDISGLSLISIEKEISEILADGSK 289
Query: 296 EEVEMNV-IRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKEVVDYVVEEIGKL 354
E E+ + + L R V + G + +G+LK + + + +L + +V ++ L
Sbjct: 290 NEEEIRMKVDDLGRTVEQSGSKS-GIVLNLGELKVLTSEANAAL------EILVSKLSDL 342
Query: 355 FGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLSLHSSSVHDS 414
E + I V +S ++Y + R PT E W L +P+ + + S+
Sbjct: 343 LKHESKQLSFIGCV--SSNETYTKLIDRFPTIEKDWDLHVLPITAS----TKPSTQGVYP 396
Query: 415 KMSISQNPSPMLESKFFSNKEE---------HEKLNCCEECVSNYEKEAQLFKPDQKNL- 464
K S+ + P FFS+ ++ L+ C C Y +E +L
Sbjct: 397 KSSLMGSFVPF--GGFFSSTSNFRVPLSSTVNQTLSRCHLCNEKYLQEVAAVLKAGSSLS 454
Query: 465 --------LPSWLQSHSTEA---------------RQKDELTQLNKKWNRLCQCLHQNKQ 501
L WL++ T+ + L KKW+ +CQ +H
Sbjct: 455 LADKCSEKLAPWLRAIETKEDKGITGSSKALDDANTSASQTAALQKKWDNICQSIH---- 510
Query: 502 PQNHWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPA--------YS 553
H+ + S P +P Q + +S + P +
Sbjct: 511 --------HTPAFPKLGFQSVSPQFPVQTEKSVRTPTSYLETPKLLNPPISKPKPMEDLT 562
Query: 554 SNLIPRFRRGQQSC-TIEFNFNDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTF 612
+++ R SC T +F A KNQ + T T+E K L NS+
Sbjct: 563 ASVTNRTVSLPLSCVTTDFGLGVIYASKNQESKT-----------TRE-KPMLVTLNSSL 610
Query: 613 SVSDQKRMENLTLQRDHIYKVLQENIPWHCETVSSIAEALVDSKS-----SKECATWLFL 667
+ QK ++L ++L + W E V++I++ + K+ ++ WL L
Sbjct: 611 EHTYQKDFKSLR-------EILSRKVAWQTEAVNAISQIICGCKTDSTRRNQASGIWLAL 663
Query: 668 QGNDSVGKKRLALAIAESVFGSVEMFSHVDMMKRENS-ETPFSEK-----VVGPLKNNEK 721
G D VGKK++A+ ++E FG + VD S + F K V G L
Sbjct: 664 LGPDKVGKKKVAMTLSEVFFGGKVNYICVDFGAEHCSLDDKFRGKTVVDYVTGELSRKPH 723
Query: 722 FVVLVENADFGDTLIRKMLADEFEIAKFGTLGQKIFILSNGGSMVSEDQKKDSVMKLVLK 781
VVL+EN + + + L++ K L ++ + N +V+ KD+ V+K
Sbjct: 724 SVVLLENVEKAEFPDQMRLSEAVSTGKIRDLHGRVISMKNVIVVVTSGIAKDNATDHVIK 783
>At2g40130 unknown protein
Length = 907
Score = 161 bits (408), Expect = 2e-39
Identities = 132/414 (31%), Positives = 212/414 (50%), Gaps = 45/414 (10%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
M + ++ LTAEA+ L+ ++ +ARRRGH+Q T LH + LLSLP+S LR+AC + +
Sbjct: 1 MPTAVNVAKQCLTAEASYALEEAVNVARRRGHSQTTSLHAISALLSLPTSVLRDACARVR 60
Query: 61 QQNNHP-LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQR--- 116
P LQ +AL+LC +V+L+R+ + L P +SN+L+AA+KR+QAHQR
Sbjct: 61 NSAYSPRLQFKALDLCLSVSLDRI---QSGHQLGSDDSPPVSNSLMAAIKRSQAHQRRLP 117
Query: 117 ---RGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSS 173
