

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148819.3 + phase: 0 /pseudo/partial
(880 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
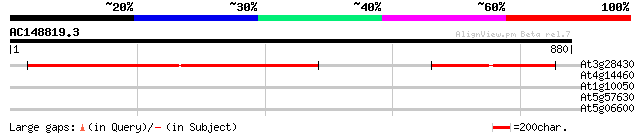
Score E
Sequences producing significant alignments: (bits) Value
At3g28430 unknown protein 511 e-145
At4g14460 retrovirus-related like polyprotein 33 0.54
At1g10050 putative xylan endohydrolase 31 2.7
At5g57630 CBL-interacting protein kinase 21 (CIPK21) 31 3.5
At5g06600 ubiquitin carboxyl-terminal hydrolase 30 4.5
>At3g28430 unknown protein
Length = 671
Score = 511 bits (1315), Expect = e-145
Identities = 270/456 (59%), Positives = 337/456 (73%), Gaps = 2/456 (0%)
Query: 29 MEKQVVGDFVRILKLSRTISIPLQLLQTVSIMIQNLQSEHAIYYMFSNEHMNYLITYSFD 88
MEKQV+G+FVRIL++S+T+++ +QLLQT+SIMIQNL+SE AIYY+FSNE++NYLITY+FD
Sbjct: 1 MEKQVMGEFVRILRVSKTVTVSVQLLQTMSIMIQNLKSEQAIYYLFSNEYVNYLITYTFD 60
Query: 89 FRNEELLSYYISFLRAIGGKLNKNTISLLVKTRNDEVVSFPLYVEAIRFAFHEESMIRAA 148
F++EELLSYYISFLRA+ GKLNK+TISLL+KT ND VVSFPLYVE I+FAFHEE+MIR A
Sbjct: 61 FQHEELLSYYISFLRAVSGKLNKHTISLLLKTENDVVVSFPLYVEGIQFAFHEENMIRTA 120
Query: 149 VRAVTLNVYHVGDDSVNRYITSEPHTDYFSNLVSFFRKQCMDLNRLISETLKNPGPDSNS 208
VRA+TLNVYHVGD+SVN Y+ S PHT+YFS L+SFF+KQCMDL+ ++ TLK+P PDS
Sbjct: 121 VRALTLNVYHVGDESVNDYVVSPPHTEYFSKLISFFQKQCMDLSAMVLNTLKSPSPDSGG 180
Query: 209 TVTTAVDEIEDNLYYFSDIISAGIPDVERLITDSILMLLIFPVLLPSLRTVADNDMQSGV 268
+ +AVD IED LYYFSD+ISAGIPD+ RLITD IL L P+LLPSL + AD +
Sbjct: 181 KLFSAVDGIEDTLYYFSDVISAGIPDIGRLITDHILQHLTLPLLLPSLCSEADISVDP-- 238
Query: 269 VTSLYLLCCILRIVKIKDLANTIVAALYYPLESFTKCSGGQVNGYIPDHGFTSKSDGIDN 328
VTSLYLL CILRIVKIKDLAN A L+ P+++F S + N + G T + D
Sbjct: 239 VTSLYLLSCILRIVKIKDLANMTAATLFCPVKAFISSSLVKPNSSLAPEGLTYVNGHPDK 298
Query: 329 DNLAENSAKCLVVNVPCSSSSSDCHPQSITILNNGSSSNVALREVLLEYVTGGADVQVLG 388
E + +C S + T + ++S++ RE LL+Y++ G DVQ G
Sbjct: 299 GVTEEANQQCSSTAAGMSDDGNSHLCSEDTPKSIFNNSHMTFRETLLQYISEGDDVQAQG 358
Query: 389 SLSVLATLLQTKELDESMLDGLGILPQRKQQKKLLLQALVGEASGEEQLFSSESSLTRDG 448
SL VLATLLQTKEL+ESMLD GILPQRKQ KKLLLQ+LVGE +GEEQLFS + RDG
Sbjct: 359 SLFVLATLLQTKELEESMLDAFGILPQRKQHKKLLLQSLVGEDTGEEQLFSPRNGSMRDG 418
Query: 449 IACGLDVYLNKIKEQYGVSFQPSNVVSSPRVPRFQV 484
++ LD YL +++EQ+GV PRV R QV
Sbjct: 419 LSSELDWYLRRLEEQFGVCCSLPGAARCPRVHRHQV 454
Score = 249 bits (635), Expect = 6e-66
Identities = 123/195 (63%), Positives = 149/195 (76%), Gaps = 6/195 (3%)
Query: 662 LVKVFVLLHQLQIFTHGRALPDQPLIYHPCDHRTNSRAHTSGL-AAVPKPGTEMNLVNAV 720
L++VFVLLHQLQIF+ GR+LP+QP IY P D SRA +GL +VPKPGTE+ LV+AV
Sbjct: 475 LIRVFVLLHQLQIFSLGRSLPEQPPIYPPADRSETSRATRAGLDVSVPKPGTELKLVDAV 534
Query: 721 PCRIAFERGKERHFCFLAISVGTSGWLVLAEELPLKKPYGVVRVAAPLAGCNPRVDDKHS 780
PCRIAFERGKER F FLA+S G SGW+VLA+ G+VRV APLAGC PR+D+KH
Sbjct: 535 PCRIAFERGKERDFSFLALSSGESGWIVLADP-----DNGIVRVTAPLAGCKPRIDEKHP 589