R E +Q Q Q L VKVEL QLILSILDDP VSRV EAGF S +K ++
Sbjct: 118 ENFRIYQEMSQSQNQNSLSCVKVELRQLILSILDDPVVSRVFGEAGFRSSELKLSIIRPV 177
Query: 174 TLINSSSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKE-DINLV 232
+ + +SS PL FL + L + + V + + S N + D +
Sbjct: 178 PHL----LRYSSQQPL----FLCN------LTGNPEPNPVRWGFTVPSLNFNGDLDYRRI 223
Query: 233 FDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKY 292
V + K +N ++VG + G+++ + E+ + + T K HGL++V++
Sbjct: 224 SAVFTKDKGRNPLLVGVS---AYGVLTSYLNSLEKNQTDGMILPT---KLHGLTAVNIGS 277
Query: 293 MKKEEVEMNVIRV-LKRKVSDYVAL-----GVGAIFYVGDLKWIVDDNDGSLNEKEVVDY 346
+++ + + + D L G G + + GDL+ + + +G++ +Y
Sbjct: 278 EISDQISVKFDKTYTDTRFHDLGKLAEQGSGPGLLLHYGDLR-VFTNGEGNV---PAANY 333
Query: 347 VVEEIGKLFGEEGNKNGKIWLV-ATASYQSYMRCQMRVPTFENQWCLQAVPVPS 399
+V I +L G ++WL+ AT S + Y + R P E W LQ + + S
Sbjct: 334 IVNRISELLRRHGR---RVWLIGATTSNEVYEKMMRRFPNVEKDWDLQLLTITS 384
Score = 57.0 bits (136), Expect = 5e-08
Identities = 80/340 (23%), Positives = 135/340 (39%), Gaps = 62/340 (18%)
Query: 630 IYKVLQENIPWHCETVSSIAEALVDS-KSSKECATWLFLQGNDSVGKKRLALAIAESVFG 688
IY+ L + + E I+ AL KS WL L G D+VGK+R++L +AE V+
Sbjct: 537 IYRRLTDMVSGQDEAARVISCALSQPPKSVTRRDVWLNLVGPDTVGKRRMSLVLAEIVYQ 596
Query: 689 SVEMFSHVDMMKRENS----ETPFS-------EKVVGPLKNNEKFVVLVENADFGDTLIR 737
S F VD+ E + P + + + N VV +EN + D ++
Sbjct: 597 SEHRFMAVDLGAAEQGMGGCDDPMRLRGKTMVDHIFEVMCRNPFCVVFLENIEKADEKLQ 656
Query: 738 KMLADEFEIAKFGT-------LGQKIFIL--SNGGSMVSEDQKKDSVMK-----LVLKIS 783
L+ E KF +G IF++ S+ GS + ++ +++ + ++I
Sbjct: 657 MSLSKAIETGKFMDSHGREVGIGNTIFVMTSSSQGSATTTSYSEEKLLRVKGRQVEIRIE 716
Query: 784 ETEKKPTFE--LSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQ 841
P P+S + K LGN + + + S ++ R
Sbjct: 717 TVSSLPMVRSVYGPTSVNKRKLMGLGNLQETKDTVESVKRLNR----------------- 759
Query: 842 SSFNNTLDLNMKADE-EDNEDYDEGENSPISSDLTRETLGEHLISNESLDSIENLFEFNQ 900
+ N LDLN+ A E E E Y ENS + +L + + L E
Sbjct: 760 -TTNGVLDLNLPAQETEIEEKYHCEENSNVWL--------------MNLKNHKRLIEVPF 804
Query: 901 SPAKNKEMTQMFMSRVKESFEE-VLGNVKFSVQDKVIEEI 939
P + + + VKE+F++ V + V K+IE +
Sbjct: 805 KPFDFEGLAEKIKKSVKENFDKCVRSDCLLEVDPKIIERL 844
>At1g74310 heat shock protein 101
Length = 911
Score = 78.2 bits (191), Expect = 2e-14
Identities = 75/284 (26%), Positives = 128/284 (44%), Gaps = 40/284 (14%)
Query: 26 LARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQCRALELCFNVALNRLPT 85
LA GHAQ TPLH+A L+S P+ A + +N ++ E N AL +LP+
Sbjct: 20 LAVNAGHAQFTPLHLAGALISDPTGIFPQAISSAGGEN----AAQSAERVINQALKKLPS 75
Query: 86 TTTSPLLQPQHVPSLSNALIAALKRAQAHQR-RGCIEQNQQQQQQPLLSVKVELDQLILS 144
+ P +P+ S++LI ++RAQA Q+ RG + +DQLI+
Sbjct: 76 QSP----PPDDIPA-SSSLIKVIRRAQAAQKSRG--------------DTHLAVDQLIMG 116
Query: 145 ILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSPLSHNHFLSSYGYGSVL 204
+L+D + ++ E G ++ VK+ +E L S S ++ L +YG ++
Sbjct: 117 LLEDSQIRDLLNEVGVATARVKSEVEK---LRGKEGKKVESASGDTNFQALKTYG-RDLV 172
Query: 205 FSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKR 264
+ K + V+ E+I V +L R+ K N V++G+ +V + +R
Sbjct: 173 EQAGKLDPVI---------GRDEEIRRVVRILSRRTKNNPVLIGEPGVGKTAVVEGLAQR 223
Query: 265 FERGEVPDEMKTTHFVKFH-GLSSVSLKYMKKEEVEMNVIRVLK 307
+G+VP+ + + G KY + E E + VLK
Sbjct: 224 IVKGDVPNSLTDVRLISLDMGALVAGAKY--RGEFEERLKSVLK 265
Score = 33.5 bits (75), Expect = 0.62
Identities = 26/88 (29%), Positives = 46/88 (51%), Gaps = 12/88 (13%)
Query: 617 QKRMENLTLQRDHIYK-VLQENIPWHCETVSSIAEALVDSKSS-----KECATWLFLQGN 670
Q E L D ++K V+ +N + V++++EA++ S++ + ++LFL G
Sbjct: 554 QNEKERLIGLADRLHKRVVGQN-----QAVNAVSEAILRSRAGLGRPQQPTGSFLFL-GP 607
Query: 671 DSVGKKRLALAIAESVFGSVEMFSHVDM 698
VGK LA A+AE +F + +DM
Sbjct: 608 TGVGKTELAKALAEQLFDDENLLVRIDM 635
>At4g14670 heat shock protein like
Length = 668
Score = 42.4 bits (98), Expect = 0.001
Identities = 57/276 (20%), Positives = 105/276 (37%), Gaps = 82/276 (29%)
Query: 17 ASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQCRALELCF 76
AS H++ L+ H Q+TPLH+ TL+S +S A
Sbjct: 15 ASARSHAMSLS----HGQVTPLHLGVTLISDLTSVFYRAI-------------------- 50
Query: 77 NVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQQQQQPLLSVKV 136
T+ + Q V ++ N + L + L KV
Sbjct: 51 --------TSAGDGDISAQSVVNVINQSLYKLTKRN------------------LGDTKV 84
Query: 137 ELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSPLSHNHFLS 196
+ L++S+L+D +S V++EAG VK+ +E + + + L
Sbjct: 85 GVAVLVISLLEDSQISDVLKEAGVVPEKVKSEVEK----LRGEVILRA----------LK 130
Query: 197 SYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVGDTVSLTEG 256
+YG ++ + K + V+ +I V +VL R+ K N V++G+
Sbjct: 131 TYG-TDLVEQAGKLDPVI---------GRHREIRRVIEVLSRRTKNNPVLIGEPGVGKTA 180
Query: 257 LVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKY 292
+V + +R +G+VP + G+ +SL++
Sbjct: 181 VVEGLAQRILKGDVP--------INLTGVKLISLEF 208
Score = 33.9 bits (76), Expect = 0.48
Identities = 23/70 (32%), Positives = 36/70 (50%), Gaps = 6/70 (8%)