Query: 781 KWLHLRIRPSALPFLDPVKYNRHGKLKTKAFVDGRWILAFRDEESCKNAFLMIREEINHL 840
+WLHLRIRPS LP LDP K + KLK+K VDGRWILAFRD+ESC +A+ M+ EI+
Sbjct: 590 RWLHLRIRPSTLPLLDPTKRGVYEKLKSKGLVDGRWILAFRDDESCHSAYSMVAGEIDLQ 649
Query: 841 CDEVDRRIKPLLKLE 855
C EV+RR++PL LE
Sbjct: 650 CSEVERRLRPLFDLE 664
>At4g14460 retrovirus-related like polyprotein
Length = 1489
Score = 33.5 bits (75), Expect = 0.54
Identities = 30/111 (27%), Positives = 51/111 (45%), Gaps = 16/111 (14%)
Query: 764 VAAPLAGCNPRVDDKHSKWLHLRIRPSALPFLDPVKYNRHGKLKTKAFVDGRWILAFRDE 823
V P+ C D KW HL+ R FL K G +T+ R IL +
Sbjct: 177 VELPVCTCGRCECDAAVKWEHLQQRSRVTKFL---KELNEGFDQTR-----RHILMLKPI 228
Query: 824 ESCKNAFLMIREEINHLCDEVDRRIKPLLKLETVLDISSSSAPVSEDSSSH 874
+ K AF M+ + DE R +KPL ++++V +++ ++ED +++
Sbjct: 229 PTIKEAFNMVTQ------DERQRNVKPLTRVDSV--AFQNTSMINEDENAY 271
>At1g10050 putative xylan endohydrolase
Length = 1063
Score = 31.2 bits (69), Expect = 2.7
Identities = 26/89 (29%), Positives = 45/89 (50%), Gaps = 11/89 (12%)
Query: 314 IPDHGFTSKSDGIDNDNLAENSAKCLVVNVPCSSSSSDCHPQSITILNNGSSSNVALREV 373
I +H F SDG+ + N N VV SS+DC+ +S ++NN S + L +
Sbjct: 185 IKNHDF---SDGLYSWNT--NGCDSFVV------SSNDCNLESNAVVNNRSETWQGLEQD 233
Query: 374 LLEYVTGGADVQVLGSLSVLATLLQTKEL 402
+ + V+ G +V S+SV +L + ++
Sbjct: 234 ITDNVSPGFSYKVSASVSVSGPVLGSAQV 262
>At5g57630 CBL-interacting protein kinase 21 (CIPK21)
Length = 416
Score = 30.8 bits (68), Expect = 3.5
Identities = 21/68 (30%), Positives = 33/68 (47%), Gaps = 1/68 (1%)
Query: 692 DHRTNSRAHTSGLAAVPKPGTEMNLVNAVPCRIAFERGKERHFCFLAISVGTSGWLVLAE 751
D + N + GL+AVPK G ++ PC IA E + + A+ V + G ++L E
Sbjct: 143 DSKGNLKVSDFGLSAVPKSGDMLSTACGSPCYIAPELIMNKGYSGAAVDVWSCG-VILFE 201
Query: 752 ELPLKKPY 759
L P+
Sbjct: 202 LLAGYPPF 209
>At5g06600 ubiquitin carboxyl-terminal hydrolase
Length = 1126
Score = 30.4 bits (67), Expect = 4.5
Identities = 21/70 (30%), Positives = 28/70 (40%), Gaps = 14/70 (20%)
Query: 67 EHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKNTISLLVKTRNDEVV 126
E I +F HMNY+ + DF++ S+Y + L VK D
Sbjct: 312 EGTIQQLFEGHHMNYIECINVDFKSTRKESFY--------------DLQLDVKGCKDVYA 357
Query: 127 SFPLYVEAIR 136
SF YVE R
Sbjct: 358 SFDKYVEVER 367
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,572,885
Number of Sequences: 26719
Number of extensions: 840354
Number of successful extensions: 2035
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 2024
Number of HSP's gapped (non-prelim): 9
length of query: 880
length of database: 11,318,596
effective HSP length: 108
effective length of query: 772
effective length of database: 8,432,944
effective search space: 6510232768
effective search space used: 6510232768
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 65 (29.6 bits)
Medicago: description of AC148819.3