Query: 634 LQENIPWHCETVSSIAEALVDSK-----SSKECATWLFLQGNDSVGKKRLALAIAESVFG 688
L E + E V ++A A++ S+ + ++LFL G VGK LA A+AE +F
Sbjct: 532 LHERVVGQDEAVKAVAAAILRSRVGLGRPQQPSGSFLFL-GPTGVGKTELAKALAEQLFD 590
Query: 689 SVEMFSHVDM 698
S + +DM
Sbjct: 591 SENLLVRLDM 600
>At4g12620 origin recognition complex subunit 1 -like protein
Length = 813
Score = 39.7 bits (91), Expect = 0.009
Identities = 25/94 (26%), Positives = 47/94 (49%), Gaps = 9/94 (9%)
Query: 776 MKLVLKISETEKKPTFELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKRE 835
M L+ K +KPT ++ PS S +++P K+ ++D F+ + R E + KK++
Sbjct: 73 MNLIRKRERAPRKPTTDVVPSKSKKTETP----KKKKKIDSFTPVSPIRSETIKKTKKKK 128
Query: 836 FSFSRQSSFNNTL-----DLNMKADEEDNEDYDE 864
+ + F+ T D+ +K E+ N D +E
Sbjct: 129 RVYYNKVEFDETEFEIGDDVYVKRREDSNSDEEE 162
>At3g28770 hypothetical protein
Length = 2081
Score = 38.5 bits (88), Expect = 0.019
Identities = 47/190 (24%), Positives = 70/190 (36%), Gaps = 15/190 (7%)
Query: 762 GGSMVSEDQKKDSVMKLVLKISETEKKPTFELSPSSSSSSKSPC-LGNKRSAELDLFS-- 818
G S+D K D +++ E+ KK E+ + SS+K N ++ S
Sbjct: 853 GNKEDSKDLKDDRSVEVKANKEESMKKKREEVQRNDKSSTKEVRDFANNMDIDVQKGSGE 912
Query: 819 KIKIPRIEENEGNKKREFSFSRQSSFNNTLDL----------NMKADEEDNEDYDEGENS 868
+K + E+ EGNK+ SS D NMK EED ++Y E
Sbjct: 913 SVKYKKDEKKEGNKEENKDTINTSSKQKGKDKKKKKKESKNSNMKKKEEDKKEYVNNELK 972
Query: 869 PISSDLTRETLGEHLISNESLDSIENLFEFNQSPAKNKEMTQMFMSRVKESFEEVLGNVK 928
+ T E+ E + E S +KN+E + K +E K
Sbjct: 973 KQEDNKKETTKSENSKLKEENKDNKEKKESEDSASKNREKKE--YEEKKSKTKEEAKKEK 1030
Query: 929 FSVQDKVIEE 938
QDK EE
Sbjct: 1031 KKSQDKKREE 1040
Score = 32.0 bits (71), Expect = 1.8
Identities = 36/162 (22%), Positives = 65/162 (39%), Gaps = 25/162 (15%)
Query: 761 NGGSMVSEDQKKDSVMKLVLKISETEKKPTFELSPSSSSSSKSPCLGNKRSAELDLFSKI 820
N S ED+K+ K + K + ++K E S S R E D K
Sbjct: 1075 NHKSKKKEDKKEHEDNKSMKKEEDKKEKKKHEESKS-------------RKKEED---KK 1118
Query: 821 KIPRIEENEGNKKREFSFSRQSSFNNTLDLNMKADEEDNEDYDEGENSPISSDLTRETLG 880
+ ++E+ NKK+E ++ S + L +E E+ ++ E I S +++
Sbjct: 1119 DMEKLEDQNSNKKKEDKNEKKKSQHVKLVKKESDKKEKKENEEKSETKEIESSKSQK--- 1175
Query: 881 EHLISNESLDSIENLFEFNQSPAKNKEMTQMFMSRVKESFEE 922
+D E +Q K KEM + ++K++ E+
Sbjct: 1176 ------NEVDKKEKKSSKDQQKKKEKEMKESEEKKLKKNEED 1211
>At5g15450 clpB heat shock protein-like
Length = 968
Score = 37.7 bits (86), Expect = 0.033
Identities = 120/677 (17%), Positives = 260/677 (37%), Gaps = 103/677 (15%)
Query: 122 QNQQQQQQPLLSVKVELDQLILSILDDPSVSR-VMREAGFSSPSVKNNLENSSTLINSSS 180
Q +Q ++ L V ++ L+L+ DD + + ++ S S+K+ +E+ + S
Sbjct: 168 QRARQFKKDLKDSYVSVEHLVLAFADDKRFGKQLFKDFQISERSLKSAIES---IRGKQS 224
Query: 181 VFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKK 240
V P L YG + + K V ++I +L R+
Sbjct: 225 VIDQDPE--GKYEALEKYGKDLTAMAREGKLDPVI--------GRDDEIRRCIQILSRRT 274
Query: 241 KKNTVIVGD----TVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFH-GLSSVSLKYMKK 295
K N V++G+ +++EGL I+ +G+VP + + G KY +
Sbjct: 275 KNNPVLIGEPGVGKTAISEGLAQRIV----QGDVPQALMNRKLISLDMGALIAGAKY--R 328
Query: 296 EEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLN-EKEVVDYVVEEIGKL 354
E E + VLK +V+D I ++ ++ +V G+ N + + + +G+
Sbjct: 329 GEFEDRLKAVLK-EVTDSEG---QIILFIDEIHTVV--GAGATNGAMDAGNLLKPMLGR- 381
Query: 355 FGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLSL-------- 406
G++ + + Y + + P E ++ V P+ +S+
Sbjct: 382 --------GELRCIGATTLDEYRKYIEKDPALERRFQQVYVDQPTVEDTISILRGLRERY 433
Query: 407 ---HSSSVHDSKMSISQNPSPMLESKFFSNKEEHEK-LNCCEECVSNYE-----KEAQLF 457
H + DS + + +L ++ S + +K ++ +E + + K L
Sbjct: 434 ELHHGVRISDSALV----EAAILSDRYISGRFLPDKAIDLVDEAAAKLKMEITSKPTALD 489
Query: 458 KPDQKNLLPSWLQSHSTEARQKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKIY 517
+ D+ + + T K +LN+ L + + W + S +++
Sbjct: 490 ELDRSVIKLEMERLSLTNDTDKASRERLNRIETELVLLKEKQAELTEQWEHERSVMSRL- 548
Query: 518 PYNSSYPYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRGQQSCTIEFNFNDEK 577
+ + ++ + Y N + G + +++ N+ +
Sbjct: 549 --------------QSIKEEIDRVNLEIQQAEREYDLNRAAELKYGSLN-SLQRQLNEAE 593
Query: 578 AQKNQVTA---TLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLTLQRDHIYKVL 634
+ N+ + ++ + + G + + V + S S++ ++ L L+ + +V+
Sbjct: 594 KELNEYLSSGKSMFREEVLGSDIAEIVSKWTGIPVSKLQQSERDKL--LHLEEELHKRVV 651
Query: 635 QENIPWHCETVSSIAEALVDSKS-----SKECATWLFLQGNDSVGKKRLALAIAESVFGS 689
+N V+++AEA+ S++ + A+++F+ G VGK LA A+A +F +
Sbjct: 652 GQN-----PAVTAVAEAIQRSRAGLSDPGRPIASFMFM-GPTGVGKTELAKALASYMFNT 705
Query: 690 VEMFSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVENADFGDTLIRK----MLADEFE 745
E +DM E E +++G +V E +T+ R+ +L DE E
Sbjct: 706 EEALVRIDM--SEYMEKHAVSRLIGAPPG---YVGYEEGGQLTETVRRRPYSVILFDEIE 760
Query: 746 IAKFGTLGQKIFILSNG 762
A + IL +G
Sbjct: 761 KAHGDVFNVFLQILDDG 777
>At1g23710 unknown protein
Length = 295
Score = 37.0 bits (84), Expect = 0.056
Identities = 27/80 (33%), Positives = 46/80 (56%), Gaps = 11/80 (13%)
Query: 799 SSSKSPCLGNKRSAELDL-FSKIKIPRIEENEGNKKREFSFSRQSSFNNTLDLNMKADEE 857
SS C+ ++R E+ + F+K+ P EE+E KK E S+S+ +++ N + DEE
Sbjct: 7 SSPPVKCIADERLTEISMDFTKLGFP--EEDEEIKKLERSWSK---LEESVEFN-EDDEE 60
Query: 858 DNEDYD----EGENSPISSD 873
+ E++ GE SPI++D
Sbjct: 61 EEEEFSFACVNGEGSPITAD 80
>At5g02390 putative protein
Length = 835
Score = 35.8 bits (81), Expect = 0.13
Identities = 51/171 (29%), Positives = 72/171 (41%), Gaps = 23/171 (13%)
Query: 781 KISETEKKPTFELSPSSSSSSKSP-CLGNKRSAELDLFSKIKIPRI------EENEG--N 831
K S +K EL S+ S ++P LG S F +KI + EE +G N
Sbjct: 500 KNSSNSEKSKMELEESALPSKRAPKFLGRILSLPQMKFHALKIDDLTVQSIEEEQDGLDN 559
Query: 832 KKREFSFSRQSSFNNTLDLNMKADEEDNEDYDEGENSPISS-DLTRETLGEHLISNES-- 888
QSS + TLD M A E+ D + ++ IS+ D+ ET S ES
Sbjct: 560 ISEISEDHSQSSEHETLDQTMDASEDSPVDAETEQDREISTLDVEHETRSLKESSEESPN 619
Query: 889 ----LDSIENLFEFNQSPAKNKEMTQMFMSRVKESFEEVLGNVKFSVQDKV 935
+D EN F+ S +++ +S K+ EEVL QDKV
Sbjct: 620 NVSTVDIDENASVFDIS----RDLDTETVSTSKQLDEEVL---NIDAQDKV 663
>At4g39040 unknown protein
Length = 296
Score = 35.0 bits (79), Expect = 0.21
Identities = 46/184 (25%), Positives = 79/184 (42%), Gaps = 26/184 (14%)
Query: 783 SETEKKPTFELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQS 842
S KP F+ S SS PC + S + FS I P +EE+E ++ +FS
Sbjct: 45 SNHRNKPRFQ----SFSSKPLPC--HSASLVVKSFSSIDEPDLEEDEESEDEDFS----- 93
Query: 843 SFNNTLDLNMKADEEDNEDYDEGENSPISSDLTRETLGEHLISNESLDSIENLFEFNQSP 902
A+E + ED +E E+ S + E E ++E + I + E ++
Sbjct: 94 -----------AEEYEYED-EEEEDEQDSGVVVSERGIEDSEASEEVSEIGDKEEKTENT 141
Query: 903 AKNKEMTQMFMSRVKESFEEVLGNVKFSVQDKVIEEIGVGCGSFTNNMFEKWLKGIFQTS 962
K K +KE E L + S+ DK+ ++ VG T+++ +L+ + +
Sbjct: 142 KKKKSRGSALKLSIKEKKE--LASYAHSLGDKLKCQL-VGKSGVTDSVVFSFLETLEKNE 198
Query: 963 LERV 966
L +V
Sbjct: 199 LLKV 202
>At5g17020 Exportin1 (XPO1) protein
Length = 1075
Score = 34.7 bits (78), Expect = 0.28
Identities = 21/74 (28%), Positives = 35/74 (46%), Gaps = 7/74 (9%)
Query: 865 GENSPISSDL------TRETLGEHLISNESLDSIENLFEFNQSPAKNKEMTQMFMSRVKE 918
GEN P S+L T + L H I + +S+ N+ + P K E Q M+ +
Sbjct: 608 GENEPFVSELLTGLATTVQDLEPHQI-HSFYESVGNMIQAESDPQKRDEYLQRLMALPNQ 666
Query: 919 SFEEVLGNVKFSVQ 932
+ E++G + SV+
Sbjct: 667 KWAEIIGQARHSVE 680
>At5g52510 scarecrow-like 8 (SCL8)
Length = 525
Score = 33.9 bits (76), Expect = 0.48
Identities = 30/109 (27%), Positives = 47/109 (42%), Gaps = 3/109 (2%)
Query: 789 PTFELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQSSFNNTL 848
P SPSSSSSS SP + S + S+ + I K E + + + T
Sbjct: 124 PVLSFSPSSSSSSSSP---STASTTTSVCSRQTVMEIATAIAEGKTEIATEILARVSQTP 180
Query: 849 DLNMKADEEDNEDYDEGENSPISSDLTRETLGEHLISNESLDSIENLFE 897
+L ++E+ + S I+S +T EHLIS + L + F+
Sbjct: 181 NLERNSEEKLVDFMVAALRSRIASPVTELYGKEHLISTQLLYELSPCFK 229
>At1g49900 Cys2/His2-type zinc finger protein, putative
Length = 917
Score = 33.9 bits (76), Expect = 0.48
Identities = 37/162 (22%), Positives = 68/162 (41%), Gaps = 23/162 (14%)
Query: 792 ELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKRE-FSFSRQSSFNNTLDL 850
+LSP+S+ S + L AE L + I + ++E R+ +S S S N LDL
Sbjct: 285 KLSPNSNKSVVTKVL----DAEQSLRASDNIHKRNQDEVVPSRDKWSLSNVSVVKNVLDL 340
Query: 851 NMKADEEDNEDYDEGENSPISSDLTRETLGEHLISNES--LDSIENLFEFNQSPA----- 903
+ E D D + + D + +++N S S+ L + N P+
Sbjct: 341 KLSLGESDGGHKDSQDEVVVHEDKCSPSSNGSIVTNVSDPEQSVRRLIDLNNPPSLEFDQ 400
Query: 904 --------KNKEMTQMFMSRVKESFEEVLGNVK---FSVQDK 934
KE+ S++K+ ++V +K F++Q++
Sbjct: 401 SRGGDVEEVEKEVVLALSSQMKQPSKDVSALIKRLEFAMQER 442
>At5g45520 putative protein
Length = 1167
Score = 33.5 bits (75), Expect = 0.62
Identities = 45/167 (26%), Positives = 70/167 (40%), Gaps = 28/167 (16%)
Query: 762 GGSMVSEDQKKDSVMKLVLK---ISETEKKPTFELSPSSSSSSKSPCLGNKRSAELDLFS 818
G + + E++K+D V K I E EKKP S + G+K +A+LD
Sbjct: 682 GKADLEEEKKQDEVEAEKSKSDEIVEGEKKP------DDKSKVEKKGDGDKENADLDEGK 735
Query: 819 KIKIPRIEENEGNKKREFSFSRQSSFNNTLDL--NM----------------KADEEDNE 860
K +++E K E ++S ++D NM K D E+ +
Sbjct: 736 KRDEVEAKKSESGKVVEGD-GKESPPQESIDTIQNMTDDQTKVEKEGDRDKGKVDPEEGK 794
Query: 861 DYDEGENSPISSDLTRETLGEHLISNESLDSIENLFEFNQSPAKNKE 907
+DE E SD E + + S ES D+IEN + +Q K +E
Sbjct: 795 KHDEVEGGIWKSDNGVEGVDKASPSRESTDAIENKPDDHQRGDKQEE 841
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,051,420
Number of Sequences: 26719
Number of extensions: 1057770
Number of successful extensions: 3449
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 56
Number of HSP's that attempted gapping in prelim test: 3312
Number of HSP's gapped (non-prelim): 112
length of query: 1009
length of database: 11,318,596
effective HSP length: 109
effective length of query: 900
effective length of database: 8,406,225
effective search space: 7565602500
effective search space used: 7565602500
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 65 (29.6 bits)
Medicago: description of AC148819.4


